Abstract
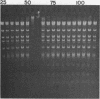
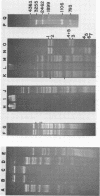
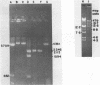
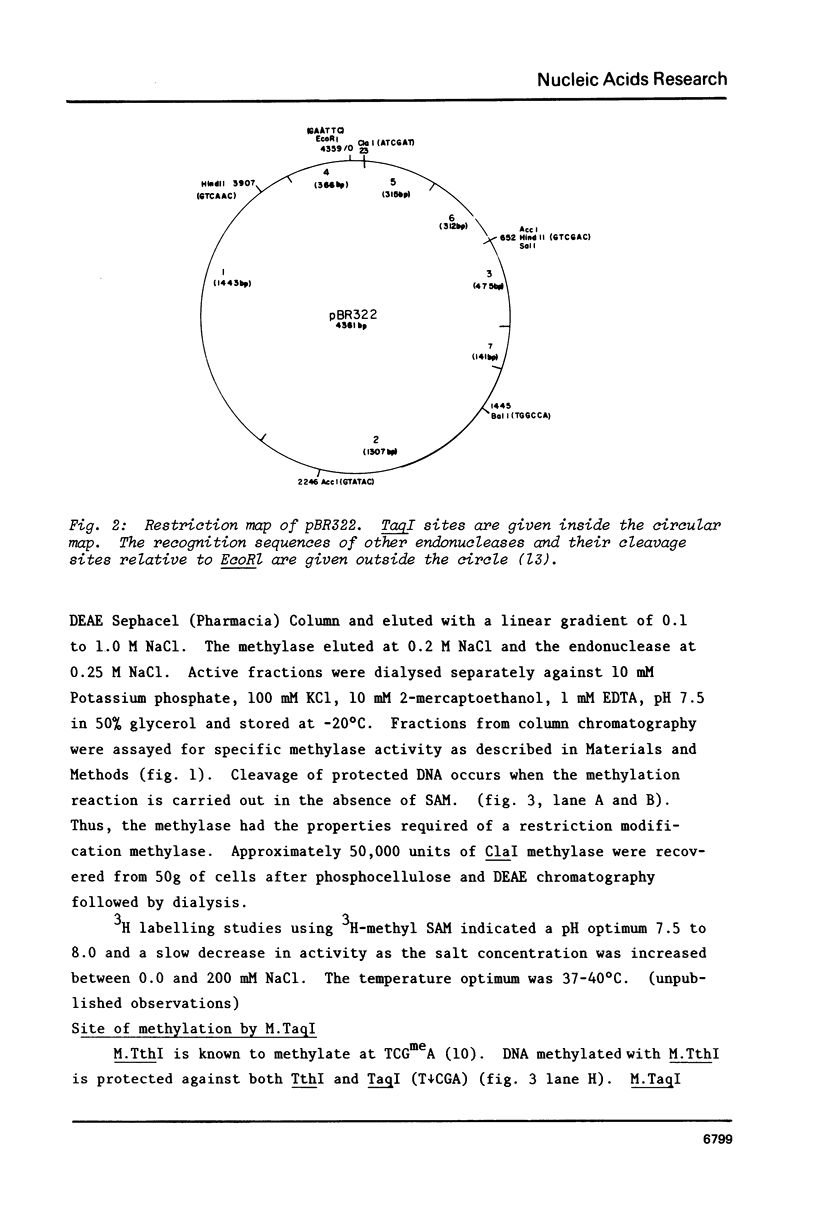
A method for detecting Type II modification methylases and determining their methylation site by assaying the ability of methylated DNA to be cleaved by heterologous restriction enzymes is described and applied to the isolation of the restriction modification methylases from Thermus thermophilus HB8, Thermus aquaticus YTI and Caryophanon latum L. M.TaqI is shown to have a methylation specificity identical to M.ThI (TCGmeA). M.ClaI methylates at adenine and protects a subset of TthI sites indicating that it methylates the sequence ATCGmeAT. Methylation by M.ThI also protects against cleavage by SalI, XhoI and at some HindII, AccI and MboI sites.
