Abstract
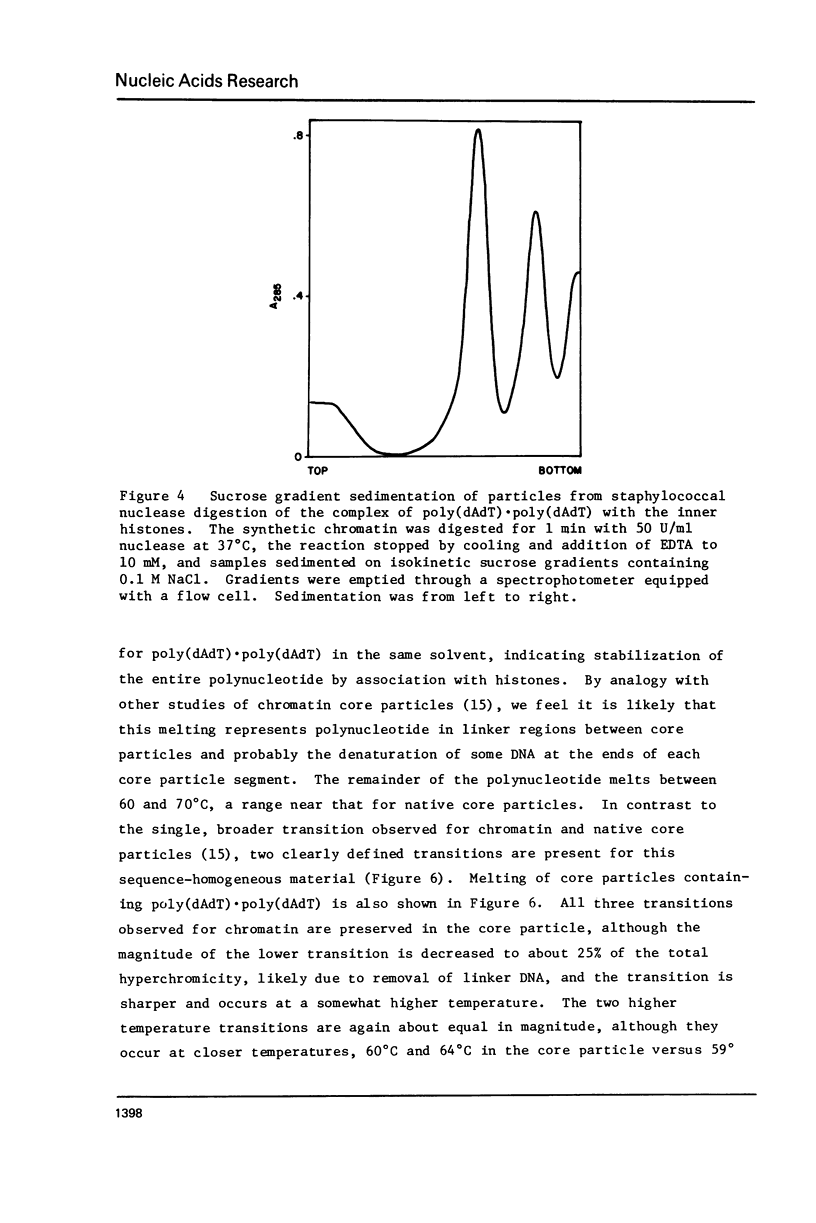
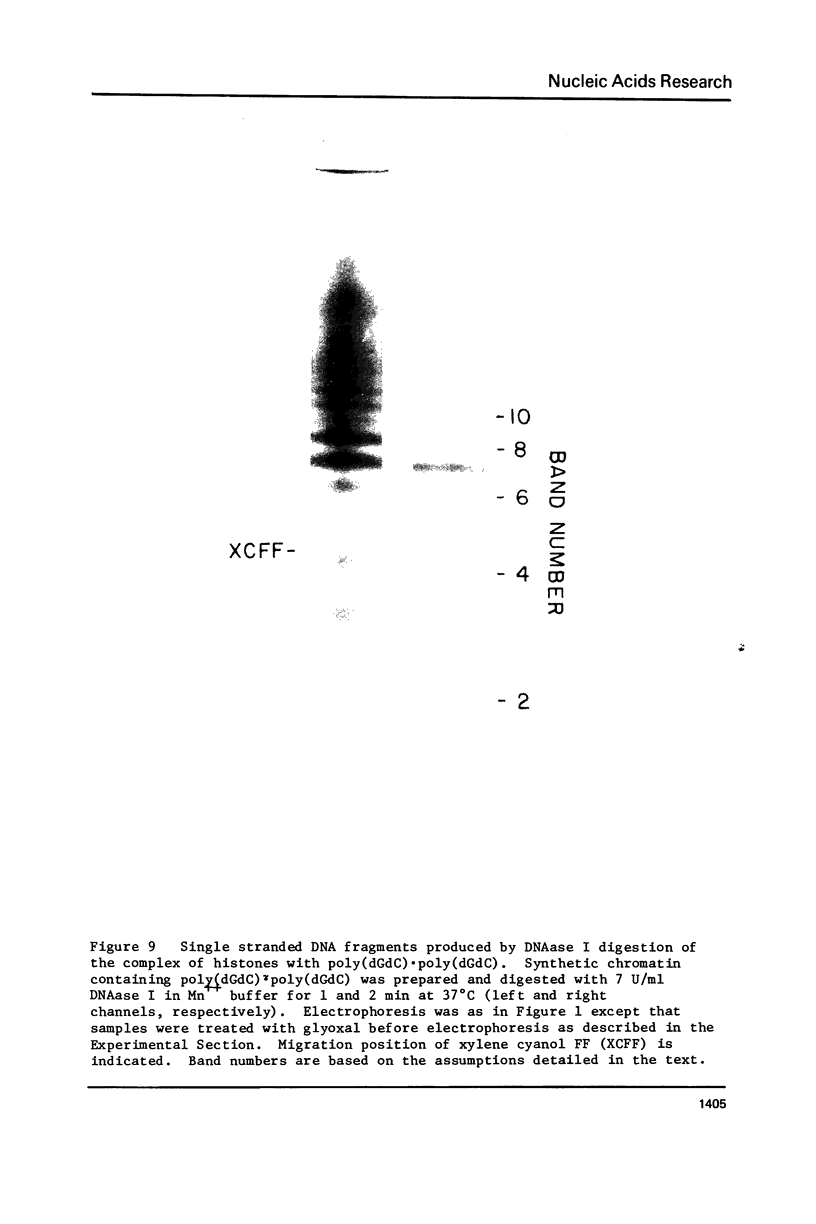
Chicken erythrocyte inner histones (H2A, H2B, H3 and H4) were associated with the two complementary homopolymeric polydeoxyribonucleotides and the two alternating copolymeric polydeoxyribonucleotides. No evidence for formation of chromatin-like structures was obtained for the complexes with poly(dG) . poly(dC) or poly(dA) . poly(dT). Both poly (dGdC) . poly(dGdC) and poly(dAdT) . poly(dAdT) could be folded by histones to yield material digested by DNAase I to multiples of about 10 and by staphylococcal nuclease to 146 bp core particles. Due to the lack of sequence heterogeniety in the complex of histones with poly(dAdT) . poly(dAdT), core particles with remarkable fine structural detail are obtained. The internal organization of DNA in the AT-containing and GC-containing core particles appears not to be identical.
