Abstract
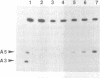
Several mutants for the minus strands of the self-cleaving domain of the satellite RNA of tobacco ringspot virus have been synthesized by joining chemically synthesized oligoribonucleotides with RNA ligase. Kinetic properties of the enzyme strands (50 nucleotides) against substrates (15-mer and 18-mer) were investigated. Structural properties of the unpaired part in the cleavage region were estimated from mutagenesis. The catalytic domain alone was proved to be responsible for the rejoining reaction of cleaved substrates. It was also found that the ribozyme could be divided into two strands without loss of activity. Effects of concentration of magnesium ion and polyamines on the cleavage reaction for the two-stranded ribozyme are also reported.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Cech T. R., Zaug A. J., Grabowski P. J. In vitro splicing of the ribosomal RNA precursor of Tetrahymena: involvement of a guanosine nucleotide in the excision of the intervening sequence. Cell. 1981 Dec;27(3 Pt 2):487–496. doi: 10.1016/0092-8674(81)90390-1. [DOI] [PubMed] [Google Scholar]
- Donis-Keller H., Maxam A. M., Gilbert W. Mapping adenines, guanines, and pyrimidines in RNA. Nucleic Acids Res. 1977 Aug;4(8):2527–2538. doi: 10.1093/nar/4.8.2527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Epstein L. M., Gall J. G. Self-cleaving transcripts of satellite DNA from the newt. Cell. 1987 Feb 13;48(3):535–543. doi: 10.1016/0092-8674(87)90204-2. [DOI] [PubMed] [Google Scholar]
- Feldstein P. A., Buzayan J. M., van Tol H., deBear J., Gough G. R., Gilham P. T., Bruening G. Specific association between an endoribonucleolytic sequence from a satellite RNA and a substrate analogue containing a 2'-5' phosphodiester. Proc Natl Acad Sci U S A. 1990 Apr;87(7):2623–2627. doi: 10.1073/pnas.87.7.2623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forster A. C., Symons R. H. Self-cleavage of plus and minus RNAs of a virusoid and a structural model for the active sites. Cell. 1987 Apr 24;49(2):211–220. doi: 10.1016/0092-8674(87)90562-9. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Gardiner K., Marsh T., Pace N., Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983 Dec;35(3 Pt 2):849–857. doi: 10.1016/0092-8674(83)90117-4. [DOI] [PubMed] [Google Scholar]
- Hampel A., Tritz R., Hicks M., Cruz P. 'Hairpin' catalytic RNA model: evidence for helices and sequence requirement for substrate RNA. Nucleic Acids Res. 1990 Jan 25;18(2):299–304. doi: 10.1093/nar/18.2.299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hampel A., Tritz R. RNA catalytic properties of the minimum (-)sTRSV sequence. Biochemistry. 1989 Jun 13;28(12):4929–4933. doi: 10.1021/bi00438a002. [DOI] [PubMed] [Google Scholar]
- Haseloff J., Gerlach W. L. Sequences required for self-catalysed cleavage of the satellite RNA of tobacco ringspot virus. Gene. 1989 Oct 15;82(1):43–52. doi: 10.1016/0378-1119(89)90028-0. [DOI] [PubMed] [Google Scholar]
- Hutchins C. J., Rathjen P. D., Forster A. C., Symons R. H. Self-cleavage of plus and minus RNA transcripts of avocado sunblotch viroid. Nucleic Acids Res. 1986 May 12;14(9):3627–3640. doi: 10.1093/nar/14.9.3627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kierzek R., Caruthers M. H., Longfellow C. E., Swinton D., Turner D. H., Freier S. M. Polymer-supported RNA synthesis and its application to test the nearest-neighbor model for duplex stability. Biochemistry. 1986 Dec 2;25(24):7840–7846. doi: 10.1021/bi00372a009. [DOI] [PubMed] [Google Scholar]
