Abstract
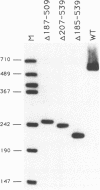
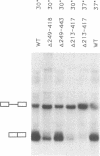
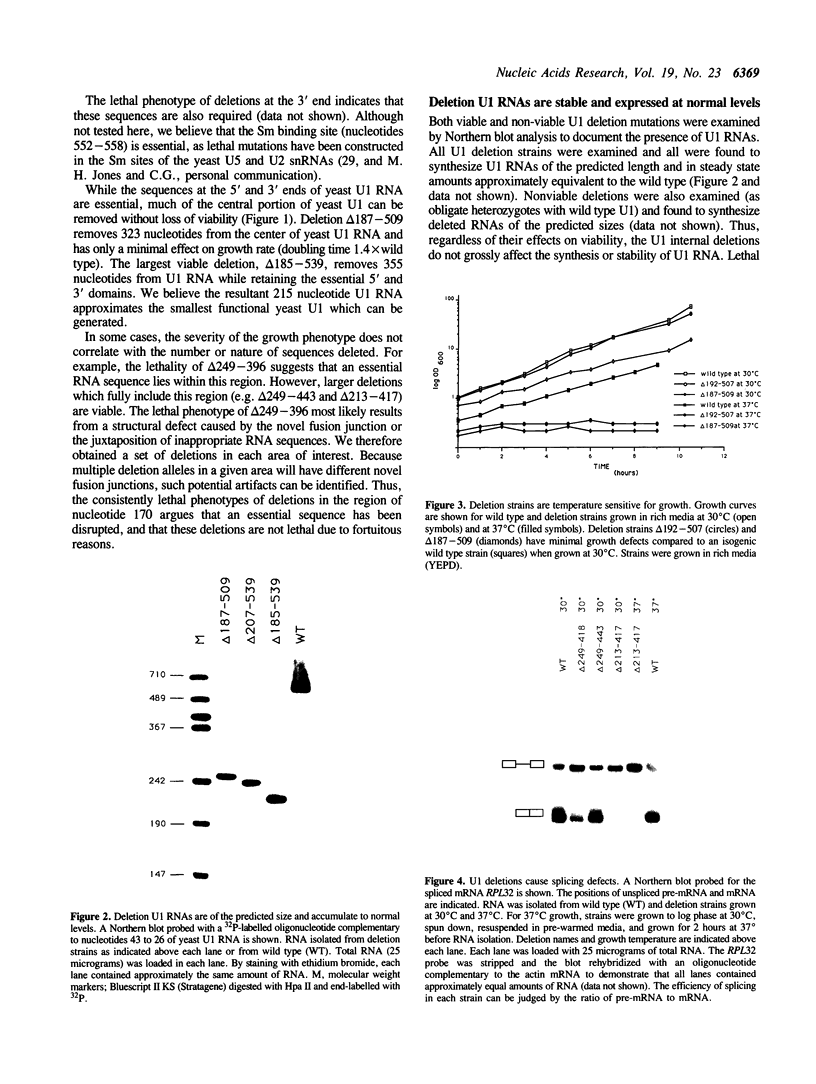
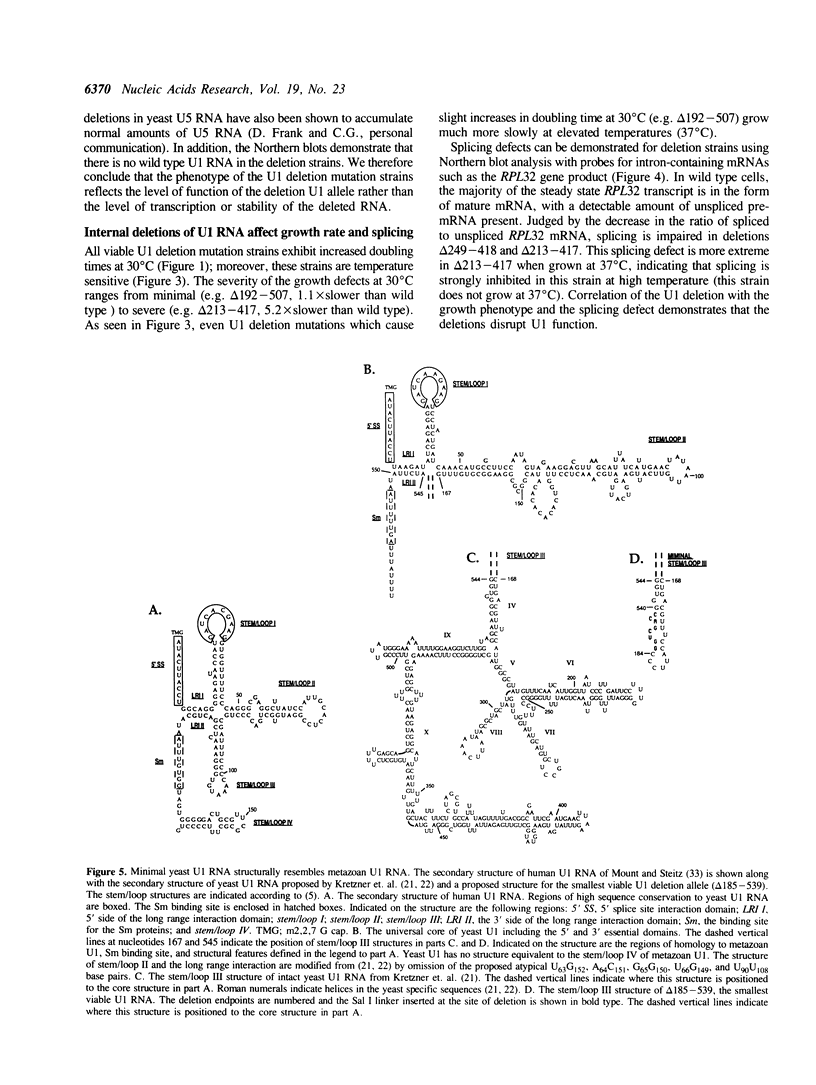
Yeast U1 snRNA (568 nucleotides) is 3.5-fold larger than its mammalian counterpart (164 nucleotides) and contains apparent sequence homology only at the 5' and 3' ends. We have used deletion analysis to determine whether the yeast-specific U1 sequences play essential roles in vivo. Yeast cells carrying a deletion of more than 60% (355 nucleotides) of the single-copy U1 gene are viable, though slow-growing, while a deletion of 316 nucleotides allows essentially wild-type growth. The boundaries of the viable deletions define a dispensable internal domain which comprises sequences unique to yeast. In contrast, the essential 5' and 3' terminal domains correspond to phylogenetically conserved sequences and/or structures previously implicated in RNA:RNA and RNA:protein interactions. The minimal essential sequences of yeast U1 can be drawn in a secondary structure which resembles metazoan U1 in four of seven structural domains.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ares M., Jr U2 RNA from yeast is unexpectedly large and contains homology to vertebrate U4, U5, and U6 small nuclear RNAs. Cell. 1986 Oct 10;47(1):49–59. doi: 10.1016/0092-8674(86)90365-x. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., Trueheart J., Natsoulis G., Fink G. R. 5-Fluoroorotic acid as a selective agent in yeast molecular genetics. Methods Enzymol. 1987;154:164–175. doi: 10.1016/0076-6879(87)54076-9. [DOI] [PubMed] [Google Scholar]
- Brody E., Abelson J. The "spliceosome": yeast pre-messenger RNA associates with a 40S complex in a splicing-dependent reaction. Science. 1985 May 24;228(4702):963–967. doi: 10.1126/science.3890181. [DOI] [PubMed] [Google Scholar]
- Couto J. R., Tamm J., Parker R., Guthrie C. A trans-acting suppressor restores splicing of a yeast intron with a branch point mutation. Genes Dev. 1987 Jul;1(5):445–455. doi: 10.1101/gad.1.5.445. [DOI] [PubMed] [Google Scholar]
- Frendewey D., Keller W. Stepwise assembly of a pre-mRNA splicing complex requires U-snRNPs and specific intron sequences. Cell. 1985 Aug;42(1):355–367. doi: 10.1016/s0092-8674(85)80131-8. [DOI] [PubMed] [Google Scholar]
- Goguel V., Liao X. L., Rymond B. C., Rosbash M. U1 snRNP can influence 3'-splice site selection as well as 5'-splice site selection. Genes Dev. 1991 Aug;5(8):1430–1438. doi: 10.1101/gad.5.8.1430. [DOI] [PubMed] [Google Scholar]
- Grabowski P. J., Seiler S. R., Sharp P. A. A multicomponent complex is involved in the splicing of messenger RNA precursors. Cell. 1985 Aug;42(1):345–353. doi: 10.1016/s0092-8674(85)80130-6. [DOI] [PubMed] [Google Scholar]
