Abstract
Approximately 30–50% of the >30 million HIV-infected subjects develop neurological complications ranging from mild symptoms to dementia. HIV does not infect neurons, and the molecular mechanisms behind HIV-associated neurocognitive decline are not understood. There are several hypotheses to explain the development of dementia in HIV+ individuals, including neuroinflammation mediated by infected microglia and neuronal toxicity by HIV proteins. A key protein associated with the neurological complications of HIV, gp120, forms part of the viral envelope and can be found in the CSF of infected individuals. HIV-1-gp120 interacts with several receptors including CD4, CCR5, CXCR4, and nicotinic acetylcholine receptors (nAChRs). However, the role of nAChRs in HIV-associated neurocognitive disorder has not been investigated. We studied the effects of gp120IIIB on the expression and function of the nicotinic receptor α7 (α7-nAChR). Our results show that gp120, through activation of the CXCR4 chemokine receptor, induces a functional up-regulation of α7-nAChRs. Because α7-nAChRs have a high permeability to Ca2+, we performed TUNEL staining to investigate the effects of receptor up-regulation on cell viability. Our data revealed an increase in cell death, which was blocked by the selective antagonist α-bungarotoxin. The in vitro data are supported by RT-PCR and Western blot analysis, confirming a remarkable up-regulation of the α7-nAChR in gp120-transgenic mice brains. Specifically, α7-nAChR up-regulation is observed in mouse striatum, a region severely affected in HIV+ patients. In summary, CXCR4 activation induces up-regulation of α7-nAChR, causing cell death, suggesting that α7-nAChR is a previously unrecognized contributor to the neurotoxicity associated with HIV infection.
Keywords: AIDS, Chemokines, HIV, Neuroblastoma, Nicotinic Acetylcholine Receptors, CHRNA7, CXCR4, HAND, NeuroAIDS, GP120
Introduction
More than 30 million people are infected with HIV worldwide (1). In addition to the health problems caused by immunosuppression, HIV infection causes HIV-associated neurocognitive disorder (HAND),4 a neurodegenerative disease that leads to severe cognitive, motor, and behavioral disturbances. Before the introduction of highly active antiretroviral therapy (HAART), 30–50% of HIV-infected patients developed HAND; after the incorporation of HAART as part of HIV treatment, the incidence decreased to ∼10% (2–5). Nevertheless, in association with the existence of HAART treatment, which prolongs the life of HIV-infected individuals, there has been an increase to ∼30% in the number of individuals who develop a milder form of neurocognitive dysfunction known as minor cognitive-motor disorder, which is characterized by neurological deficits that do not interfere with everyday functioning (6). Therefore, despite the reduction in HAND incidence, its prevalence is expected to increase due the improved care of HIV-infected patients. When evaluating the alterations in the incidence of HAND as a result of HAART, it is important to consider that of the ∼30 million HIV-infected individuals worldwide, only 2 million have access to HAART (1). In addition, HIV infection continues to be the most common cause of dementia in young adults in the United States (7, 8). This suggests that HAND will continue to be an important health care problem in the United States and worldwide.
The presence of neurological symptoms in HIV-infected patients is an interesting finding, considering that HIV does not infect neurons directly. The molecular mechanisms by which HIV infection leads to neurocognitive decline are not fully understood, but several hypotheses have emerged to explain the development of neurocognitive impairment in HIV+ individuals. Two potential mechanisms by which HIV infection could interfere with normal brain functioning are (i) chemokine-induced neuroinflammation mediated by infected macrophages/microglia and (ii) direct neuronal toxicity induced by soluble HIV proteins (9). HIV-1-gp120, a glycoprotein that forms part of the envelope of HIV-1 particles, can be found in the cerebrospinal fluid of HIV+ individuals, and it has been shown to have neurotoxic effects in cell cultures (9–13). HIV-1-gp120 interacts with several receptors found in the central nervous system (CNS), including CD4, CCR5, and CXCR4, as well as nicotinic acetylcholine receptors (nAChRs) (14–16). Several lines of evidence suggest the potential involvement of the nicotinic acetylcholine receptor α7-nAChR in HIV neuropathology (17–19); however, the role that nAChRs may play in the development of HAND has not been investigated.
For HIV-1 to infect cells, it must bind the CD4 receptor and either the CCR5 or CXCR4 co-receptor. Interestingly, during the course of infection, HIV-1 evolves from an M-tropic (CCR5-dependent) variant that primarily infects macrophages to a T-tropic (CXCR4-dependent) variant that primarily infects T cells (15, 20). The increase in HIV particles with tropism for CXCR4 receptor correlates with the development of HAND (20, 21). Studies have shown that gp120 not only binds to CXCR4, but also activates its signaling pathway (22). CXCR4 activation by stromal cell-derived growth factor (SDF-1α), the endogenous agonist of CXCR4, has been shown to rapidly up-regulate the early growth response gene 1 (Egr-1), a transcription factor known to drive the expression of the α7-nAChR (23, 24). Furthermore, it has been shown that SDF-1α is secreted by astrocytes during inflammation and is increased in response to macrophage activation by HIV infection (25). Consequently, we designed experiments to study the effects of gp120 on the expression and function of α7-nAChRs in SH-SY5Y neuroblastoma cells, which endogenously express α7-nAChRs and CXCR4 receptors (26). Our results show that CXCR4 activation, by gp120 or SDF-1α, leads to an increase in α7-nAChR activity that can culminate in cell death. In addition, Western blotting and quantitative RT-PCR analysis confirm up-regulation of α7-nAChR in transgenic mice expressing the HIV-gp120 gene.
