Abstract
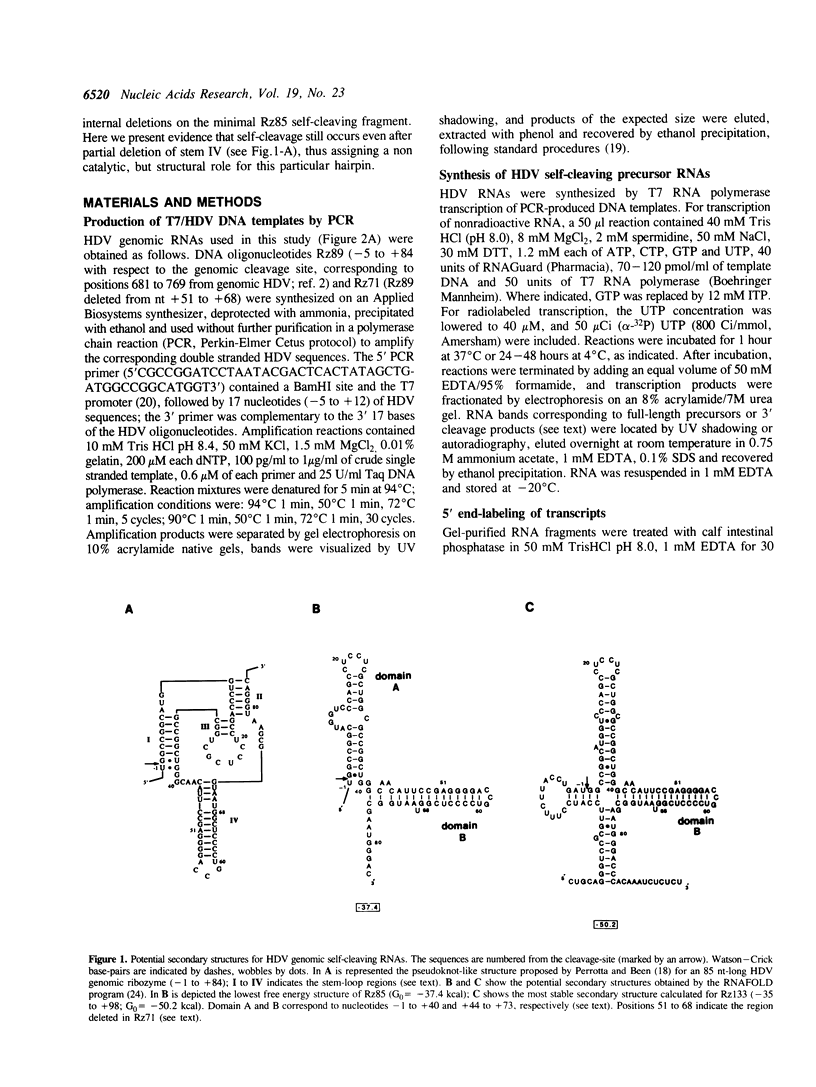
Self-cleavage efficiency of ribozymes derived from hepatitis delta virus (HDV) has been shown to depend on the RNA structure, which in turn may be determined by the length of the considered sequences. Here we describe the construction and functional analysis of a 71 nucleotide-long RNA genomic fragment, Rz71, which carries an 18 nucleotide deletion in a very stable GC-rich stem-loop (stem IV), predicted to be present in several computer-derived secondary structure models. Rz71 is able to undergo self-cleavage under non-denaturing conditions (the t1/2 of the reaction at 37 degrees C is 3 min). The deletion, however, is not neutral, since under the same conditions the non-deleted ribozyme cleaves to 50% in less than 15 sec. Therefore, stem-loop IV seems to play a structural role, not being directly involved in the catalytic reaction, but contributing to the correct positioning of the catalytic core of the HDV ribozyme. Rz71 is the smallest self-cleaving sub-fragment of HDV genomic RNA reported so far.