- Guthrie C., Patterson B. Spliceosomal snRNAs. Annu Rev Genet. 1988;22:387–419. doi: 10.1146/annurev.ge.22.120188.002131. [DOI] [PubMed] [Google Scholar]
- Hamm J., van Santen V. L., Spritz R. A., Mattaj I. W. Loop I of U1 small nuclear RNA is the only essential RNA sequence for binding of specific U1 small nuclear ribonucleoprotein particle proteins. Mol Cell Biol. 1988 Nov;8(11):4787–4791. doi: 10.1128/mcb.8.11.4787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Igel A. H., Ares M., Jr Internal sequences that distinguish yeast from metazoan U2 snRNA are unnecessary for pre-mRNA splicing. Nature. 1988 Aug 4;334(6181):450–453. doi: 10.1038/334450a0. [DOI] [PubMed] [Google Scholar]
- Jones M. H., Guthrie C. Unexpected flexibility in an evolutionarily conserved protein-RNA interaction: genetic analysis of the Sm binding site. EMBO J. 1990 Aug;9(8):2555–2561. doi: 10.1002/j.1460-2075.1990.tb07436.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kretzner L., Krol A., Rosbash M. Saccharomyces cerevisiae U1 small nuclear RNA secondary structure contains both universal and yeast-specific domains. Proc Natl Acad Sci U S A. 1990 Jan;87(2):851–855. doi: 10.1073/pnas.87.2.851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kretzner L., Rymond B. C., Rosbash M. S. cerevisiae U1 RNA is large and has limited primary sequence homology to metazoan U1 snRNA. Cell. 1987 Aug 14;50(4):593–602. doi: 10.1016/0092-8674(87)90032-8. [DOI] [PubMed] [Google Scholar]
- Liao X. L., Kretzner L., Seraphin B., Rosbash M. Universally conserved and yeast-specific U1 snRNA sequences are important but not essential for U1 snRNP function. Genes Dev. 1990 Oct;4(10):1766–1774. doi: 10.1101/gad.4.10.1766. [DOI] [PubMed] [Google Scholar]
- Lutz-Freyermuth C., Keene J. D., Lutz-Reyermuth C. The U1 RNA-binding site of the U1 small nuclear ribonucleoprotein (snRNP)-associated A protein suggests a similarity with U2 snRNPs. Mol Cell Biol. 1989 Jul;9(7):2975–2982. doi: 10.1128/mcb.9.7.2975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mount S. M., Steitz J. A. Sequence of U1 RNA from Drosophila melanogaster: implications for U1 secondary structure and possible involvement in splicing. Nucleic Acids Res. 1981 Dec 11;9(23):6351–6368. doi: 10.1093/nar/9.23.6351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker R., Siliciano P. G., Guthrie C. Recognition of the TACTAAC box during mRNA splicing in yeast involves base pairing to the U2-like snRNA. Cell. 1987 Apr 24;49(2):229–239. doi: 10.1016/0092-8674(87)90564-2. [DOI] [PubMed] [Google Scholar]
