Abstract
Sla1 is a Schizosaccharomyces pombe homolog of the human La protein. La proteins are known to be RNA-binding proteins that bear conserved RNA recognition motifs (La and RRMs), but their biological functions still have not been fully resolved. In this study, we show that the S. pombe La homolog (Sla1) is involved in regulating sexual development. Sla1 truncated in the C terminus (Sla1ΔC) induced ectopic sporulation in the ras1Δ strain and several other sporulation-deficient mutants. The C terminus contains a nuclear localization signal. While full-length Sla1 localizes in the nucleus, Sla1ΔC is found throughout the cell, suggesting the cytoplasmic localization of Sla1ΔC is involved in its sporulation-inducing activity. Further deletion analysis of Sla1 indicated that a small region (35 amino acids) that includes a portion of RRM2 is sufficient to induce sporulation. The La motif (RRM1) is not involved in this activity. Strikingly, Sla1ΔC induced haploid meiosis in a heterothallic strain, similar to the pat1-114 or mei2-SATA mutation. Sla1ΔC induced sporulation in a mei3 disruptant but not in a mei2 disruptant, indicating that Sla1ΔC requires Mei2 to induce haploid meiosis. Deletion of the chromosomal sla1 gene lowered the temperature sensitivity of the pat1-114 mutant. Two-hybrid analysis indicated that Pat1 interacts with Sla1ΔC but not full-length Sla1. Thus, Sla1ΔC may block Pat1 activity. This block would remove the inhibition on Mei2, which would then drive the cell into haploid meiosis. Finally, Sla1 was degraded prior to the start of meiosis when we monitored Sla1 in cells in which meiosis was synchronously induced. The ability of truncated Sla1 to induce ectopic meiosis represents a very novel function that has hitherto not been suspected for the La family of proteins.
Sexual development in the fission yeast Schizosaccharomyces pombe requires two simultaneously occurring conditions, namely, the presence of pheromone from cells of the opposite mating types and nutrient starvation. Under these conditions, cells of the opposite mating types, h+ and h−, conjugate to form diploid zygotes. The zygotes subsequently undergo the various steps of meiosis, i.e., karyogamy, premeiotic DNA synthesis, meiosis I, meiosis II, and sporulation. The switching of the cell cycle from mitosis to meiosis is regulated by many factors with complex mechanisms (64, 65). One of the many regulatory components involved in sexual development is Ste11, a key transcription factor that is the target of the signaling pathways that are initiated separately by nutrient starvation and the presence of mating pheromones. Ste11 is regulated mainly by the cyclic AMP-dependent signaling pathway (12, 52) and the stress-responsive pathway (65). The elevation of the levels of cyclic AMP, which is synthesized by adenylyl cyclase (17), negatively regulates Ste11 through Rst2 (12), which is phosphorylated by protein kinase A (26). Meanwhile, the activation of the stress-responsive mitogen-activated protein (MAP) kinase pathway composed of the MAP kinase cascade (Wis4/Wak1-Wis1-Spc1) positively regulates Ste11 through Atf1 (49, 54, 65).
Upon starvation and the presence of mating pheromone, Ste11 activates a number of genes required for mating and meiosis, including mei2+, mat1-P, and mat1-M (21, 23). The regions upstream of these genes all bear the TR box sequence motif, which contains the specific DNA sequence to which Ste11 binds (52). Mei2 is a key RNA-binding protein that is required for the initiation of meiosis in S. pombe (4, 48, 60, 61, 64). In vegetative cells, Mei2 is inactive due to the phosphorylation of its Ser438 and Thr527 amino acids by Pat1 kinase (62). In diploid cells, mei3+ expression is induced (32), and Mei3 inhibits Pat1 kinase (24, 33, 59). The dephosphorylated Mei2 then initiates the switch from the mitotic cell cycle to the meiotic cell cycle (62).
The temperature-sensitive pat1-114 mutant undergoes ectopic haploid meiosis at the restrictive temperature because Pat1 kinase is inactivated at that temperature (4). Cells that also undergo ectopic meiosis are those that bear the mutant mei2-SATA gene, which encodes a Mei2 protein whose Ser438 and Thr527 phosphorylation sites have been replaced by alanine. In these cells, constitutively active Mei2 that cannot be phosphorylated by Pat1 kinase accumulates, resulting in the initiation of meiosis (62). Other factors known to affect Mei2 activity includes its proteolysis through the ubiquitin system. It also appears to be regulated negatively by the 14-3-3 proteins in a manner that is dependent on the phosphorylation status of Mei2 (22, 47).
To signal the mating pheromone, S. pombe employs the MAP kinase pathway composed of Byr2 (MAP kinase kinase kinase), Byr1 (MAP kinase kinase), and Spk1 (MAP kinase). This signaling pathway is essential for conjugation and meiosis (36, 37, 56, 58) and is regulated by the Ras1 protein, a unique homolog of the mammalian Ras protein (35). Sterility is one of the phenotypes of the ras1Δ strain. Mutations in the other components in this pathway also lead to sterility (36, 37, 58). The 14-3-3 proteins were found to be negative regulators of this pathway through the binding of Byr2 (40).
To explore the Ras pathway further, we isolated two dominant mutations (sam3 and sam9) that bypass the sterile phenotype of ras1Δ cells (16). In this study, we report that when we screened for the genes that suppress the sterility of the ras1Δ strain by using a library constructed from the sam3 genome, we isolated a truncated form of sla1 that encodes a unique homolog of the La protein in S. pombe. The human La protein has been identified as being a major target of the autoimmune response in patients with autoimmune diseases (3, 27, 63). La is also known to be a multifunctional RNA-binding protein implicated in a number of transcriptional and posttranscriptional processes. For example, through its affinity for the 3′ oligo(U) tail and 5′ ends, La binds transcripts from RNA polymerase III and thus facilitates the initiation and termination of transcription (8, 9, 10, 13, 28, 41, 51). It has also been reported that Sla1 binds to the 3′ end of pre-tRNA and thereby facilitates its processing (57). However, it has never been reported that Sla1 or any of its homologs is involved in the sexual development of fission yeast or any other organisms. To further investigate this issue, we examined here the function of sla1+ in fission yeast sexual development. We found that the sporulation-inducing truncated form of Sla1 localizes in the whole cell and induces Mei2-dependent haploid meiosis in S. pombe.
MATERIALS AND METHODS
Strains, media, genetic methods, and S. pombe transformation.
The S. pombe strains used in this study are listed in Table 1. Yeast media YE (0.5% yeast extract, 2% glucose, 75 μg of adenine per ml) and YES, PM, and SD (5, 34) were used to routinely culture S. pombe strains. PM and SSA (7) were used to induce sporulation. We used a high-efficiency lithium acetate method with slight modifications to transform S. pombe (39). Briefly, cells that had been grown in 50 ml of PM minimal medium were collected by centrifugation (1,000 × g, 30 s), washed once in 10 ml of 0.1 M lithium acetate, and resuspended in 200 μl of 0.1 M lithium acetate. Cells were mixed with sample DNA and 300 μl of 50% polyethylene glycol 3350 and incubated at 30°C for at least 30 min. The cells were washed once with distilled H2O, resuspended in distilled H2O, and plated on a selective medium.
TABLE 1.
Fission yeast strains used in this study
| Strain | Genotype | Source |
|---|---|---|
| HS416RU | h90ura4-D18 ade6-216 sam3 ras1::ura4+ | Lab stock |
| SP870 | h90leu1-32 ura4-D18ade6-210 | D. Beach |
| YM71 | h−leu1-32 ura4-D18 | Lab stock |
| HM123 | h−leu1− | Lab stock |
| SPRUDA | h90leu1-32 ura4-D18 ade6-210 ras1::ura4+/h90leu1-32 ura4-D18 ade6-216 ras1::ura4+ | This study |
| SPRN1 | h90leu1-32 ura4-D18 ade6-210 ras1-ΔBglII-NheI | D. Beach |
| SPRN1A | h90leu1-32 ura4-D18 ade6-21 ras1-ΔBglII-NheI | D. Beach |
| SPRN1DA | h90leu1-32 ura4-D18 ade6-210 ras1-ΔBglII-NheI/h90leu1-32 ura4-D18 ade6-216 ras1-ΔBglII-NheI | D. Beach |
| SPBUD | h90leu1-32 ura4-D18 ade6-210 byr1::ura4+/h90leu1-32 ura4-D18 ade6-216 byr1::ura4+ | M. Wigler |
| SPSUD | h90leu1-32 ura4-D18 ade6-210 byr2::ura4+/h90leu1-32 ura4-D18 ade6-216 byr2::ura4+ | M. Wigler |
| SPKUD | h90leu1-32 ura4-D18 ade6-210 spk1::ura4+/h90leu1-32 ura4-D18 ade6-216 spk1::ura4+ | M. Wigler |
| TK105 | h90leu1-32 ura4-D18 phh1::ura4+ | T. Kato |
| JZ127 | h90leu1-32 ura4-D18 ade6-216 mei2::ura4+ | M. Yamamoto |
| JZ497 | h90leu1-32 ura4-D18 ade6-216 mei3::ura4+ | M. Yamamoto |
| KJ33-1A | h90leu1-32 ura4-D18 ade6-210 ste11::ura4+ | K. Kitamura |
| TW1 | h90leu1-32 ura4-D18 ade6-210 sla1::ura4+ | This study |
| TW5 | h90leu1-32 ura4-D18 ade6-210 ras1-ΔBglII-NheI sla1::ura4+/h90leu1-32 ura4-D18 ade6-216 ras1-ΔBglII-NheI sla1::ura4+ | This study |
| NOR 1 | h−leu1-32 ura4-D18 sla1+-gfp<<LEU2a | This study |
| NOR 12 | h90leu1-32 ura4-D18 sla1+-gfp<<LEU2 | This study |
| S964 | h−leu1-32 ade6-210 pat1-114/h−leu1-32 ade6-216 pat1-114 | P. A. San-Segundo |
| HT148 | h−leu1-32 ura4-D18 ade6-216 pat1-114 | Lab stock |
| KT101 | h90leu1-32 ura4-D18 ade6-216 pat1-114 sla1::ura4+ | This study |
| KT102 | h−leu1-32 ura4-D18 ade6-216 pat1-114 sla1::ura4+ | This study |
sla1+-gfp<<LEU2 indicates that LEU2 is close to sla1+-gfp.