Full text
PDF

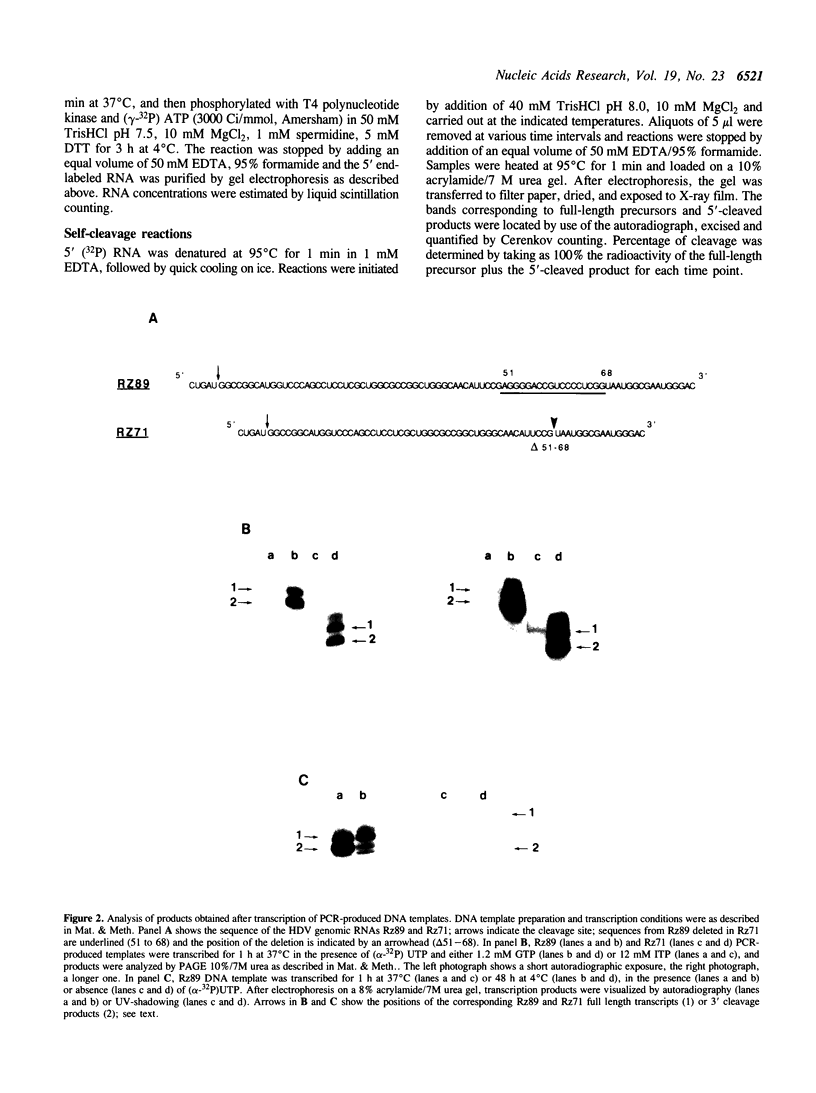




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Belinsky M. G., Dinter-Gottlieb G. Non-ribozyme sequences enhance self-cleavage of ribozymes derived from Hepatitis delta virus. Nucleic Acids Res. 1991 Feb 11;19(3):559–564. doi: 10.1093/nar/19.3.559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen P. J., Kalpana G., Goldberg J., Mason W., Werner B., Gerin J., Taylor J. Structure and replication of the genome of the hepatitis delta virus. Proc Natl Acad Sci U S A. 1986 Nov;83(22):8774–8778. doi: 10.1073/pnas.83.22.8774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davies R. W., Waring R. B., Ray J. A., Brown T. A., Scazzocchio C. Making ends meet: a model for RNA splicing in fungal mitochondria. Nature. 1982 Dec 23;300(5894):719–724. doi: 10.1038/300719a0. [DOI] [PubMed] [Google Scholar]
- Forster A. C., Symons R. H. Self-cleavage of plus and minus RNAs of a virusoid and a structural model for the active sites. Cell. 1987 Apr 24;49(2):211–220. doi: 10.1016/0092-8674(87)90562-9. [DOI] [PubMed] [Google Scholar]
- Hampel A., Tritz R., Hicks M., Cruz P. 'Hairpin' catalytic RNA model: evidence for helices and sequence requirement for substrate RNA. Nucleic Acids Res. 1990 Jan 25;18(2):299–304. doi: 10.1093/nar/18.2.299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konarska M. M., Sharp P. A. Replication of RNA by the DNA-dependent RNA polymerase of phage T7. Cell. 1989 May 5;57(3):423–431. doi: 10.1016/0092-8674(89)90917-3. [DOI] [PubMed] [Google Scholar]
- Kos A., Dijkema R., Arnberg A. C., van der Meide P. H., Schellekens H. The hepatitis delta (delta) virus possesses a circular RNA. Nature. 1986 Oct 9;323(6088):558–560. doi: 10.1038/323558a0. [DOI] [PubMed] [Google Scholar]
- Kuo M. Y., Sharmeen L., Dinter-Gottlieb G., Taylor J. Characterization of self-cleaving RNA sequences on the genome and antigenome of human hepatitis delta virus. J Virol. 1988 Dec;62(12):4439–4444. doi: 10.1128/jvi.62.12.4439-4444.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milligan J. F., Groebe D. R., Witherell G. W., Uhlenbeck O. C. Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res. 1987 Nov 11;15(21):8783–8798. doi: 10.1093/nar/15.21.8783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milligan J. F., Uhlenbeck O. C. Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol. 1989;180:51–62. doi: 10.1016/0076-6879(89)80091-6. [DOI] [PubMed] [Google Scholar]
- Perrotta A. T., Been M. D. A pseudoknot-like structure required for efficient self-cleavage of hepatitis delta virus RNA. Nature. 1991 Apr 4;350(6317):434–436. doi: 10.1038/350434a0. [DOI] [PubMed] [Google Scholar]