Full text
PDF


























Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arnott S., Fuller W., Hodgson A., Prutton I. Molecular conformations and structure transitions of RNA complementary helices and their possible biological significance. Nature. 1968 Nov 9;220(5167):561–564. doi: 10.1038/220561a0. [DOI] [PubMed] [Google Scholar]
- Arnott S., Selsing E. Letter: The structure of polydeoxyguanylic acid with polydeoxycytidylic acid. J Mol Biol. 1974 Sep 15;88(2):551–552. doi: 10.1016/0022-2836(74)90502-6. [DOI] [PubMed] [Google Scholar]
- Arnott S., Selsing E. Structures for the polynucleotide complexes poly(dA) with poly (dT) and poly(dT) with poly(dA) with poly (dT). J Mol Biol. 1974 Sep 15;88(2):509–521. doi: 10.1016/0022-2836(74)90498-7. [DOI] [PubMed] [Google Scholar]
- BOLLUM F. J. DEGRADATION OF THE HOMOPOLYMER COMPLEXES POLYDEOXYADENYLATE-POLYDEOXYTHYMIDYLATE, POLYDEOXYINOSINATE-POLYDEOXYCYTIDYLATE, AND POLYDEOXYGUANYLATE-POLYDEOXYCYTIDYLATE BY DEOXYRIBONUCLEASE I. J Biol Chem. 1965 Jun;240:2599–2601. [PubMed] [Google Scholar]
- Crick F. H., Klug A. Kinky helix. Nature. 1975 Jun 12;255(5509):530–533. doi: 10.1038/255530a0. [DOI] [PubMed] [Google Scholar]
- Laskey R. A., Honda B. M., Mills A. D., Finch J. T. Nucleosomes are assembled by an acidic protein which binds histones and transfers them to DNA. Nature. 1978 Oct 5;275(5679):416–420. doi: 10.1038/275416a0. [DOI] [PubMed] [Google Scholar]
- Liu L. F., Wang J. C. DNA-DNA gyrase complex: the wrapping of the DNA duplex outside the enzyme. Cell. 1978 Nov;15(3):979–984. doi: 10.1016/0092-8674(78)90281-7. [DOI] [PubMed] [Google Scholar]
- Lutter L. C. Kinetic analysis of deoxyribonuclease I cleavages in the nucleosome core: evidence for a DNA superhelix. J Mol Biol. 1978 Sep 15;124(2):391–420. doi: 10.1016/0022-2836(78)90306-6. [DOI] [PubMed] [Google Scholar]
- Maniatis T., Jeffrey A., van deSande H. Chain length determination of small double- and single-stranded DNA molecules by polyacrylamide gel electrophoresis. Biochemistry. 1975 Aug 26;14(17):3787–3794. doi: 10.1021/bi00688a010. [DOI] [PubMed] [Google Scholar]
- Milman G., Langridge R., Chamberlin M. J. The structure of a DNA-RNA hybrid. Proc Natl Acad Sci U S A. 1967 Jun;57(6):1804–1810. doi: 10.1073/pnas.57.6.1804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noll M. DNA folding in the nucleosome. J Mol Biol. 1977 Oct 15;116(1):49–71. doi: 10.1016/0022-2836(77)90118-8. [DOI] [PubMed] [Google Scholar]
- Peacock A. C., Dingman C. W. Resolution of multiple ribonucleic acid species by polyacrylamide gel electrophoresis. Biochemistry. 1967 Jun;6(6):1818–1827. doi: 10.1021/bi00858a033. [DOI] [PubMed] [Google Scholar]
- ROBERTS W. K., DEKKER C. A., RUSHIZKY G. W., KNIGHT C. A. Studies on the mechanism of action of micrococcal nuclease. 1. Degradation of thymus deoxyribonucleic acid. Biochim Biophys Acta. 1962 May 14;55:664–673. doi: 10.1016/0006-3002(62)90844-2. [DOI] [PubMed] [Google Scholar]
- Reddy V. B., Thimmappaya B., Dhar R., Subramanian K. N., Zain B. S., Pan J., Ghosh P. K., Celma M. L., Weissman S. M. The genome of simian virus 40. Science. 1978 May 5;200(4341):494–502. doi: 10.1126/science.205947. [DOI] [PubMed] [Google Scholar]