Full text
PDF




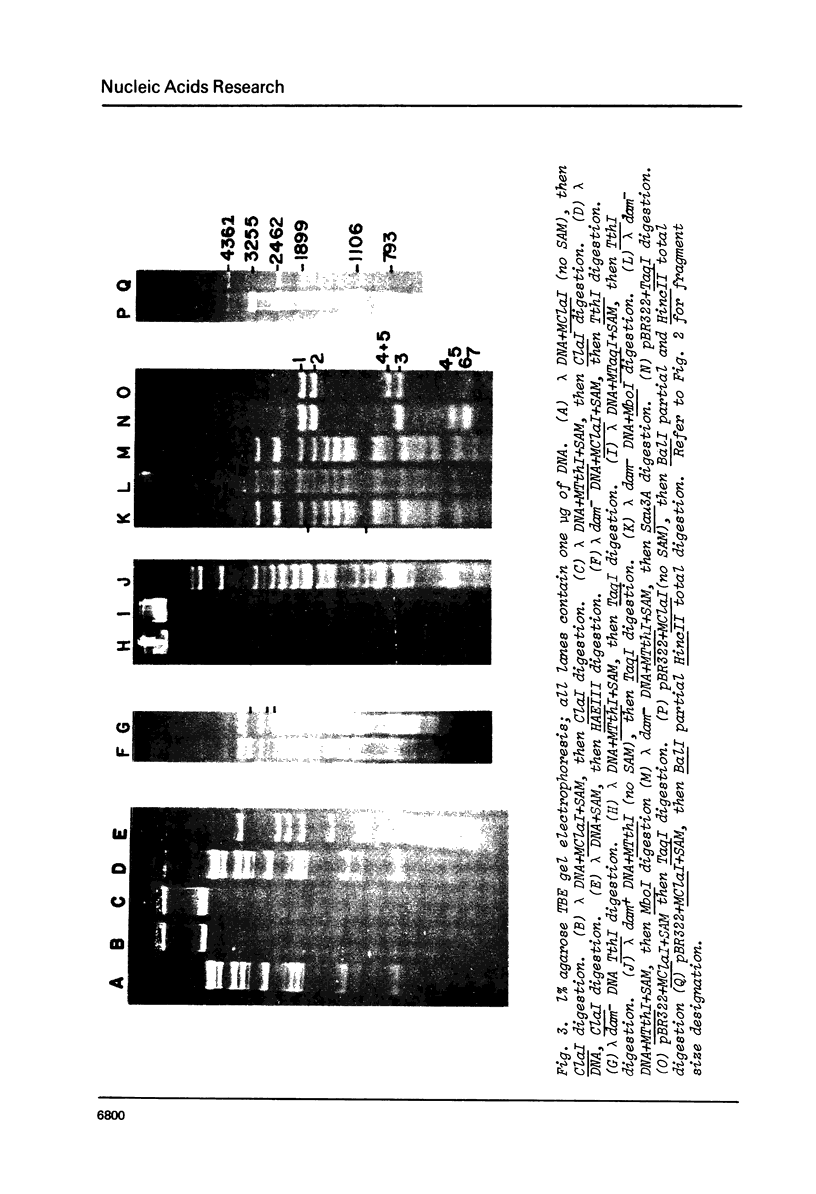


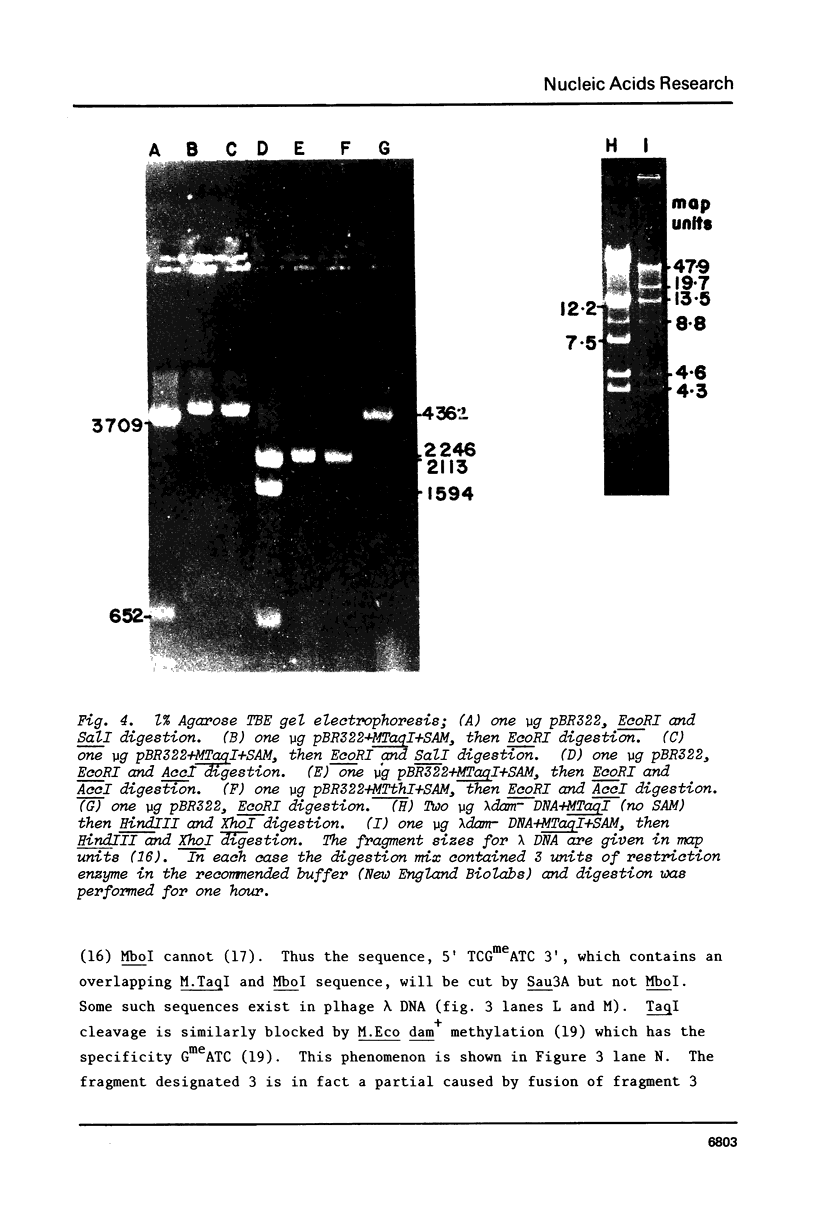

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arber W. Host-controlled modification of bacteriophage. Annu Rev Microbiol. 1965;19:365–378. doi: 10.1146/annurev.mi.19.100165.002053. [DOI] [PubMed] [Google Scholar]
- Backman K. A cautionary note on the use of certain restriction endonucleases with methylated substrates. Gene. 1980 Oct;11(1-2):169–171. doi: 10.1016/0378-1119(80)90097-9. [DOI] [PubMed] [Google Scholar]
- Bird A. P., Southern E. M. Use of restriction enzymes to study eukaryotic DNA methylation: I. The methylation pattern in ribosomal DNA from Xenopus laevis. J Mol Biol. 1978 Jan 5;118(1):27–47. doi: 10.1016/0022-2836(78)90242-5. [DOI] [PubMed] [Google Scholar]
- Bird A. P., Southern E. M. Use of restriction enzymes to study eukaryotic DNA methylation: I. The methylation pattern in ribosomal DNA from Xenopus laevis. J Mol Biol. 1978 Jan 5;118(1):27–47. doi: 10.1016/0022-2836(78)90242-5. [DOI] [PubMed] [Google Scholar]
- Dreiseikelmann B., Eichenlaub R., Wackernagel W. The effect of differential methylation by Escherichia coli of plasmid DNA and phage T7 and lambda DNA on the cleavage by restriction endonuclease MboI from Moraxella bovis. Biochim Biophys Acta. 1979 May 24;562(3):418–428. doi: 10.1016/0005-2787(79)90105-9. [DOI] [PubMed] [Google Scholar]
- Dugaiczyk A., Hedgpeth J., Boyer H. W., Goodman H. M. Physical identity of the SV40 deoxyribonucleic acid sequence recognized by the Eco RI restriction endonuclease and modification methylase. Biochemistry. 1974 Jan 29;13(3):503–512. doi: 10.1021/bi00700a016. [DOI] [PubMed] [Google Scholar]
- Ehrlich M., Wang R. Y. 5-Methylcytosine in eukaryotic DNA. Science. 1981 Jun 19;212(4501):1350–1357. doi: 10.1126/science.6262918. [DOI] [PubMed] [Google Scholar]
