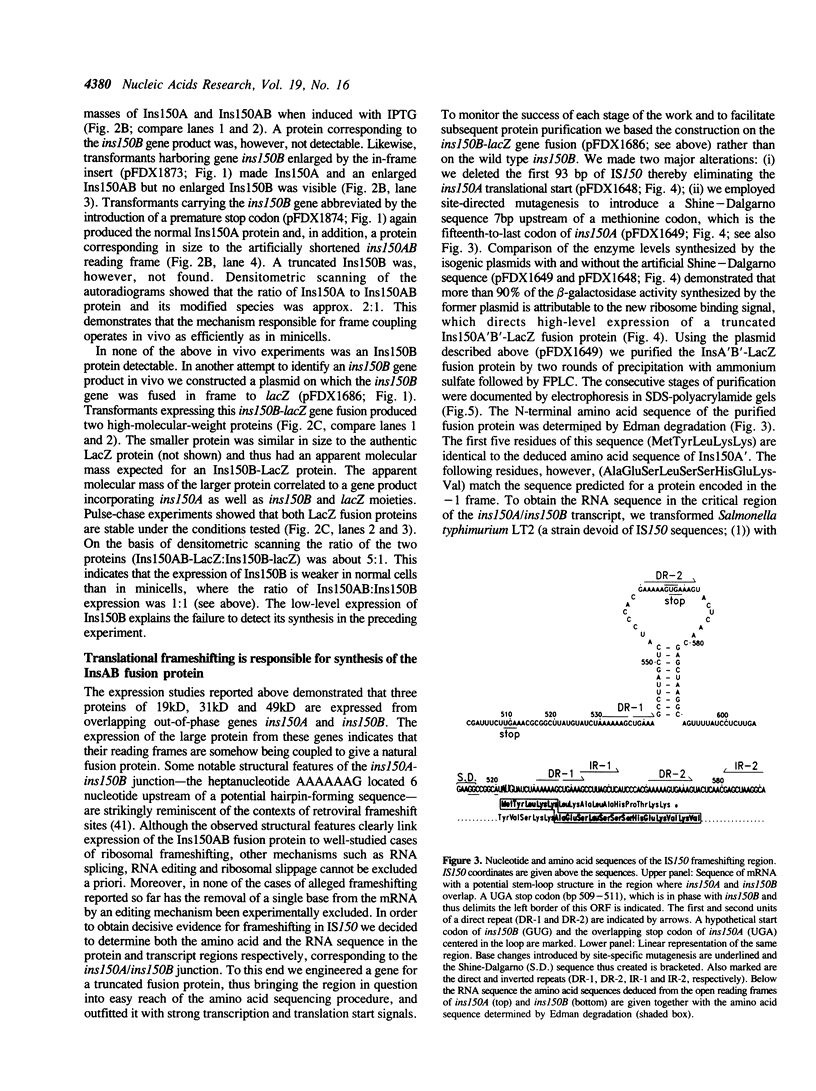
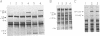Abstract
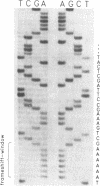
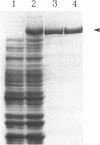
IS150 contains two tandem, out-of-phase, overlapping genes, ins150A and ins150B, which are controlled by the same promoter. These genes encode proteins of 19 and 31 kD, respectively. A third protein of 49 kD is a transframe gene product consisting of domains encoded by both genes. Specific -1 ribosomal frameshifting is responsible for the synthesis of the large protein. Expression of ins150B also involves frameshifting. The IS150 frameshifting signals operate with a remarkably high efficiency, causing about one third of the ribosomes to switch frame. All of the signals required for this process are encoded in a 83-bp segment of the element. The heptanucleotide A AAA AAG and a potential stem-loop-forming sequence mark the frameshifting site. Similar sequence elements are found in -1 frameshifting regions of bacterial and retroviral genes. A mutation within the stem-loop sequence reduces the rate of frameshifting by about 80%. Artificial transposons carrying this mutation transpose at a normal frequency, but form cointegrates at a approximately 100-fold reduced rate.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Amann E., Brosius J. "ATG vectors' for regulated high-level expression of cloned genes in Escherichia coli. Gene. 1985;40(2-3):183–190. doi: 10.1016/0378-1119(85)90041-1. [DOI] [PubMed] [Google Scholar]
- Bachmann B., Lüke W., Hunsmann G. Improvement of PCR amplified DNA sequencing with the aid of detergents. Nucleic Acids Res. 1990 Mar 11;18(5):1309–1309. doi: 10.1093/nar/18.5.1309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blinkowa A. L., Walker J. R. Programmed ribosomal frameshifting generates the Escherichia coli DNA polymerase III gamma subunit from within the tau subunit reading frame. Nucleic Acids Res. 1990 Apr 11;18(7):1725–1729. doi: 10.1093/nar/18.7.1725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F. Construction and characterization of new cloning vehicles. III. Derivatives of plasmid pBR322 carrying unique Eco RI sites for selection of Eco RI generated recombinant DNA molecules. Gene. 1978 Oct;4(2):121–136. doi: 10.1016/0378-1119(78)90025-2. [DOI] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Brierley I., Digard P., Inglis S. C. Characterization of an efficient coronavirus ribosomal frameshifting signal: requirement for an RNA pseudoknot. Cell. 1989 May 19;57(4):537–547. doi: 10.1016/0092-8674(89)90124-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bölker M., Kahmann R. The Escherichia coli regulatory protein OxyR discriminates between methylated and unmethylated states of the phage Mu mom promoter. EMBO J. 1989 Aug;8(8):2403–2410. doi: 10.1002/j.1460-2075.1989.tb08370.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Charlier D., Piette J., Glansdorff N. IS3 can function as a mobile promoter in E. coli. Nucleic Acids Res. 1982 Oct 11;10(19):5935–5948. doi: 10.1093/nar/10.19.5935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craigen W. J., Caskey C. T. Translational frameshifting: where will it stop? Cell. 1987 Jul 3;50(1):1–2. doi: 10.1016/0092-8674(87)90652-0. [DOI] [PubMed] [Google Scholar]
- Escoubas J. M., Prère M. F., Fayet O., Salvignol I., Galas D., Zerbib D., Chandler M. Translational control of transposition activity of the bacterial insertion sequence IS1. EMBO J. 1991 Mar;10(3):705–712. doi: 10.1002/j.1460-2075.1991.tb08000.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fayet O., Ramond P., Polard P., Prère M. F., Chandler M. Functional similarities between retroviruses and the IS3 family of bacterial insertion sequences? Mol Microbiol. 1990 Oct;4(10):1771–1777. doi: 10.1111/j.1365-2958.1990.tb00555.x. [DOI] [PubMed] [Google Scholar]
