Abstract
Background
Weevils of the genus Otiorhynchus are regarded as devastating pests in a wide variety of horticultural crops worldwide. So far, little is known on the presence of endosymbionts in Otiorhynchus spp.. Investigation of endosymbiosis in this genus may help to understand the evolution of different reproductive strategies in these weevils (parthenogenesis or sexual reproduction), host-symbiont interactions, and may provide a future basis for novel pest management strategy development. Here, we used a multitag 454 pyrosequencing approach to assess the bacterial endosymbiont diversity in larvae of four economically important Otiorhynchus species.
Results
High-throughput tag-encoded FLX amplicon pyrosequencing of a bacterial 16S rDNA fragment was used to characterise bacterial communities associated with different Otiorhynchus spp. larvae. By sequencing a total of ~48,000 PCR amplicons, we identified 49 different operational taxonomic units (OTUs) as bacterial endosymbionts in the four studied Otiorhynchus species. More than 90% of all sequence reads belonged either to the genus Rickettsia or showed homology to the phylogenetic group of “Candidatus Blochmannia” and to endosymbionts of the lice Pedicinus obtusus and P. badii. By using specific primers for the genera Rickettsia and “Candidatus Blochmannia”, we identified a new phylogenetic clade of Rickettsia as well as “Candidatus Nardonella” endosymbionts in Otiorhynchus spp. which are closely related to “Candidatus Blochmannia” bacteria.
Conclusions
Here, we used multitag 454 pyrosequencing for assessment of insect endosymbiotic communities in weevils. As 454 pyrosequencing generates only quite short sequences, results of such studies can be regarded as a first step towards identifying respective endosymbiotic species in insects. In the second step of our study, we analysed sequences of specific gene regions for a more detailed phylogeny of selected endosymbiont genera. As a result we identified the presence of Rickettsia and “Candidatus Nardonella” endosymbionts in Otiorhynchus spp.. This knowledge is an important step in exploring bacteria-insect associations for potential use in insect pest control.
Background
It is estimated that more than 65% of insects are associated with symbiotic bacteria, among them Wolbachia spp. being the most common genus [1,2]. The range of the symbiotic relationships between insect hosts and bacteria varies from being mutualistic and commensal to a pathogenic one [3-5]. Accordingly, intracellular symbionts in insects are usually referred to as primary or secondary endosymbionts (P- and S-symbionts, respectively), with P-symbionts being obligate for the insect e.g. due to providing nutrients, while S-symbionts have a beneficial but not essential role for host insect survival (for reviews see [3] and [6]). In many insects, endosymbionts are located in specialized organs (referred to as bacteriomes or mycetomes) and their inheritance usually follows a strict vertical transmission from mother to offspring.
Understanding relationships between insect hosts and their endosymbiotic bacteria is not only relevant from an evolutionary point of view, but can also aid in the identification of new targets for insect pest control [7] as well as for biotechnology and biomedicine [3]. Yet, since many of the relevant microorganisms cannot be cultured, their identification and functional characterization was so far difficult or not possible at all. Lately, the accessibility of novel genomic techniques, in particular next generation sequencing (NGS) technologies represent new, cost-efficient and fast strategies to depict microbial diversity without the need for culturing the respective organisms [8]. With these techniques thousands of sequence reads can be analysed in parallel allowing an extensive assessment of bacterial diversity within insects. As a target for bacterial NGS projects, ribosomal DNA genes (rDNA) like the 16S rDNA, also used for the taxonomic classification of bacterial species [9], have frequently been applied for analysing the bacterial microbial community in metagenomic studies of soil [10,11], mines [12], the deep sea [13] or oral human microflora [14].
In this study, we used high-throughput tag-encoded FLX amplicon pyrosequencing [15] to characterise bacterial communities associated with four different weevil species of the genus Otiorhynchus Germar (Coleoptera: Curculionidae). Members of this genus are polyphagous and are regarded as pests of a variety of ornamental and nursery plants worldwide. Their soilborne larvae feed on the host plants’ roots which may be lethal in particular for younger plants or recently transplanted cuttings. Further, feeding damage of adults on the plants foliage may reduce the market value of ornamentals. For these reasons weevils are often controlled by intensive insecticide applications [16]. Moreover, Otiorhynchus spp. can serve as a model genus for understanding the evolution of asexual reproduction, since it includes species both reproducing mostly parthenogenetically (like O. sulcatus and O. rugosostriatus) as well as sexually (like O. salicicola and O. armadillo) [17,18].
Here, by applying 454 sequencing technology, we show that weevils of the genus Otiorhynchus are associated with several endosymbiotic bacteria. This study is the first to report Rickettsia and “Candidatus Nardonella” endosymbionts - the ancestral endosymbiont of weevils - in Otiorhynchus spp.. Identifying endosymbionts in the genus Otiorhynchus can expand to our understanding of the evolution of both endosymbiont-host insect interactions as well as of different reproductive strategies of insects and may provide a future basis for novel pest management approaches.
Results and discussion
454 pyrosequencing and identification of endosymbionts in Otiorhynchus spp
A total of ~48,000 PCR amplicons were sequenced via GS FLX titanium 454 sequencing, of which ~27,000 reads were assembled after having passed the additional quality controls. These sequences were summarized into 49 consensus sequences (Table 1), representing the total retrieved endosymbiotic bacterial diversity in the four different Otiorhynchus species. Sequence abundances of the respective OTUs were different in each weevil species analysed. We expect these differences in sequence abundance within the 16S rDNA amplicons to reflect the respective bacterial abundances in the sample.
Table 1.