- Patterson B., Guthrie C. An essential yeast snRNA with a U5-like domain is required for splicing in vivo. Cell. 1987 Jun 5;49(5):613–624. doi: 10.1016/0092-8674(87)90537-x. [DOI] [PubMed] [Google Scholar]
- Query C. C., Bentley R. C., Keene J. D. A common RNA recognition motif identified within a defined U1 RNA binding domain of the 70K U1 snRNP protein. Cell. 1989 Apr 7;57(1):89–101. doi: 10.1016/0092-8674(89)90175-x. [DOI] [PubMed] [Google Scholar]
- Seraphin B., Rosbash M. Identification of functional U1 snRNA-pre-mRNA complexes committed to spliceosome assembly and splicing. Cell. 1989 Oct 20;59(2):349–358. doi: 10.1016/0092-8674(89)90296-1. [DOI] [PubMed] [Google Scholar]
- Shuster E. O., Guthrie C. Human U2 snRNA can function in pre-mRNA splicing in yeast. Nature. 1990 May 17;345(6272):270–273. doi: 10.1038/345270a0. [DOI] [PubMed] [Google Scholar]
- Shuster E. O., Guthrie C. Two conserved domains of yeast U2 snRNA are separated by 945 nonessential nucleotides. Cell. 1988 Oct 7;55(1):41–48. doi: 10.1016/0092-8674(88)90007-4. [DOI] [PubMed] [Google Scholar]
- Siliciano P. G., Guthrie C. 5' splice site selection in yeast: genetic alterations in base-pairing with U1 reveal additional requirements. Genes Dev. 1988 Oct;2(10):1258–1267. doi: 10.1101/gad.2.10.1258. [DOI] [PubMed] [Google Scholar]
- Siliciano P. G., Jones M. H., Guthrie C. Saccharomyces cerevisiae has a U1-like small nuclear RNA with unexpected properties. Science. 1987 Sep 18;237(4821):1484–1487. doi: 10.1126/science.3306922. [DOI] [PubMed] [Google Scholar]
- Surowy C. S., van Santen V. L., Scheib-Wixted S. M., Spritz R. A. Direct, sequence-specific binding of the human U1-70K ribonucleoprotein antigen protein to loop I of U1 small nuclear RNA. Mol Cell Biol. 1989 Oct;9(10):4179–4186. doi: 10.1128/mcb.9.10.4179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Séraphin B., Kretzner L., Rosbash M. A U1 snRNA:pre-mRNA base pairing interaction is required early in yeast spliceosome assembly but does not uniquely define the 5' cleavage site. EMBO J. 1988 Aug;7(8):2533–2538. doi: 10.1002/j.1460-2075.1988.tb03101.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu J., Manley J. L. Mammalian pre-mRNA branch site selection by U2 snRNP involves base pairing. Genes Dev. 1989 Oct;3(10):1553–1561. doi: 10.1101/gad.3.10.1553. [DOI] [PubMed] [Google Scholar]
- Zhuang Y., Weiner A. M. A compensatory base change in U1 snRNA suppresses a 5' splice site mutation. Cell. 1986 Sep 12;46(6):827–835. doi: 10.1016/0092-8674(86)90064-4. [DOI] [PubMed] [Google Scholar]
- Zhuang Y., Weiner A. M. A compensatory base change in human U2 snRNA can suppress a branch site mutation. Genes Dev. 1989 Oct;3(10):1545–1552. doi: 10.1101/gad.3.10.1545. [DOI] [PubMed] [Google Scholar]