- Koizumi M., Hayase Y., Iwai S., Kamiya H., Inoue H., Ohtsuka E. Design of RNA enzymes distinguishing a single base mutation in RNA. Nucleic Acids Res. 1989 Sep 12;17(17):7059–7071. doi: 10.1093/nar/17.17.7059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koizumi M., Iwai S., Ohtsuka E. Cleavage of specific sites of RNA by designed ribozymes. FEBS Lett. 1988 Nov 7;239(2):285–288. doi: 10.1016/0014-5793(88)80935-9. [DOI] [PubMed] [Google Scholar]
- Koizumi M., Iwai S., Ohtsuka E. Construction of a series of several self-cleaving RNA duplexes using synthetic 21-mers. FEBS Lett. 1988 Feb 15;228(2):228–230. doi: 10.1016/0014-5793(88)80004-8. [DOI] [PubMed] [Google Scholar]
- Koizumi M., Ohtsuka E. Effects of phosphorothioate and 2-amino groups in hammerhead ribozymes on cleavage rates and Mg2+ binding. Biochemistry. 1991 May 28;30(21):5145–5150. doi: 10.1021/bi00235a005. [DOI] [PubMed] [Google Scholar]
- Lockard R. E., Alzner-Deweerd B., Heckman J. E., MacGee J., Tabor M. W., RajBhandary U. L. Sequence analysis of 5'[32P] labeled mRNA and tRNA using polyacrylamide gel electrophoresis. Nucleic Acids Res. 1978 Jan;5(1):37–56. doi: 10.1093/nar/5.1.37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prody G. A., Bakos J. T., Buzayan J. M., Schneider I. R., Bruening G. Autolytic processing of dimeric plant virus satellite RNA. Science. 1986 Mar 28;231(4745):1577–1580. doi: 10.1126/science.231.4745.1577. [DOI] [PubMed] [Google Scholar]
- Quigley G. J., Teeter M. M., Rich A. Structural analysis of spermine and magnesium ion binding to yeast phenylalanine transfer RNA. Proc Natl Acad Sci U S A. 1978 Jan;75(1):64–68. doi: 10.1073/pnas.75.1.64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugimoto N., Kierzek R., Turner D. H. Kinetics for reaction of a circularized intervening sequence with CU, UCU, CUCU, and CUCUCU: mechanistic implications from the dependence on temperature and on oligomer and Mg2+ concentrations. Biochemistry. 1988 Aug 23;27(17):6384–6392. doi: 10.1021/bi00417a029. [DOI] [PubMed] [Google Scholar]
- Sugimoto N., Tomka M., Kierzek R., Bevilacqua P. C., Turner D. H. Effects of substrate structure on the kinetics of circle opening reactions of the self-splicing intervening sequence from Tetrahymena thermophila: evidence for substrate and Mg2+ binding interactions. Nucleic Acids Res. 1989 Jan 11;17(1):355–371. doi: 10.1093/nar/17.1.355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Usher D. A., McHale A. H. Nonenzymic joining of oligoadenylates on a polyuridylic acid template. Science. 1976 Apr 2;192(4234):53–54. doi: 10.1126/science.1257755. [DOI] [PubMed] [Google Scholar]
- Verma I. M. The reverse transcriptase. Biochim Biophys Acta. 1977 Mar 21;473(1):1–38. doi: 10.1016/0304-419x(77)90005-1. [DOI] [PubMed] [Google Scholar]
- Wu H. N., Lai M. M. Reversible cleavage and ligation of hepatitis delta virus RNA. Science. 1989 Feb 3;243(4891):652–654. doi: 10.1126/science.2492677. [DOI] [PubMed] [Google Scholar]
- van Tol H., Buzayan J. M., Feldstein P. A., Eckstein F., Bruening G. Two autolytic processing reactions of a satellite RNA proceed with inversion of configuration. Nucleic Acids Res. 1990 Apr 25;18(8):1971–1975. doi: 10.1093/nar/18.8.1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Veen R., Arnberg A. C., van der Horst G., Bonen L., Tabak H. F., Grivell L. A. Excised group II introns in yeast mitochondria are lariats and can be formed by self-splicing in vitro. Cell. 1986 Jan 31;44(2):225–234. doi: 10.1016/0092-8674(86)90756-7. [DOI] [PubMed] [Google Scholar]