Endosymbiotic bacterial diversity and abundance in the four analysed Otiorhynchus species.
| Bacteria from weevil species | GenBank accession No. | Number of reads | % of total reads | Closest phylogenetic match and 16S rDNA accession number | Class |
|---|---|---|---|---|---|
| O. salicicola (in total 6073 reads) | JN563736 | 5516 | 90.83 | AB478978, endosymbiont of Pedicinus obtusus and AJ245596 endosymbiont of Camponotus balzanii (referred to as “Candidatus Blochmanni” endosymbionts throughout the text) | γ-Proteobacteria |
| JN563737 | 121 | 1.99 | DQ417336, Schlegelella aquatica | β-Proteobacteria | |
| JN563738 | 96 | 1.58 | FJ268988, uncultured Acinetobacter | γ-Proteobacteria | |
| JN563739 | 69 | 1.14 | CU927677, uncultured bacterium | - | |
| JN563740 | 48 | 0.79 | FJ534956, uncultured bacterium | - | |
| JN563741 | 44 | 0.72 | EF210100, Enterobacter hormaechei | γ-Proteobacteria | |
| JN563742 | 34 | 0.56 | AY923125, Streptococcus sp. | Bacilli | |
| JN563743 | 26 | 0.43 | EU464962, uncultured bacterium | - | |
| JN563744 | 25 | 0.41 | EU766013, uncultured bacterium | - | |
| JN563745 | 23 | 0.38 | FJ393126, uncultured Bacteroides sp. | Bacteroidetes | |
| JN563746 | 18 | 0.30 | EU721814, uncultured epsilon proteobacterium | ε-Proteobacteria | |
| JN563747 | 17 | 0.28 | AY953252, Prevotella sp. | Bacteroidetes | |
| JN563748 | 15 | 0.25 | FJ799146, bacterium enrichment culture clone LA29 | - | |
| JN563749 | 11 | 0.18 | EU802152, uncultured bacterium | - | |
| JN563750 | 10 | 0.16 | AY568512, Burkholderia fungorum | β-Proteobacteria | |
| O. rugosostriatus (in total 8584 reads) | JN563751 | 7800 | 90.87 | AB021128, Rickettsia sp. | α-Proteobacteria |
| JN563752 | 396 | 4.61 | EF633744, Candidatus Neoehrlichia lotoris | α-Proteobacteria | |
| JN563753 | 338 | 3.94 | AB478978, endosymbiont of Pedicinus obtusus and AJ245596 endosymbiont of Camponotus balzanii (referred to as “Candidatus Blochmanni” endosymbionts throughout the text) | γ-Proteobacteria | |
| JN563754 | 17 | 0.20 | AB021128, Rickettsia sp. | α-Proteobacteria | |
| JN563755 | 11 | 0.13 | EF633744, Candidatus Neoehrlichia lotoris | α-Proteobacteria | |
| JN563756 | 7 | 0.08 | AB021128, Rickettsia sp. | α-Proteobacteria | |
| JN563757 | 6 | 0.07 | AB021128, Rickettsia sp. | α-Proteobacteria | |
| JN563758 | 5 | 0.06 | FJ868862, uncultured bacterium | - | |
| JN563759 | 4 | 0.05 | GQ845011, Nevskia sp. | γ-Proteobacteria | |
| O. sulcatus (in total 6412 reads) | JN563760 | 6358 | 99.16 | AB021128, Rickettsia sp. | α-Proteobacteria |
| JN563761 | 35 | 0.55 | EF633744, Candidatus Neoehrlichia lotoris | α-Proteobacteria | |
| JN563762 | 19 | 0.30 | EF633744, Candidatus Neoehrlichia lotoris | α-Proteobacteria | |
| O. armadillo (in total 6311 reads) | JN563763 | 5900 | 93.49 | AB478978, endosymbiont of Pedicinus obtusus and AJ245596 endosymbiont of Camponotus balzanii (referred to as “Candidatus Blochmanni” endosymbionts throughout the text) | γ-Proteobacteria |
| JN563764 | 60 | 0.95 | FJ823944, uncultured Comamonas sp. | β-Proteobacteria | |
| JN563765 | 54 | 0.86 | FJ868862, uncultured bacterium | - | |
| JN563766 | 43 | 0.68 | FJ823944, uncultured Comamonas sp. | β-Proteobacteria | |
| JN563767 | 35 | 0.55 | FJ544375, Comamonas aquatica | β-Proteobacteria | |
| JN563768 | 31 | 0.49 | EU560802, uncultured bacterium | - | |
| JN563769 | 23 | 0.36 | DQ407746, primary endosymbiont of Liposcelis decolor | - | |
| JN563770 | 21 | 0.33 | DQ469223, uncultured bacterium | - | |
| JN563771 | 21 | 0.33 | GQ845011, Nevskia sp. | γ-Proteobacteria | |
| JN563772 | 20 | 0.32 | DQ860049, uncultured bacterium | - | |
| JN563773 | 11 | 0.17 | AF006670, Shewanella putrefaciens | γ-Proteobacteria | |
| JN563774 | 11 | 0.17 | X82133, Shewanella putrefaciens | γ-Proteobacteria | |
| JN563775 | 11 | 0.17 | EU801479, uncultured bacterium | - | |
| JN563776 | 10 | 0.16 | EF019306, uncultured proteobacterium | - | |
| JN563777 | 9 | 0.14 | AY953252, Prevotella sp. | Bacteroidetes | |
| JN563778 | 8 | 0.13 | EU464962, uncultured bacterium | - | |
| JN563779 | 8 | 0.13 | EU536078, uncultured bacterium | - | |
| JN563780 | 8 | 0.13 | GQ068015, uncultured bacterium | - | |
| JN563781 | 8 | 0.13 | L16490, Porphyromonas asaccharolytica | Bacteroidetes | |
| JN563782 | 8 | 0.13 | AY351787, uncultured marine bacterium | - | |
| JN563783 | 6 | 0.10 | EF648074, uncultured Azoarcus sp., | β-Proteobacteria | |
| JN563784 | 5 | 0.08 | EF648074, uncultured Azoarcus sp., | β-Proteobacteria | |
Only the closest relatives and their 16S rDNA accession numbers (see additional file 1: 16S rDNA gene-based phylogeny of endosymbionts in four different Otiorhynchus spp. larvae) are mentioned.
In addition to the most abundant reads, which belonged either to the genus Rickettsia or were similar to “Candidatus Blochmannia” bacteria and endosymbionts of the lice Pedicinus obtusus and P. badii, numerous reads with low sequence frequency were detected (Table 1). Indeed, we can not fully exclude the possibility that these sequences of putative rare endosymbionts are rather artefacts e.g. due to PCR contaminations.
