Abstract
We report the presence and diversity of Bartonella spp. in bats of 13 insectivorous and frugivorous species collected from various locations across Kenya. Bartonella isolates were obtained from 23 Eidolon helvum, 22 Rousettus aegyptiacus, 4 Coleura afra, 7 Triaenops persicus, 1 Hipposideros commersoni, and 49 Miniopterus spp. bats. Sequence analysis of the citrate synthase gene from the obtained isolates showed a wide assortment of Bartonella strains. Phylogenetically, isolates clustered in specific host bat species. All isolates from R. aegyptiacus, C. afra, and T. persicus bats clustered in separate monophyletic groups. In contrast, E. helvum and Miniopterus spp. bats harbored strains that clustered in several groups. Further investigation is needed to determine whether these agents are responsible for human illnesses in the region.
Keywords: Bacteria, Bartonella, bats, zoonoses, Kenya, research
An unprecedented, increasing interest in bats as reservoirs of infectious diseases occurred during the past decade. Mounting evidence indicates an association of bats with various emerging infections, some with high mortality rates, including lyssaviruses, severe acute respiratory syndrome and other coronaviruses, and henipa, Ebola, and Marburg viruses (1–6). However, the list of pathogens discovered in bats is even more extensive and includes other representatives from various taxonomic groups (6–8). Most infectious agents in bats are viruses; bacterial species have been rarely reported (9). Excluding reports resulting from serologic and microscopic observations before the 1990s, only a few recent publications describe bacterial species in bats. One publication reports fatal borreliosis in a bat caused by a relapsing fever spirochete in the United Kingdom (10). Another study on the molecular detection of hemoparasites infecting bats in southwest England showed the presence of Bartonella spp. DNA in the blood of 5 of 60 tested bats (11). This report on bat infection potentially caused by bacteria of the genus Bartonella is consistent with studies showing detection of Bartonella spp.–specific DNA in ectoparasites collected from bats in Egypt and the United States (12–14).
Bartonella spp. are mainly hemotropic, facultative intracellular parasites associated with erythrocytes and endothelial cells of mammals (15,16). Bartonella spp. organisms are highly adapted to a wide variety of mammals, including rodents, insectivores, carnivores, ungulates, and marine mammals such as dolphins. New insights into the natural history of various Bartonella spp. suggest that these bacteria have adapted to their mammalian reservoir hosts in unique ways with frequently restricted host species ranges (17). The bacteria can cause chronic intraerythrocyte infections that sometimes result in a large proportion of the reservoir host population being bacteremic simultaneously (18). Infections usually cause few or no clinical signs in the reservoir hosts. Host adaptation is evident as some Bartonella species and genotypes are found in very specific mammalian species.
Available data on Bartonella spp. have expanded rapidly during recent years, as this group of organisms has been found to be associated with a growing spectrum of emerging and reemerging diseases. In addition to cat-scratch disease, trench fever, and Carrión disease, other illnesses linked to Bartonella spp. infection range from a self-limiting, short-term fever to potentially fatal systemic diseases with cardiovascular, nervous system, or hepatosplenic involvement (19). Some Bartonella spp. that have been implicated as human pathogens are linked to rodent species; these species include B. elizabethae, B. grahamii, B. washoensis, and B. vinsonii subsp. arupensis (20–23). Other Bartonella spp. are linked to wild and pet carnivores. For example, B. henselae, carried by cats, causes cat-scratch disease in immunocompetent persons and bacillary angiomatosis in immunocompromised persons (19,24); B. vinsonii subsp. berkhoffii has been carried by dogs and is responsible for endocarditis in a human patient (24,25). For some Bartonella spp. recently implicated as human pathogens (such as B. rochalimae, which was isolated from an American tourist traveling to Peru, or B. tamiae, isolated from 3 patients in Thailand), a mammalian reservoir has not been determined despite a wide range of tested animals collected in these countries (26,27). These unidentified reservoirs indicate a need for extensive surveillance among diverse groups of animals for Bartonella strains, especially among bats, which represent around 20% of all mammalian species (6).