- Flower A. M., McHenry C. S. The gamma subunit of DNA polymerase III holoenzyme of Escherichia coli is produced by ribosomal frameshifting. Proc Natl Acad Sci U S A. 1990 May;87(10):3713–3717. doi: 10.1073/pnas.87.10.3713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fowler A. V. High-level production of -galactosidase by Escherichia coli merodiploids. J Bacteriol. 1972 Nov;112(2):856–860. doi: 10.1128/jb.112.2.856-860.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hall B. G., Parker L. L., Betts P. W., DuBose R. F., Sawyer S. A., Hartl D. L. IS103, a new insertion element in Escherichia coli: characterization and distribution in natural populations. Genetics. 1989 Mar;121(3):423–431. doi: 10.1093/genetics/121.3.423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harley C. B., Lawrie J., Boyer H. W., Hedgpeth J. Reiterative copying by E.coli RNA polymerase during transcription initiation of mutant pBR322 tet promoters. Nucleic Acids Res. 1990 Feb 11;18(3):547–552. doi: 10.1093/nar/18.3.547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hashimoto-Gotoh T., Franklin F. C., Nordheim A., Timmis K. N. Specific-purpose plasmid cloning vectors. I. Low copy number, temperature-sensitive, mobilization-defective pSC101-derived containment vectors. Gene. 1981 Dec;16(1-3):227–235. doi: 10.1016/0378-1119(81)90079-2. [DOI] [PubMed] [Google Scholar]
- Hatfield D., Oroszlan S. The where, what and how of ribosomal frameshifting in retroviral protein synthesis. Trends Biochem Sci. 1990 May;15(5):186–190. doi: 10.1016/0968-0004(90)90159-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu W. S., Wang R. Y., Liou R. S., Shih J. W., Lo S. C. Identification of an insertion-sequence-like genetic element in the newly recognized human pathogen Mycoplasma incognitus. Gene. 1990 Sep 1;93(1):67–72. doi: 10.1016/0378-1119(90)90137-g. [DOI] [PubMed] [Google Scholar]
- Kearney B., Staskawicz B. J. Characterization of IS476 and its role in bacterial spot disease of tomato and pepper. J Bacteriol. 1990 Jan;172(1):143–148. doi: 10.1128/jb.172.1.143-148.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khan E., Mack J. P., Katz R. A., Kulkosky J., Skalka A. M. Retroviral integrase domains: DNA binding and the recognition of LTR sequences. Nucleic Acids Res. 1991 Feb 25;19(4):851–860. doi: 10.1093/nar/19.4.851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kramer W., Drutsa V., Jansen H. W., Kramer B., Pflugfelder M., Fritz H. J. The gapped duplex DNA approach to oligonucleotide-directed mutation construction. Nucleic Acids Res. 1984 Dec 21;12(24):9441–9456. doi: 10.1093/nar/12.24.9441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkel T. A., Roberts J. D., Zakour R. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lüthi K., Moser M., Ryser J., Weber H. Evidence for a role of translational frameshifting in the expression of transposition activity of the bacterial insertion element IS1. Gene. 1990 Mar 30;88(1):15–20. doi: 10.1016/0378-1119(90)90054-u. [DOI] [PubMed] [Google Scholar]
- Matsutani S., Ohtsubo H., Maeda Y., Ohtsubo E. Isolation and characterization of IS elements repeated in the bacterial chromosome. J Mol Biol. 1987 Aug 5;196(3):445–455. doi: 10.1016/0022-2836(87)90023-4. [DOI] [PubMed] [Google Scholar]
- McAdam R. A., Hermans P. W., van Soolingen D., Zainuddin Z. F., Catty D., van Embden J. D., Dale J. W. Characterization of a Mycobacterium tuberculosis insertion sequence belonging to the IS3 family. Mol Microbiol. 1990 Sep;4(9):1607–1613. doi: 10.1111/j.1365-2958.1990.tb02073.x. [DOI] [PubMed] [Google Scholar]