- Riley D., Weintraub H. Nucleosomal DNA is digested to repeats of 10 bases by exonuclease III. Cell. 1978 Feb;13(2):281–293. doi: 10.1016/0092-8674(78)90197-6. [DOI] [PubMed] [Google Scholar]
- Rushizky G. W., Shaternikov V. A., Mozejko J. H., Sober H. A. S1 nuclease hydrolysis of single-stranded nucleic acids with partial double-stranded configuration. Biochemistry. 1975 Sep 23;14(19):4221–4226. doi: 10.1021/bi00690a011. [DOI] [PubMed] [Google Scholar]
- Scheffler I. E., Elson E. L., Baldwin R. L. Helix formation by dAT oligomers. I. Hairpin and straight-chain helices. J Mol Biol. 1968 Sep 28;36(3):291–304. doi: 10.1016/0022-2836(68)90156-3. [DOI] [PubMed] [Google Scholar]
- Simpson R. T. Structure of chromatin containing extensively acetylated H3 and H4. Cell. 1978 Apr;13(4):691–699. doi: 10.1016/0092-8674(78)90219-2. [DOI] [PubMed] [Google Scholar]
- Simpson R. T. Structure of the chromatosome, a chromatin particle containing 160 base pairs of DNA and all the histones. Biochemistry. 1978 Dec 12;17(25):5524–5531. doi: 10.1021/bi00618a030. [DOI] [PubMed] [Google Scholar]
- Simpson R. T., Whitlock J. P. Mapping DNAase l-susceptible sites in nucleosomes labeled at the 5' ends. Cell. 1976 Oct;9(2):347–353. doi: 10.1016/0092-8674(76)90124-0. [DOI] [PubMed] [Google Scholar]
- Sobell H. M., Tsai C. C., Gilbert S. G., Jain S. C., Sakore T. D. Organization of DNA in chromatin. Proc Natl Acad Sci U S A. 1976 Sep;73(9):3068–3072. doi: 10.1073/pnas.73.9.3068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stein A., Bina-Stein M., Simpson R. T. Crosslinked histone octamer as a model of the nucleosome core. Proc Natl Acad Sci U S A. 1977 Jul;74(7):2780–2784. doi: 10.1073/pnas.74.7.2780. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tatchell K., Van Holde K. E. Compact oligomers and nucleosome phasing. Proc Natl Acad Sci U S A. 1978 Aug;75(8):3583–3587. doi: 10.1073/pnas.75.8.3583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varshavsky A. J., Sundin O. H., Bohn M. J. SV40 viral minichromosome: preferential exposure of the origin of replication as probed by restriction endonucleases. Nucleic Acids Res. 1978 Oct;5(10):3469–3477. doi: 10.1093/nar/5.10.3469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weischet W. O., Tatchell K., Van Holde K. E., Klump H. Thermal denaturation of nucleosomal core particles. Nucleic Acids Res. 1978 Jan;5(1):139–160. doi: 10.1093/nar/5.1.139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wells R. D., Blakesley R. W., Hardies S. C., Horn G. T., Larson J. E., Selsing E., Burd J. F., Chan H. W., Dodgson J. B., Jensen K. F. The role of DNA structure in genetic regulation. CRC Crit Rev Biochem. 1977;4(3):305–340. doi: 10.3109/10409237709102561. [DOI] [PubMed] [Google Scholar]
- Whitlock J. P., Jr, Simpson R. T. Localization of the sites along nucleosome DNA which interact with NH2-terminal histone regions. J Biol Chem. 1977 Sep 25;252(18):6516–6520. [PubMed] [Google Scholar]
- Whitlock J. P., Jr, Simpson R. T. Preparation and physical characterization of a homogeneous population of monomeric nucleosomes from HeLa cells. Nucleic Acids Res. 1976 Sep;3(9):2255–2266. doi: 10.1093/nar/3.9.2255. [DOI] [PMC free article] [PubMed] [Google Scholar]