This study was conducted within the framework of the Centers for Disease Control and Prevention (CDC) Global Disease Detection program, which is designed to estimate the health and financial effects and transmission patterns of emerging infectious pathogens associated with humans and animals (including bats, rodents, and other likely reservoirs for human infection) in Kenya and other locations. This study had 5 objectives: 1) to estimate prevalence of Bartonella spp. infections among diverse chiropteran species in Kenya; 2) to isolate and identify detected Bartonella spp. and create a reference collection of Bartonella isolates from East Africa with further characterization and diagnostic investigation; 3) to evaluate genetic heterogeneity of circulating Bartonella strains by using the partial sequence variability of the citrate synthase gene (gltA), proven to be an excellent genetic marker for analysis of Bartonella strains (28); 4) to compare Bartonella strains obtained from bats in this study with strains obtained from other animal reservoirs and available from the public domain; and 5) potentially to identify new species of Bartonella.
Material and Methods
Blood Samples Collection
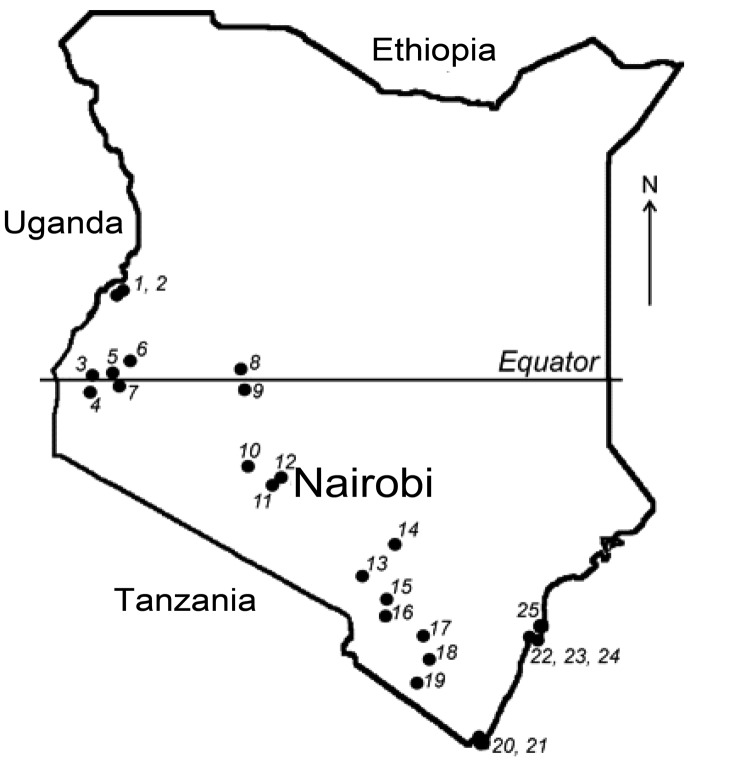
More than 500 bats were collected from 25 locations across Kenya (Figure 1). Sampling sites were chosen on the basis of available information about bat roosts and by using field observations of flying and foraging bats. The number of samples and the collection protocol were justified and approved by the National Museums of Kenya and the Kenyan Wildlife Service. Detailed information on the collection procedure has been described elsewhere (5). Briefly, bats were collected by using hand nets or collected manually in caves and human dwellings and were captured in mist nets around roosts or in locations of nocturnal foraging. Captured bats were anesthetized by an intramuscular injection of ketamine hydrochloride (0.05–0.1 mg/g bodyweight) and euthanized under sedation in compliance with the field protocol approved by CDC’s Institutional Animal Care and Use Committee. The bats were measured, and their sex and species were identified.
Figure 1.
Field sites where bats were collected in Kenya. Numbers identify collection sites (5).
