Abstract
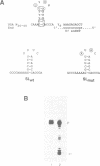
Specific protein-RNA complexes are formed by incubating a synthetic histone mRNA 3' end (a 30 nucleotide stem-loop structure) RNA with extracts of either nuclei or polyribosomes. The complex formed between the stem-loop and nuclear proteins has a lower electrophoretic mobility than the complex formed between the stem-loop and polyribosomal proteins. Binding of the synthetic 3' end by both polyribosomal and nuclear proteins is abolished when two of the conserved uridine residues in the loop are replaced with adenosines. UV crosslinking of the protein complexes to the synthetic RNA resulted in transferring radiolabel to similar sized proteins, 50 kD, in both the nuclear and polyribosomal extracts.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adam S. A., Nakagawa T., Swanson M. S., Woodruff T. K., Dreyfuss G. mRNA polyadenylate-binding protein: gene isolation and sequencing and identification of a ribonucleoprotein consensus sequence. Mol Cell Biol. 1986 Aug;6(8):2932–2943. doi: 10.1128/mcb.6.8.2932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernstein P., Peltz S. W., Ross J. The poly(A)-poly(A)-binding protein complex is a major determinant of mRNA stability in vitro. Mol Cell Biol. 1989 Feb;9(2):659–670. doi: 10.1128/mcb.9.2.659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernstein P., Ross J. Poly(A), poly(A) binding protein and the regulation of mRNA stability. Trends Biochem Sci. 1989 Sep;14(9):373–377. doi: 10.1016/0968-0004(89)90011-x. [DOI] [PubMed] [Google Scholar]
- Birnstiel M. L., Busslinger M., Strub K. Transcription termination and 3' processing: the end is in site! Cell. 1985 Jun;41(2):349–359. doi: 10.1016/s0092-8674(85)80007-6. [DOI] [PubMed] [Google Scholar]
- Brewer G. An A + U-rich element RNA-binding factor regulates c-myc mRNA stability in vitro. Mol Cell Biol. 1991 May;11(5):2460–2466. doi: 10.1128/mcb.11.5.2460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Casey J. L., Hentze M. W., Koeller D. M., Caughman S. W., Rouault T. A., Klausner R. D., Harford J. B. Iron-responsive elements: regulatory RNA sequences that control mRNA levels and translation. Science. 1988 May 13;240(4854):924–928. doi: 10.1126/science.2452485. [DOI] [PubMed] [Google Scholar]
- Dreyfuss G., Philipson L., Mattaj I. W. Ribonucleoprotein particles in cellular processes. J Cell Biol. 1988 May;106(5):1419–1425. doi: 10.1083/jcb.106.5.1419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gick O., Krämer A., Keller W., Birnstiel M. L. Generation of histone mRNA 3' ends by endonucleolytic cleavage of the pre-mRNA in a snRNP-dependent in vitro reaction. EMBO J. 1986 Jun;5(6):1319–1326. doi: 10.1002/j.1460-2075.1986.tb04362.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graves R. A., Pandey N. B., Chodchoy N., Marzluff W. F. Translation is required for regulation of histone mRNA degradation. Cell. 1987 Feb 27;48(4):615–626. doi: 10.1016/0092-8674(87)90240-6. [DOI] [PubMed] [Google Scholar]
- Greenberg J. R. Proteins crosslinked to messenger RNA by irradiating polyribosomes with ultraviolet light. Nucleic Acids Res. 1980 Dec 11;8(23):5685–5701. doi: 10.1093/nar/8.23.5685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greenberg J. R. Ultraviolet light-induced crosslinking of mRNA to proteins. Nucleic Acids Res. 1979 Feb;6(2):715–732. doi: 10.1093/nar/6.2.715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris M. E., Böhni R., Schneiderman M. H., Ramamurthy L., Schümperli D., Marzluff W. F. Regulation of histone mRNA in the unperturbed cell cycle: evidence suggesting control at two posttranscriptional steps. Mol Cell Biol. 1991 May;11(5):2416–2424. doi: 10.1128/mcb.11.5.2416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hentschel C. C., Birnstiel M. L. The organization and expression of histone gene families. Cell. 1981 Aug;25(2):301–313. doi: 10.1016/0092-8674(81)90048-9. [DOI] [PubMed] [Google Scholar]