Phylogenetic analysis of Otiorhynchus spp. endosymbionts
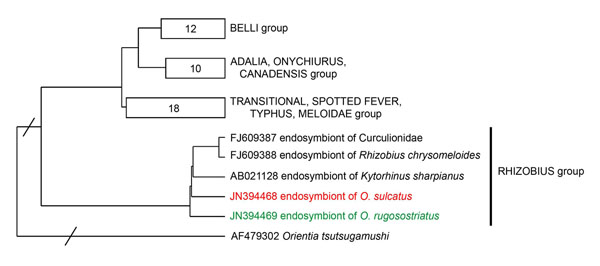
Phylogenetic analysis of 454 sequence data was performed to establish the relationship of the partial 16S rDNA sequences to each other and to related sequences gained from public databases. Among all studied weevil species, O. sulcatus showed the lowest bacterial endosymbiotic diversity (Table 1). The vast majority of sequences in O. sulcatus (~99% of the total reads) and O. rugosostriatus (~91% of the total reads) belonged to the genus Rickettsia (see additional file 1: 16S rDNA gene-based phylogeny of endosymbionts in four different Otiorhynchus spp. larvae, Table 1). Bacteria similar to the endosymbionts of the lice Pedicinus obtusus and P. badii [19,20] and the genus “Candidatus Blochmannia” were dominant in O. salicicola (~91% of the total reads) and O. armadillo (~93% of the total reads) (see additional file 1: 16S rDNA gene-based phylogeny of endosymbionts in four different Otiorhynchus spp. larvae, Table 1). These bacteria were also found in a less dominant manner in O. rugosostriatus (~4% of the total reads). To determine the phylogenetic position of Rickettsia and putative “Candidatus Blochmannia” like endosymbionts detected via 454 pyrosequencing in a more precise way, genus specific primers [21,22] were used to amplify a ~750 bp fragment of the Rickettsia and “Candidatus Blochmannia” specific 16S rDNA and a ~800 bp fragment of the Rickettsia cytochome C subunit I (coxA) gene, respectively. Phylogenetic analysis of these sequences placed the Otiorhynchus spp. specific Rickettsia into a new clade within the genus Rickettsia (Figure 1 and 2). Sequences gained by using “Candidatus Blochmannia” specific primers were grouped within the clade of “Candidatus Nardonella” bacteria, which are closely related to “Candidatus Blochmannia” endosymbionts (Figure 3). Accordingly, the additional analysis of these endosymbionts using gene specific primers revealed for the first time the presence of Rickettsia and “Candidatus Nardonella” bacteria within the genus Otiorhynchus spp..
Figure 1.
Neighbour joining tree of Rickettsia endosymbionts using sequences of 16S rDNA. Sequences obtained in the present study are coloured and phylogenetic groups were constructed according to Weinert et al [22]. The amount of sequences included in the groups are indicated by numbers. Branch lengths were reduced in two positions (marked with diagonal slashes).
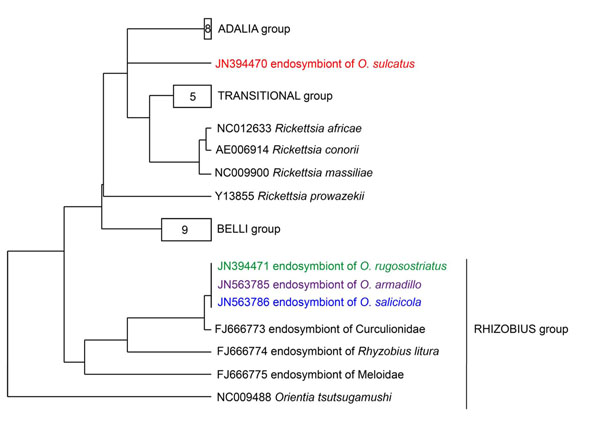
Figure 2.
Neighbour joining tree of Rickettsia endosymbionts using sequences of coxA gene. Sequences obtained in the present study are coloured. Sequences were combined in groups according to Weinert et al [22]. The amount of sequences included in the groups are indicated by numbers.
Figure 3.
Neighbour joining tree of “Candidatus Nardonella” endosymbionts using sequences of 16S rDNA. Sequences obtained in the present study are coloured. Branch lengths were reduced in four positions (marked with diagonal slashes). The amount of sequences included in the groups are indicated by numbers.
Phylogenetic analysis and putative biological function of Rickettsia endosymbionts
In the parthenogenetically reproducing species O. sulcatus and O. rugosostriatus, Rickettsia endosymbionts were the most dominant group found via 454 pyrosequencing. By using Rickettsia specific primers for the 16S rDNA and the coxA gene these results were strengthened, however, a fragment of the Rickettsia specific coxA gene was also amplified in O. armadillo and O. salicicola, respectively, while 454 pyrosequencing previously indicated that these two species were missing Rickettsia endosymbionts (Table 1). Yet, at the same time it was not possible to amplify the Rickettsia specific 16S rDNA fragment in the same two species. We thus suppose that the coxA gene sequence is rather conserved among bacteria and may not be adequate for precise species determination. Supplementary sequence analysis of a range of additional bacterial genes may resolve this issue.
Phylogenetic analysis of the Rickettsia endosymbiontic 16S rDNA and coxA gene fragments amplified from Otiorhynchus spp. revealed the relatedness to the rhizobius and/or adalia Rickettsia group as defined by Weinert et al [22]. These subgroups contain Rickettsia bacteria identified in various beetles, including members of the Curculionidae [22]. Rickettsia endosymbionts act as male-killing agents in leaf mining beetles and ladybirds [23,24] and play an essential role in the early development of the oocyte and egg production in parthenogenetic book lice [25,26]. Thus it could be speculated that Rickettsia endosymbionts may also manipulate host reproduction in Otiorhynchus species.
Phylogenetic analysis and putative biological function of “Candidatus Nardonella” endosymbionts
454 pyrosequencing detected endosymbionts similar to “Candidatus Blochmannia” and bacterial endosymbionts of the lice Pedicinus obtusus and P. badii in O. armadillo, O. salicicola and to a lesser extent in O. rugosostriatus. The presence of these putative “Candidatus Blochmannia” like bacteria was verified in these species by using primers specific for the “Candidatus Blochmannia” 16S rDNA [21], which indicated that the obtained sequences are similar to “Candidatus Nardonella”. In addition, a fragment of the same size and sequence was also amplified in O. sulcatus, even though 454 pyrosequencing did not reveal the presence of these bacteria in this weevil species (Table 1). “Candidatus Nardonella” bacteria are often localized in the bacteriome whereas Rickettsia endosymbionts may infect as well different tissues. As we used whole larvae for DNA extraction, the amount of overall isolated DNA might have been lower for “Candidatus Nardonella” than for Rickettsia. Therefore we assume that respective bacterial DNA might have not been amplified in O. sulcatus with the universal primers used for 454 pyrosequencing due to competition for PCR reagents with taxa such as Rickettsia, having a higher template abundance [27]. However, these results also demonstrate that studies using 454 pyrosequencing can be regarded as a first step towards identifying respective endosymbiotic species in insects, but that for a detailed phylogeny and a more comprehensive insight into endosymbiont-insect-associations, the amplification of specific gene regions is still indispensable.