MATERIALS AND METHODS
Fluorescent α-Bungarotoxin (Bgtx) Binding
SH-SY5Y cells were grown in 4-well tissue culture slides (Nalge Nunc International, Rochester, NY). Cell culture medium was removed, the cells were washed with phosphate-buffered saline (PBS), fixed by incubation with 4% paraformaldehyde for 10 min followed by incubation with 1% bovine serum albumin (BSA) for 10 min. The cells were washed with PBS and then incubated with Alexa Fluor 488-conjugated α-bgtx (Invitrogen) 1:50 for 1 h at room temperature. Cells were then washed three times with PBS, and coverslips were mounted with 90% glycerol and analyzed with an Axiovert 200M confocal microscope.
Electrophysiology
Whole cell currents were measured in the whole cell configuration of the patch clamp technique using an Axopatch 200B amplifier (Axon Instruments, Foster City, CA). The bath solution contained 140 mm NaCl, 4 mm KCl, 2 mm CaCl2, 1 mm MgCl2, 5 mm HEPES, 10 mm glucose, pH 7.4, ∼280 mosmol/kg. The pipette solution contained 145 mm KCl, 6 mm MgCl2, 7.2 mm K2HPO4, 2.8 mm KH2PO4, 5 mm EDTA, pH 7.4, ∼270 mosmol/kg. Pipettes were pulled from thick wall borosilicate glass (World Precision Instruments, Sarasota, FL) with a multistage P-87 Flaming-Brown micropipette puller (Sutter Instruments, San Rafael, CA) and fire-polished. Pipette resistance was 2–4 megohms, and a 1–2% agar bridge with composition similar to the bath solution was utilized as the reference electrode. Chemicals were purchased from Sigma. Whole cell current traces were filtered at 2 kHz and acquired at 10 kHz. Currents were measured in response to a 1-s pulse of 500 μm acetylcholine (ACh) at a holding potential of −100 mV. Pulse generation, data collection, and analyses were carried out with Clampex 10.1 (Molecular Devices, Sunnyvale, CA).
Calcium Flux Assay
SH-SY5Y cells were grown on coverslips and treated with 0.15 nm gp120 overnight followed by incubation on nonsupplement medium with 10 μm Fluo4-AM for 30 min in the dark. Cells were then washed with PBS and incubated with acini buffer (120 mm NaCl, 4 mm KCl, 1 mm CaCl2, 1.2 mm KH2PO4, 1 mm MgSO4, 15 mm HEPES, 0.1% BSA, 10 mm glucose, pH 7.4) for 3 min at 37 °C in the dark for de-esterification. Cells were then stimulated with 1 mm ACh for 1.5 min while exciting at a wavelength of 488 nm using an Argon/2 laser. Emission was acquired at 520 nm using a BP 500-550 filter on a Zeiss LSM 510 confocal microscope. Images were acquired in a time series of 180 frames at 0.5-s intervals for 90 s at 40× magnification. To calculate the maximal intensity (Fmax), incubation with 10 μm ionomicin in presence of 10 mm CaCl2 was performed. Ca2+ concentration was determined as described (27).
Cell Death Assay
SH-SY5Y cells were treated with gp120 (0.15 nm) overnight. Cell death was measured using the APO-BrdU TUNEL Assay kit (Invitrogen) following the manufacturer's instructions with slight modifications. All centrifugations steps were done at 6,000 rpm. In brief, all cells were detached, centrifuged for 6 min, fixed with 2% paraformaldehyde for 20 min, and centrifuged for 6 min. Cells were then permeabilized with 70% ethanol for 24 h at −20 °C, centrifuged for 6 min, resuspended in wash buffer, and centrifuged again. Then, cells were incubated with labeling solution for 24 h at room temperature, washed with 1 ml of rinse buffer, and centrifuged. This step was followed by incubation with antibody staining solution for 1 h at room temperature. After this, cells were washed with PBS, mounted in glass slides, and visualized with an Axiovert 200M confocal microscope.
Real-time PCR
Total RNA was isolated from SH-SY5Y cells or mouse brains using the TRIzol reagent (Invitrogen). The cDNA synthesis was carried out using 1 μg of total RNA with the iScriptTM cDNA Synthesis kit (Bio-Rad) following the manufacturer's instructions. Real-time PCR experiments were done using the iQTM SYBR® Green Supermix (Bio-Rad) in a Mastercycler® ep realplex Thermal Cycler (Eppendorf, Hauppauge, NY). The following primer pairs were used: forward, 5′-GCTCCGGGACTCAACATG-3′ and reverse, 5-GGGATTGTAGTTCTTGACCAGC-3′ for CHRNA7; forward, 5′-AGCACCTTCAACCCTCA-3′ and reverse, 5′-AGTCGAGTGGTTTGGCT-3′ for EGR1; and forward, 5′-GCTCTCTGCTCCTCCTGTTC-3′ and reverse, 5′-GACTCCGACCTTCACCTTCC-3′ for GAPDH.
Animals
Transgenic mice expressing the HIV-1 coat protein gp120 under the regulatory control of a modified murine glial fibrillary acidic protein gene were obtained from a previously established line (Toggas et al., Ref. 49) and housed in clear plastic cages, maintained in a temperature- and humidity-controlled room on a 12-h light/dark cycle with food and water provided ad libitum. Animals from the background strain B6SJLF were used as control.