Species were identified phenotypically by using available field keys. Additionally, representative tissues of each collected species were submitted for confirmation to Guelph University (Guelph, Ontario, Canada), where partial sequences of the cytochrome oxidase gene were generated and compared with those available from the database of the Barcode of Life Data Systems (BOLD; www.barcodinglife.org). Because the taxonomy of African bats is under development and reference sequences for several species were unavailable in the BOLD database, the examined sequences were identified to the genus level only.
For microbiologic studies, selected organs and swabs were collected by using sterile plastic tubes. Serum was separated from the blood clots by centrifugation. All samples were transported on dry ice and stored at –80°C until testing.
Culture
Preliminary attempts to culture Bartonella spp. from bat blood showed that the technique developed for isolation of these bacteria from rodent blood (18) is appropriate, with some minor modifications, for processing bat samples. Specifically, we used agar plates supplemented by a 10% addition of rabbit blood. Blood from bats was resuspended in brain–heart infusion (BHI) broth supplemented with 5% amphotericin B. The ratio between blood and BHI was determined to be 1:4. However, this ratio was difficult to adhere to for samples with a limited amount of blood (obtained from small bats). Although dilution of tested blood reduces our ability to detect bacteria in cases with a low level of bacteremia, this approach allowed us to reduce the likelihood that bacterial and fungal contaminants would overgrow the fastidious and slow-growing Bartonella spp. bacteria. Bacterial colonies were presumptively identified as Bartonella spp. on the basis of their morphology and later were confirmed by PCR amplification and sequencing of a specific fragment of the gltA. Subcultures of Bartonella spp. colonies from the original agar plate were streaked onto secondary agar plates, also supplemented by a 10% addition of rabbit blood, and, in case of confluent and pure cultures, harvested and stored in 10% glycerol. Agar plates inoculated with bat blood were incubated at 35°C in an aerobic atmosphere of 5% carbon dioxide for ≤30 days postinoculation, and plates with subcultures were incubated with the same conditions until sufficient growth was observed, usually 5–7 days.
Amplification of the gltA Fragment
A heavy suspension of the microorganisms was heated for 10 min at 95°C followed by 1-min centrifugation at 8,000 rpm to precipitate the lysed cells. The supernant containing the genomic DNA was then moved to a clean centrifuge tube to be examined. PCR amplifications were performed in a 25-μL reaction mixture containing 5 μL 5× Green GoTaq PCR buffer (Promega, Madison, WI, USA), 0.4 μmol of each primer, 200 μM each dNTP, 1 U Taq DNA polymerase (Promega), and ≈20 ng of template DNA. Two oligonucleotides, BhCS781.p (5′-GGGGACCAGCTCATGGTGG-3′) and BhCS1137.n (5′-AATGCAAAAAGAACAGTAAACA-3′) were used as PCR primers to generate a 379-bp amplicon of the Bartonella gltA gene. Positive and negative controls were included with each PCR to evaluate the presence of appropriately sized amplicons and contamination, respectively. Each PCR was performed in a PTC 200 Peltier thermal cycler (MJ Research, Inc., Waltham, MA, USA) or in an Eppendorf Mastercycler Gradient (Eppendorf, Westbury, NY, USA). PCR products were separated by 1.5% agarose gel electrophoresis and visualized by ethidium bromide staining.
Sequencing and Analysis of DNA
PCR products of correct size were purified with the QIAquick PCR Purification Kit (QIAGEN, Valencia, CA, USA) according to manufacturer’s instructions and sequenced in both directions by using an Applied Biosystems Model 3130 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA). Sequencing reactions were performed in a PTC 200 Peltier Thermal cycler by using the same primers as the initial PCR with a concentration of 1.6 µM. Sequences were analyzed by using Lasergene (DNASTAR, Madison, WI, USA) sequence analysis software to determine consensus of sequences for the amplified region of the gltA gene. The Clustal V program within MegAlign (DNASTAR) was used to align and compare homologous Bartonella spp. gltA sequences obtained from bat samples and from the GenBank database.