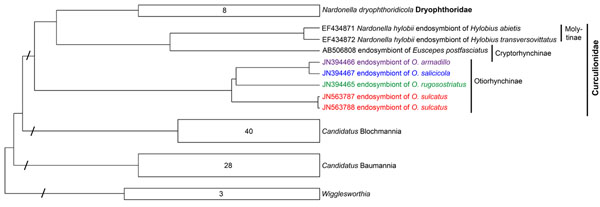
Phylogenetic analysis of the putative “Candidatus Blochmannia” specific 16S rDNA sequence amplified from the four studied Otiorhynchus weevils showed a close relatedness of these bacteria to the genus “Candidatus Nardonella”. Sequences generated in the present study build a separate branch next to endosymbionts from molytine, cryptorhynchine and dryophthorid weevils [28-30] (Figure 3). The biological function of “Candidatus Nardonella” endosymbionts in their host weevils is unknown so far, except for the cryptorhynchine West Indian sweet potato weevil, Euscepes postfasciatus. Within this species “Candidatus Nardonella” endosymbionts are involved in growth and development of the host weevil [31].
Implications and future directions of endosymbiosis in Otiorhynchus spp
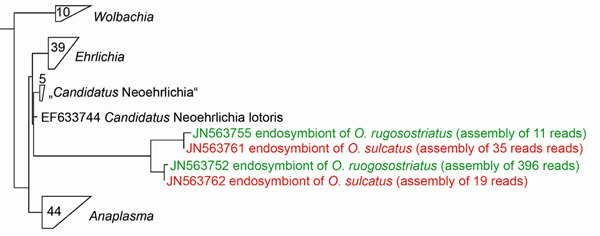
For several Otiorhynchus species, an association with bacteria of the genus Wolbachia has been proven in previous studies [32-34]. Wolbachia cause several reproductive alterations in insects, including cytoplasmic incompatibility, feminization of genetic males or parthenogenesis [35]. In Otiorhynchus species Wolbachia are assumed to rather play a role in normal development of e.g. O. sulcatus eggs [34] rather than in the evolution of parthenogenesis or polyploidy [32,33,36]. Unexpectedly, in the present 454 pyrosequencing approach, none of the bacterial sequence reads obtained from four different Otiorhynchus spp. weevil larvae corresponded to Wolbachia. Instead, bacterial sequences similar to “Candidatus Neoehrlichia”, a close relative to Wolbachia, were found in however low frequencies in O. sulcatus (~1% of the total reads) and O. rugosostriatus (~5% of the total reads) (Table 1, Figure 4). Species of that genus are known as tick-borne bacterial pathogens [37] and have been isolated from raccoons and rats [38,39] but their biological function in insects is unclear so far. As the presence of different Wolbachia strains may differ within a given species between geographical regions [40] further studies are required using Wolbachia specific PCR primers to shed light on the prevalence and distribution of Wolbachia within Otiorhynchus species and between populations, respectively.
Figure 4.
Phylogenetic analysis of endosymbionts under “Candidatus Neoehrlichia” subregion in Otiorhynchus spp. The tree represents the “Candidatus Neoehrlichia” subregion of the complete tree (see additional file 1: 16S rDNA gene-based phylogeny of endosymbionts in four different Otiorhynchus spp. larvae) and was constructed by using parsimony algorithm. Sequences obtained in the present study are coloured. The amount of sequences included in the groups of Wolbachia, Ehrlichia, „Candidatus Neoehrlichia” and Anaplasma are indicated by numbers.
Recent microbiological characterization of bacterial endosymbionts in the Curculionoidea of the family Molytinae and Dryophthoridae has demonstrated that endosymbiosis with “Candidatus Nardonella” bacteria is ~125 Myr old in curculionids and is most of the times evolutionary stable, except for a few clades where respective endosymbionts have been lost and were replaced by different microbes during evolution (endosymbiont replacement; [29]). Our study broadens the range of weevils associated with “Candidatus Nardonella” endosymbionts and indicates a benefit for Otiorhynchus weevils due to the long-lasting bacterial inheritance.
In a number of weevil species it has been shown that endosymbionts are frequently found within specialized host cells (so-called bacteriocytes) sometimes forming a distinctive organ, the bacteriome, which is often associated with the larval midgut [29,30,41-43]. As Buchner [44] has described a bacteriome in Otiorhynchus spp., we assume that the four Otiorhynchus species analysed in the present study also harbour their endosymbiotic bacteria intracellularly in a bacteriome. However, this assumption has to be confirmed via microscopic examinations of the respective organs.
For a couple of insects and their associated microorganisms it has been shown, that endosymbiotic bacteria are known to be involved in protecting their host insect against natural antagonists such as predators and pathogens or are even implicated in insecticide resistance mechanisms (for a review see Zindel et al [45]). Moreover, particularly obligatory endosymbionts are essential for central functions of their host insect [3]. Accordingly, endosymbiotic bacteria are an interesting target for direct or indirect manipulation, thus offering new possibilities for designing insect control strategies [45-47]. Identification of respective endosymbiotic organisms of the target insect is an important step in exploring these associations for potential use in insect pest control. Thanks to the agar-based artificial diet for rearing of O. sulcatus [48], physiological, nutritional and reproductive studies will be carried out to analyse the respective effects of symbionts on the host development and reproduction.
Conclusions
In this study, endosymbiotic bacterial diversity in weevil larvae was assessed via multitag 454 pyrosequencing of a bacterial 16S rRNA fragment. Pyrosequencing is therefore a promising, fast and economic alternative to other culture-independent methods in metagenomics like DGGE (Denaturing Gradient Gel Electrophoresis) or SSCP (Single Strand Conformation Polymorphism), which have been used in bacterial community studies of the red turpentine beetle [49] or for diversity assessment of gut microbiota in bees [50], respectively. However, as 454 pyrosequencing generates only quite short sequences, results of such studies can just be regarded as a first step towards identifying respective endosymbiotic species in insects. Accordingly, a subsequent analysis of sequences of specific gene regions of selected endosymbiont genera detected via 454 pyrosequencing revealed the presence of endosymbionts of the genera Rickettsia and “Candidatus Nardonella” in Otiorhynchus spp.. Further studies are now required to clarify the biological function of these endosymbiotic bacteria in Otiorhynchus spp. and their potential as novel targets for weevil pest control.