- Priefer U. B., Kalinowski J., Rüger B., Heumann W., Pühler A. ISR1, a transposable DNA sequence resident in Rhizobium class IV strains, shows structural characteristics of classical insertion elements. Plasmid. 1989 Mar;21(2):120–128. doi: 10.1016/0147-619x(89)90055-3. [DOI] [PubMed] [Google Scholar]
- Prère M. F., Chandler M., Fayet O. Transposition in Shigella dysenteriae: isolation and analysis of IS911, a new member of the IS3 group of insertion sequences. J Bacteriol. 1990 Jul;172(7):4090–4099. doi: 10.1128/jb.172.7.4090-4099.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rak B. A control system which causes alternating turn-off/turn-on of transcription on insertion element IS1. Mol Gen Genet. 1983;192(1-2):61–65. doi: 10.1007/BF00327647. [DOI] [PubMed] [Google Scholar]
- Rak B., von Reutern M. Insertion element IS5 contains a third gene. EMBO J. 1984 Apr;3(4):807–811. doi: 10.1002/j.1460-2075.1984.tb01889.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubens C. E., Heggen L. M., Kuypers J. M. IS861, a group B streptococcal insertion sequence related to IS150 and IS3 of Escherichia coli. J Bacteriol. 1989 Oct;171(10):5531–5535. doi: 10.1128/jb.171.10.5531-5535.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnetz K., Rak B. Regulation of the bgl operon of Escherichia coli by transcriptional antitermination. EMBO J. 1988 Oct;7(10):3271–3277. doi: 10.1002/j.1460-2075.1988.tb03194.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnetz K., Sutrina S. L., Saier M. H., Jr, Rak B. Identification of catalytic residues in the beta-glucoside permease of Escherichia coli by site-specific mutagenesis and demonstration of interdomain cross-reactivity between the beta-glucoside and glucose systems. J Biol Chem. 1990 Aug 15;265(23):13464–13471. [PubMed] [Google Scholar]
- Schwartz E., Herberger C., Rak B. Second-element turn-on of gene expression in an IS1 insertion mutant. Mol Gen Genet. 1988 Feb;211(2):282–289. doi: 10.1007/BF00330605. [DOI] [PubMed] [Google Scholar]
- Schwartz E., Kröger M., Rak B. IS150: distribution, nucleotide sequence and phylogenetic relationships of a new E. coli insertion element. Nucleic Acids Res. 1988 Jul 25;16(14B):6789–6802. doi: 10.1093/nar/16.14.6789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sekine Y., Ohtsubo E. Frameshifting is required for production of the transposase encoded by insertion sequence 1. Proc Natl Acad Sci U S A. 1989 Jun;86(12):4609–4613. doi: 10.1073/pnas.86.12.4609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. A bacteriophage T7 RNA polymerase/promoter system for controlled exclusive expression of specific genes. Proc Natl Acad Sci U S A. 1985 Feb;82(4):1074–1078. doi: 10.1073/pnas.82.4.1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thierry D., Cave M. D., Eisenach K. D., Crawford J. T., Bates J. H., Gicquel B., Guesdon J. L. IS6110, an IS-like element of Mycobacterium tuberculosis complex. Nucleic Acids Res. 1990 Jan 11;18(1):188–188. doi: 10.1093/nar/18.1.188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsuchihashi Z., Kornberg A. Translational frameshifting generates the gamma subunit of DNA polymerase III holoenzyme. Proc Natl Acad Sci U S A. 1990 Apr;87(7):2516–2520. doi: 10.1073/pnas.87.7.2516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Weiner A. M., Maizels N. RNA editing: guided but not templated? Cell. 1990 Jun 15;61(6):917–920. doi: 10.1016/0092-8674(90)90053-h. [DOI] [PubMed] [Google Scholar]
- Weiss R. B., Dunn D. M., Shuh M., Atkins J. F., Gesteland R. F. E. coli ribosomes re-phase on retroviral frameshift signals at rates ranging from 2 to 50 percent. New Biol. 1989 Nov;1(2):159–169. [PubMed] [Google Scholar]