- Hurt M. M., Chodchoy N., Marzluff W. F. The mouse histone H2a.2 gene from chromosome 3. Nucleic Acids Res. 1989 Nov 11;17(21):8876–8876. doi: 10.1093/nar/17.21.8876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jackson R. J., Standart N. Do the poly(A) tail and 3' untranslated region control mRNA translation? Cell. 1990 Jul 13;62(1):15–24. doi: 10.1016/0092-8674(90)90235-7. [DOI] [PubMed] [Google Scholar]
- Koeller D. M., Casey J. L., Hentze M. W., Gerhardt E. M., Chan L. N., Klausner R. D., Harford J. B. A cytosolic protein binds to structural elements within the iron regulatory region of the transferrin receptor mRNA. Proc Natl Acad Sci U S A. 1989 May;86(10):3574–3578. doi: 10.1073/pnas.86.10.3574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levine B. J., Chodchoy N., Marzluff W. F., Skoultchi A. I. Coupling of replication type histone mRNA levels to DNA synthesis requires the stem-loop sequence at the 3' end of the mRNA. Proc Natl Acad Sci U S A. 1987 Sep;84(17):6189–6193. doi: 10.1073/pnas.84.17.6189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malter J. S. Identification of an AUUUA-specific messenger RNA binding protein. Science. 1989 Nov 3;246(4930):664–666. doi: 10.1126/science.2814487. [DOI] [PubMed] [Google Scholar]
- Miller D. J., McMillan J., Miles A., ten Lohuis M., Mahony T. Nucleotide sequence of the histone H3-encoding gene from the scleractinian coral Acropora formosa (Cnidaria: Scleractinia). Gene. 1990 Sep 14;93(2):319–320. doi: 10.1016/0378-1119(90)90243-k. [DOI] [PubMed] [Google Scholar]
- Milligan J. F., Groebe D. R., Witherell G. W., Uhlenbeck O. C. Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res. 1987 Nov 11;15(21):8783–8798. doi: 10.1093/nar/15.21.8783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mowry K. L., Oh R., Steitz J. A. Each of the conserved sequence elements flanking the cleavage site of mammalian histone pre-mRNAs has a distinct role in the 3'-end processing reaction. Mol Cell Biol. 1989 Jul;9(7):3105–3108. doi: 10.1128/mcb.9.7.3105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. Both conserved signals on mammalian histone pre-mRNAs associate with small nuclear ribonucleoproteins during 3' end formation in vitro. Mol Cell Biol. 1987 May;7(5):1663–1672. doi: 10.1128/mcb.7.5.1663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. Identification of the human U7 snRNP as one of several factors involved in the 3' end maturation of histone premessenger RNA's. Science. 1987 Dec 18;238(4834):1682–1687. doi: 10.1126/science.2825355. [DOI] [PubMed] [Google Scholar]
- Munroe D., Jacobson A. mRNA poly(A) tail, a 3' enhancer of translational initiation. Mol Cell Biol. 1990 Jul;10(7):3441–3455. doi: 10.1128/mcb.10.7.3441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller K., Lindauer A., Brüderlein M., Schmitt R. Organization and transcription of Volvox histone-encoding genes: similarities between algal and animal genes. Gene. 1990 Sep 14;93(2):167–175. doi: 10.1016/0378-1119(90)90221-c. [DOI] [PubMed] [Google Scholar]
- Müllner E. W., Kühn L. C. A stem-loop in the 3' untranslated region mediates iron-dependent regulation of transferrin receptor mRNA stability in the cytoplasm. Cell. 1988 Jun 3;53(5):815–825. doi: 10.1016/0092-8674(88)90098-0. [DOI] [PubMed] [Google Scholar]