Materials and methods
Insect rearing, bacterial DNA extraction and species determination of larvae
All experiments were performed with four different Otiorhynchus species (O. sulcatus, O. rugosostriatus, O. salicicola and O. armadillo) collected in the field and kept in the laboratory until egg deposition. During that period of time weevils were fed with leaves of Prunus sp., Potentilla sp. or Fragaria sp.. Freshly laid weevil eggs (at most 10 days old) were collected and surface sterilized according to the method developed by Hosokawa et al [51]. The eggs were air dried under the clean bench and transferred individually with sterile featherweight forceps in Petri dishes filled with sterile TSA (40,0 g/l DifcoTM Tryptic Soy Agar, pH 7.3 ± 0.2; Voigt Global Distribution Inc, Lawrence, Kansas). In order to enlarge the contact of egg and TSA agar and to check the success of surface sterilisation, eggs were rolled several times over the agar plate. For further analysis only eggs with no bacterial growth on TSA were included. Eggs were kept usually at 21-24°C until eclosion. Freshly emerged larvae (approximately 24-72 hours old) without egg material were individually collected from the TSA agar plates, and were stored frozen at -80°C until further processing. Total metagenomic DNA (~20-40 ng/µl DNA per larva) was extracted from the complete larvae using the MasterPureTM DNA Purification Kit (Epicentre® Biotechnologies, Madison, Wisconsin). Taxonomic identity of each larva was confirmed according to a diagnostic PCR-RFLP pattern of the COII region [52]. For metagenomic analysis seven individuals of each Otiorhynchus species were included.
Bacterial 16S rDNA PCR amplification and 454 pyrosequencing
Universal bacteria primers (fwd: 5’-MGAGTTTGATCCTGGCTCAG-3’ and rev: 5’-GCTGCCTCCCGTAGGAGT-3’; [53]), amplifying an approximately 450 bp fragment of the 16S rDNA, were used in the present study. These primers are covering the V1-V2 regions of the 16S rDNA gene and showed good phylogenetic resolution from phylum to family level in a recent study by Hamp et al [53]. Primers were modified by the addition of a GS FLX Titanium Key-Primer A and B (A: CGTATCGCCTCCCTCGCGCCA and B: CTATGCGCCTTGCCAGCCCGC), a four-base library “key” sequence (TCAG) and a multiplex identifier (MID) sequence specific to each Otiorhynchus species. The MID sequences (forward/reverse) were as follows for the respective weevil species: O. salicicola (ATCGCG / CGCGAT), O. rugosostriatus (ATAGCC / GGCTAT), O. sulcatus (CCATAG / CTATGG) and O. armadillo (CTTGAG / CTCAAG). PCR reaction mixture consisted of 0.1 µl of Phire® Hot Start II DNA Polymerase (Finnzymes Oy, Espoo, Finland), 0,2 mM dNTPs (Metabion, Martinsried, Germany), 10 pmol primers and 40-80 ng of DNA template in a final volume of 20 µl. The PCR parameters (C1000TM Thermal Cycler, Bio-Rad Laboratories GmbH, München, Germany) were 95°C for 3 min followed by 35 cycles of 93°C for 60 s, 50°C for 60 s and 72°C for 70 s. A final extension step at 72°C for 5 min was added. An aliquot of 4 µl of each PCR product was checked for correct size (~450 bp) on a 1% agarose gel and was afterwards purified with HiYield PCR Clean-up/Gel Extraction Kit (Süd-Laborbedarf GmbH, Gauting, Germany). Bacterial 16S rDNA PCR products generated from all 28 Otiorhynchus individuals were mixed at equal molar concentrations according to species, and next generation 454 pyrosequencing was performed commercially (LGC Genomics GmbH, Berlin, Germany). The GenBank accession numbers for sequences obtained via 454 pyrosequencing are listed in Table 1.
Sample assignment and analysis of 454 sequencing data
Sequence reads were assembled independently by Geneious Pro Version 5.0 [54] and WiMSeEx (Window Match Seed Extension)-Algorithm (unpublished). Results of both procedures for diversity and sequence identity were compared. Only high quality reads that did accurately match the four-base library “key” sequence (TCAG) and the multiplex identifier (MID) sequence were used for Geneious Pro assembly. Geneious Pro assembly was performed with medium sensitivity, a maximum of 120 contigs and default settings. Consensus sequences were extracted manually from all contigs. WiMSeEx assembly was performed for each tag with all raw data reads and the following parameters: minimum seed size: 200 bp, window size: 60 bp. The four-base identifier and 20 bp of the primer were chosen for seed detection. Each assembly run was stopped by reaching 500 kb sequence data. Resulting sequences of both procedures were then aligned independently using MAFFT version 5 [55] and consensus sequences were extracted manually from clustered sequences and redundant sequence data were removed. Afterwards the sequence identifier and the primer sequence were eliminated from each consensus sequence. All consensus sequences extracted from Geneious Pro contigs were found in the WiMSeEx consensus sequences assembly data and vice versa.
Amplification of selected genes of most dominant endosymbionts
For accurate phylogenetic analysis of the most dominant endosymbionts in Otiorhynchus spp., specific 16S rDNA and cytochrome C oxidase subunit I (coxA) primers for the genus Rickettsia [22] as well as 16S rDNA primers for “Candidatus Blochmannia” bacteria [21] were used for amplification of the respective sequences from 2-4 Otiorhynchus individuals per species. PCR reactions were set up in a final volume of 20 µl consisting of 0.1 µl of Phire® Hot Start II DNA Polymerase (Finnzymes Oy, Espoo, Finland), 0.25 mM dNTPs (Fermentas GmbH, St. Leon-Rot, Germany), 10 pmol primers and 40-80 ng of DNA template. The PCR parameters (C1000TM Thermal Cycler, Bio-Rad Laboratories GmbH, München, Germany) were 95°C for 2 min followed by 40 cycles of 95°C for 30 s, 55°C for 30 s and 72°C for 1 min. A final extension step at 72°C for 10 min was added. An aliquot of 4 µl of each PCR product was checked for correct size on a 1% agarose gel and was afterwards purified with HiYield PCR Clean-up/Gel Extraction Kit (Süd-Laborbedarf GmbH, Gauting, Germany). Direct sequencing of the resulting PCR product was performed commercially (LGC Genomics GmbH, Berlin, Germany). As we did not detect any bacterial sequence variation within one weevil species (except for O. sulcatus and the 16S rDNA amplified with “Candidatus Blochmannia” specific primers), only one sequence per Otiorhynchus species and gene region was submitted to GenBank (accession numbers JN394465-JN394471, JN563785-JN563788).