- Perrotta A. T., Been M. D. The self-cleaving domain from the genomic RNA of hepatitis delta virus: sequence requirements and the effects of denaturant. Nucleic Acids Res. 1990 Dec 11;18(23):6821–6827. doi: 10.1093/nar/18.23.6821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rizzetto M., Canese M. G., Aricò S., Crivelli O., Trepo C., Bonino F., Verme G. Immunofluorescence detection of new antigen-antibody system (delta/anti-delta) associated to hepatitis B virus in liver and in serum of HBsAg carriers. Gut. 1977 Dec;18(12):997–1003. doi: 10.1136/gut.18.12.997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenstein S. P., Been M. D. Self-cleavage of hepatitis delta virus genomic strand RNA is enhanced under partially denaturing conditions. Biochemistry. 1990 Sep 4;29(35):8011–8016. doi: 10.1021/bi00487a002. [DOI] [PubMed] [Google Scholar]
- Schenborn E. T., Mierendorf R. C., Jr A novel transcription property of SP6 and T7 RNA polymerases: dependence on template structure. Nucleic Acids Res. 1985 Sep 11;13(17):6223–6236. doi: 10.1093/nar/13.17.6223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharmeen L., Kuo M. Y., Dinter-Gottlieb G., Taylor J. Antigenomic RNA of human hepatitis delta virus can undergo self-cleavage. J Virol. 1988 Aug;62(8):2674–2679. doi: 10.1128/jvi.62.8.2674-2679.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharmeen L., Kuo M. Y., Taylor J. Self-ligating RNA sequences on the antigenome of human hepatitis delta virus. J Virol. 1989 Mar;63(3):1428–1430. doi: 10.1128/jvi.63.3.1428-1430.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith J. B., Dinter-Gottlieb G. Antigenomic Hepatitis delta virus ribozymes self-cleave in 18 M formamide. Nucleic Acids Res. 1991 Mar 25;19(6):1285–1289. doi: 10.1093/nar/19.6.1285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor J. M. Human hepatitis delta virus. Curr Top Microbiol Immunol. 1991;168:141–166. doi: 10.1007/978-3-642-76015-0_7. [DOI] [PubMed] [Google Scholar]
- Wang K. S., Choo Q. L., Weiner A. J., Ou J. H., Najarian R. C., Thayer R. M., Mullenbach G. T., Denniston K. J., Gerin J. L., Houghton M. Structure, sequence and expression of the hepatitis delta (delta) viral genome. Nature. 1986 Oct 9;323(6088):508–514. doi: 10.1038/323508a0. [DOI] [PubMed] [Google Scholar]
- Wu H. N., Lai M. M. RNA conformational requirements of self-cleavage of hepatitis delta virus RNA. Mol Cell Biol. 1990 Oct;10(10):5575–5579. doi: 10.1128/mcb.10.10.5575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu H. N., Lin Y. J., Lin F. P., Makino S., Chang M. F., Lai M. M. Human hepatitis delta virus RNA subfragments contain an autocleavage activity. Proc Natl Acad Sci U S A. 1989 Mar;86(6):1831–1835. doi: 10.1073/pnas.86.6.1831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuker M. On finding all suboptimal foldings of an RNA molecule. Science. 1989 Apr 7;244(4900):48–52. doi: 10.1126/science.2468181. [DOI] [PubMed] [Google Scholar]