Results
Bartonella spp. Cultures
Culturing from bat blood pellets, especially from small-sized bats, presented a challenge because of the limited sample volume and the potential for contamination with other bacteria and fungi, problems hard to avoid during field sampling. Consequently, of >500 processed blood samples, the presence or absence of Bartonella spp. cultures have been conclusively evaluated in samples from 331 bats of 13 species from 9 genera. Further estimation of the infection rate was determined exclusively on the basis of the specimens from these 331 animals. The time required for growth of Bartonella spp. colonies on agar greatly varied from 3 days, observed for several samples from Triaenops persicus bats to 28 days in one of the samples from Rousettus aegyptiacus bats.
Bartonella spp. Prevalence
Bartonella isolates were cultured from the blood of 30.2% (106/331) of the bats tested. All isolates were confirmed genetically by amplification and sequence of the gltA. Prevalence of bats positive for Bartonella spp. by culture was as follows: Eidolon helvum (straw-colored fruit bat), 23/88 (26.1%); R. aegyptiacus (Egyptian fruit bat), 22/105 (21.0%); Coleura afra (African sheath-tailed bat), 4/9 (44.4%); T. persicus (Persian trident bat), 7/8 (87.5%); Hipposideros commersoni (giant leaf-nosed bat), 1/4 (25.0%); and Miniopterus spp. (long-fingered bats), 49/87 (56.3%) (Table). Miniopterus spp. and T. persicus bats were at significantly higher risk (p<0.0001 and p<0.001, respectively) for being infected with Bartonella spp. when each was compared with all other bats, and R. aegyptiacus and Epomophorus spp. bats were at significantly lower risk for infection (p<0.01 for both) (Table).
Table. Prevalence of culture-positive test results for Bartonella spp. among bat species, Kenya*.
| Bat species | No. tested | No. (%) positive | Relative risk | p value |
|---|---|---|---|---|
| Miniopterus spp. | 87 | 49 (56) | 2.41 | <0.0001 |
| Eidolon helvum | 88 | 23 (26) | NS | |
| Rousettus aegyptiacus | 105 | 22 (21) | 0.6 | <0.01 |
| Coleura afra | 9 | 4 (44) | NS | |
| Triaenops persicus | 8 | 7 (88) | 2.85 | <0.001 |
| Hipposideros commersoni | 4 | 1 (25) | NS | |
| Epomophorus spp. | 23 | 0 | Not defined† | <0.01 |
| Rhinolophus spp. | 6 | 0 | Not defined† | |
| Chaerephon sp. | 1 | 0 | Not defined† |
*Relative risk was calculated by comparing proportions positive for each bat species compared with that for all other bats. NS, not statistically significant. †Relative risk is <1 but is not defined because of 0 in 1 of the cells.
Genetic Heterogeneity and Sequence Clustering
Sequence analyses of DNA from the 94 Bartonella isolates obtained from bats revealed 58 gltA genotypes (unique sequence variants with ≥1 nucleotide difference) that represented 11 genogroups with a sequence identity value of >96% between genotypes within each group, as proposed by La Scola et al. (28), as a cutoff for this specific gene fragment for species definition within the Bartonella genus. The 11 novel gltA sequences representing each genogroup were submitted to GenBank and assigned accession nos. HM363764–363768 and HM545136–545141.
