Abstract
The c-myc proto-oncogene encodes a highly unstable mRNA. Stabilized, truncated myc transcripts have been found in several human and murine tumors of hematopoietic origin. Recently, two tumors expressing 3' truncated c-myc mRNAs that were five times more stable than normal myc transcripts, were described. We have tried to determine the cause of the increased stability of the 3' truncated myc transcripts by studying the half-life of mutated c-myc mRNAs. The c-myc 3' untranslated region has been shown to contain sequences that confer mRNA instability. Possible candidates for such sequences are two (A + U)-rich regions in the 3' end of the c-myc RNA that resemble RNA destabilizing elements present in the c-fos and GMCSF mRNAs. We show that deletions in the (A + U)-rich regions do not stabilize c-myc messengers, and that hybrid mRNAs containing SV40 sequences at their 3' ends and terminating at an SV40 polyadenylation signal decay as quickly as normal c-myc transcripts. Our results indicate that neither the loss of (A + U)-rich sequences nor the mere addition of non-myc sequences to the 3' end of the mRNA lead to stabilization. We also show that rapid degradation of c-myc mRNA does not require complete translation of the coding sequences.
Full text
PDF



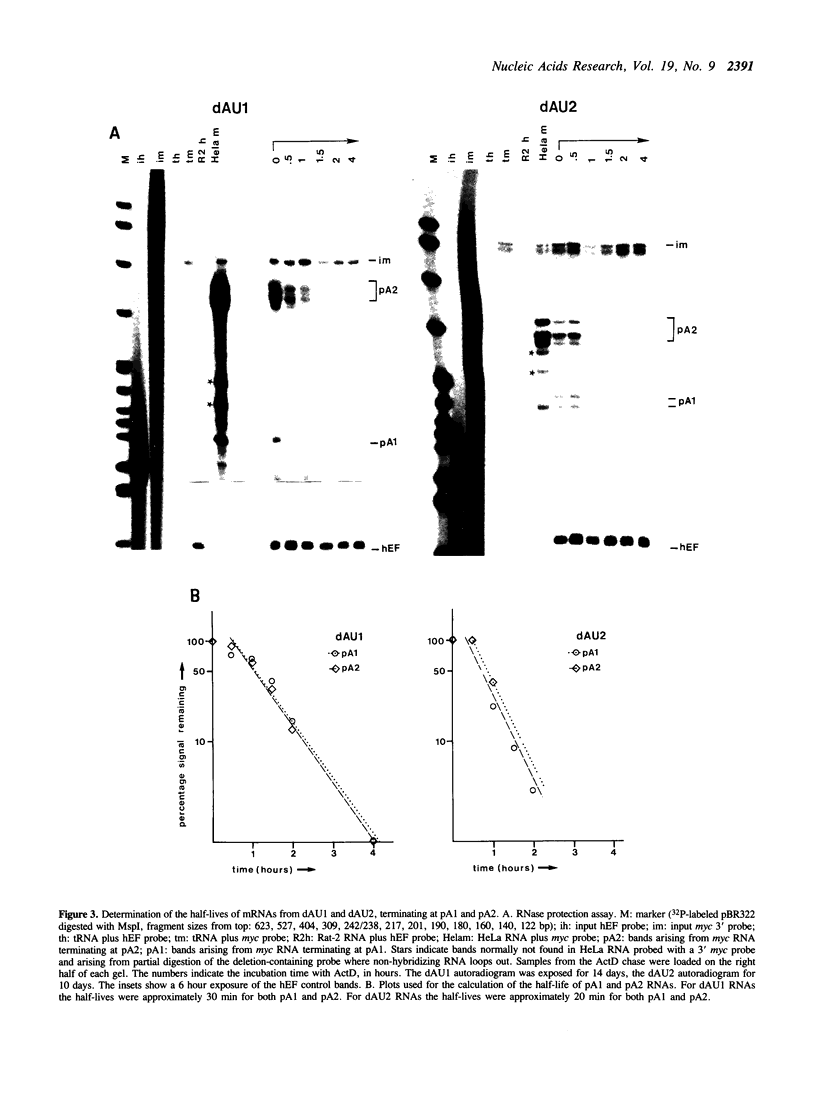



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abrahams J. P., van den Berg M., van Batenburg E., Pleij C. Prediction of RNA secondary structure, including pseudoknotting, by computer simulation. Nucleic Acids Res. 1990 May 25;18(10):3035–3044. doi: 10.1093/nar/18.10.3035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adams J. M., Gerondakis S., Webb E., Corcoran L. M., Cory S. Cellular myc oncogene is altered by chromosome translocation to an immunoglobulin locus in murine plasmacytomas and is rearranged similarly in human Burkitt lymphomas. Proc Natl Acad Sci U S A. 1983 Apr;80(7):1982–1986. doi: 10.1073/pnas.80.7.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aghib D. F., Bishop J. M., Ottolenghi S., Guerrasio A., Serra A., Saglio G. A 3' truncation of MYC caused by chromosomal translocation in a human T-cell leukemia increases mRNA stability. Oncogene. 1990 May;5(5):707–711. [PubMed] [Google Scholar]
- Alitalo K., Koskinen P., Mäkelä T. P., Saksela K., Sistonen L., Winqvist R. myc oncogenes: activation and amplification. Biochim Biophys Acta. 1987 Apr 20;907(1):1–32. doi: 10.1016/0304-419x(87)90016-3. [DOI] [PubMed] [Google Scholar]
- Bauer S. R., Piechaczyk M., Nordan R. P., Owens J. D., Nepveu A., Marcu K. B., Mushinski J. F. Altered myc gene transcription and intron-induced stabilization of myc RNAs in two mouse plasmacytomas. Oncogene. 1989 May;4(5):615–623. [PubMed] [Google Scholar]
- Bonnieu A., Piechaczyk M., Marty L., Cuny M., Blanchard J. M., Fort P., Jeanteur P. Sequence determinants of c-myc mRNA turn-over: influence of 3' and 5' non-coding regions. Oncogene Res. 1988 Sep;3(2):155–166. [PubMed] [Google Scholar]
- Bonnieu A., Rech J., Jeanteur P., Fort P. Requirements for c-fos mRNA down regulation in growth stimulated murine cells. Oncogene. 1989 Jul;4(7):881–888. [PubMed] [Google Scholar]