- Pandey N. B., Marzluff W. F. The stem-loop structure at the 3' end of histone mRNA is necessary and sufficient for regulation of histone mRNA stability. Mol Cell Biol. 1987 Dec;7(12):4557–4559. doi: 10.1128/mcb.7.12.4557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ratnasabapathy R., Hwang S. P., Williams D. L. The 3'-untranslated region of apolipoprotein II mRNA contains two independent domains that bind distinct cytosolic factors. J Biol Chem. 1990 Aug 15;265(23):14050–14055. [PubMed] [Google Scholar]
- Roberts S. B., Emmons S. W., Childs G. Nucleotide sequences of Caenorhabditis elegans core histone genes. Genes for different histone classes share common flanking sequence elements. J Mol Biol. 1989 Apr 20;206(4):567–577. doi: 10.1016/0022-2836(89)90566-4. [DOI] [PubMed] [Google Scholar]
- Ross J., Kobs G. H4 histone messenger RNA decay in cell-free extracts initiates at or near the 3' terminus and proceeds 3' to 5'. J Mol Biol. 1986 Apr 20;188(4):579–593. doi: 10.1016/s0022-2836(86)80008-0. [DOI] [PubMed] [Google Scholar]
- Ross J., Peltz S. W., Kobs G., Brewer G. Histone mRNA degradation in vivo: the first detectable step occurs at or near the 3' terminus. Mol Cell Biol. 1986 Dec;6(12):4362–4371. doi: 10.1128/mcb.6.12.4362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rouault T. A., Tang C. K., Kaptain S., Burgess W. H., Haile D. J., Samaniego F., McBride O. W., Harford J. B., Klausner R. D. Cloning of the cDNA encoding an RNA regulatory protein--the human iron-responsive element-binding protein. Proc Natl Acad Sci U S A. 1990 Oct;87(20):7958–7962. doi: 10.1073/pnas.87.20.7958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sachs A. B., Davis R. W. The poly(A) binding protein is required for poly(A) shortening and 60S ribosomal subunit-dependent translation initiation. Cell. 1989 Sep 8;58(5):857–867. doi: 10.1016/0092-8674(89)90938-0. [DOI] [PubMed] [Google Scholar]
- Sellos D., Krawetz S. A., Dixon G. H. Organization and complete nucleotide sequence of the core-histone-gene cluster of the annelid Platynereis dumerilii. Eur J Biochem. 1990 May 31;190(1):21–29. doi: 10.1111/j.1432-1033.1990.tb15540.x. [DOI] [PubMed] [Google Scholar]
- Sittman D. B., Graves R. A., Marzluff W. F. Histone mRNA concentrations are regulated at the level of transcription and mRNA degradation. Proc Natl Acad Sci U S A. 1983 Apr;80(7):1849–1853. doi: 10.1073/pnas.80.7.1849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soldati D., Schümperli D. Structural and functional characterization of mouse U7 small nuclear RNA active in 3' processing of histone pre-mRNA. Mol Cell Biol. 1988 Apr;8(4):1518–1524. doi: 10.1128/mcb.8.4.1518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasserot A. P., Schaufele F. J., Birnstiel M. L. Conserved terminal hairpin sequences of histone mRNA precursors are not involved in duplex formation with the U7 RNA but act as a target site for a distinct processing factor. Proc Natl Acad Sci U S A. 1989 Jun;86(12):4345–4349. doi: 10.1073/pnas.86.12.4345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wahle E. A novel poly(A)-binding protein acts as a specificity factor in the second phase of messenger RNA polyadenylation. Cell. 1991 Aug 23;66(4):759–768. doi: 10.1016/0092-8674(91)90119-j. [DOI] [PubMed] [Google Scholar]
- Wells D., Brown D. Histone and histone gene compilation and alignment update. Nucleic Acids Res. 1991 Apr 25;19 (Suppl):2173–2188. doi: 10.1093/nar/19.suppl.2173. [DOI] [PMC free article] [PubMed] [Google Scholar]