Western Blotting
Transgenic and wild-type (WT) mice were killed by cervical dislocation, and brains were placed on ice and dissected to separate the striatum and hippocampus. Pulverized tissue was transferred to 15-ml tubes containing cold radioimmune precipitation assay buffer (10 ml/g) supplemented with a protease inhibitor mixture. The sample was agitated for 30 s and incubated for 1 h at 4°C, shaking. Then, the sample was centrifuged at 14,000 rpm for 15 min at 4 °C, and protein in the lysates was quantified with a BCA kit (Pierce). Samples containing 50 μg of total protein were loaded on a 4–20% linear gradient polyacrylamide gel (Bio-Rad), and electrophoresis was done at 100 mV for 1 h at room temperature. Proteins were transferred to a PVDF membrane (Amersham Biosciences) at 100 mV for 1 h. Nonspecific binding was blocked with 5% nonfat dry milk in TBS-T (1×) for 1 h at room temperature. The membrane was then incubated with a polyclonal rabbit anti-α7-nAChR (1:1,000; Millipore), diluted in 5% skim milk, overnight at 4 °C, shaking. Three consecutive washes with TBS-T (5 min each) were done to eliminate excess antibody, and the membrane was incubated with an anti-rabbit antibody conjugated to HRP (1:5,000; Abcam) for 1 h at room temperature. Antibody binding was detected with the ECL Plus Western blotting detection system (Amersham Biosciences) and developed using BioMax MS film (Kodak).
Cell Culture and Treatment with Gp120, SDF-1α, or AMD3100
SH-SY5Y cells were grown in DMEM/F12 medium supplemented with 10% FBS and 1% antibiotic/antimicotic solution (Sigma-Aldrich) and treated for 24 h. Several compounds were used at various concentrations: gp120IIIB (Fitzgerald Industries International, Concord, MA), SDF-1α (EMD Chemicals, Inc., Gibbstown, NJ) at 0.3 μg/ml; AMD3100 (EMD Chemicals, Gibbstown, NJ) at 1 μm, where pretreatment was carried out 10 min before the addition of gp120 or SDF-1α; and α-bgtx (Invitrogen) at 1 μm, where pretreatment was carried out 10 min before the addition of gp120 or SDF-1α.
RESULTS
Gp120 Increases α-Bgtx Binding and Acetylcholine-stimulated Currents in SH-SY5Y Cells
To study the effects of gp120 on α7-nAChR expression, SH-SY5Y cells were treated with different concentrations of gp120. Alexa Fluor 488-α-bgtx was used to detect α7-nAChRs. As shown in Fig. 1, A and B, cells treated with various concentrations of gp120 showed an increase in α-bgtx binding, consistent with higher levels of α7-nAChRs. To determine whether the increase in α-bgtx binding correlates with an increase in functional nAChRs, we measured currents in response to stimulation with 1 mm ACh using the whole cell configuration of the patch clamp technique. As shown in Fig. 1, C and D, cells treated with different concentrations of gp120 presented an increase in whole cell current density in response to ACh. These results show that gp120 treatment leads to an increase in functional α7-nAChRs.
FIGURE 1.
Increased α-bgtx binding and ACh-stimulated currents in SH-SY5Y cells after treatment with gp120. A, Alexa Fluor 488-conjugated α-bgtx binding to SH-SY5Y was assessed in cells treated with 0.15, 15, 150, or 1,500 nm gp120, or control cells. B, normalized fluorescence intensity for control SH-SY5Y cells was 100.0 ± 6.0% (n = 16). Cells treated with 0.15, 15, 150, and 1,500 nm showed a percentage fluorescence intensity of 163.5 ± 26.7% (n = 12), 174.6 ± 10.7% (n = 13), 150.4 ± 9.1% (n = 12), and 167.5 ± 26.3% (n = 17), respectively. C, whole cell representative current traces recorded from control SH-SY5Y cells or cells treated with 0.15, 15, 150, and 1,500 nm gp120 are shown. Currents were recorded in response to application of a 1 mm ACh pulse at a holding potential of −100 mV. D, average density for ACh-stimulated currents recorded from control SH-SY5Y cells was 18.9 ± 3.9 pA/pF (n = 15). For cells treated with 0.15, 15, 150, and 1,500 nm gp120, the average current density was 32.1 ± 13.1 pA/pF (n = 4), 48.5 ± 8.1 pA/pF (n = 7), 41.7 ± 16.9 pA/pF (n = 7), and 44.7 ± 9.5 pA/pF (n = 12), respectively. *, p < 0.05.
Gp120 Effects Are Mediated by CXCR4 Receptor
It has been shown that gp120 activates the chemokine receptor CXCR4, which is expressed in SH-SY5Y cells (10). We hypothesized that this receptor was involved in the gp120-induced up-regulation of α7-nAChRs on the basis of findings by Luo et al. (23) and Criado and del Toro (24), which showed that SDF-1α, the endogenous CXCR4 agonist, up-regulates Egr-1, a known transcription factor for the α7-nAChR. As shown in Fig. 2A, the CXCR4 antagonist, AMD3100, prevented the increase in α-bgtx binding observed after treatment with gp120. Pretreatment with AMD3100 also prevented the gp120-induced increase in ACh-stimulated currents (Fig. 2B). These results reveal that CXCR4 activation by gp120 is necessary for the up-regulation of the α7-nAChR. On the basis of these findings, we proceeded to test whether activation of the CXCR4 receptor by its endogenous ligand, SDF-1, would have similar effects on α-bgtx binding and ACh-stimulated currents. Fig. 2B shows that SDF-1α treatment resulted in increased α-bgtx binding and ACh-stimulated currents, similar to what was observed after gp120 treatment. The α7-nAChR up-regulation can be prevented by the MEK inhibitor PD98059, suggesting that the effect of SDF1 requires activation of the MAP kinase pathway (supplemental Fig. S2).
FIGURE 2.
