Abstract
DNA methylation, a widely used epigenetic mark, has been associated with many tumors. However, few studies have addressed the role of cell-free plasma DNA methylation in discriminating aggressive prostate cancer (PCa) from indolent cases. We conducted a case series and a case–control study among histologically confirmed stage II/III cases and matched controls recruited at Columbia University Medical Center. The aim of this study was to investigate whether plasma DNA methylation levels are appropriate surrogate biomarker of PCa tumor tissue levels and whether these markers are associated with worse clinicopathological tumor characteristics, which correlate with poorer prognosis. Quantitative pyrosequencing was used to detect methylation levels of p16 (CDKN4A), APC, GSTP1, and LINE-1 in 24 pairs of prostate tumor and adjacent tissues, as well as 27 plasma samples of PCa patients and 24 of controls. DNA methylation levels were significantly higher in tumor tissue than in adjacent nontumor tissue for p16 (CDKN4A), GSTP1, and APC; GSTP1 had a higher average percentage methylation in tumor tissue (38.9%) compared with p16 (CDKN4A) (5.9%) and APC (14.5%). GSTP1, p16 (CDKN4A), and APC methylation in tumor tissue was statistically significantly higher for cases with Gleason score ≥7 compared with those with Gleason score <7 [49.0% vs. 21.9% (p=0.01), 6.6% vs. 4.5% (p=0.04), and 19.1% vs. 7.4% (p=0.02), respectively]. Plasma LINE-1 methylation levels were higher in those with higher Gleason (67.6%) than in those with Gleason's below 7 (64.6%, p=0.03). Significant plasma–tissue correlations were observed for GSTP1 and LINE-1 methylation. These data, although preliminary, suggest that aberrant methylation may be a useful marker to identify PCa patients with clinically aggressive disease.
Introduction
Prostate cancer (PCa) is the most commonly diagnosed cancer and the second leading cause of cancer death in men in the United States (Jemal et al., 2010). Current diagnosis of PCa relies on widespread prostate-specific antigen (PSA) screening and/or digital rectal examination (Cook and Nelson, 2011). However, conflicting results have been found when investigating the benefit of PSA screening on PCa-associated mortality (Andriole et al., 2009; Schroder et al., 2009; Hugosson et al., 2010). Results from the European Randomized Study of Screening for Prostate Cancer showed that 43 PCa cases need to be treated to prevent one death from the disease (Loeb et al., 2010). Therefore, one of the research challenges is how to discriminate patients with nonfatal disease from those at high risk of PCa-related death (Brawley et al., 2009; Tang et al., 2010). There is an urgent need to find biomarkers to accurately distinguish aggressive PCa from the overwhelming majority of cases.
DNA methylation is a widely recognized epigenetic mark and it has been associated with prognosis for many malignancies (Deng et al., 2010; Veeck and Esteller, 2010). One of the epigenetic mechanisms involved in cancer progression is the silencing of relevant tumor suppressor genes by hypermethylation of their promoter regions. Previous studies of DNA methylation and PCa risk have found that specific promoter sequences are hypermethylated at a higher frequency in PCa tumors than in nontumor tissues (Nelson et al., 2007). The most frequent epigenetic mark in PCa is GSTP1 promoter hypermethylation, which has been found in cancerous and precancerous prostate tissues (Meiers et al., 2007) as well as in serum, urine, and ejaculate of PCa patients (Dobosy et al., 2007; Ahmed, 2010). Methylation of promoter regions of other genes has also been reported, but results are inconsistent throughout different studies (Ahmed, 2010; Phe et al., 2010). Few studies have addressed the most relevant question of DNA methylation in discriminating aggressive from indolent PCa. We carried out a case series analysis to explore whether DNA methylation in cell-free plasma can be used as a surrogate for prostate tumor tissue and to investigate the correlations between methylation markers and clinicopathological variables. Candidate genes were selected on the basis of prior data suggesting differences in plasma DNA methylation between PCa cases and controls (Li, 2007; Phe et al., 2010), and LINE-1, a marker of global methylation levels, was also investigated (Yegnasubramanian et al., 2008).