Construction of sam3 genomic DNA library and cloning of sla1.
To construct the genomic DNA library based on the sam3 mutant (HS416RU), genomic DNA was prepared from a 20-liter culture of HS416RU (16) and purified with a CsCl gradient. Genomic DNA was partially digested with SacI or HindIII and fractionated in a 10 to 40% sucrose gradient. The DNA fractions larger than 4 kb were ligated with SacI- or HindIII-digested two-copy vector pYC11 3002 + 3 (6). The diploid ras1Δ strain SPRUHDA was transformed with the genomic library plasmids. The transformants were spread onto PM (+N) plates and incubated at 30°C for 3 days and then at 25°C for 2 days. Colonies were then stained with I2 vapor for several seconds. The dark sporulating colonies were selected, and their plasmids were rescued. That the recovered DNA clones could overcome the sterilizing effects of the ras1 gene loss in SPRUHDA was checked by transforming SPRUHDA with the clones again. The plasmid p416SacI was isolated in the screening of the library constructed by chromosomal digestion with SacI.
The full-length genomic sla1+ gene was isolated by colony hybridization from a genomic library based on the vector pWH5 (19), and sla1+ cDNA was obtained by plaque hybridization of a pBluescript KS(+) cDNA library (18). The resulting plasmids were designated pWH5-genomic sla1+ and pBluescript KS(+)-cDNA sla1+, respectively.
Gene disruption.
One-step gene disruption of sla1+ was performed as follows (44). The 800-bp HincII-HincII fragment was replaced by the 1.8-kb ura4+ cassette derived from pHSG398-ura4 (55). Cells of the wild-type strain (SP870) and the ras1− strains (SPRN1 and SPRN1A) (40) were transformed with the 3.8-kb sla1::ura4+ DNA fragment and then spread on SD plates in the absence of uracil. Stable Ura+ transformants were then selected by allowing the strains to grow in rich medium (YE). Southern blot analysis, performed as described before (46) with the sla1 and ura4 probes, confirmed that the sla1 allele was properly disrupted in these stable Ura+ cells (data not shown). To obtain a diploid ras1Δ sla1Δ strain, we employed a protoplast fusion method as described previously (1). To obtain a pat1-114 sla1Δ strain, TW1 (h90 sla1::ura4) was crossed with HT148 (h− pat1-114) on MEA medium and incubated at 25°C for 3 days. The cells were diluted in 30% ethanol to kill vegetative cells, plated on PMAL medium containing Phloxine B, and incubated at 25°C for 5 days. The white colonies that appeared on these plate were regrown on YES plate and then replica plated to grow at 25 and 34°C. The cells that retained temperature sensitivity at 34°C were, selected and the strains were named KT102 (h− pat1-114 sla1::ura4) and KT103 (h90 pat1-114 sla1::ura4). The sla1 deletion of KT102 was confirmed by the absence of the Sla1 protein as determined by Western blotting with an anti-Sla1 antibody.
Plasmids.
pBluescript KS(+)-cDNA sla1+ was used as the template for PCR cloning. pREP1-Sla1FL, pSLF173L-Sla1FL, and pSla1FL-GFP were constructed as follows. A 1.0-kb sla1 FL fragment was amplified by PCR with the oligonucleotide primers (sla1FL) shown in Table 2. The PCR product was ligated into pT7 Blue(R) vector (Novagen). The SalI-SalI fragment of the resulting plasmid was then inserted into the same site of pREP1 (no tag) (30, 31). Alternatively, the NotI-NotI fragment of the resulting plasmid was inserted into the same sites of either pSLF173L (the LEU2 version of pSLF173), which contains the 3HA epitope (11, 55), or pSLF172L-GFP. pSLF172L-GFP is derived from pSLF172L (pSLF172L is the LEU2 version of pSLF172) (11) in which the NotI-SalI fragment, including the 3HA tag, has been replaced with the NotI-SalI fragment containing GFPS65A. pSFL173L-Sla1ΔC, pSFL173L-Sla1RRM, pSFL173L-Sla1La, and pSFL173L-Sla1CT were constructed by a method similar to that used for pSFL173L-Sla1FL. To make pSFL173L-Sla1ΔRNP1, pSFL173L-Sla1ΔRNP21, pSFL173L-Sla1ΔRNP2, and pSFL173L-Sla1RRM (FS), the relevant portions of the sla1 fragments were amplified by PCR with oligonucleotide primers sla1ΔRNP1, sla1ΔRNP21, sla1ΔRNP2, and sla1RRM(FS), respectively (Table 2). The PCR products were ligated into the EcoRV site of pBluescript KS(+), and then the NotI-NotI fragments were inserted into the same site of pSLF173L. pSla1FL-GFP, pSla1ΔC-GFP, pSla1RRM-GFP, pSla1La-GFP, and pSla1CT-GFP in Fig. 4 were derived from pSLF172L-GFP. They were constructed by using the same fragments used to construct the pSLFL173-derived plasmids in Fig. 3. pSla1RERL-GFP and pSla1RERL-CT-GFP were constructed by PCR-based site-directed mutagenesis according to the Takara manual. Replacement of two Rs (CGC and AGA) by E (GAG) and L (CTC) was performed by using primer KELR (Table 2). The PCR fragment with the putative nuclear localization signal (NLS) was ligated into pT7 Blue(R), and the NotI-NotI fragment was inserted into the same site of pSLF172L-GFP.
TABLE 2.
Oligonucleotide primers used in this study
| Primer | Sequencea |
|---|---|
| sla1FL | 5′TAGTCGAC(SalI)ATGCGGCCGGC(NotI)ATGTCAACTGAAGAACAGAAGG3′ |
| 5′TAGTCGAC(SalI) TTAGCGGCCGC (NotI)ACTCCGAAGCGGCCGAAGGCT3′ | |
| sla1ΔC | 5′-TAGTCGAC(SalI) ATGCGGCCGGC (NotI)ATGTCAACTGAAGAACAGAAGG3′ |
| 5′-TAGTCGAC(SalI) TTAGCGGCCGC (NotI) TGAGCTCGTCCTCGCCCC3′ | |
| sla1La | 5′TAGTCGAC(SalI) ATGCGGCCGGC (NotI) ATGTCAACTGAAGAACAGAAGG3′ |
| 5′TTTGTCGAC(SalI) TCAGCGGCCGC (NotI) TTTCCATGACGCTCTTG3′ | |
| sla1RRM | 5′ATGTCGAC(SalI) TAGCGGCCGC (NotI)ATGAGTCCTGAACTTCTT3′ |
| 5′TAGTCGAC(SalI) TTAGCGGCCGC (NotI) TGAGCTCGTCCTCGCCCC3′ | |
| sla1CT | 5′AAAGTCGAC(SalI) TGCGGCCGC(NotI) ATGGAGCTCACTATAATG3′ |
| 5′-TAGTCGAC(SalI) TTAGCGGCCGC (NotI) ACTCCGAAGCGGCCGAAGGCT3′ | |
| sla1ΔRNP1 | 5′ATGTCGAC(SalI) TAGCGGCCGC (NotI) ATGAGTCCTGAACTTCTT3′ |
| 5′-TTAGCGGCCGC(NotI) TAAATTTTTTATCGTC3′ | |
| sla1ΔRNP21 | 5′AAAGCGGCCGC(NotI) ATGGGTGATGAAAAAGAT3′ |
| 5′TTAGCGGCCGC(NotI) TAAATTTTTTATCGTC3′ | |
| sla1ΔRNP2 | 5′AAAGCGGCCGC(NotI) ATGGGTGATGAAAAAGAT3′ |
| 5′TAGTCGAC(SalI) TTAGCGGCCGC (NotI)TGAGCTCGTCCTCGCCCC3′ | |
| sla1RRM(FS) | 5′TAGCGGCCGC(NotI)ATGGTCCTGAACTTCTT3′ |
| 5′TAGTCGAC(SalI) TTAGCGGCCGC (NotI)TGAGCTCGTCCTCGCCCC3′ | |
| Mutagenesis primer (KRRR to KELR) | 5′AAAGAGCTCCGTTTTGACGCTTTT3′ |
| 5′AAAGAGCTCTTTGCTGCTAAACTT3′ |
The upper and lower sequences indicate the sense and antisense primers, respectively. The boldface and underlined sequences in the mutagenesis primers indicate the mutated region. Restriction enzyme sites are underlined.
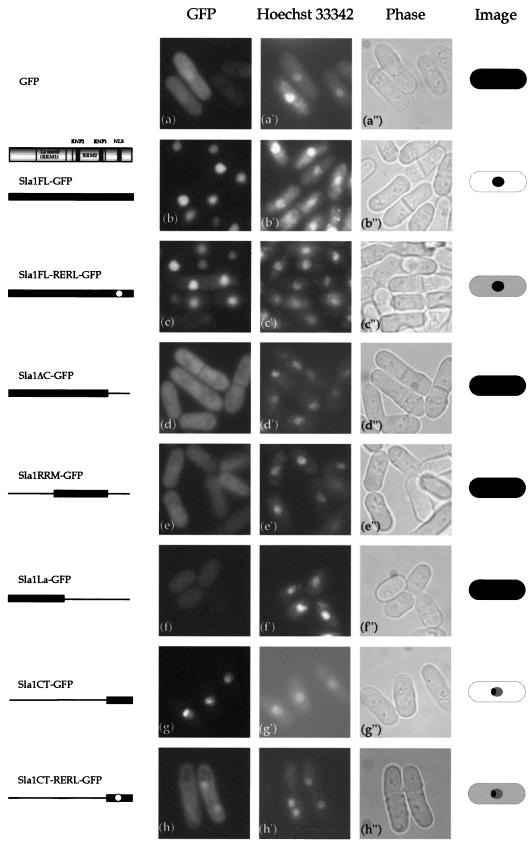
FIG. 4.