CXCR4 receptor mediates gp120-induced up-regulation of α7-nAchR. A, Alexa Fluor 488-conjugated α-bgtx binding to control SH-SY5Y cells or cells treated with 1,500 nm gp120, 1,500 nm gp120 plus AMD3100 (0.1 μm), SDF-1 (0.3 μg/ml), or SDF-1 (0.3 μg/ml) plus AMD3100 (0.1 μm) was assessed. B, percentage fluorescence intensity for control SH-SY5Y cells was 100.0 ± 6.5% (n = 14). Cells treated with 1,500 nm gp120 showed a percentage fluorescence intensity of 153.9 ± 13.1% (n = 12). The percentage fluorescence for cells treated with 1,500 nm gp120 plus AMD3100 was 90.2 ± 9.7% (n = 15). Cells treated with SDF-1 had a percentage fluorescence intensity of 207.1 ± 17.3% (n = 11). C, representative whole cell current traces of control SH-SY5Y cells, cells treated with 1,500 nm gp120, gp120 plus AMD3100, SDF-1, or SDF-1 plus AMD3100 are shown. D, average current density for control cells was 18.9 ± 3.9 pA/pF (n = 15). Cells treated with gp120, gp120 plus AMD3100, SDF-1, and SDF-1 plus AMD3100 showed an average current density of 44.7 ± 9.4 pA/pF (n = 12), 20.7 ± 5.9 pA/pF (n = 9), 58.9 ± 13.1 pA/pF (n = 5), and 16.0 ± 4.1 (n = 4), respectively. Currents were recorded as described for Fig. 1. *, p < 0.05.
Gp120 Increases ACh-stimulated Calcium Movement
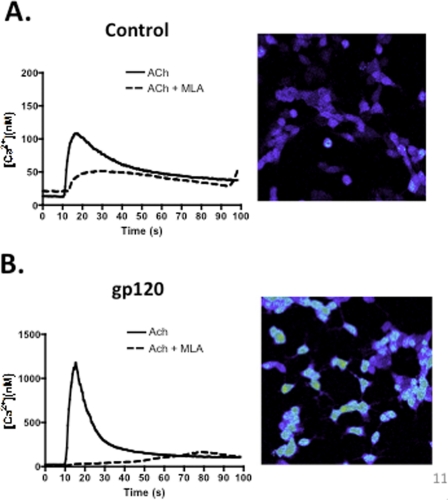
Of all the nicotinic receptors, the α7-nAChR has the highest permeability to Ca2+ ions (28, 29). Taking this fact into consideration, we decided to measure the intracellular Ca2+ concentration after stimulating SH-SY5Y cells with ACh. As shown in Fig. 3, control cells stimulated with ACh had an increase in intracellular Ca2+ concentration ( [Ca2+]i) that peaked around 100 nm (supplemental Video S1). This increase in [Ca2+]i can be blocked by the addition of methyllycaconitine (supplemental Video S3), a selective antagonist for the α7-nAChR, suggesting its role in mediating this effect. In contrast, cells treated with gp120 (0.15 nm) had an increase in [Ca2+]i that peaked at around 1,100 nm (supplemental Video S2), 11 times more than the [Ca2+]i observed in control cells. Once more, this effect can be inhibited by addition of MLA (supplemental Video S4), confirming the involvement of the α7-nAChR in mediating the increase in [Ca2+]i.
FIGURE 3.
Increased Ca2+ mobilization after gp120 treatment. Gp120 exposure that induced up-regulation of the α7-nAChR led to a remarkable Ca2+ mobilization in SH-SY5Y cells. Our data using confocal microscopy suggest that neuroblastoma cells exposed to gp120 (0.15 nm) reach a higher [Ca2+]i (1,179 ± 222.0 nm) (B) in response to ACh than do control cells (108.6 ± 9.2 nm) (A). The Ca2+ mobilization can be blocked by preincubation with methyllycaconitine, an antagonist of the α7-nAChR in gp120-treated (167.0 ± 48.0 nm) and control (51.7 ± 3.3 nm) cells.
Gp120-induced Up-regulation of α7-nAChRs Leads to Cell Death
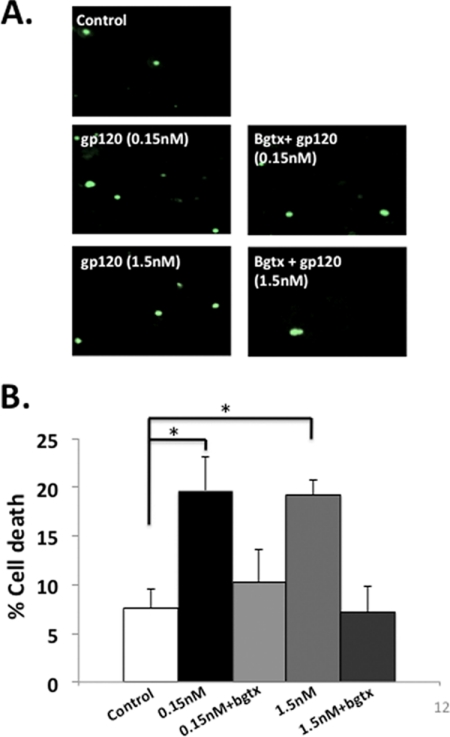
When considering the effects of the increase in α7-nAChRs for cellular function, it is important to keep in mind that large increases in intracellular Ca2+ have been shown to induce cell death (30). Moreover, treatment with gp120 has been shown to induce cell death in neuroblastoma cells (10). To test the hypothesis that α7-nAChR up-regulation plays a role in the gp120-induced cell death observed in SH-SY5Y cells, we performed TUNEL staining after treatment with gp120 with and without the addition of α-bgtx, an α7-nAChR antagonist. Fig. 4 shows that antagonizing α7-nAChRs with α-bgtx reduced the percentage of cells undergoing cell death after treatment with gp120. These data suggest that increased activity of α7-nAChRs contributes to gp120-induced cell death.
FIGURE 4.