Materials and Methods
Patients and sample collection
This study was approved by the Institutional Review Board of Columbia University Medical Center (CUMC). A total of 27 PCa patients who underwent radical prostatectomy at CUMC during the period of December 2008 to March 2009 were enrolled in this study and blood samples were obtained prior to treatment. All cases were histologically confirmed and had tumors with clinical stage II or III, no clinical evidence of lymph node or distant metastases, available specimens, and complete clinical and serum PSA data. Prostate tissue samples from 24 of these cases were obtained at the time of radical prostatectomy; microdissection of the tissue yielded tumor and nontumor tissue, from which DNA was separately extracted. Additional plasma fractions were obtained from 24 age-matched males who participated in a prior vitamin intervention study [described in detail elsewhere (Jacobson et al., 2000)].
Pyrosequencing measurements
DNA was extracted from plasma using Qiagen Viral Nucleic Acid Kits (Qiagen, Valencia, CA) and from tissue by traditional phenol–chloroform extraction methods. DNA samples were quantitated using a Nanodrop spectophotometer (Thermo Fisher Scientific, Wilmington, DE). Approximately 200 ng of extracted DNA were bisulfite converted using ZYMO Gold (ZYMO, Orange, CA) following the conditions suggested by the manufacturer. Methylation levels of the promoter regions of p16 (CDKN4A), APC, GSTP1, and LINE-1 were determined by pyrosequencing after bisulfite conversion. Briefly, 30–50 ng of bisulfite-converted DNAs were used as a template for PCR amplification using Qiagen PyroMark PCR mix (Qiagen). Twenty-five microliters of the PCR product was subjected to quantitative pyrosequencing analysis using a PyroMark™Q24 system (Qiagen). This is a quantitative measure and percentage of DNA methylation is given for each CpG site and is reported as an average of all measured sites. The number of CpG sites per gene are as follows: APC and LINE-1, three sites; GSTP1, four sites; and p16 (CDKN4A), seven CpG sites. All values reported are the average of duplicate samples. Primers and amplification conditions for p16(CDKN4A), GSTP1, and LINE-1 have been previously described (Bollati et al., 2007). APC primers were specifically designed for this study as follows: forward primer 5′ GGTATGGGGTTAGGGTTAGGTAGG 3′, reverse primer 5′ Biotin-CCCACAACACCTCCATTCTATC 3′, and sequencing primer 5′ GAGAGAAGTA GTTGTGTAAT 3′.
Statistical analysis
Quantitative methylation data obtained from the pyrosequencing assay allows us not only to compare the correlations of continuous methylation variables between groups, but also to examine the impact of dichotomized methylation variables on disease outcomes. As methylation data were skewed, we log transformed the data. Methylation levels are reported as mean±SD, and comparisons between tumor and nontumor tissues from PCa cases and plasma DNAs from the cases and controls used the Student's t-test. Spearman correlation coefficients were calculated to evaluate the associations between methylation markers in tumor tissues and plasma DNAs, as well as associations between methylation markers and clinicopathological outcomes. Pathological tumor stage was categorized as pT2 versus pT3. Gleason score was categorized as 6 versus 7–9, and presurgery PSA level was categorized as ≤4 ng/mL versus >4 ng/mL. A p-value of <0.05 was considered statistically significant. All statistical analyses were performed using Statistical Analysis System 9.0 (SAS Institute, Cary, NC).
Results
The clinicopathologic characteristics of 27 PCa cases are shown in Table 1. The mean age of cases was 60 years, with an average PSA level of 9.4 (ng/mL). Most of our cases have tumors with stage 2 or below 2 (77%) and Gleason score 7 or below 7 (88%).
Table 1.
Demographic and Clinicopathologic Characteristics of the Prostate Cancer Cases Included in This Study
| Cases | |
|---|---|
| Age (years), mean±SD | 60±7 |
| Ethnicity, n (%) | |
| Caucasian | 15 (56%) |
| African American | 6 (22%) |
| Other | 6 (22%) |
| PSA levels (ng/mL), mean±SD | 9.4±10.7 |
| Stage, n (%) | |
| pT2a and b | 4 (15%) |
| pT2c | 16 (62%) |
| pT3a and b | 6 (23%) |
| Gleason sum score, n (%) | |
| 6 | 7 (27%) |
| 7 | 16 (61%) |
| 8–9 | 3 (12%) |
PSA, prostate-specific antigen.