Subcellular localization of full-length or truncated Sla1-GFP proteins by live observation of cells. GFP fusion proteins were expressed under control of the nmt1 promoter in the vector pSLF172L-GFP. h90 wild-type cells (SP870) carrying each plasmid were grown in PM (+N) medium and incubated at 25°C. Cells were examined for GFP fluorescence, and the same cells were stained with the DNA-binding dye Hoechst 33342. Images of the cells are depicted on the right.
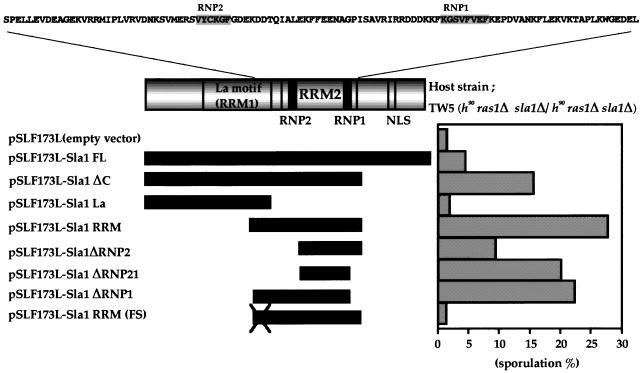
FIG. 3.
C-terminally truncated Sla1 and RRM2 induce sporulation in S. pombe TW5. The effect on sporulation of transforming the TW5 strain with constructs expressing Sla1 or a truncated variant is shown. The TW5 (h90/h90 ras1Δ/ras1Δ sla1Δ /sla1Δ) strain was transformed with the vector pSLF173L, with pSLF173L-Sla1FL (which overexpress sla1+), or with various pSLF173L constructs encoding truncated Sla1. The transformants were streaked on PM (+N) medium and incubated for 72 h at 25°C, and the percentage of sporulated cells was determined by counting at least 1,000 cells.
GFP tagging.
Green fluorescent protein (GFP) carrying the S65A mutation was fused to the C terminus of the genomic sla1+ gene in the integration vector pYC11, a derivative of pBluescript KS(+) that retains the LEU2 marker (53). The wild-type haploid strain YM71 was transformed with pYC11-Sla1-GFP, and stable Leu+ integrants were selected and checked by Southern blot analysis with sla1 and GFP probes (data not shown). The resulting strain was named NOR 1. To obtain an h90 isogenic strain with this tag, NOR 1 was crossed with SP870. The h90 strain that retained sla1+-gfp on its chromosome was named NOR 12 (see Fig. 2A).
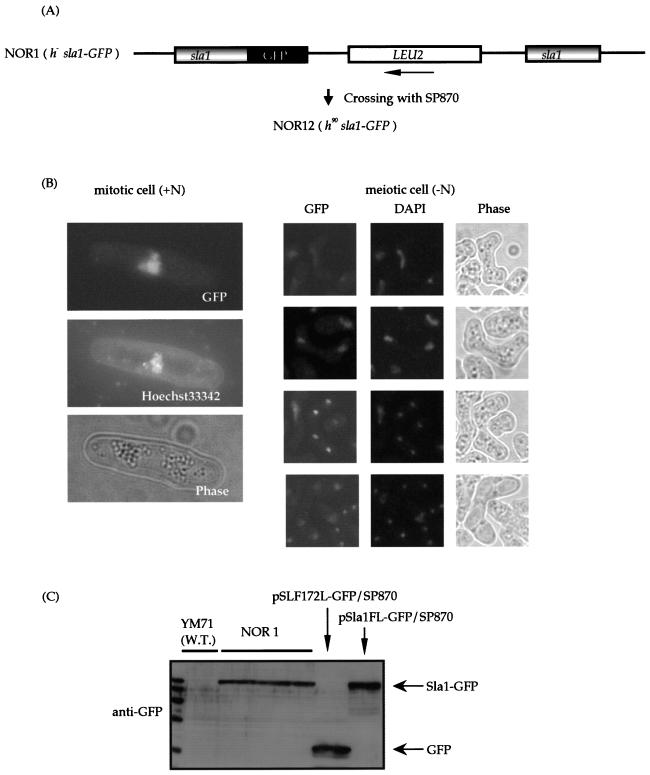
FIG. 2.
Sla1 localizes in the nucleus. (A) Strategy used to construct the genomic sla1-gfp fusion gene. (B) Subcellular localization of Sla1 tagged with GFP. NOR 12 was grown in PM (+N) minimal medium and shifted to PM (−N) medium to induce meiosis. Cells right after the shift (mitotic cell, +N) and cells incubated on PM (meiotic cell, −N) were examined for GFP fluorescence. Cells were counterstained with the DNA-binding dye Hoechst 33342 or DAPI to observe their nuclei. (C) Western blot analysis of Sla1-GFP. The Sla1-GFP proteins from NOR 1 cells and the wild-type (W.T.) strain harboring the plasmid were detected without degradation.
Fluorescence microscopy of GFP fusion proteins.
Wild-type strains harboring various plasmids containing GFP-fused sla1 and the strains containing the genomically GFP-fused sla1 were cultured in PM at 25°C to the mid-log phase. GFP fluorescence images of living cells were taken with a Hamamatsu C5985 charge-coupled device camera. Cells were counterstained with Hoechst 33342 or DAPI (4′,6′-diamidino-2-phenylindole) to visualize the nuclei.
Cell extracts and Western blotting.
About 108cells of S. pombe were harvested. Pellets were washed with STOP buffer (150 mM NaCl, 50 mM NaF, 10 mM EDTA, 1 mM NaN3 [pH 8.0]) and stored at −80°C. The pellets were diluted in 100 μl of distilled water (dH2O) and boiled at 95°C for 5 min, and then 120 μl of 2× Laemmli buffer (4% sodium dodecyl sulfate [SDS], 20% glycerol, 0.6 M β-mercaptoethanol, 0.12 M Tris-HCl [pH 6.8]) containing 8 M urea and 0.02% bromophenol blue was added to the samples, which were vigorously vortexed with an equal volume of zirconium-silica beads for 3 min and then heated again 95°C for 5 min. The zirconium-silica beads and large debris were then removed by centrifugation at 10,000 × g for 15 min. Approximately equal amounts of each sample were analyzed by SDS-polyacrylamide gel electrophoresis with a 10 to 15% polyacrylamide gel and then transferred to Immobilon transfer membranes (Millipore) by use of a wet-type transfer system. For detection of GFP fusion proteins, membranes were incubated with an anti-GFP polyclonal antibody (Molecular Probes, Inc.) diluted 1:3,000 in 5% dry milk in TBS-T (15 mM Tris, 137 mM NaCl, 0.1% Tween20), washed, and then incubated with horseradish peroxidase-conjugated anti-rabbit secondary antibody (Bio-Rad Laboratories) diluted 1:5,000 in 5% dry milk in TBS-T. The secondary antibodies were detected with the ECL system as described by the manufacturer (Amersham). For detection of untagged Sla1p, membranes were incubated with an anti-Sla1 polyclonal antibody diluted 1:500 in 2% dry milk in PBS-T (137 mM NaCl, 8 mM Na2HPO4·12H2O, 2.7 mM KCl, 1.5 mM KH2PO4, 0.1% Tween20), washed, and then incubated with horseradish peroxidase-conjugated anti-rabbit secondary antibody diluted 1:5,000 in 2% dry milk in PBS-T. The secondary antibodies were detected with the ECL-plus system (Amersham). For detection of Cdc2p, membranes were incubated with an anti-PSTAIRE polyclonal antibody (Santa Cruz Biotechnology, Inc.) diluted 1:3,000 in 2% dry milk in PBS-T, washed and then incubated with horseradish peroxidase-conjugated anti-rabbit secondary antibody diluted 1:2,000 in 2% dry milk in PBS-T. The secondary antibodies were detected with the ECL system (Amersham).
RNA preparation and Northern blotting.
Total RNA from S. pombe cells was prepared as follows. About 108 cells grown in an appropriate medium were washed with dH2O, resuspended in 0.5 ml of Isogen RNA isolation reagent (Nippon Gene), and vigorously vortexed with an equal volume of zirconium-silica beads for 5 min. After centrifugation at 10,000 × g for 15 min, nucleic acid in the supernatant was precipitated with isopropanol. The RNA was resolved on formaldehyde-agarose gels and transferred to a membrane (Hybond N+). PCR fragments of sla1and leu1 were used as probes. The probe was labeled with [α-32P]dCTP (Amersham) by using the BcaBEST labeling kit (Takara).
Yeast two-hybrid assay.
Saccharomyces cerevisiae strain L40 was transformed with pBTM116 carrying the SmaI-PstI fragment of full-length sla1+ or sla1ΔC fused to the LexA DNA binding domain together with pACT II carrying pat1+ fused to the GAL4 activating domain (29). The combination of pat1 and mei2 was used as a positive control. The β-galactosidase activity was measured as described previously (42) with o-nitrophenyl-β-d-galactopyranoside as a substrate.
Production of antibody against the Sla1 protein.