Phylogenetic analysis
Consensus sequences gained from 454 pyrosequencing were included into an alignment of more than 260,000 (SSURef_102_SILVA_NR_99_18_02_10_opt.ARF) bacterial 16S rDNA sequences [56] and best positions in the resulting phylogenetic tree were found including all nucleotides (positions) from the 454 assemblies using the Parsimony algorithm of the ARB 5.1 software package [57]. The here presented trees are subregions of the complete tree (see additional file 1: 16S rDNA gene-based phylogeny of endosymbionts in four different Otiorhynchus spp. larvae) including the sequences assembled from the 454 sequencing approach reported in this paper and the most similar sequences available from public databases. More distantly related or unrelated sequences were included in the calculation but are not shown.
Additional 16S rDNA sequences amplified with specific primers for “Candidatus Blochmannia” and Rickettsia endosymbionts were included in the above mentioned alignment and a Neighbour joining analysis was inferred using the Neighbour joining algorithm included in the software package ARB 5.1 like described above. In addition, sequences of part of the coxA gene amplified in Otiorhynchus spp. were included in an alignment of sequences used by Weinert et al [22] and a Neighbour joining tree was calculated accordingly.
Authors’ contributions
JH and AR conceived the study design; JH performed sample collection and template preparation for pyrosequencing analysis; JH, SS, and MP performed phylogenetic analysis, and all authors contributed to the writing of the manuscript.
Competing interests
The authors declare that they have no competing interests.
Supplementary Material
16S rDNA gene-based phylogeny of endosymbionts in four different Otiorhynchus spp. larvae. Sequences obtained in the present study are coloured and accession numbers of 16S rDNA sequences are shown for related bacterial species. More distantly related or unrelated sequences are not shown. Sequences from this work were added using the parsimony algorithm. This tree results from a phylogenetic calculation including more than 26,0000 bacterial 16S rDNA sequences. Only the nearest relatives are shown in this tree.
Contributor Information
Jacqueline Hirsch, Email: hirsch@fa-gm.de.
Stephan Strohmeier, Email: strohmeier@freshx.de.
Martin Pfannkuchen, Email: pfannkuchen@cim.irb.hr.
Annette Reineke, Email: reineke@fa-gm.de.
Acknowledgements
We are grateful to the Federal Ministry of Food, Agriculture and Consumer Protection, Germany for providing financial support. We thank Gerlinde Michaelis, Diana Schneider and Peter Sprick for supplying us with Otiorhynchus spp. eggs and larvae for pretests. The authors thank two anonymous reviewers for their helpful comments on an earlier version of the manuscript.
This article has been published as part of BMC Microbiology Volume 11 Supplement 1, 2012: Arthropod symbioses: from fundamental studies to pest and disease mangement. The full contents of the supplement are available online at http://www.biomedcentral.com/1471-2180/12?issue=S1.
References
- Duron O, Bouchon D, Boutin S, Bellamy L, Zhou L, Engelstädter J, Hurst GD. The diversity of reproductive parasites among arthropods: Wolbachia do not walk alone. BMC Biol. 2008;6:27. doi: 10.1186/1741-7007-6-27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hilgenboecker K, Hammerstein P, Schlattmann P, Telschow A, Werren JH. How many species are infected with Wolbachia?--A statistical analysis of current data. FEMS Microbiol Lett. 2008;281(2):215–220. doi: 10.1111/j.1574-6968.2008.01110.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moya A, Pereto J, Gil R, Latorre A. Learning how to live together: genomic insights into prokaryote-animal symbioses. Nature Rev Genet. 2008;9(3):218–229. doi: 10.1038/nrg2319. [DOI] [PubMed] [Google Scholar]
- Kikuchi Y. Endosymbiotic bacteria in insects: their diversity and culturability. Microbes Environ. 2009;24(3):195–204. doi: 10.1264/jsme2.ME09140S. [DOI] [PubMed] [Google Scholar]
- Gil R, Latorre A, Moya A. Bacterial endosymbionts of insects: insights from comparative genomics. Environ Microbiol. 2004;6(11):1109–1122. doi: 10.1111/j.1462-2920.2004.00691.x. [DOI] [PubMed] [Google Scholar]
- Moran NA, McCutcheon JP, Nakabachi A. Genomics and evolution of heritable bacterial symbionts. Annu Rev Genet. 2008;42:165–190. doi: 10.1146/annurev.genet.41.110306.130119. [DOI] [PubMed] [Google Scholar]
- Douglas AE. Symbiotic microorganisms: untapped resources for insect pest control. Trends Biotechnol. 2007;25(8):338–342. doi: 10.1016/j.tibtech.2007.06.003. [DOI] [PubMed] [Google Scholar]
- Harkins T, Jarvie T. Metagenomics analysis using the Genome Sequencer™ FLX system. Nature Methods. 2007;4:6. [Google Scholar]
- Head IM, Saunders JR, Pickup RW. Microbial evolution, diversity, and ecology: A decade of ribosomal RNA analysis of uncultivated microorganisms. Microb Ecol. 1998;35(1):1–21. doi: 10.1007/s002489900056. [DOI] [PubMed] [Google Scholar]
- Acosta-Martinez V, Dowd S, Sun Y, Allen V. Tag-encoded pyrosequencing analysis of bacterial diversity in a single soil type as affected by management and land use. Soil Biol Biochem. 2008;40(11):2762–2770. doi: 10.1016/j.soilbio.2008.07.022. [DOI] [Google Scholar]
- Teixeira L, Peixoto RS, Cury JC, Sul WJ, Pellizari VH, Tiedje J, Rosado AS. Bacterial diversity in rhizosphere soil from Antarctic vascular plants of Admiralty Bay, maritime Antarctica. ISME J. 2010;4(8):989–1001. doi: 10.1038/ismej.2010.35. [DOI] [PubMed] [Google Scholar]