Association of Bartonella Genotypes with Particular Bat Species
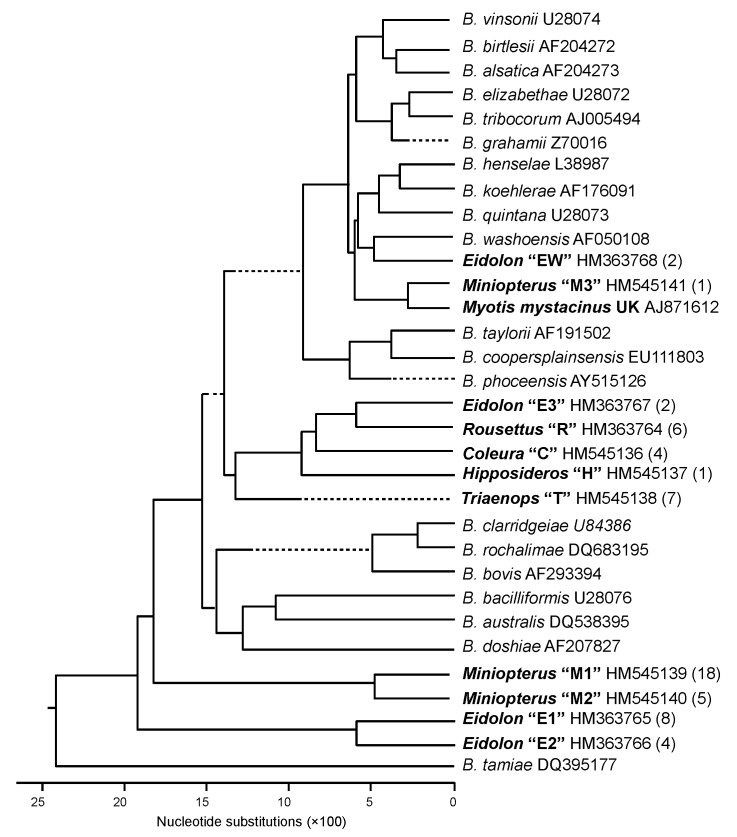
All 15 Bartonella spp. gltA sequences obtained from R. aegyptiacus bats were similar to each other (>96%) and distant from all sequences found in other bat species (<91% identity) and previously described Bartonella species (<84% identity). Six unique genetic variants identified in R. aegyptiacus bats were clustered in a monophyletic genogroup, marked as R in Figure 2.
Figure 2.
Phylogenetic relations among the citrate synthase sequences of Bartonella spp. genotypes detected in bats from Kenya and previously described Bartonella spp. The phylogenetic tree was constructed by the neighbor-joining method. Each Bartonella spp. genogroup detected in bats was provided with the Latin name of the bat genus from which the Bartonella strains were obtained (boldface), the proposed name of genogroup (quotation marks), the GenBank accession number, and the number of genotypes assigned to the genogroup (parentheses).
The level of gltA sequence identity between 16 Bartonella genotypes identified in E. helvum bats varied greatly, ranging from 78.6%–100%. Eight genotypes (sequence variants) were clustered around 4 clades (genogroups), which are marked as E1, E2, E3, and EW in Figure 2. Intergroup identity values ranged from 78.6%–87.2%, much higher values than the sufficient criteria proposed for species demarcation within the Bartonella genus (28).
All 4 gltA genotypes of Bartonella spp. identified in C. afra bats differed slightly from one another but clustered tightly within genogroup C with a sequence identity range of 98.2%–99.7%, compared with a <90% identity between these sequences and Bartonella strains found in other bat species or other sources. Similarly, all 7 gltA sequences identified in T. persicus bats formed a monophyletic genogroup, marked as T in Figure 2, with the identity range of 98.2%–99.7% within the group and <90% identity to any Bartonella sequence outside the group. The only isolate obtained from H. commersoni bats also differed from all described Bartonella strains or sequences (identity <85%).
More complex phylogeny was observed when isolates obtained from bats of the genus Miniopterus were compared by using a similarity between the gltA sequences. In total, 51 Bartonella spp. gltA sequences, obtained from 3 identified Miniopterus spp. (M. africanus, M. minor, and M. natalensis) and from Miniopterus spp. bats that have not been identified at the species level, were analyzed. Among the 51 bats, 24 unique gltA genotypes represented at least 3 genogroups with a level of divergence ranging from 6.6%–18.5%, higher than has been recommended previously for differentiation of Bartonella spp (28). The first genogroup, M1, was composed of 30 sequences obtained from M. minor bats, 1 sequence from M. natalensis bats, and 1 sequence from an unidentified species within the genus Miniopterus. The second genogroup, M2, consisted of 5 sequences from M. minor bats, 2 sequences from M. africanus bats, 1 sequence from M. natalensis bats, and 2 sequences from an unidentified species of Miniopterus spp. bats. Although the second genogroup looked more diverse, an insufficient number of isolates were available to describe an association of specific Bartonella spp. lineages to certain Miniopterus spp. bats. In addition, 1 genotype, M3, from M. natalensis bats, was distant from both groups (identity <90%), but was relatively close (identity 94.2%) to the genotype identified in a whiskered bat (Myotis mystacinus) from the United Kingdom (11).