α-Bgtx treatment reduces gp120-induced cell death in SH-SY5Y cells. A, TUNEL staining of control SH-SY5Y cells (top left), cells treated with 0.15 nm gp120 with or without incubation with 1 μm α-Bgtx (middle), or cells treated with 1.5 nm gp120 with or without incubation with 1 μm α-Bgtx (bottom). B, the percent of cell death in control cells was 7.4 ± 1.8% (N = 6), whereas cells treated with 0.15 nm and 1.5 nm gp120 presented 19.6 ± 3.5% (N = 6) and 19.2 ± 1.5% (N = 4), respectively. Cells treated with 0.15 nm gp120 + α-Bgtx or 1.5 nm gp120 + α-Bgtx presented 10.2 ± 2.5% (N = 4) and 7.2 ± 2.6% (N = 4) cell death, respectively. *, p-value <0.05.
α7-nAChR and Egr-1 mRNA Levels Are Up-regulated in SH-SY5Y Cells Treated with Gp120
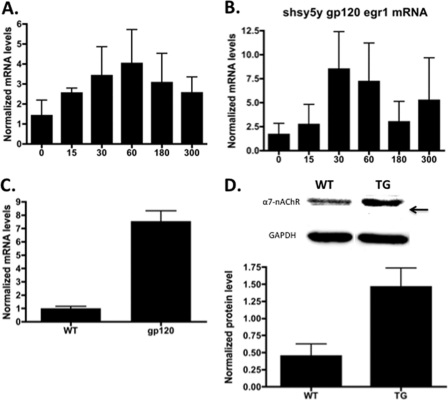
The experiments performed in SH-SY5Y cells suggest that CXCR4 activation by either gp120 or SDF-1 results in increased levels of functional α7-nAChRs. The CXCR4 receptor has been shown to activate the early growth response protein, Egr-1 (23). Egr-1, binds to the promoter region of the α7 gene and increases transcription (24, 31). Real-time PCR on SH-SY5Y cells treated with gp120 for different periods of time showed a fast and transient up-regulation of Egr-1 with a peak of 8.4-fold increase after 30 min. To test whether α7-nAChRs mRNA levels are increased after treatment with gp120, real-time PCR was performed using CHRNA7-specific primers. Fig. 5 shows that the transient up-regulation of Egr-1 was accompanied by an ∼4-fold increase in α7 mRNA levels that peaked 60 min after gp120 treatment. Consistently, increased α7-nAChR currents were observed as early as 1 h after gp120 treatment and persisted for at least 48 h (supplemental Fig. S1).
FIGURE 5.
Real-time PCR and Western blot analysis of α7-nAChR on SH-SY5Y and gp120-transgenic mice. A, SH-SY5Y cells were treated with gp120 at different time points. Up-regulation of α7-nAChR is observed after exposure to gp120 (n = 3). B, levels of Egr-1 mRNA increased transiently in SH-SY5Y cells after gp120 treatment (n = 3). C, increase in the levels of α7-nAChR mRNA was observed in the striatum of gp120-transgenic mice (7.55 ± 0.77, n = 3) compared with WT (1.0 ± 0.1, n = 4) animals. D, Western blot analysis of protein extracts from striatum revealed increase in α7-nAChR protein in gp120-transgenic (TG) mice compared with WT.
α7-nAChR mRNA and Protein Levels Are Increased in Gp120-transgenic Mice
Quantitative RT-PCR and Western blotting experiments showed that mRNA and protein levels of α7-nAChRs are increased in the CNS of gp120-transgenic mice (Fig. 5). However, the up-regulation of α7-nAChRs was observed in the striatum, a region affected in HAND patients, but not in the hippocampus (supplemental Fig. S3). Interestingly, the striatum expresses high levels of the CXCR4 receptor and, furthermore, the expression of this receptor is increased in HIV+ patients (32). These findings are in agreement with our data from SH-SY5Y cells, and previous studies that implicate the CXCR4 in gp120-induced neurotoxicity (33, 34).
DISCUSSION
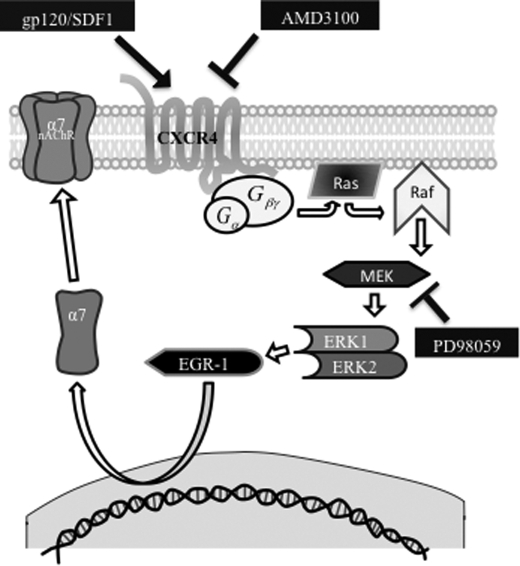
We have developed a model for gp120 neurotoxicity, in which activation of the CXCR4 receptor by gp120 or SDF1 leads to activation of the MAPK pathway, increased EGR1 levels, and up-regulation of α7-nAChRs that triggers cell death (Fig. 6). This model is supported by data showing that treatment of SH-SY5Y cells with gp120 results in higher expression of α7-nAChRs, which are highly permeable to Ca2+. Increased intracellular Ca2+ can lead to cell death and has been associated with neuronal death in neurodegenerative diseases (35). Along these lines, overexpression of α7-nAChRs has been shown to play a role in neurological disorders (36). Studies have shown that in neuronal cultures, gp120 can induce an increase in Ca2+ levels, leading to neuronal injury, an effect that can be blocked with memantine (37). Interestingly, besides being an antagonist for the NMDA receptor, memantine has been shown to also act as an antagonist of α7-nAChRs (38). Moreover, subsequent studies have shown involvement of CXCR4 in the neurotoxic effects of gp120 (38). These results suggest that other proteins, besides the NMDA receptor, contribute to gp120 neurotoxicity.