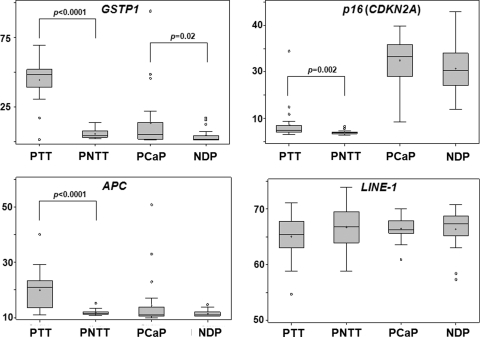
Gene-specific promoter DNA methylation levels were significantly higher in prostate tumor tissue than in adjacent nontumor tissue for the three candidate genes (p16 (CDKN4A), GSTP1, and APC) (Fig. 1). Of the individual promoter sequence studied, GSTP1 had a higher average % methylation in tumor tissue (38.9%) compared with p16 (CDKN4A) (5.9%) and APC (14.5%). There was no significant difference in LINE-1 methylation used as an indicator of global DNA methylation levels.
FIG. 1.
Box plot graph of DNA methylation percentages in plasma and prostate tissue. p-values of significant differences are indicated. PTT, prostate tumor tissue; PNTT, prostate non-tumor tissue; PCaP, cell-free plasma of prostate cancer patients; NDP, cell-free plasma of healthy individuals.
Cell-free plasma DNA methylation levels in p16 (CDKN4A), GSTP1, and APC were higher in PCa cases compared with cancer-free controls (Fig. 1). Only GSTP1 displayed a statistically significant case–control difference (p=0.02) with 6.0% methylation in cases and 2.9% in controls. Significant tumor tissue–plasma correlations for DNA methylation were observed for two markers (GSTP1 and LINE-1). The Spearman's correlation coefficients for GSTP1 and LINE-1 were 0.446 (p=0.049) and 0.424 (p=0.038), respectively (Fig. 2).
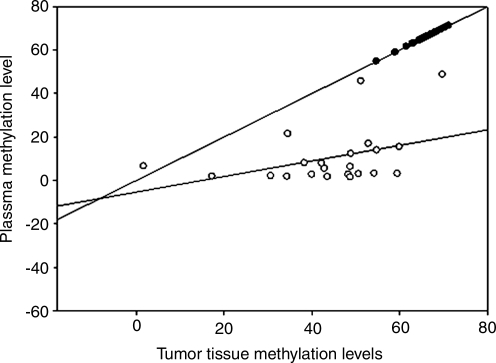
FIG. 2.
The Spearman's correlation coefficients for DNA methylation levels between prostate tumor tissue and matched plasma samples. Solid circles indicate the results for LINE-1, and open circles for GSTP1. The Spearman's correlation coefficients for LINE-1 and GSTP1 were 0.424 (p=0.038) and 0.446 (p=0.049), respectively.
We evaluated the associations between DNA methylation in prostate tumor tissue and cell-free plasma DNA and clinicopathological outcomes (Gleason score, tumor stage, and preoperative PSA levels). GSTP1, p16 (CDKN4A), and APC DNA methylation levels in tumor tissue were statistically significantly higher for cases with Gleason score ≥7 (49.0%, 6.6%, and 19.1%, respectively) compared with those with Gleason score <7 [21.9% (p=0.01), 4.5% (p=0.04), and 7.4% (p=0.02), respectively] (Table 2). In addition, APC methylation level was significantly higher in tumor tissues of stage pT3 or above (24.0%), compared with those with tumor stage pT2 (12.3%). Plasma but not tissue LINE-1 methylation level statistically significantly increased in cases with Gleason score ≥7 compared with those with Gleason score <7 (67.6% vs. 64.6%, p=0.03). There were no other significant associations for the studied DNA methylation markers in either tissue or plasma DNA, tumor stage, and preoperative PSA levels.
Table 2.