To raise a polyclonal antibody against the Sla1 protein, the glutathione S-transferase (GST)-fused Sla1 protein was expressed in Escherichia coli and purified. The 897-bp cDNA fragment corresponding to the full length of sla1+ was amplified by PCR, cloned into the SmaI-BamHI site of pGEX-1 (Amersham) (a GST fusion protein expression vector), and expressed in E. coli BL21(DE3). The soluble GST-Sla1 fusion protein was purified from E. coli extract by using glutathione-Sepharose 4B (Amersham) and separated on a 10% SDS-acrylamide gel. The purified GST-Sla1 protein eluted from the gel was used as an antigen for generation of a polyclonal antibody in rabbit. About 2 mg of protein was used to raise an antibody. Steps from the injection of antigen into a rabbit to create antiserum were ordered to Sawady technology. Western blotting was performed with an affinity-purified anti-Sla1 antibody. The bacterially expressed GST-Sla1 fusion protein was bound to a polyvinylidene difluoride membrane in advance, and it was used to purify an anti-Sla1 antibody from crude serum by absorbing and releasing.
pat1-driven synchronous meiosis.
h−/h− pat1-114/pat1-114 diploid cells (S964) were cultured in YES medium at 25°C for 1 day and transferred to PM plus leucine (100 μg/ml) the next day. The cells were washed three times with dH2O and transferred to PM minus NH4Cl plus leucine (50 μg/ml) at a density of 1 × 106 to 2 × 106 cells/ml. After the cells were incubated for 15 h at 25°C, most cells were arrested at G1 phase. The culture was shifted to 34°C, in the presence of 0.5 g of NH4Cl per liter and 50 μg of leucine per ml, to inactivate the Pat1 kinase to induce meiosis. Samples were taken every 1 h or 30 min for 10 h.
RESULTS
Isolation of the sla1 gene.
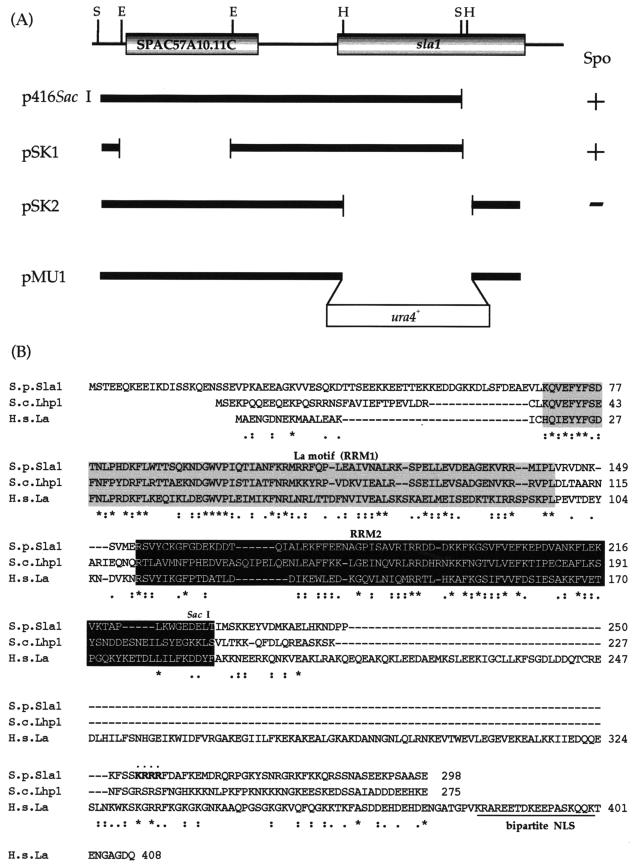
We found earlier that the presence of the sam3 mutation is dominant and can bypass the sporulation-deficient (Spo−) phenotype in ras1 null diploid cells (16). To isolate the sam3 mutant allele, we transformed ras1 null diploid cells with a library constructed from the chromosomal DNA of the sam3 mutant and screened for genes that converted the ras1 null diploid cells to Spo+. We obtained a clone named p416SacI that contained two genes. One was a full-length open reading frame of an unknown gene (SPAC57A10.11c), and the other was the sla1 (S. pombe La) gene encoding a protein truncated in its C terminus (sla1ΔC) (Fig. 1A). When we deleted the sla1 gene fragment from p416SacI and transformed ras1− cells with the resulting fragment, the host cells remained Spo− (Fig. 1A). In contrast, when SPAC57A10.11c was deleted from p416SacI, the ras1− cells became Spo+ (Fig. 1A). Thus, sla1ΔC is involved in suppressing the ras1− phenotype.
FIG.1.
Structure of the sla1+ gene. (A) Restriction map, subcloning, and construction of null mutants. Restriction enzyme sites: S, SacI; E, Eco T221; H, HincII. Spo indicates the effect of transforming the ras1Δ diploid strain with the subclones. (B) Comparison of the amino acid sequences of Sla1, human La, and S. cerevisiae Lhp1. Identical amino acids are marked with asterisks. The La protein-specific motif (RRM1) is shaded. RRM2 is shown in white against black. The putative NLS of Sla1 is indicated in boldface. The presumable basic bipartite NLS of human La is underlined. S.p., S. pombe; S.c., S. cerevisiae; H.s., Homo sapiens.
Comparison of the predicted protein sequences of sla1ΔC and the wild type showed that the slaΔC gene in p416SacI had the same sequence as the wild type. Furthermore, when we cloned and sequenced the entire sla1 allele from the sam3 mutant, sequence differences were not found. Thus, we concluded that sla1 is not the relevant allele of sam3. As the sla1ΔC gene in p416SacI was truncated, we obtained the full-length genomic sla1 gene by colony hybridization. The sla1+ cDNA (GenBank accession number AB011371) was obtained by plaque hybridization. Interestingly, while sla1ΔC could suppress the Spo− phenotype of the ras1 null diploid strain, the full-length sla1 had almost no effect (see Fig. 4). Thus, Sla1 must be truncated at its C terminus to suppress the Spo− phenotype of the ras1Δ strain.
sla1+ encodes a homolog of La protein in S. pombe.
The sla1+ gene encodes a 298-amino-acid protein that is a unique La protein homolog in S. pombe. The overall homologies of Sla1 with human La and the S. cerevisiae homolog Lhp1 are 37 and 35%, respectively (Fig. 1B). The La protein family is known to bear RNA recognition motifs (La and RRMs). The N-terminal motif is particularly conserved between the La protein homologs and is designated the La motif (38). Like the other La proteins, Sla1 bears a La motif (RRM1), RRM2, and a highly charged C terminus. We originally named this gene sla1 and registered it in the National Center for Biotechnology Information database before finishing our analysis. Several studies of sla1 have appeared in the meantime (15, 43, 57). With regard to function, Sla1 is now known to be a nonessential phosphorylated RNA-binding protein (57). However, the possibility that it is involved in the sexual development of S. pombe has not been previously reported. Consequently, we concentrated on analysis of the functional roles that sla1+ plays in sexual development.
Phenotype of a sla1+ disruptant.
To further assess the cellular function of sla1+, we independently made a sla1 disruption strain. Using the ura4+ cassette, the sla1::ura4+ gene was constructed and used to disrupt the chromosomal sla1 gene in the haploid wild-type strain SP870 (Fig. 1A). That sla1+ had been disrupted was verified by Southern blotting with both sla1 and ura4 as probes (data not shown). The resulting sla1Δ strain was named TW1 (sla1::ura4+). TW1 grew as well as the wild type, indicating that sla1+ is not an essential gene, as shown by Van Horn et al. (57). Despite the fact that we isolated sla1+ because it could reconstitute the sporulation deficiency of the ras1 null diploid strain, the conjugation and sporulation rate of the sla1 disruptant appeared to be almost normal. However, when we more carefully examined this, we noticed that the sla1 disruptant sporulated slightly faster than the wild-type strain. The sla1 disruptant formed spores with a 38.1% ratio at 24 h after cells were plated on SSA medium at 30°C, while the parental wild-type SP870 formed spores with a 25.5% ratio.
Sla1p localizes in the nucleus in fission yeast.
It was previously reported that the human, rat, fruit fly, and budding yeast La proteins bear the NLS and localize in the nucleus, where they can associate with newly synthesized RNA polymerase III transcripts such as the tRNA precursors (20, 25, 43, 50, 66). The localization of S. pombe Sla1 in the nucleus of budding yeast was shown by Rosenblum et al. (43). To assess the localization of Sla1p in fission yeast, we constructed a haploid wild-type strain, designated NOR 12, that carries a single copy of the sla1+ gene tagged with GFP (Fig. 2A). To obtain an h90 isogenic strain with this tag, NOR 1 was crossed with SP870. The h90 strain that retained sla1+-gfp on the chromosome was named NOR 12. In vegetative NOR 1 cells (+N) as well as in meiotic NOR 12 cells (−N) at all stages of meiosis, Sla1-GFP is localized predominantly in the nucleus (Fig. 2B). Thus, Sla1p localizes in the nucleus throughout the vegetative and meiotic cell cycles. Western blotting of NOR 1 as well as SP870 cells harboring pSLF172-Sla1-GFP revealed that Sla1-GFP is present in the cells at the expected molecular weight without any degradation (Fig. 2C). The nuclear localization of the Sla1 protein could also be observed by indirect immunostaining with an antibody against Sla1 (data not shown).
Sla1 induces sporulation when its C terminus is deleted.
Full-length sla1+ and various truncated forms were inserted into the pSLF173L vector (hemagglutinin [HA] tagged). The ras1 null diploid strain was transfected with each plasmid to investigate the effect on its sporulation efficiency. Full-length sla1+ and the truncated variants were also fused to the N terminus of GFP in the pSLF172L-GFP vector to investigate the subcellular localization of Sla1p and the truncated proteins. Consistent with the previous observations, full-length Sla1 (pSLF173L-Sla1FL) could not efficiently induce sporulation (Fig. 3). In contrast, when C-terminally truncated Sla1ΔC was expressed under the control of the nmt1 promoter (pSLF173L-Sla1ΔC), it induced efficient sporulation (Fig. 3). When the GFP-linked proteins were examined, Sla1FL-GFP was found to localize in the nucleus but Sla1ΔC-GFP was diffused throughout the entire cell (Fig. 4). Thus, Sla1 induces sporulation only when its C terminus is deleted. In addition, we observed that the GFP tag did not disturb the sporulation-inducible effect of Sla1ΔC (data not shown).