- Edwards RA, Rodriguez-Brito B, Wegley L, Haynes M, Breitbart M, Peterson DM, Saar MO, Alexander S, Alexander EC, Rohwer F. Using pyrosequencing to shed light on deep mine microbial ecology. BMC Genomics. 2006;7:57. doi: 10.1186/1471-2164-7-57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sogin ML, Morrison HG, Huber JA, Mark Welch D, Huse SM, Neal PR, Arrieta JM, Herndl GJ. Microbial diversity in the deep sea and the underexplored "rare biosphere". Proc Natl Acad Sci USA. 2006;103(32):12115–12120. doi: 10.1073/pnas.0605127103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keijser BJF, Zaura E, Huse SM, van der Vossen J, Schuren FHJ, Montijn RC, ten Cate JM, Crielaard W. Pyrosequencing analysis of the oral microflora of healthy adults. J Dent Res. 2008;87(11):1016–1020. doi: 10.1177/154405910808701104. [DOI] [PubMed] [Google Scholar]
- Meyer M, Stenzel U, Hofreiter M. Parallel tagged sequencing on the 454 platform. Nat Protoc. 2008;3(2):267–278. doi: 10.1038/nprot.2007.520. [DOI] [PubMed] [Google Scholar]
- Moorhouse ER, Charnley AK, Gillespie AT. A review of the biology and control of the vine weevil, Otiorhynchus sulcatus (Coleoptera: Curculionidae) Ann Appl Biol. 1992;121(2):431–454. doi: 10.1111/j.1744-7348.1992.tb03455.x. [DOI] [Google Scholar]
- Suomalainen E. Zur Zytologie der parthenogenetischen Curculioniden der Schweiz. Chromosoma. 1954;6:627–655. [PubMed] [Google Scholar]
- Magnano L, Heijerman T, Germann C. On the species status of Otiorhynchus armadillo (Rossi, 1792) and Otiorhynchus salicicola Heyden, 1908 (Coleoptera, Curculionidae, Entimini) Mitt Schweiz Entomol Ges. 2008;81:155–163. [Google Scholar]
- Allen JM, Light JE, Perotti MA, Braig HR, Reed DL. Mutational meltdown in primary endosymbionts: selection limits Muller's ratchet. PLoS One. 2009;4(3):e4969. doi: 10.1371/journal.pone.0004969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukatsu T, Hosokawa T, Koga R, Nikoh N, Kato T, Hayama S, Takefushi H, Tanaka I. Intestinal endocellular symbiotic bacterium of the macaque louse Pedicinus obtusus: distinct endosymbiont origins in anthropoid primate lice and the old world monkey louse. Appl Environ Microbiol. 2009;75(11):3796–3799. doi: 10.1128/AEM.00226-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wernegreen JJ, Kauppinen SN, Brady SG, Ward PS. One nutritional symbiosis begat another: phylogenetic evidence that the ant tribe Camponotini acquired Blochmannia by tending sap-feeding insects. BMC Evol Biol. 2009;9:292. doi: 10.1186/1471-2148-9-292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinert LA, Werren JH, Aebi A, Stone GN, Jiggins FM. Evolution and diversity of Rickettsia bacteria. BMC Biol. 2009;7:6. doi: 10.1186/1741-7007-7-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Majerus MEN, Hurst GDD. Ladybirds as a model system for the study of male-killing symbionts. Entomophaga. 1997;42(1-2):13–20. doi: 10.1007/BF02769875. [DOI] [Google Scholar]
- Fukatsu T, Shimada M. Molecular characterization of Rickettsia sp. in a bruchid beetle, Kytorhinus sharpianus (Coleoptera: Bruchidae) Appl Entomol Zool. 1999;34(3):391–397. [Google Scholar]
- Perotti MA, Clarke HK, Turner BD, Braig HR. Rickettsia as obligate and mycetomic bacteria. FASEB J. 2006;20:2372–2374. doi: 10.1096/fj.06-5870fje. [DOI] [PubMed] [Google Scholar]
- Yusuf M, Turner B. Characterisation of Wolbachia-like bacteria isolated from the parthenogenetic stored-product pest psocid Liposcelis bostrychophila (Badonnel) (Psocoptera) J Stored Prod Res. 2004;40(2):207–225. doi: 10.1016/S0022-474X(02)00098-X. [DOI] [Google Scholar]
- Amend AS, Seifert KA, Bruns TD. Quantifying microbial communities with 454 pyrosequencing: does read abundance count? Mol Ecol. 2010;19(24):5555–5565. doi: 10.1111/j.1365-294X.2010.04898.x. [DOI] [PubMed] [Google Scholar]
- Hosokawa T, Fukatsu T. Nardonella endosymbiont in the West Indian sweet potato weevil Euscepes postfasciatus (Coleoptera: Curculionidae) Appl Entomol Zool. 2010;45:115–120. doi: 10.1303/aez.2010.115. [DOI] [Google Scholar]
- Conord C, Despres L, Vallier A, Balmand S, Miquel C, Zundel S, Lemperiere G, Heddi A. Long-term evolutionary stability of bacterial endosymbiosis in Curculionoidea: additional evidence of symbiont replacement in the Dryophthoridae family. Mol Biol Evol. 2008;25(5):859–868. doi: 10.1093/molbev/msn027. [DOI] [PubMed] [Google Scholar]
- Lefevre C, Charles H, Vallier A, Delobel B, Farrell B, Heddi A. Endosymbiont phylogenesis in the Dryophthoridae weevils: evidence for bacterial replacement. Mol Biol Evol. 2004;21(6):965–973. doi: 10.1093/molbev/msh063. [DOI] [PubMed] [Google Scholar]
- Kuriwada T, Hosokawa T, Kumano N, Shiromoto K, Haraguchi D, Fukatsu T. Biological role of Nardonella endosymbiont in its weevil host. PLoS One. 2010;5(10):e13101. doi: 10.1371/journal.pone.0013101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lachowska D, Kajtoch L, Knutelski S. Occurrence of Wolbachia in central European weevils: correlations with host systematics, ecology, and biology. Entomol Expl Appl. 2010;135(1):105–118. doi: 10.1111/j.1570-7458.2010.00974.x. [DOI] [Google Scholar]
- Stenberg P, Lundmark M. Distribution, mechanisms and evolutionary significance of clonality and polyploidy in weevils. Agri For Entomol. 2004;6(4):259–266. doi: 10.1111/j.1461-9555.2004.00231.x. [DOI] [Google Scholar]
- Son Y, Luckhart S, Zhang X, Lieber MJ, Lewis EE. Effects and implications of antibiotic treatment on Wolbachia-infected vine weevil (Coleoptera: Curculionidae) Agri For Entomol. 2008;10(2):147–155. doi: 10.1111/j.1461-9563.2008.00369.x. [DOI] [Google Scholar]
- Werren JH, Baldo L, Clark ME. Wolbachia: master manipulators of invertebrate biology. Nature Rev Microbiol. 2008;6(10):741–751. doi: 10.1038/nrmicro1969. [DOI] [PubMed] [Google Scholar]