Quantitative DNA Methylation Levels for GSTP1, APC, P16 (CDKN2A), and LINE-1 by Gleason Score, Tumor Stage, and Preoperative Prostate-Specific Antigen Status
|
Clinical characteristics |
n |
p16 (CDKN4A) |
APC |
GSTP1 |
LINE-1 |
|---|---|---|---|---|---|
| Tumor tissue DNA methylation (%±SD) | |||||
| Gleason score | |||||
| <7 | 7 | 4.5±1.3 | 7.4±2.8 | 21.9±3.3 | 66.1±1.0 |
| ≥7 | 17 | 6.6±1.9 | 19.1±2.0 | 49.0±1.2 | 64.6±1.1 |
| p | 0.04 | 0.02 | 0.01 | 0.62 | |
| Tumor stage | |||||
| pT2 | 18 | 5.9±1.7 | 12.3±2.7 | 35.5±2.2 | 66.1±1.0 |
| pT3 | 6 | 5.8±1.8 | 24.0±1.3 | 51.3±1.2 | 61.7±1.1 |
| p | 0.94 | 0.02 | 0.09 | 0.20 | |
| Preoperative PSA | |||||
| <4 ng/mL | 5 | 5.6±1.7 | 25.1±1.8 | 43.7±1.1 | 67.6±1.0 |
| ≥4 ng/mL | 19 | 5.9±1.8 | 12.6±2.6 | 38.0±2.3 | 64.6±1.1 |
| p | 0.90 | 0.13 | 0.36 | 0.17 | |
| Cell-free plasma DNA methylation (%±SD) | |||||
|---|---|---|---|---|---|
| Gleason score | |||||
| <7 | 7 | 33.1±1.3 | 3.6±4.1 | 4.7±2.6 | 64.6±1.0 |
| ≥7 | 19 | 33.1±1.5 | 3.2±3.9 | 7.1±3.9 | 67.6±1.0 |
| p | 0.84 | 0.95 | 0.48 | 0.03 | |
| Tumor stage | |||||
| pT2 | 20 | 33.1±1.5 | 4.2±3.6 | 6.2±3.5 | 66.1±1.0 |
| pT3 | 6 | 31.6±1.4 | 1.7±4.1 | 6.8±3.8 | 67.6±1.0 |
| p | 0.74 | 0.17 | 0.88 | 0.10 | |
| Preoperative PSA | |||||
| <4 ng/mL | 8 | 34.7±1.3 | 3.6±5.0 | 9.8±4.2 | 66.1±1.0 |
| ≥4 ng/mL | 19 | 33.1±1.5 | 3.0±3.7 | 5.4±3.2 | 66.1±1.0 |
| p | 0.76 | 0.80 | 0.34 | 0.49 | |
Discussion
We applied a quantitative pyrosequencing approach to determine DNA methylation in cell-free plasma and prostate tumor/nontumor tissue DNA from PCa cases, as well as DNA from cancer-free individuals. The selected loci were those previously shown to discriminate presence of PCa in cases when compared with controls using urine, plasma, and serum samples (Hoque et al., 2005; Roupret et al., 2007, 2008; Li, 2007; Baden et al., 2009; Hoque, 2009; Phe et al., 2010). Our data included paired tumor, nontumor tissues, and plasma samples from cases, which allowed us to investigate not only tumor/nontumor differences, but also the tissue–plasma correlations. The frequency of hypermethylation in PCa tissues for GSTP1, p16 (CDKN4A), and APC ranged from 83.3% to 95.8%, similar to previous studies, although different approaches were used for methylation detection (Jeronimo et al., 2004; Kang et al., 2004; Maruyama et al., 2002). Consistent with previous research (Jarrard et al., 1997; Chu et al., 2002; Kang et al., 2004; Chuang et al., 2007; Phe et al., 2010), we also found significant tumor/nontumor differences in gene-specific methylation levels (Fig. 1). These data provide further support for the relevance of these epigenetic markers in prostate carcinogenesis.
Cell-free nucleic acids present in plasma are released by tumor cells either as a result of necrosis/apoptosis or by active release (Pinzani et al., 2010). Therefore, it is possible to measure plasma DNA methylation biomarker as surrogate of target prostate tissue. Our findings of significant correlations for methylation markers (GSTP1 and LINE-1) between tumor tissue and plasma samples support this hypothesis. Neoplasia is not the only contributor to an altered cell-free nucleic acid profile. Other diseases have been shown to affect cell-free plasma DNA levels and composition (Levenson, 2010). This might partly explain the difference between the prevalence of the different epigenetic marks in cell-free plasma DNA, which suggests that markers should be individually investigated. The circulating DNA levels may also be impacted by the efficiencies of phagocyte, namely, removing apoptotic cells from tissue, releasing them into the blood, and subsequently lysing those cells (Schwarzenbach et al., 2009; Thierry et al., 2010), which might partly explain why DNA methylation marks in cell-free plasma have a wide range even among the individuals free of disease.