The C terminus must contain the NLS, as nuclear localization is observed only when the C terminus is intact. Examination of the amino acid sequence of Sla1 indeed revealed the presence of the basic monopartite NLS-like sequence K-R-R-R in its C terminus (Fig. 1B). To investigate the role of this sequence, we mutated it in Sla1FL and Sla1CT (the C-terminal region of Sla1) into K-E-L-R by site-directed mutagenesis, generating Sla1FL-RERL and Sla1CT-RERL, respectively. Sla1FL-RERL-GFP localized in both the nucleus and the cytoplasm (Fig. 4). The change of localization due to the mutated sequence was particularly obvious when the C-terminal regions bearing the wild-type K-R-R-R and the mutant K-E-L-R were compared (Sla1CT versus Sla1CT-RERL) (Fig. 4). Thus, it appears that the C-terminal sequence K-R-R-R is a part of the NLS of Sla1.
To further analyze the functional domains of Sla1, Sla1ΔC was divided into two domains, one that included the N terminus and the La motif (Sla1La) and one that contained the second RRM (Sla1RRM). Like the Sla1ΔC protein, both constructs localized throughout the entire cell (Fig. 4). However, Sla1La could not induce sporulation in ras1Δ diploid cells (Fig. 3). One of the functions of Sla1 is to stabilize the 3′ end of newly synthesized RNA polymerase III transcripts (57). This function is mediated by the La motif (RRM1) and RRM2. That Sla1La does not induce sporulation suggests that this stabilization function of Sla1p is not involved in sporulation in S. pombe. In contrast, Sla1RRM (which contains RRM2) induced sporulation in the ras1Δ strain as efficiently as Sla1ΔC (Fig. 3). Thus, the effect of Sla1 on sporulation is dependent on RRM2.
To further investigate the role of RRM2, we constructed deletion mutants of Sla1RRM that lacked either or both of the RNP motifs in RRM2. Surprisingly, even when both RNP1 and RNP2 had been deleted (pSLF173L-Sla1ΔRNP21), leaving only a small 35-amino-acid region, ectopic sporulation was still induced (Fig. 3). Thus, it appears that the sporulation-inducible effect of Sla1ΔC depends on a domain contained in RRM2 but does not require the entire RRM. To test whether Sla1ΔRNP21 functions as a protein product, a frameshift was introduced into the 5′ terminus of Sla1RRM. This mutant (Sla1RRM-FS) could not induce sporulation in the ras1 null diploid strain (Fig. 3). Thus, we conclude that the sporulation-inducing truncated forms of Sla1, including Sla1ΔC and Sla1RRM, act in sporulation as protein products rather than as RNA molecules. In all cases of the truncated variants, we used Western blotting to check the amount of the fusion protein produced. These observations indicated that Sla1 variants that could not induce sporulation were properly expressed, indicating that protein instability is not the cause of the ineffective sporulation (data not shown).
Sla1ΔC induces sporulation in other sporulation-defective strains.
We transformed the byr2Δ, byr1Δ, and spk1Δ strains with pREP1-Sla1ΔC to investigate whether Sla1ΔC could induce sporulation in mutants that are defective in sporulation due to mutations in genes operating downstream of the Ras pathway. We found that pREP1-Sla1ΔC induced sporulation in all three mutant strains (Table 3). pREP1-Sla1ΔC also induced sporulation in the stress-responsive MAP kinase mutant strain (spc1/phh1Δ) (Table 3). Thus, the action of Sla1ΔC is not limited to the ras1 pathway.
TABLE 3.
Effect of Sla1ΔC expression on sporulation of various sporulation-defective mutants
| Genotype | % Sporulation with:
|
||
|---|---|---|---|
| pREP1 | pREP1-Sla1FL | pREP1-Sla1ΔC | |
| byr2Δ/byr2Δa | 0 | 0 | 15.3 |
| byr1Δ/byr1Δa | 0 | 0 | 17.6 |
| spk1Δ/spk1Δa | 0 | 0 | 11.0 |
| phh1Δa | 0 | 0 | 9.8 |
| mei3Δb | 0 | NTc | 22.2 |
| mei2Δb | 0 | NT | 0 |
| ste11Δb | 0 | NT | 0 |
Each transformant was streaked onto a PM (+N) plate and incubated at 25°C for 72 h. Under these conditions wild-type cells sporulate more than 50%. The sporulation efficiency of at least 1,000 cells was then examined by counting the sporulating cells.
Each transformant was streaked onto a PM(+N)-adenine (75 mg/liter) plate and incubated at 25°C for 72 h. The sporulation efficiency of at least 1,000 cells was then examined by counting the sporulating cells.
NT, not tested.
Sla1ΔC induces haploid meiosis in the h− haploid strain.
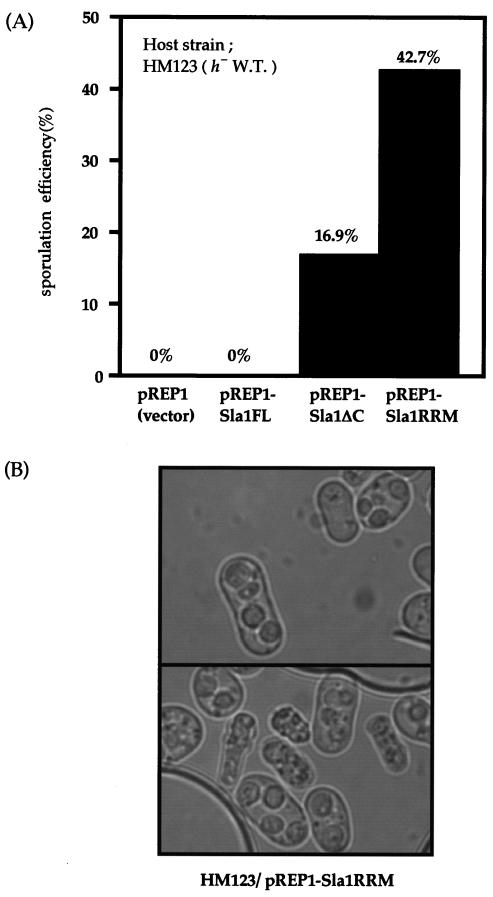
As expression of Sla1ΔC can compensate for the sporulation-blocking mutations of many of the components in the signal transduction pathway related to sexual development, we examined whether Sla1ΔC could also induce meiosis in an h− haploid strain. HM123 (h−) was transformed with pREP1 (empty vector), pREP1-Sla1ΔC, or pREP1-Sla1RRM. Transformants were streaked onto a PM (+N) plate to examine the sporulation efficiency. Both Sla1ΔC and Sla1RRM induced haploid meiosis (Fig. 5). When pREP1-Sla1RRM was transformed in the heterothallic strain, the induction of haploid meiosis was extremely high (42.7%), and aberrant spores were observed in those cells (Fig. 5B). Such haploid meiosis has been previously observed in the pat1-114 strain (4) and in a strain that expresses mei2-SATA (62). The high induction of haploid meiosis by Sla1RRM is comparable to that by pat1-114, and it suggests that Sla1ΔC is involved in regulating Mei2 or Pat1 rather than in regulating the Ras pathway that operates upstream of these proteins.
FIG. 5.
Sla1ΔC induces haploid meiosis in the h− wild-type strain. (A) h− wild-type (W.T.) strain HM123 was transformed with vector pREP1, pREP1-Sla1ΔC, and pREP1-Sla1RRM. The transformants were streaked on PM (+N) medium and incubated for 72 h at 25°C, and the percentage of sporulated cells was determined by counting at least 1,000 cells. (B) Cell morphology of the haploid meiosis induced in HM123 due to Sla1ΔC expression. The plasmid pREP1-Sla1ΔC or pREP1-Sla1RRM caused the haploid cells to undergo a meiotic catastrophe. The resulting zygotes, including the spore-like bodies, are shown.
Sla1ΔC acts downstream of Mei3 and upstream of Mei2.
To locate where Sla1ΔC acts, we investigated whether or not Sla1ΔC can induce the mei2, mei3, and ste11 null strains to sporulate. As is well known, Ste11 is the essential transcription factor for mei2, and Mei3 inhibits Pat1, a negative regulator of Mei2 (65). Sla1ΔC induced sporulation in the mei3Δ strain but not in the mei2Δ or ste11Δ strain (Table 3). Thus, Sla1ΔC functions upstream of Mei2 and downstream of Mei3.
Pat1p interacts with Sla1ΔC but not Sla1FL in the yeast two-hybrid assay.
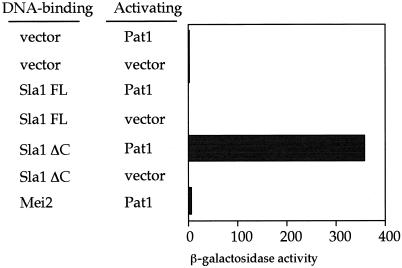
The observations described above suggest that Sla1ΔC may function to inhibit Pat1. This is also how Mei3 inactivates Pat1 (59). This idea was assessed by use of the yeast two-hybrid assay. Sla1FL and Sla1ΔC were fused to the LexA DNA binding domain. S. cerevisiae strain L40 was transformed with either of these plasmids together with Pat1 fused to the GAL4 activating domain (29), and the β-galactosidase activity was assayed. The combination of Sla1ΔC and Pat1 showed high-level β-galactosidase activity, whereas the Sla1FL-Pat1 pair did not (Fig. 6). Another two-hybrid experiment failed to show that Sla1ΔC or Sla1FL interacts with Mei2 (data not shown). Thus, Pat1 and Sla1ΔC interact, and the truncation of the C-terminal portion of Sla1 is necessary for this interaction.