FIGURE 6.
Proposed mechanism for gp120-induced up-regulation of the α7-nAChR. Activation of CXCR4 by SDF-1 or gp120 causes activation of the Ras-Raf-MEK pathway. This leads to activation of Egr-1, a known transcription factor for the α7-nAChR gene (CHRNA7). Activation of Egr-1 leads to an increase in α7-nAChR mRNA levels.
TUNEL staining showed that exposure to gp120 increases cell death in SH-SY5Y cells, consistent with previous studies (37, 39). Our experiments show that pretreatment with α-bgtx reduced the percentage of cells undergoing cell death after gp120 treatment. These data suggest that the gp120-induced up-regulation of functional α7-nAChRs may be detrimental to neurons and implicates the α7-nAChR as a potential key player in HIV pathogenesis. It is important to mention that desensitization of α7-nAChRs by chronic exposure to different agonists, such as nicotine, has been associated to receptor up-regulation and neuroprotection (40, 41). This scenario differs from the gp120-induced α7-nAChR up-regulation and toxicity in that the observed gp120 effects involve activation of the CXCR4 receptor. It is conceivable that chronic nicotine exposure leads to an increase in α7-nAChRs to compensate for the reduced receptor activity due to desensitization. In contrast, our data suggest that gp120 activates CXCR4 receptors, leading to an increase in functional α7-nAChRs without involving desensitization. Whereas small influxes in Ca2+ associated with tonic small levels of activation of α7-nAChRs might be neuroprotective, large influxes of Ca2+ as consequence of receptor up-regulation appear to be cytotoxic (42, 43).
Many studies have shown that gp120 binding to the CXCR4 or CCR5 receptors elicits intracellular responses that include the activation of the MAPK pathway, transcription factors, and ion channels (32, 44–46). In the course of HIV infection, the virus exhibits changes in its tropism: from an R5 variant to an X4 variant (15). By the time HIV-1 becomes capable of infecting cells through the use of the CXCR4, there is an associated CD4 cell decline and disease progression (20). This rapid decrease in CD4 cell count associated with CXCR4 tropism couples with an increased risk of developing HAND (18). This correlates with the fact that hippocampal and basal ganglia neurons express CXCR4 but not CCR5, which is present in glial cells (32). Previous studies have demonstrated that activation of CCR5 can lead to cell death in the presence of gp120 (51); however, our experiments show that gp120IIIB as well as SDF-1α, both specific to CXCR4, induce an up-regulation of the α7-nAChR. Moreover, this effect is abolished by pretreatment with the CXCR4 antagonist AMD3100, ruling out the contribution of CCR5 in our experimental setting. These results show that CXCR4 activation is necessary for the up-regulation of α7-nAChRs, and CXCR4-expressing neurons would be more susceptible to the effects of HIV infection.
CXCR4 activates the MAPK pathway, and its activation by SDF-1α induces the up-regulation of Egr-1, a transcription factor capable of stimulating the expression of the α7 gene (CHRNA7) (23, 24). Our experiments show that treating SH-SY5Y cells with SDF-1α induced a functional up-regulation of α7-nAChRs, as seen by increased α-bgtx binding and larger ACh-stimulated whole cell currents. Furthermore, quantitative RT-PCR experiments showed that gp120 and SDF-1α treatment of SH-SY5Y cells induced a rapid and transient increase in Egr-1 mRNA expression levels concomitant with an increase in α7-nAChRs mRNA expression levels. Together, these results provide a molecular mechanism as to how the α7-nAChR could be up-regulated in neurons in the context of an HIV-infected CNS.
In HAND, the sustained immune response can produce neuronal injury despite viral control (47). In the brain, viral particles such as gp120 can cause macrophages and/or astrocytes to release various cytokines and chemokines (nitric oxide, TNF-α, IL-1, IL-6, monocyte chemoattractant protein 1), resulting in further neuronal stress and cell death. Production of IL-1 by astrocytes leads to an increase in SDF-1 (25). The α7-nAChR has been shown to contribute to the regulation of inflammation in macrophages (the cholinergic anti-inflammatory pathway); however, the sustained immune response and increased production of SDF-1 could cause alterations in α7-nAChR function. Immune responses depend on the equilibrium between pro- and anti-inflammatory cytokines, and alterations of this equilibrium could convert a beneficial inflammatory response into a pathologic process (48). In this context, gp120 and SDF-1 could have synergistic effects, leading to up-regulation of the α7-nAChR and subsequent neuronal death. Interestingly, a similar up-regulation of α7-nAChR has been observed in macrophages derived from HIV+ donors,5 but the implications of these findings for macrophage function and inflammation remain to be explored.
To validate our results in an in vivo model, we used transgenic mice that express HIV-gp120 under the glial fibrillary acidic protein promoter. HIV-gp120 has been shown to induce neuropathological changes in mice similar to those seen in HAND patients (49, 50), making it a suitable model to study gp120-mediated neurotoxicity. Consistent with the results from in vitro experiments, we observed that α7-nAChR mRNA and protein levels are increased in the CNS of gp120-transgenic mice. However, the α7-nAChR up-regulation appears to be restricted to the striatum, a component of the basal ganglia and a region greatly affected in HAND patients (47).
Our results are in agreement with experiments performed by Kaul et al. (51) showing that gp120IIIB and SDF-1 are neurotoxic only in the presence of the CXCR4 receptor. In these experiments it was shown that inhibiting p38 prevented the gp120 and SDF-1 neurotoxicity. Interestingly, it has been shown that induction of Egr-1 requires p38 activity (52). These data, in conjunction with our results, suggest the existence of a neurotoxic pathway in which CXCR4 activation leads to a p38-dependent Egr-1 induction that increases α7-nAChR expression levels and causes cell death.