- Bonnieu A., Roux P., Marty L., Jeanteur P., Piechaczyk M. AUUUA motifs are dispensable for rapid degradation of the mouse c-myc RNA. Oncogene. 1990 Oct;5(10):1585–1588. [PubMed] [Google Scholar]
- Cole M. D. The myc oncogene: its role in transformation and differentiation. Annu Rev Genet. 1986;20:361–384. doi: 10.1146/annurev.ge.20.120186.002045. [DOI] [PubMed] [Google Scholar]
- Dani C., Blanchard J. M., Piechaczyk M., El Sabouty S., Marty L., Jeanteur P. Extreme instability of myc mRNA in normal and transformed human cells. Proc Natl Acad Sci U S A. 1984 Nov;81(22):7046–7050. doi: 10.1073/pnas.81.22.7046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eick D., Piechaczyk M., Henglein B., Blanchard J. M., Traub B., Kofler E., Wiest S., Lenoir G. M., Bornkamm G. W. Aberrant c-myc RNAs of Burkitt's lymphoma cells have longer half-lives. EMBO J. 1985 Dec 30;4(13B):3717–3725. doi: 10.1002/j.1460-2075.1985.tb04140.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fort P., Rech J., Vie A., Piechaczyk M., Bonnieu A., Jeanteur P., Blanchard J. M. Regulation of c-fos gene expression in hamster fibroblasts: initiation and elongation of transcription and mRNA degradation. Nucleic Acids Res. 1987 Jul 24;15(14):5657–5667. doi: 10.1093/nar/15.14.5657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graham F. L., van der Eb A. J. A new technique for the assay of infectivity of human adenovirus 5 DNA. Virology. 1973 Apr;52(2):456–467. doi: 10.1016/0042-6822(73)90341-3. [DOI] [PubMed] [Google Scholar]
- Graves R. A., Pandey N. B., Chodchoy N., Marzluff W. F. Translation is required for regulation of histone mRNA degradation. Cell. 1987 Feb 27;48(4):615–626. doi: 10.1016/0092-8674(87)90240-6. [DOI] [PubMed] [Google Scholar]
- Hayashi K., Makino R., Kawamura H., Arisawa A., Yoneda K. Characterization of rat c-myc and adjacent regions. Nucleic Acids Res. 1987 Aug 25;15(16):6419–6436. doi: 10.1093/nar/15.16.6419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hollis G. F., Gazdar A. F., Bertness V., Kirsch I. R. Complex translocation disrupts c-myc regulation in a human plasma cell myeloma. Mol Cell Biol. 1988 Jan;8(1):124–129. doi: 10.1128/mcb.8.1.124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones T. R., Cole M. D. Rapid cytoplasmic turnover of c-myc mRNA: requirement of the 3' untranslated sequences. Mol Cell Biol. 1987 Dec;7(12):4513–4521. doi: 10.1128/mcb.7.12.4513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King M. W., Roberts J. M., Eisenman R. N. Expression of the c-myc proto-oncogene during development of Xenopus laevis. Mol Cell Biol. 1986 Dec;6(12):4499–4508. doi: 10.1128/mcb.6.12.4499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laird-Offringa I. A., Elfferich P., Knaken H. J., de Ruiter J., van der Eb A. J. Analysis of polyadenylation site usage of the c-myc oncogene. Nucleic Acids Res. 1989 Aug 25;17(16):6499–6514. doi: 10.1093/nar/17.16.6499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laird-Offringa I. A., de Wit C. L., Elfferich P., van der Eb A. J. Poly(A) tail shortening is the translation-dependent step in c-myc mRNA degradation. Mol Cell Biol. 1990 Dec;10(12):6132–6140. doi: 10.1128/mcb.10.12.6132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linial M., Gunderson N., Groudine M. Enhanced transcription of c-myc in bursal lymphoma cells requires continuous protein synthesis. Science. 1985 Dec 6;230(4730):1126–1132. doi: 10.1126/science.2999973. [DOI] [PubMed] [Google Scholar]