Gleason score is the clinicopathological variable that most accurately predicts aggressiveness of PCa (Epstein, 2010). Consistent with most previous observations in prostate tumor tissue and blood (Jeronimo et al., 2004; Yegnasubramanian et al., 2004; Reibenwein et al., 2007; Sunami et al., 2009; Steiner et al., 2010), we found that elevated DNA methylation levels in prostate tumor tissues were related to worse clinicopathological outcomes. Specifically, GSTP1, p16 (CDKN4A), and APC methylation levels were significantly associated with higher Gleason score. In addition, we found that APC methylation level was significantly higher in tumor tissues of stage pT3 or above (24.0%), compared with those with tumor stage pT2 (12.3%). However, Bastian et al. (2005) did not find a link between GSTP1 or APC methylation and tumor progressive markers using methylation-specific PCR to measure DNA methylation.
Hypomethylation of LINE-1 is an epigenetic change that occurs later in prostate carcinogenesis (Schulz and Hoffmann, 2009). Hence, we did not anticipate detecting a case/control difference in this marker, as our cases have mostly early stage disease. We found that LINE-1 showed significant differences in plasma DNA methylation by Gleason score (Table 2). The direction of the methylation change is unexpected and our results suggest that a worse outcome might be reflected by hypermethylation of this marker in plasma DNA. This has not been previously reported. Therefore, further studies are warranted to clarify the role of this and other epigenetic markers of plasma DNA methylation in predicting PCa prognosis.
One strength of the present study is the inclusion of paired tumor, nontumor tissues, and plasma samples from the same PCa patients as well as unrelated control plasma samples. The quantitative pyrosequencing approach utilized offers quantitative methylation information of individual CpG site, allowing an in-depth analysis of associations between the biomarkers and clinicopathological variables (Tost and Gut, 2007). Our study overall has several limitations. One is the fact that we only have one-time measurement of the selected methylation markers. The amounts and composition of cell-free plasma DNA can vary over time in individuals suffering from different types of cancer (Jung et al., 2010). This is of particular importance in detecting plasma DNA methylation that may be affected by disease status (Levenson, 2010) at different time points. In addition, we used microdissection and cannot rule out the presence of some nontumor DNA in the tumor samples. This would lead to lower tumor DNA methylation levels and result in underestimation of the observed significant difference in the present study. Another limitation is the small sample size, which might have limited our ability to detect other significant associations. However, we analyzed multiple samples (tumor, nontumor, and plasma) from the same subjects. This allowed us to obtain data on the correlations of tissue–plasma and methylation–clinicopathological variables. Assuming the type I error probability for a two-sided test is 0.05 and the standard deviation of the methylation difference is 0.5 after logarithm transformation, with a sample of 24 pairs, we have a statistical power of 79% to detect a minimal 0.4 mean methylation difference. Larger prospective studies with serial measurements of methylation markers are clearly necessary to improve the validation of dynamic plasma DNA methylation alterations in PCa development and prognosis.
In conclusion, our results, although preliminary, suggest that aberrant methylation may serve as a useful marker to identify PCa patients with clinically aggressive disease.
Acknowledgment
This work was supported in part by NIH (grants P30 ES009089 and P30 CA013696).
Disclosure Statement