- Roberts R. J. Restriction and modification enzymes and their recognition sequences. Nucleic Acids Res. 1981 Jan 10;9(1):r75–r96. doi: 10.1093/nar/9.1.213-c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roy P. H., Smith H. O. DNA methylases of Hemophilus influenzae Rd. I. Purification and properties. J Mol Biol. 1973 Dec 25;81(4):427–444. doi: 10.1016/0022-2836(73)90515-9. [DOI] [PubMed] [Google Scholar]
- Sato S., Hutchinson C. A., 3rd, Harris J. I. A thermostable sequence-specific endonuclease from Thermus aquaticus. Proc Natl Acad Sci U S A. 1977 Feb;74(2):542–546. doi: 10.1073/pnas.74.2.542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato S., Nakazawa K., Shinomiya T. A DNA methylase from Thermus thermophilus HB8. J Biochem. 1980 Sep;88(3):737–747. doi: 10.1093/oxfordjournals.jbchem.a133026. [DOI] [PubMed] [Google Scholar]
- Sato S., Shinomiya T. An isochizomer of TaqI from Thermus thermophilus HB8. J Biochem. 1978 Nov;84(5):1319–1321. doi: 10.1093/oxfordjournals.jbchem.a132252. [DOI] [PubMed] [Google Scholar]
- Stobberingh E. E., Schiphof R., Sussenbach J. S. Occurrence of a class II restriction endonuclease in Staphylococcus aureus. J Bacteriol. 1977 Aug;131(2):645–649. doi: 10.1128/jb.131.2.645-649.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waalwijk C., Flavell R. A. DNA methylation at a CCGG sequence in the large intron of the rabbit beta-globin gene: tissue-specific variations. Nucleic Acids Res. 1978 Dec;5(12):4631–4634. doi: 10.1093/nar/5.12.4631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Ploeg L. H., Flavell R. A. DNA methylation in the human gamma delta beta-globin locus in erythroid and nonerythroid tissues. Cell. 1980 Apr;19(4):947–958. doi: 10.1016/0092-8674(80)90086-0. [DOI] [PubMed] [Google Scholar]