- Stenberg P, Lundmark M, Knutelski S, Saura A. Evolution of clonality and polyploidy in a weevil system. Mol Biol Evol. 2003;20(10):1626–1632. doi: 10.1093/molbev/msg180. [DOI] [PubMed] [Google Scholar]
- Fehr JS, Bloemberg GV, Ritter C, Hombach M, Luscher TF, Weber R, Keller PM. Septicemia caused by tick-borne bacterial pathogen Candidatus Neoehrlichia mikurensis. Emerg Infect Diseases. 2010;16(7):1127–1129. doi: 10.3201/eid1607.091907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yabsley MJ, Murphy SM, Luttrell MP, Wilcox BR, Ruckdeschel C. Raccoons (Procyon lotor), but not rodents, are natural and experimental hosts for an ehrlichial organism related to "Candidatus Neoehrlichia mikurensis". Vet Microbiol. 2008;131(3-4):301–308. doi: 10.1016/j.vetmic.2008.04.004. [DOI] [PubMed] [Google Scholar]
- Kawahara M, Rikihisa Y, Isogai E, Takahashi M, Misumi H, Suto C, Shibata S, Zhang CB, Tsuji M. Ultrastructure and phylogenetic analysis of "Candidatus Neoehrlichia mikurensis" in the family Anaplasmataceae, isolated from wild rats and found in Ixodes ovatus ticks. Int J Sys Evol Microbiol. 2004;54:1837–1843. doi: 10.1099/ijs.0.63260-0. [DOI] [PubMed] [Google Scholar]
- Arthofer W, Riegler M, Schneider D, Krammer M, Miller WJ, Stauffer C. Hidden Wolbachia diversity in field populations of the European cherry fruit fly, Rhagoletis cerasi (Diptera, Tephritidae) Mol Ecol. 2009;18(18):3816–3830. doi: 10.1111/j.1365-294X.2009.04321.x. [DOI] [PubMed] [Google Scholar]
- Toju H, Hosokawa T, Koga R, Nikoh N, Meng XY, Kimura N, Fukatsu T. "Candidatus Curculioniphilus buchneri," a novel clade of bacterial endocellular symbionts from weevils of the genus Curculio. Appl Environl Microbiol. 2010;76(1):275–282. doi: 10.1128/AEM.02154-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nardon P. Oogenesis and transmission of symbiotic bacteria in the weevil Sitophilus oryzae L. (Coleoptera: Dryophthoridae) Ann Soc Entomol Fr. 2006;42(2):129–164. [Google Scholar]
- Anselme C, Vallier A, Balmand S, Fauvarque MO, Heddi A. Host PGRP gene expression and bacterial release in endosymbiosis of the weevil Sitophilus zeamais. Appl Environ Microbiol. 2006;72(10):6766–6772. doi: 10.1128/AEM.00942-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buchner P. Endosymbiose der Tiere mit pflanzlichen Mikroorganismen. Birkhäuser Verlag Basel. 1953.
- Zindel R, Gottlieb Y, Aebi A. Arthropod symbioses: a neglected parameter in pest- and disease-control programmes. J Appl Ecol. 2011;48(4):864–872. doi: 10.1111/j.1365-2664.2011.01984.x. [DOI] [Google Scholar]
- Zabalou S, Riegler M, Theodorakopoulou M, Stauffer C, Savakis C, Bourtzis K. Wolbachia-induced cytoplasmic incompatibility as a means for insect pest population control. Proc Natl Acad Sci USA. 2004;101(42):15042–15045. doi: 10.1073/pnas.0403853101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hurst GDD, Jiggins FM. Male-killing bacteria in insects: mechanisms, incidence, and implications. Emerg Infect Diseases. 2000;6(4):329–336. doi: 10.3201/eid0604.000402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher JR, Bruck DJ. A technique for continuous mass rearing of the black vine weevil, Otiorhynchus sulcatus. Entomol Exp Appl. 2004;113(1):71–75. doi: 10.1111/j.0013-8703.2004.00206.x. [DOI] [Google Scholar]
- Adams AS, Adams SM, Currie CR, Gillette NE, Raffa KF. Geographic variation in bacterial communities associated with the Red Turpentine Beetle (Coleoptera: Curculionidae) Environ Entomol. 2010;39:406–414. doi: 10.1603/EN09221. [DOI] [PubMed] [Google Scholar]
- Mohr KI, Tebbe CC. Diversity and phylotype consistency of bacteria in the guts of three bee species (Apoidea) at an oilseed rape field. Environ Microbiol. 2006;8(2):258–272. doi: 10.1111/j.1462-2920.2005.00893.x. [DOI] [PubMed] [Google Scholar]
- Hosokawa T, Kikuchi Y, Shimada M, Fukatsu T. Obligate symbiont involved in pest status of host insect. Proc R Soc Lond [Biol] 2007;274(1621):1979–1984. doi: 10.1098/rspb.2007.0620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirsch J, Sprick P, Reineke A. Molecular identification of larval stages of Otiorhynchus (Coleoptera: Curculionidae) species based on polymerase chain reaction-restriction fragment length polymorphism analysis. J Econ Entomol. 2010;103(3):898–907. doi: 10.1603/EC09381. [DOI] [PubMed] [Google Scholar]
- Hamp TJ, Jones WJ, Fodor AA. Effects of experimental choices and analysis noise on surveys of the "rare biosphere". Appl Environ Microbiol. 2009;75(10):3263–3270. doi: 10.1128/AEM.01931-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drummond A, Ashton B, Cheung M, Heled J, Kearse M, Moir R, Stones-Havas S, Thierer T, Wilson A. Geneious v4.8. http://www.geneious.com. 2009.
- Katoh K, Kuma K, Toh H, Miyata T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 2005;33:511–518. doi: 10.1093/nar/gki198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig WG, Peplies J, Glockner FO. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 2007;35:7188–7196. doi: 10.1093/nar/gkm864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar Buchner A, Lai T, Steppi S, Jobb G. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 2004;32(4):1363–1371. doi: 10.1093/nar/gkh293. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
16S rDNA gene-based phylogeny of endosymbionts in four different Otiorhynchus spp. larvae. Sequences obtained in the present study are coloured and accession numbers of 16S rDNA sequences are shown for related bacterial species. More distantly related or unrelated sequences are not shown. Sequences from this work were added using the parsimony algorithm. This tree results from a phylogenetic calculation including more than 26,0000 bacterial 16S rDNA sequences. Only the nearest relatives are shown in this tree.