FIG. 6.
Yeast two-hybrid assay. S. cerevisiae strain L40 was transformed with plasmids harboring Sla1FL, Sla1ΔC, or Pat1 fused with the DNA binding domain or the DNA activating domain. The β-galactosidase activity (in nanomoles per minute per milligram of protein) of the resulting transformants was assayed by using o-nitrophenyl-β-d-galactopyranoside.
Suppression of the pat1-114 mutant by deletion of chromosomal sla1.
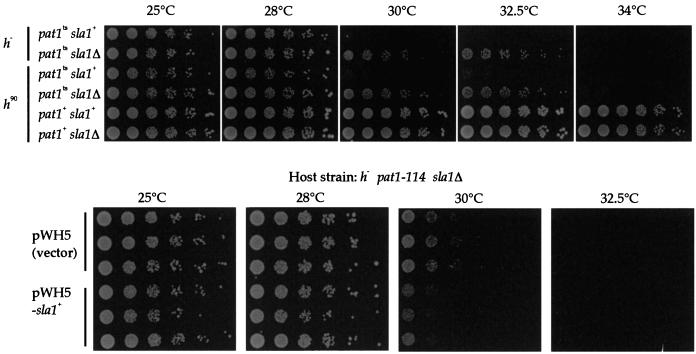
We thought that if the target of truncated Sla1 is Pat1 kinase, the temperature sensitivity of the pat1-114 mutant might be rescued by deletion of the chromosomal sla1 gene. We made a pat1-114 sla1Δ double mutant as described in Materials and Methods and compared its temperature sensitivity with that of the parental pat1-114 mutant by serial dilution of cells. While the pat1-114 mutant could not grow at 34°C and grew slowly at 30°C and the pat1+ strain grew at any temperature, the pat1-114 sla1Δ double mutant grew better at 30 and 32.5°C than the pat1-114 mutant and grew slightly at 34°C (Fig. 7, upper panels). Thus, the suppression of the growth retardation of the pat1-114 mutant at the restrictive temperature (32.5°C) was observed in the pat1-114 sla1Δ double mutant. To confirm this observation, the growth of the pat1-114 sla1Δ strain with the vector alone and the plasmid retaining sla1 was compared by serial dilution (Fig. 7, lower panels). The reacquisition of temperature sensitivity at 30°C was observed. We also tested the effect of overexpression of the sla1 gene in the pat1-114 strain, but we did not see any apparent effect on temperature-sensitive growth of the pat1-114 strain (data not shown).
FIG. 7.
Lowering of the temperature sensitivity of the pat1-114 mutant by sla1 deletion. (Upper panel) Fivefold serial dilutions of pat1-114, pat1-114 sla1Δ, and sla1Δ cultures were spotted on YES medium and incubated at 25, 28, 30, 32.5, and 34°C. Two independently isolated strains with different mating type were tested. (Lower panel) The pat1-114 sla1Δ strain was transformed with pWH5 or pWH5-sla1+. Fivefold serial dilution of transformants were spotted on PMA medium and incubated at 25, 28, 30, and 32.5°C.
Detection of the Sla1 protein during meiosis.
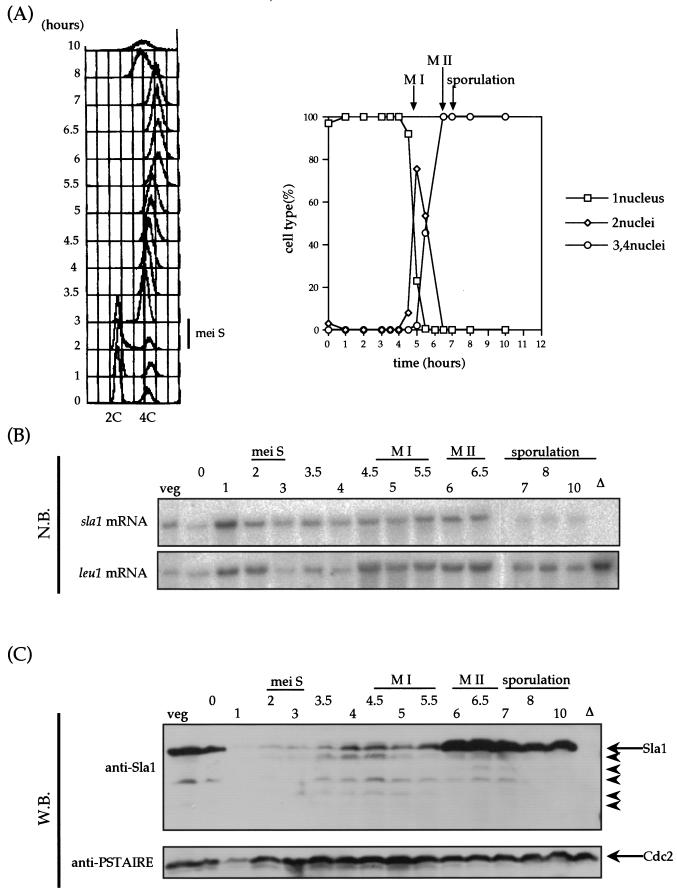
Because the truncated version of Sla1p induces ectopic meiosis, we next asked whether Sla1 is processed or altered during meiosis. Meiosis was synchronously induced by using the pat1-114 diploid, and its synchrony was verified both by fluorescence-activated cell sorter (FACS) analysis and monitoring of ploidy (Fig. 8A). In this synchronously induced meiotic cycle, we monitored Sla1 by using an antibody raised against the purified Sla1 protein as described in Materials and Methods. We also monitored the expression of the sla1+ gene by Northern blot analysis. While the sla1+ mRNA did not obviously fluctuate throughout the meiotic events (Fig. 8B), the condition of Sla1 was dramatically altered (Fig. 8C). In vegetative cells, Sla1 accumulated to high levels, which declined immediately after Pat1 kinase was inactivated. The low level of Sla1 was maintained during the premeiotic S phase and meiosis I. The high level of Sla1 was restored immediately when cells entered the second meiotic division. Note that there is no background protein in the sla1 disruptant when this antibody is used. The breakdown of Sla1 from the point 1 h after inactivation of Pat1 kinase through meiosis I may indicate a role of the truncated Sla1 as a trigger of meiosis. Because only a small region of Sla1 is sufficient for induction of meiosis, we could not distinguish which pattern of degradation is important for meiosis. We also detected some degradation of the Sla1 protein when the samples were taken from homothallic wild-type strain nonsynchronously grown at meiotic phase, but it was not as clear as that observed in pat1-114-based synchronized cells (data not shown).
FIG.8.
Degradation of the Sla1 protein during meiosis. (A) Synchronous meiosis in the S964 strain. Left panel, DNA content measured by FACS analysis. Right panel, meiotic progression was monitored by DAPI staining of nuclei. The peaks of meiosis I (M I), meiosis II (M II), and spore formation (sporulation) are indicated. (B) Northern blot (N.B.) analysis of sla1+ expression. The leu1 mRNA level is shown as an internal control. veg, vegetative growing cell; Δ, sla1 disruptant. (C) Western blot analysis of Sla1 production. Cdc2 is present as an internal control. Arrowheads indicate truncated versions of Sla1p.
DISCUSSION
When we sought a gene(s) that could reverse the sporulation deficiency of the ras1 null diploid strain, we isolated a truncated version of the sla1+ gene which encodes the S. pombe La homolog. The human La protein is known to be an antigen that is a target for immune responses that lead to autoimmune disease, but the exact cause of disease is not known (13, 63). Many groups have found that La and its homologs are RNA-binding proteins that are involved in mRNA processing, stabilization, and tRNA maturation (27, 41, 51). Our observations of the La homolog in S. pombe suggest that this protein may have additional or different functions from those previously postulated.
Sla1 has at least three motifs, based on previous results (15, 43, 57) and ours. One is the La motif, an RRM that is structurally conserved among the known La homologs, located in the N terminus. The second is another RRM motif (RRM2), located in the central region, and the third is the NLS located in the C-terminal region (14). From various deletion analyses of Sla1 we concluded that to induce sporulation in a ras1 null diploid strain, deletion of the NLS from Sla1 is required and only a small 35-amino-acid subregion of RRM2 is sufficient. The cytoplasmic localization of the truncated Sla1 proteins was always associated with sporulation-inducing activity. Curiously, in contrast to our finding, the NLS region of Sla1 is essential in its function of tRNA maturation (15, 57). Both results combined suggest that the Sla1 protein has at least two different functions in two separate loci. Sla1 functions as a mature protein in the nucleus as the regulator of tRNA transcription or maturation, and the truncated Sla1 in cytoplasm involves meiosis.
A heterothallic strain that expressed truncated Sla1 induced ectopic (haploid) meiosis and reached a point at which 42.7% of the cells showed signs of meiosis. This effect of the truncated Sla1 protein is very striking, as to date only a few examples of mutations that induce heterothallic haploids to enter haploid meiosis are known. These include the pat1ts mutant (4) and the mei2-SATA mutant (62). We suggested that the target of truncated Sla1 is Pat1, from the results that truncated sla1 could induce ectopic meiosis in the mei3 mutant but not the mei2 and ste11 mutant and from yeast two-hybrid analysis indicating that the truncated Sla1p, but not the full-length Sla1p, could interact with Pat1. We have attempted to demonstrate that truncated Sla1p and Pat1 interact in vivo, but technical problems are currently hampering these efforts. It was proposed that a sequence motif called RKDIII in Mei3 is essential for inhibition of Pat1 (24, 59), but we did not find a similar sequence motif in Sla1. The absence of apparent similarity alone does not exclude the possibility that Sla1p works as an inhibitor of Pat1 in a manner similar to that for Mei3.