- Marsh J. L., Erfle M., Wykes E. J. The pIC plasmid and phage vectors with versatile cloning sites for recombinant selection by insertional inactivation. Gene. 1984 Dec;32(3):481–485. doi: 10.1016/0378-1119(84)90022-2. [DOI] [PubMed] [Google Scholar]
- Piechaczyk M., Bonnieu A., Eick D., Remmers E., Yang J. Q., Marcu K., Jeanteur P., Blanchard J. M. Altered c-myc RNA metabolism in Burkitt's lymphomas and mouse plasmacytomas. Curr Top Microbiol Immunol. 1986;132:331–338. doi: 10.1007/978-3-642-71562-4_49. [DOI] [PubMed] [Google Scholar]
- Piechaczyk M., Yang J. Q., Blanchard J. M., Jeanteur P., Marcu K. B. Posttranscriptional mechanisms are responsible for accumulation of truncated c-myc RNAs in murine plasma cell tumors. Cell. 1985 Sep;42(2):589–597. doi: 10.1016/0092-8674(85)90116-3. [DOI] [PubMed] [Google Scholar]
- Rabbitts P. H., Forster A., Stinson M. A., Rabbitts T. H. Truncation of exon 1 from the c-myc gene results in prolonged c-myc mRNa stability. EMBO J. 1985 Dec 30;4(13B):3727–3733. doi: 10.1002/j.1460-2075.1985.tb04141.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rahmsdorf H. J., Schönthal A., Angel P., Litfin M., Rüther U., Herrlich P. Posttranscriptional regulation of c-fos mRNA expression. Nucleic Acids Res. 1987 Feb 25;15(4):1643–1659. doi: 10.1093/nar/15.4.1643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw G., Kamen R. A conserved AU sequence from the 3' untranslated region of GM-CSF mRNA mediates selective mRNA degradation. Cell. 1986 Aug 29;46(5):659–667. doi: 10.1016/0092-8674(86)90341-7. [DOI] [PubMed] [Google Scholar]
- Shyu A. B., Greenberg M. E., Belasco J. G. The c-fos transcript is targeted for rapid decay by two distinct mRNA degradation pathways. Genes Dev. 1989 Jan;3(1):60–72. doi: 10.1101/gad.3.1.60. [DOI] [PubMed] [Google Scholar]
- Stanton L. W., Fahrlander P. D., Tesser P. M., Marcu K. B. Nucleotide sequence comparison of normal and translocated murine c-myc genes. Nature. 1984 Aug 2;310(5976):423–425. doi: 10.1038/310423a0. [DOI] [PubMed] [Google Scholar]
- Topp W. C. Normal rat cell lines deficient in nuclear thymidine kinase. Virology. 1981 Aug;113(1):408–411. doi: 10.1016/0042-6822(81)90168-9. [DOI] [PubMed] [Google Scholar]
- Watson D. K., Reddy E. P., Duesberg P. H., Papas T. S. Nucleotide sequence analysis of the chicken c-myc gene reveals homologous and unique coding regions by comparison with the transforming gene of avian myelocytomatosis virus MC29, delta gag-myc. Proc Natl Acad Sci U S A. 1983 Apr;80(8):2146–2150. doi: 10.1073/pnas.80.8.2146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson T., Treisman R. Removal of poly(A) and consequent degradation of c-fos mRNA facilitated by 3' AU-rich sequences. Nature. 1988 Nov 24;336(6197):396–399. doi: 10.1038/336396a0. [DOI] [PubMed] [Google Scholar]
- Wisdom R., Lee W. The protein-coding region of c-myc mRNA contains a sequence that specifies rapid mRNA turnover and induction by protein synthesis inhibitors. Genes Dev. 1991 Feb;5(2):232–243. doi: 10.1101/gad.5.2.232. [DOI] [PubMed] [Google Scholar]