In conclusion, our experiments suggest that the α7-nAChR is a previously unrecognized contributor to HIV neurotoxicity. Drugs that antagonize the CXCR4 or α7-nACh receptors might be of therapeutic benefit in the treatment of HAND.
Supplementary Material
This work was supported, in whole or in part, by National Institutes of Health Grant SNRP-U54N54301.

This article contains supplemental Figs. S1–S3 and Videos S1–S4.
L. Y. Ballester, C. M. Capó-Vélez, W. F. García-Beltrán, F. M. Ramos, E. Vázquez-Rosa, R. Ríos, J. R. Mercado, R. I. Meléndez, and J. A. Lasalde-Dominicci, unpublished data.
- HAND
- HIV-associated neurocognitive disorder
- α-bgtx
- α-bungarotoxin
- ACh
- acetylcholine
- Egr-1
- early growth response gene 1
- gp
- glycoprotein
- HAART
- highly active antiretroviral therapy
- nAChR
- nicotinic ACh receptor
- pF
- pico-Farad
- SDF
- stromal cell-derived growth factor.
REFERENCES
- 1. UNAIDS/WHO (2008) Report on Global AIDS Epidemic [Google Scholar]
- 2. Tozzi V., Balestra P., Bellagama R., Corpolongo A., Salvatori M. F., Visco-Comandini U., Vlassi C., Giulianelli M., Galgani S., Antinori A., Narciso P. (2007) J. Acquir. Immune Defic. Syndr. 45, 174–1820 [DOI] [PubMed] [Google Scholar]
- 3. Sacktor N., Lyles R. H., Skolasky R., Kleeberger C., Selnes O. A., Miller E. N., Becker J. T., Cohen B., McArthur J. C. (2001) Neurology 56, 257–260 [DOI] [PubMed] [Google Scholar]
- 4. Njamnshi A. K., Bissek A. C., Ongolo-Zogo P., Tabah E. N., Lekoubou A. Z., Yepnjio F. N., Fonsah J. Y., Kuate C. T., Angwafor S. A., Dema F., Njamnshi D. M., Kouanfack C., Djientcheu Vde P., Muna W. F., Kanmogne G. D. (2009) J. Neurol. Sci. 285, 149–153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Sacktor N. (2002) J. Neurovirol. 8, 115–121 [DOI] [PubMed] [Google Scholar]
- 6. Wong M. H., Robertson K., Nakasujja N., Skolasky R., Musisi S., Katabira E., McArthur J. C., Ronald A., Sacktor N. (2007) Neurology 68, 350–355 [DOI] [PubMed] [Google Scholar]
- 7. Kaul M. (2009) Curr. Opin. Neurol. 22, 315–320 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Ghafouri M., Amini S., Khalili K., Sawaya B. E. (2006) Retrovirology 3, 28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Mattson M. P., Haughey N. J., Nath A. (2005) Cell Death Differ. 12, Suppl. 1, 893–904 [DOI] [PubMed] [Google Scholar]
- 10. Bardi G., Sengupta R., Kahn M. Z., Patel J. P., Meucci O. (2006) J. Neurovirol. 12, 211–218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Dreyer E. B., Kaiser P. K., Offerman J. T., Lipton S. A. (1990) Science 248, 364–367 [DOI] [PubMed] [Google Scholar]
- 12. Okamoto S., Kang Y. J., Brechtel C. W., Siviglia E., Russo R., Clemente A., Harrop A., McKercher S., Kaul M., Lipton S. A. (2007) Cell Stem Cell 1, 230–236 [DOI] [PubMed] [Google Scholar]
- 13. Buzy J., Brenneman D. E., Pert C. B., Martin A., Salazar A., Ruff M. R. (1992) Brain Res. 598, 10–18 [DOI] [PubMed] [Google Scholar]
- 14. Bracci L., Lozzi L., Rustici M., Neri P. (1992) FEBS Lett. 311, 115–118 [DOI] [PubMed] [Google Scholar]
- 15. Berger E. A., Murphy P. M., Farber J. M. (1999) Annu. Rev. Immunol. 17, 657–700 [DOI] [PubMed] [Google Scholar]
- 16. Lee C., Liu Q. H., Tomkowicz B., Yi Y., Freedman B. D., Collman R. G. (2003) J. Leukocyte Biol. 74, 676–682 [DOI] [PubMed] [Google Scholar]
- 17. Conejero-Goldberg C., Davies P., Ulloa L. (2008) Neurosci. Biobehav. Rev. 32, 693–706 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Giunta B., Ehrhart J., Townsend K., Sun N., Vendrame M., Shytle D., Tan J., Fernandez F. (2004) Brain Res. Bull. 64, 165–170 [DOI] [PubMed] [Google Scholar]
- 19. Rock R. B., Gekker G., Aravalli R. N., Hu S., Sheng W. S., Peterson P. K. (2008) J. Neuroimmune Pharmacol. 3, 143–149 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Stalmeijer E. H., Van Rij R. P., Boeser-Nunninck B., Visser J. A., Naarding M. A., Schols D., Schuitemaker H. (2004) J. Virol. 78, 2722–2728 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Nieves D. M., Plaud M., Wojna V., Skolasky R., Meléndez L. M. (2007) J. Neurovirol. 13, 315–327 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Missé D., Cerutti M., Noraz N., Jourdan P., Favero J., Devauchelle G., Yssel H., Taylor N., Veas F. (1999) Blood 93, 2454–2462 [PubMed] [Google Scholar]