It may be that the truncated Sla1 protein sequesters Pat1 in the cytoplasm, thereby preventing Pat1 from exerting its activities in the nucleus. We investigated the possibility that Sla1p is processed or altered during meiosis and found that Sla1p is degraded from 1 h after meiosis was induced until meiosis I (Fig. 8). Supporting this notion, we often observed that the fusion protein of Sla1 with GFP was degraded during meiosis (data not shown). The result that Sla1p is degraded during meiosis well explains a potential role of truncated Sla1p as an inducer of the meiotic cycle.
Although its exact meaning was unclear, truncation of the mammalian La protein was also reported. When the C-terminal portion of mammalian La is cleaved off, the cells undergo apoptosis and the cleaved La protein localizes in the cytoplasm (2, 45). Considering that the La protein was originally found as an antigen in patients suffering from autoimmune disease and the meiotic function of truncated Sla1 in fission yeast, truncated La may have something to do with the cause or the result of autoimmune disease. It will also be interesting to study the role of truncated La in mammalian meiosis.
The dispensability of sla1 in S. pombe and of its homolog Lhp1 in S. cerevisiae and the lack of apparent phenotypes in their disruptant mutants of both yeasts have been reported (15, 57, 66). We independently isolated the sla1 mutant and observed only a slight difference in sporulation efficiency between wild-type and sla1 disruption cells despite a prominent effect of the truncated Sla1 on meiosis. However, we found that deletion of the chromosomal sla1 gene lowered the temperature sensitivity of the pat1-114 mutant. This result supports the idea that Sla1 is involved in meiosis through the function of Pat1. The commitment of Sla1 in ordinary meiosis is probably limited or well coordinated, because the action of truncated Sla1 to induce ectopic meiosis is strong enough to be harmful for cells. If unexpected meiosis in heterothallic cells occurred by truncation of Sla1, cells could not survive.
Taking these findings together, we hypothesize that when it is processed, Sla1 works as an inducer of meiosis, possibly through Pat1 when cells undergo meiosis. Our observations in this study suggest that La proteins may be involved in activities that have hitherto not been suspected for this family of proteins.
Acknowledgments
We thank S. L. Forsburg for the HA-tagged plasmid vectors, J. A. Huberman for the pYC11-3002 + 3 plasmid, M. Yamamoto and Y. Watanabe for the mei2 and mei3 strains and the pACTII (pat1) and pBTM116 (mei2) plasmids, T. Kato for the phh1 mutant, K. Kitamura for the ste11 mutant, and P. A. San-Segundo for the pat1-114 mutant. We thank Y. Murakami and H. Kato for FACS analysis and M. Sato for technical advice.
This work was supported by Grants-in-Aid from the Kato Memorial Foundation and the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
REFERENCES
- 1.Alfa, C., P. Fantes, J. Hyams, M. McLeod, and E. Warbrick. 1993. Experiments with fission yeast: a laboratory course manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 2.Ayukawa, K., S. Taniguchi, J. Masumoto, S. Hashimoto, H. Sarvotham, A. Hara, T. Aoyama, and J. Sagara. 2000. La autoantigen is cleaved in the COOH terminus and loses the nuclear localization signal during apoptosis. J. Biol. Chem. 275:34465-34470. [DOI] [PubMed] [Google Scholar]
- 3.Bachmann, M., D. Falke, and W. E. Muller. 1990. Is La protein involved in autoimmunization and inflammatory events during disease? Characterization of La protein as an unwinding enzyme. Mol. Biol. Rep. 14:49-50. [DOI] [PubMed] [Google Scholar]
- 4.Beach, D., L. Rodgers, and J. Gould. 1985. ran1+ controls the transition from mitotic division to meiosis in fission yeast. Curr. Genet. 10:297-311. [DOI] [PubMed] [Google Scholar]
- 5.Burke, D., D. Dawson, and T. Stearns. 2000. Methods in yeast genetics; a Cold Spring Harbor Laboratory course manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 6.Dubey, D. D., J. Zhu, D. L Carlson, K. Sharma, and J. A. Huberman. 1994. Three ARS elements contribute to the ura4 replication origin region in the fission yeast, Schizosaccharomyces pombe. EMBO J. 13:3638-3647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Egel, R., and M. Egel-Mitani. 1974. Premeiotic DNA synthesis in fission yeast. Exp. Cell Res. 88:127-134. [DOI] [PubMed] [Google Scholar]
- 8.Fan, H., A. L. Sakulich, J. L. Goodier, X. Zhang, J. Qin, and R. J. Maraia. 1997. Phosphorylation of the human La antigen on serine 366 can regulate recycling of RNA polymerase III transcription complexes. Cell 88:707-715. [DOI] [PubMed] [Google Scholar]
- 9.Fan, H., J. L. Goodier, J. R. Chamberlain, D. R. Engelke, and R. J. Maraia. 1998. 5′ processing of tRNA precursors can be modulated by the human La antigen phosphoprotein. Mol. Cell. Biol. 18:3201-3211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ford, L. P., and J. Wilusz. 1999. 3′-terminal RNA structures and poly(U) tracts inhibit initiation by a 3′→5′ exonuclease in vitro. Nucleic Acids Res. 27:1159-1167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Forsburg, S. L., and D. A. Sherman. 1997. General purpose tagging vectors for fission yeast. Gene 191:191-195. [DOI] [PubMed] [Google Scholar]
- 12.Higuchi, T., Y. Watanabe, and M. Yamamoto. 2002. Protein kinase A regulates sexual development and gluconeogenesis through phosphorylation of the Zn finger transcriptional activator Rst2p in fission yeast. Mol. Cell. Biol. 22:1-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hoch, S. O., and P. B. Billings. 1984. Characterization of the La (SS-B) antigen from several mammalian sources. J. Immunol. 133:1397-1403. [PubMed] [Google Scholar]
- 14.Intine, R. V., M. Dundr, T. Misteli, and R. J. Maraia. 2002. Aberrant nuclear trafficking of La protein leads to disordered processing of associated precursor tRNAs. Mol. Cell 9:1113-1123. [DOI] [PubMed] [Google Scholar]
- 15.Intine, R. V., A. L. Sakulich, S. B. Koduru, Y. Huang, E. Pierstorff, J. L. Goodier, L. Phan, and R. J. Maraia. 2000. Control of transfer RNA maturation by phosphorylation of the human La antigen on serine 366. Mol. Cell 6:339-348. [DOI] [PubMed] [Google Scholar]
- 16.Katayama, S., F. Ozoe, R. Kurokawa, K. Tanaka, T. Nakagawa, H. Matsuda, and M. Kawamukai. 1996. Genetic analysis of the sam mutations, which induce sexual development with no requirement for nutritional starvation in fission yeast. Biosci. Biotechnol. Biochem. 60:994-999. [DOI] [PubMed] [Google Scholar]
- 17.Kawamukai, M., K. Ferguson, M. Wigler, and D. Young. 1991. Genetic and biochemical analysis of the adenylyl cyclase of Schizosaccharomyces pombe. Cell Regul. 2:155-164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kawamukai, M., J. Gerst, J. Field, M. Riggs, L. Rodgers, M. Wigler, and D. Young. 1992. Genetic and biochemical analysis of the adenylyl cyclase-associated protein, cap, in Schizosaccharomyces pombe. Mol. Biol. Cell 3:167-180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kawamukai, M. 1999. Isolation of a novel gene, moc2, encoding a putative RNA helicase as a suppressor of sterile strains in Schizosaccharomyces pombe. Biochim. Biophys. Acta 1446:93-101. [DOI] [PubMed] [Google Scholar]
- 20.Keech, C. L., T. P. Gordon, P. Reynolds, and J. McCluskey. 1993. Expression and functional conservation of the human La (SS-B) nuclear autoantigen in murine cell lines. J. Autoimmun. 6:543-555. [DOI] [PubMed] [Google Scholar]
- 21.Kelly, M., J. Burke, M. Smith, A. Klar, and D. Beach. 1988. Four mating-type genes control sexual differentiation in the fission yeast. EMBO J. 7:1537-1547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kitamura, K., S. Katayama, S. Dhut, M. Sato, Y. Watanabe, M. Yamamoto, and T. Toda. 2001. Phosphorylation of Mei2 and Ste11 by Pat1 kinase inhibits sexual differentiation via ubiquitin proteolysis and 14-3-3 protein in fission yeast. Dev. Cell 1:389-399. [DOI] [PubMed] [Google Scholar]
- 23.Kjaerulff, S., D. Dooijes, H. Clevers, and O. Nielsen. 1997. Cell differentiation by interaction of two HMG-box proteins: Mat1-Mc activates M cell-specific genes in S. pombe by recruiting the ubiquitous transcription factor Ste11 to weak binding sites. EMBO J. 16:4021-4033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Li, P., and M. McLeod. 1996. Molecular mimicry in development: identification of ste11+ as a substrate and mei3+ as a pseudosubstrate inhibitor of ran1+ kinase. Cell 87:869-880. [DOI] [PubMed] [Google Scholar]
- 25.Lin-Marq, N., and S. G. Clarkson. 1998. Efficient synthesis, termination and release of RNA polymerase III transcripts in Xenopus extracts depleted of La protein. EMBO J. 17:2033-2041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Maeda, T., Y. Watanabe, H. Kunitomo, and M. Yamamoto. 1994. Cloning of the pka1 gene encoding the catalytic subunit of the cAMP-dependent protein kinase in Schizosaccharomyces pombe. J. Biol. Chem. 269:9632-9637. [PubMed] [Google Scholar]