The authors declare that no competing financial interests exist.
References
- Ahmed H. Promoter methylation in prostate cancer and its application for the early detection of prostate cancer using serum and urine samples. Biomark Cancer. 2010;2010:17–33. doi: 10.4137/BIC.S3187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andriole G.L. Crawford E.D. Grubb R.L., III Buys S.S. Chia D. Church T.R. Fouad M.N. Gelmann E.P. Kvale P.A. Reding D.J. Weissfeld J.L. Yokochi L.A. O'Brien B. Clapp J.D. Rathmell J.M. Riley T.L. Hayes R.B. Kramer B.S. Izmirlian G. Miller A.B. Pinsky P.F. Prorok P.C. Gohagan J.K. Berg C.D. Mortality results from a randomized prostate-cancer screening trial. N Engl J Med. 2009;360:1310–1319. doi: 10.1056/NEJMoa0810696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baden J. Green G. Painter J. Curtin K. Markiewicz J. Jones J. Astacio T. Canning S. Quijano J. Guinto W. Leibovich B.C. Nelson J.B. Vargo J. Wang Y. Wuxiong C. Multicenter evaluation of an investigational prostate cancer methylation assay. J Urol. 2009;182:1186–1193. doi: 10.1016/j.juro.2009.05.003. [DOI] [PubMed] [Google Scholar]
- Bastian P.J. Ellinger J. Wellmann A. Wernert N. Heukamp L.C. Muller S.C. von Ruecker A. Diagnostic and prognostic information in prostate cancer with the help of a small set of hypermethylated gene loci. Clin Cancer Res. 2005;11:4097–4106. doi: 10.1158/1078-0432.CCR-04-1832. [DOI] [PubMed] [Google Scholar]
- Bollati V. Baccarelli A. Hou L. Bonzini M. Fustinoni S. Cavallo D. Byun H.M. Jiang J. Marinelli B. Pesatori A.C. Bertazzi P.A. Yang A.S. Changes in DNA methylation patterns in subjects exposed to low-dose benzene. Cancer Res. 2007;67:876–880. doi: 10.1158/0008-5472.CAN-06-2995. [DOI] [PubMed] [Google Scholar]
- Brawley O.W. Ankerst D.P. Thompson I.M. Screening for prostate cancer. CA Cancer J Clin. 2009;59:264–273. doi: 10.3322/caac.20026. [DOI] [PubMed] [Google Scholar]
- Chu D.C. Chuang C.K. Fu J.B. Huang H.S. Tseng C.P. Sun C.F. The use of real-time quantitative polymerase chain reaction to detect hypermethylation of the CpG islands in the promoter region flanking the GSTP1 gene to diagnose prostate carcinoma. J Urol. 2002;167:1854–1858. [PubMed] [Google Scholar]
- Chuang C.K. Chu D.C. Tzou R.D. Liou S.I. Chia J.H. Sun C.F. Hypermethylation of the CpG islands in the promoter region flanking GSTP1 gene is a potential plasma DNA biomarker for detecting prostate carcinoma. Cancer Detect Prev. 2007;31:59–63. doi: 10.1016/j.cdp.2006.11.001. [DOI] [PubMed] [Google Scholar]
- Cook E.D. Nelson A.C. Prostate cancer screening. Curr Oncol Rep. 2011;13:57–62. doi: 10.1007/s11912-010-0136-x. [DOI] [PubMed] [Google Scholar]
- Deng D. Liu Z. Du Y. Epigenetic alterations as cancer diagnostic, prognostic, and predictive biomarkers. Adv Genet. 2010;71:125–176. doi: 10.1016/B978-0-12-380864-6.00005-5. [DOI] [PubMed] [Google Scholar]
- Dobosy J.R. Roberts J.L. Fu V.X. Jarrard D.F. The expanding role of epigenetics in the development, diagnosis and treatment of prostate cancer and benign prostatic hyperplasia. J Urol. 2007;177:822–831. doi: 10.1016/j.juro.2006.10.063. [DOI] [PubMed] [Google Scholar]
- Epstein J.I. An update of the Gleason grading system. J Urol. 2010;183:433–440. doi: 10.1016/j.juro.2009.10.046. [DOI] [PubMed] [Google Scholar]
- Hoque M.O. DNA methylation changes in prostate cancer: current developments and future clinical implementation. Expert Rev Mol Diagn. 2009;9:243–257. doi: 10.1586/erm.09.10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoque M.O. Topaloglu O. Begum S. Henrique R. Rosenbaum E. Van Criekinge W. Westra W.H. Sidransky D. Quantitative methylation-specific polymerase chain reaction gene patterns in urine sediment distinguish prostate cancer patients from control subjects. J Clin Oncol. 2005;23:6569–6575. doi: 10.1200/JCO.2005.07.009. [DOI] [PubMed] [Google Scholar]