- 23. Luo Y., Lathia J., Mughal M., Mattson M. P. (2008) J. Biol. Chem. 283, 24789–24800 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Criado M., Domínguez del Toro E., Carrasco-Serrano C., Smillie F. I., Juíz J. M., Viniegra S., Ballesta J. J. (1997) J. Neurosci. 17, 6554–6564 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Peng H., Erdmann N., Whitney N., Dou H., Gorantla S., Gendelman H. E., Ghorpade A., Zheng J. (2006) Glia 54, 619–629 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Geminder H., Sagi-Assif O., Goldberg L., Meshel T., Rechavi G., Witz I. P., Ben-Baruch A. (2001) J. Immunol. 167, 4747–4757 [DOI] [PubMed] [Google Scholar]
- 27. Navedo M. F., Amberg G. C., Nieves M., Molkentin J. D., Santana L. F. (2006) J. Gen. Physiol. 127, 611–622 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. McGehee D. S., Role L. W. (1995) Annu. Rev. Physiol. 57, 521–546 [DOI] [PubMed] [Google Scholar]
- 29. Séguéla P., Wadiche J., Dineley-Miller K., Dani J. A., Patrick J. W. (1993) J. Neurosci. 13, 596–604 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Mattson M. P., Chan S. L. (2003) Nat. Cell Biol. 5, 1041–1043 [DOI] [PubMed] [Google Scholar]
- 31. Nagavarapu U., Danthi S., Boyd R. T. (2001) J. Biol. Chem. 276, 16749–16757 [DOI] [PubMed] [Google Scholar]
- 32. van der Meer P., Ulrich A. M., González-Scarano F., Lavi E. (2000) Exp. Mol. Pathol. 69, 192–201 [DOI] [PubMed] [Google Scholar]
- 33. Zheng J., Thylin M. R., Ghorpade A., Xiong H., Persidsky Y., Cotter R., Niemann D., Che M., Zeng Y. C., Gelbard H.A., Shepard R. B., Swartz J. M., Gendelman H. E. (1999) J. Neuroimmunol. 98, 185–200 [DOI] [PubMed] [Google Scholar]
- 34. Hesselgesser J., Taub D., Baskar P., Greenberg M., Hoxie J., Kolson D. L., Horuk R. (1998) Curr. Biol. 8, 595–598 [DOI] [PubMed] [Google Scholar]
- 35. Marambaud P., Dreses-Werringloer U., Vingtdeux V. (2009) Mol. Neurodegener. 4, 20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Counts S. E., He B., Che S., Ikonomovic M. D., DeKosky S. T., Ginsberg S. D., Mufson E. J. (2007) Arch. Neurol. 64, 1771–1776 [DOI] [PubMed] [Google Scholar]
- 37. Lipton S. A. (1992) Neurology 42, 1403–1405 [DOI] [PubMed] [Google Scholar]
- 38. Aracava Y., Pereira E. F., Maelicke A., Albuquerque E. X. (2005) J. Pharmacol. Exp. Ther. 312, 1195–1205 [DOI] [PubMed] [Google Scholar]
- 39. Catani M. V., Corasaniti M. T., Navarra M., Nisticò G., Finazzi-Agrò A., Melino G. (2000) J. Neurochem. 74, 2373–2379 [DOI] [PubMed] [Google Scholar]
- 40. Jonnala R. R., Buccafusco J. J. (2001) J. Neurosci. Res. 66, 565–572 [DOI] [PubMed] [Google Scholar]
- 41. De Rosa M. J., Esandi Mdel C., Garelli A., Rayes D., Bouzat C. (2005) J. Neuroimmunol. 160, 154–161 [DOI] [PubMed] [Google Scholar]
- 42. Johnson E. M., Jr., Koike T., Franklin J. (1992) Exp. Neurol. 115, 163–166 [DOI] [PubMed] [Google Scholar]
- 43. Uteshev V. V., Meyer E. M., Papke R. L. (2002) Brain Res. 948, 33–46 [DOI] [PubMed] [Google Scholar]
- 44. Liu Q. H., Williams D. A., McManus C., Baribaud F., Doms R. W., Schols D., De Clercq E., Kotlikoff M. L., Collman R. G., Freedman B. D. (2000) Proc. Natl. Acad. Sci. U.S.A. 97, 4832–4837 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Stantchev T. S., Broder C. C. (2001) Cytokine Growth Factor Rev. 12, 219–243 [DOI] [PubMed] [Google Scholar]
- 46. Del Corno M., Liu Q. H., Schols D., de Clercq E., Gessani S., Freedman B. D., Collman R. G. (2001) Blood 98, 2909–2916 [DOI] [PubMed] [Google Scholar]
- 47. Woods S. P., Moore D. J., Weber E., Grant I. (2009) Neuropsychol. Rev. 19, 152–168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Tyagi E., Agrawal R., Nath C., Shukla R. (2010) Innate Immun. 16, 3–13 [DOI] [PubMed] [Google Scholar]
- 49. Toggas S. M., Masliah E., Rockenstein E. M., Rall G. F., Abraham C. R., Mucke L. (1994) Nature 367, 188–193 [DOI] [PubMed] [Google Scholar]
- 50. D'Hooge R., Franck F., Mucke L., De Deyn P. P. (1999) Eur. J. Neurosci. 11, 4398–4402 [DOI] [PubMed] [Google Scholar]
- 51. Kaul M., Ma Q., Medders K. E., Desai M. K., Lipton S. A. (2007) Cell Death Differ. 14, 296–305 [DOI] [PubMed] [Google Scholar]
- 52. Kenzel S., Santos-Sierra S., Deshmukh S. D., Moller I., Ergin B., Fitzgerald K. A., Lien E., Akira S., Golenbock D. T., Henneke P. (2009) Infect. Immun. 77, 2474–2481 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.