- 27.Maraia, R. J., and R. V. Intine. 2001. Recognition of nascent RNA by the human La antigen: conserved and divergent features of structure and function Mol. Cell. Biol. 21:367-379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Maraia, R. J., D. J. Kenan, and J. D. Keene. 1994. Eukaryotic transcription termination factor La mediates transcript release and facilitates reinitiation by RNA polymerase III. Mol. Cell. Biol. 14:2147-2158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Matsuyama, A., N. Yabana, Y. Watanabe, and M. Yamamoto. 2000. Schizosaccharomyces pombe Ste7p is required for both promotion and withholding of the entry to meiosis. Genetics 155:539-549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Maundrell, K. 1990. nmt1 of fission yeast: a highly transcribed gene completely repressed by thiamine. J. Biol. Chem. 265:10857-10864. [PubMed] [Google Scholar]
- 31.Maundrell, K. 1993. Thiamine-repressible expression vectors prep and prip for fission yeast. Gene 123:127-130. [DOI] [PubMed] [Google Scholar]
- 32.McLeod, M., M., Stein, and D. Beach. 1987. The product of the mei3+ gene, expressed under control of the mating-type locus, induces meiosis and sporulation in fission yeast. EMBO J. 6:729-736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.McLeod, M., and D. Beach. 1988. A specific inhibitor of the ran1+ protein kinase regulates entry into meiosis in Schizosaccharomyces pombe. Nature 332:509-514. [DOI] [PubMed] [Google Scholar]
- 34.Moreno, S., A. Klar, and P. Nurse. 1991. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194:795-823. [DOI] [PubMed] [Google Scholar]
- 35.Nadin-Davis, S. A., R. C. Yang, S. A. Narang, and A. Nasim. 1986. The cloning and characterization of a RAS gene from Schizosaccharomyces pombe. J. Mol. Evol. 23:41-51. [DOI] [PubMed] [Google Scholar]
- 36.Nadin-Davis, S. A., and A. Nasim. 1988. A gene which encodes a predicted protein kinase can restore some functions of the ras gene in fission yeast. EMBO J. 7:985-993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Nadin-Davis, S. A., and A. Nasim. 1990. Schizosaccharomyces pombe ras1 and byr1 are functionally related genes of the ste family that affect starvation-induced transcription of mating-type genes. Mol. Cell. Biol. 10:549-560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ohndorf, U. M., C. Steegborn, R. Knijff, and P. Sondermann. 2001. Contributions of the individual domains in human La protein to its RNA 3′-end binding activity. J. Biol. Chem. 276:27188-27196. [DOI] [PubMed] [Google Scholar]
- 39.Okazaki, K., N. Okazaki, K. Kume, S. Jinno, K. Tanaka, and H. Okayama. 1990. High-frequency transformation method and library transducing vectors for cloning mammalian cDNAs by trans-complementation of Schizosaccharomyces pombe. Nucleic Acids Res. 18:6485-6489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Ozoe, F., R. Kurokawa, Y. Kobayashi, H. T. Jeong, K. Tanaka, K. Sen, T. Nakagawa, H. Matsuda, and M. Kawamukai. 2002. The 14-3-3 proteins Rad24 and Rad25 negatively regulate Byr2 by affecting its localization in Schizosaccharomyces pombe. Mol. Cell. Biol. 22:7105-7119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Rinke, J., and J. A. Steitz. 1982. Precursor molecules of both human 5S ribosomal RNA and transfer RNAs are bound by a cellular protein reactive with anti-La lupus antibodies. Cell. 29:149-159. [DOI] [PubMed] [Google Scholar]
- 42.Rose, M., and D. Botstein. 1983. Construction and use of gene fusions to lacZ (β-galactosidase) that are expressed in yeast. Methods Enzymol. 101:167-180. [DOI] [PubMed] [Google Scholar]
- 43.Rosenblum, J., L. Pemberton, N. Bonifaci, and G. Blobel. 1998. Nuclear import and the evolution of a multifunctional RNA-binding protein. J. Cell Biol. 143:887-899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Rothstein, R. J. 1983. One-step gene disruption in yeast. Methods Enzymol. 101:202-211. [DOI] [PubMed] [Google Scholar]
- 45.Rutjes, S. A., P. J. Utz, A. van der Heijden, C., Broekhuis, W. J. van Venrooij, and G. J. Pruijn. 1999. The La (SS-B) autoantigen, a key protein in RNA biogenesis, is dephosphorylated and cleaved early during apoptosis. Cell Death Differ. 6:976-986. [DOI] [PubMed] [Google Scholar]
- 46.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 47.Sato, M., Y. Watanabe, Y. Akiyoshi, and M. Yamamoto. 2002. 14-3-3 protein interferes with the binding of RNA to the phosphorylated form of fission yeast meiotic regulator Mei2p. Curr. Biol. 12:141-145. [DOI] [PubMed] [Google Scholar]
- 48.Shimoda, C., M. Uehira, M. Kishida, H. Fujioka, Y. Iino, Y. Watanabe, and M. Yamamoto. 1987. Cloning and analysis of transcription of the mei2 gene responsible for initiation of meiosis in the fission yeast Schizosaccharomyces pombe. J. Bacteriol. 169:93-96. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Shiozaki, K., and P. Russell. 1996. Conjugation, meiosis, and the osmotic stress response are regulated by Spc1 kinase through Atf1 transcription factor in fission yeast. Genes Dev. 10:2276-2288. [DOI] [PubMed] [Google Scholar]
- 50.Simons, F. H., F. J. Broers, W. J. Van Venrooij, and G. J. Pruijn. 1996. Characterization of cis-acting signals for nuclear import and retention of the La (SS-B) autoantigen. Exp. Cell Res. 224:224-236. [DOI] [PubMed] [Google Scholar]
- 51.Stefano, J. E. 1984. Purified lupus antigen La recognizes an oligouridylate stretch common to the 3′ termini of RNA polymerase III transcripts. Cell 36:145-154. [DOI] [PubMed] [Google Scholar]
- 52.Sugimoto, A., Y. Iino, T. Maeda, Y. Watanabe, and M. Yamamoto. 1991. Schizosaccharomyces pombe ste11+ encodes a transcription factor with an HMG motif that is a critical regulator of sexual development. Genes Dev. 5:1990-1999. [DOI] [PubMed] [Google Scholar]
- 53.Takahashi, K., S. Murakami, Y. Chikashige, H. Funabiki, O. Niwa, and M. Yanagida. 1992. A low copy number central sequence with strict symmetry and unusual chromatin structure in fission yeast centromere. Mol. Biol. Cell. 3:819-835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Takeda, T., T. Toda, K. Kominami, A. Kohnosu, M. Yanagida, and N. Jones. 1995. Schizosaccharomyces pombe atf1+ encodes a transcription factor required for sexual development and entry into stationary phase. EMBO J. 14:6193-6208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Tanaka, K., J. Nishide, K. Okazaki, H. Kato, O. Niwa, T. Nakagawa H. Matsuda, M. Kawamukai, and Y. Murakami. 1999. Characterization of a fission yeast SUMO-1 homologue, pmt3p, required for multiple nuclear events, including the control of telomere length and chromosome segregation. Mol. Cell. Biol. 19:8660-8672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Toda, T., M. Shimanuki, and M. Yanagida. 1991. Fission yeast genes that confer resistance to staurosporine encode an AP-1-like transcription factor and a protein kinase related to the mammalian ERK1/MAP2 and budding yeast FUS3 and KSS1 kinases. Genes Dev. 5:60-73. [DOI] [PubMed] [Google Scholar]
- 57.Van Horn, D. J., C. J. Yoo, D. Xue, H. Shi, and S. L. Wolin. 1997. The La protein in Schizosaccharomyces pombe: a conserved yet dispensable phosphoprotein that functions in tRNA maturation. RNA 3:1434-1443. [PMC free article] [PubMed] [Google Scholar]
- 58.Wang, Y., H. P. Xu, M. Riggs, L. Rodgers, and M. Wigler. 1991. byr2, a Schizosaccharomyces pombe gene encoding a protein kinase capable of partial suppression of the ras1 mutant phenotype. Mol. Cell. Biol. 11:3554-3563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Wang, W., P. Li, A. Schettino, Z. Peng, and M. McLeod. 1998. Characterization of functional regions in the Schizosaccharomyces pombe mei3 developmental activator. Genetics 150:1007-1018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Watanabe, Y., Y. Lino, K. Furuhata, C. Shimoda, and M. Yamamoto. 1988. The S. pombe mei2 gene encoding a crucial molecule for commitment to meiosis is under the regulation of cAMP. EMBO J. 7:761-767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Watanabe, Y., and M. Yamamoto. 1994. S. pombe mei2 encodes an RNA-binding protein essential for premeiotic DNA synthesis and meiosis I, which cooperates with a novel RNA species meiRNA. Cell 12:487-498. [DOI] [PubMed] [Google Scholar]
- 62.Watanabe, Y., S. Shinozaki-Yabana, Y. Chikashige, Y. Hiraoka, and M. Yamamoto. 1997. Phosphorylation of RNA-binding protein controls cell cycle switch from mitotic to meiotic in fission yeast. Nature 386:187-190. [DOI] [PubMed] [Google Scholar]
- 63.Wolin, S. L., and T. Cedervall. 2002. The La protein. Annu. Rev. Biochem. 71:375-403. [DOI] [PubMed] [Google Scholar]
- 64.Yamamoto, M. 1996. The molecular control mechanisms of meiosis in fission yeast. Trends Biochem. Sci. 21:18-22. [PubMed] [Google Scholar]
- 65.Yamamoto, M., Y. Imai, and Y. Watanabe. 1997. Mating and sporulation in Schizosaccharomyces pombe. In J. R. Pringle, J. R. Broach, and E. W. Jones (ed.), The molecular and cellular biology of the yeast Saccharomyces, vol. 3, p. 1037-1106. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 66.Yoo, C. J., and S. L. Wolin. 1997. The yeast La protein is required for the 3′ endonucleolytic cleavage that matures tRNA precursors. Cell 89:393-402. [DOI] [PubMed] [Google Scholar]