- Hugosson J. Carlsson S. Aus G. Bergdahl S. Khatami A. Lodding P. Pihl C.G. Stranne J. Holmberg E. Lilja H. Mortality results from the Goteborg randomised population-based prostate-cancer screening trial. Lancet Oncol. 2010;11:725–732. doi: 10.1016/S1470-2045(10)70146-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobson J.S. Begg M.D. Wang L.W. Wang Q. Agarwal M. Norkus E.P. Singh V.N. Young T.L. Yang D. Santella R.M. Effects of a six-month intervention on DNA damage in heavy smokers. Cancer Epidemiol Biomark Prevent. 2000;9:1303–1311. [PubMed] [Google Scholar]
- Jarrard D.F. Bova G.S. Ewing C.M. Pin S.S. Nguyen S.H. Baylin S.B. Cairns P. Sidransky D. Herman J.G. Isaacs W.B. Deletional, mutational, and methylation analyses of CDKN2 (p16/MTS1) in primary and metastatic prostate cancer. Genes Chromosomes Cancer. 1997;19:90–96. [PubMed] [Google Scholar]
- Jemal A. Siegel R. Xu J. Ward E. Cancer Statistics, 2010. CA Cancer J Clin. 2010;60:277–300. doi: 10.3322/caac.20073. [DOI] [PubMed] [Google Scholar]
- Jeronimo C. Henrique R. Hoque M.O. Mambo E. Ribeiro F.R. Varzim G. Oliveira J. Teixeira M.R. Lopes C. Sidransky D. A quantitative promoter methylation profile of prostate cancer. Clin Cancer Res. 2004;10:8472–8478. doi: 10.1158/1078-0432.CCR-04-0894. [DOI] [PubMed] [Google Scholar]
- Jung K. Fleischhacker M. Rabien A. Cell-free DNA in the blood as a solid tumor biomarker—a critical appraisal of the literature. Clin Chim Acta. 2010;411:1611–1624. doi: 10.1016/j.cca.2010.07.032. [DOI] [PubMed] [Google Scholar]
- Kang G.H. Lee S. Lee H.J. Hwang K.S. Aberrant CpG island hypermethylation of multiple genes in prostate cancer and prostatic intraepithelial neoplasia. J Pathol. 2004;202:233–240. doi: 10.1002/path.1503. [DOI] [PubMed] [Google Scholar]
- Levenson V.V. DNA methylation as a universal biomarker. Expert Rev Mol Diagn. 2010;10:481–488. doi: 10.1586/erm.10.17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li L.C. Epigenetics of prostate cancer. Front Biosci. 2007;12:3377–3397. doi: 10.2741/2320. [DOI] [PubMed] [Google Scholar]
- Loeb S. Vonesh E.F. Metter E.J. Carter H.B. Gann P.H. Catalona W.J. What is the true number needed to screen and treat to save a life with prostate-specific antigen testing? J Clin Oncol. 2010;29:464–467. doi: 10.1200/JCO.2010.30.6373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maruyama R. Toyooka S. Toyooka K.O. Virmani A.K. Zochbauer-Muller S. Farinas A.J. Minna J.D. McConnell J. Frenkel E.P. Gazdar A.F. Aberrant promoter methylation profile of prostate cancers and its relationship to clinicopathological features. Clin Cancer Res. 2002;8:514–519. [PubMed] [Google Scholar]
- Meiers I. Shanks J.H. Bostwick D.G. Glutathione S-transferase pi (GSTP1) hypermethylation in prostate cancer: review 2007. Pathology. 2007;39:299–304. doi: 10.1080/00313020701329906. [DOI] [PubMed] [Google Scholar]
- Nelson W.G. Yegnasubramanian S. Agoston A.T. Bastian P.J. Lee B.H. Nakayama M. De Marzo A.M. Abnormal DNA methylation, epigenetics, and prostate cancer. Front Biosci. 2007;12:4254–4266. doi: 10.2741/2385. [DOI] [PubMed] [Google Scholar]
- Phe V. Cussenot O. Roupret M. Methylated genes as potential biomarkers in prostate cancer. BJU Int. 2010;105:1364–1370. doi: 10.1111/j.1464-410X.2009.09167.x. [DOI] [PubMed] [Google Scholar]
- Pinzani P. Salvianti F. Pazzagli M. Orlando C. Circulating nucleic acids in cancer and pregnancy. Methods. 2010;50:302–307. doi: 10.1016/j.ymeth.2010.02.004. [DOI] [PubMed] [Google Scholar]
- Reibenwein J. Pils D. Horak P. Tomicek B. Goldner G. Worel N. Elandt K. Krainer M. Promoter hypermethylation of GSTP1, AR, and 14–3-3sigma in serum of prostate cancer patients and its clinical relevance. Prostate. 2007;67:427–432. doi: 10.1002/pros.20533. [DOI] [PubMed] [Google Scholar]
- Roupret M. Hupertan V. Catto J.W. Yates D.R. Rehman I. Proctor L.M. Phillips J. Meuth M. Cussenot O. Hamdy F.C. Promoter hypermethylation in circulating blood cells identifies prostate cancer progression. Int J Cancer. 2008;122:952–956. doi: 10.1002/ijc.23196. [DOI] [PubMed] [Google Scholar]
- Roupret M. Hupertan V. Yates D.R. Catto J.W. Rehman I. Meuth M. Ricci S. Lacave R. Cancel-Tassin G. de la T.A. Rozet F. Cathelineau X. Vallancien G. Hamdy F.C. Cussenot O. Molecular detection of localized prostate cancer using quantitative methylation-specific PCR on urinary cells obtained following prostate massage. Clin Cancer Res. 2007;13:1720–1725. doi: 10.1158/1078-0432.CCR-06-2467. [DOI] [PubMed] [Google Scholar]
- Schroder F.H. Hugosson J. Roobol M.J. Tammela T.L. Ciatto S. Nelen V. Kwiatkowski M. Lujan M. Lilja H. Zappa M. Denis L.J. Recker F. Berenguer A. Maattanen L. Bangma C.H. Aus G. Villers A. Rebillard X. van der K.T. Blijenberg B.G. Moss S.M. de Koning H.J. Auvinen A. Screening and prostate-cancer mortality in a randomized European study. N Engl J Med. 2009;360:1320–1328. doi: 10.1056/NEJMoa0810084. [DOI] [PubMed] [Google Scholar]
- Schulz W.A. Hoffmann M.J. Epigenetic mechanisms in the biology of prostate cancer. Semin Cancer Biol. 2009;19:172–180. doi: 10.1016/j.semcancer.2009.02.006. [DOI] [PubMed] [Google Scholar]
- Schwarzenbach H. Alix-Panabieres C. Muller I. Letang N. Vendrell J.P. Rebillard X. Pantel K. Cell-free tumor DNA in blood plasma as a marker for circulating tumor cells in prostate cancer. Clin Cancer Res. 2009;15:1032–1038. doi: 10.1158/1078-0432.CCR-08-1910. [DOI] [PubMed] [Google Scholar]
- Steiner I. Jung K. Schatz P. Horns T. Wittschieber D. Lein M. Dietel M. Erbersdobler A. Gene promoter methylation and its potential relevance in early prostate cancer diagnosis. Pathobiology. 2010;77:260–266. doi: 10.1159/000318017. [DOI] [PubMed] [Google Scholar]
- Sunami E. Shinozaki M. Higano C.S. Wollman R. Dorff T.B. Tucker S.J. Martinez S.R. Singer F.R. Hoon D.S. Multimarker circulating DNA assay for assessing blood of prostate cancer patients. Clin Chem. 2009;55:559–567. doi: 10.1373/clinchem.2008.108498. [DOI] [PubMed] [Google Scholar]
- Tang P. Sun L. Uhlman M.A. Polascik T.J. Freedland S.J. Moul J.W. Baseline PSA as a predictor of prostate cancer-specific mortality over the past 2 decades: Duke University experience. Cancer. 2010;116:4711–4717. doi: 10.1002/cncr.25447. [DOI] [PubMed] [Google Scholar]
- Thierry A.R. Mouliere F. Gongora C. Ollier J. Robert B. Ychou M. Del Rio M. Molina F. Origin and quantification of circulating DNA in mice with human colorectal cancer xenografts. Nucleic Acids Res. 2010;38:6159–6175. doi: 10.1093/nar/gkq421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tost J. Gut I.G. DNA methylation analysis by pyrosequencing. Nat Protoc. 2007;2:2265–2275. doi: 10.1038/nprot.2007.314. [DOI] [PubMed] [Google Scholar]
- Veeck J. Esteller M. Breast cancer epigenetics: from DNA methylation to microRNAs. J Mammary Gland Biol Neoplasia. 2010;15:5–17. doi: 10.1007/s10911-010-9165-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yegnasubramanian S. Haffner M.C. Zhang Y. Gurel B. Cornish T.C. Wu Z. Irizarry R.A. Morgan J. Hicks J. DeWeese T.L. Isaacs W.B. Bova G.S. De Marzo A.M. Nelson W.G. DNA hypomethylation arises later in prostate cancer progression than CpG island hypermethylation and contributes to metastatic tumor heterogeneity. Cancer Res. 2008;68:8954–8967. doi: 10.1158/0008-5472.CAN-07-6088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yegnasubramanian S. Kowalski J. Gonzalgo M.L. Zahurak M. Piantadosi S. Walsh P.C. Bova G.S. De Marzo A.M. Isaacs W.B. Nelson W.G. Hypermethylation of CpG islands in primary and metastatic human prostate cancer. Cancer Res. 2004;64:1975–1986. doi: 10.1158/0008-5472.can-03-3972. [DOI] [PubMed] [Google Scholar]