Discussion
This investigation has resulted in the identification and characterization of Bartonella strains in bats and describes the prevalence and genetic characteristics of Bartonella spp. in bats in Africa. Detection of viable bacteria in a high proportion of bats in ≥6 bat species suggests that Bartonella spp. infection is highly prevalent in bat communities in eastern Africa. Some bat species tested negative for the bacteria; however, we had few samples from these species, making speculation difficult concerning whether those species are truly free from Bartonella spp. infection. However, we identified some bat species that were statistically more likely to be infected with Bartonella spp.
A high prevalence of Bartonella spp. in bats, ranging from 21%–88% in various species, is especially surprising considering the life spans of bats. Numerous investigations have shown that prevalence of Bartonella spp. infection can reach high rates in rodent populations, but rodents usually live no longer than 1–2 years, whereas bats can live for >20 years (6). Explaining such high prevalence is difficult without assuming persistent infection. This hypothesis was not confirmed for some rodent species, specifically for cotton rats; a high prevalence of Bartonella spp. in that population could be explained by replacement of diverse Bartonella strains sequentially colonizing an individual rat rather than by a long-term bacteremia (29). This scenario is unlikely in bats because they have much longer lives than rodents.
The lifestyle of bats, such as the colonial structure of their populations, close physical contact, aggressive interactions, and typically heavy ectoparasite infestations also might contribute to the frequent transmission of Bartonella spp. among individual animals. All Bartonella spp. are widely regarded as vector-transmitted agents, and diverse arthropods, such as sandflies, lice, fleas, ticks, and mites, have been implicated as potential vectors (30). Bats carry a wide range of ectoparasites, including bat flies, fleas, soft ticks, and mites, some of which are highly specific to bats (31). Limited information is available to suggest that alternative mechanisms beyond vector transmission may be responsible for the spread of Bartonella spp. infections among animals. For example, identification of viable Bartonella spp. bacteria in the blood of cotton rat embryos and neonates indicated a possibility of vertical transmission from a pregnant female to offspring (32). Detection of Bartonella spp. DNA in the saliva of dogs suggests a potential possibility of transmission through biting (33), and transmission of Bartonella henselae from cats to humans by cat scratch is well documented (24).
Comparative analyses of the gltA sequences obtained from Bartonella spp. cultures showed that bats in Kenya harbor a diverse assemblage of Bartonella strains, some of which appear to represent distinct species. Although we used only a portion of the citrate synthase gene for phylogenetic analysis, this gene has been shown to be a reliable tool for distinguishing between closely related Bartonella genotypes (28,34). By using this gene, we were able to compare the variety of Bartonella genotypes isolated from bats with homologous sequences of Bartonella strains found in other mammals. Finding considerable sequence diversity is typical for different species of Bartonella, although more characteristics are needed to describe novel Bartonella species.
Evidence for cospeciation of Bartonella spp. with natural hosts varies among studies and around the world. A strong association exists between specific Bartonella species and their mammalian hosts, whereas some observations have indicated that 1 species of Bartonella can infect a variety of rodent species at a given site in Europe (35). Our investigation indicated a definite host-specificity for Bartonella strains in bat species. All isolates obtained from R. aegyptiacus, C. afra, and T. persicus bats clearly belonged to the specific Bartonella spp. group found exclusively in the particular bat species. By contrast, straw-colored fruit bats (E. helvum) and long-fingered bats (Miniopterus spp.) harbored strains clustered around 3 and 4 different groups of Bartonella spp., respectively, based on their gltA identity. Nevertheless, all strains of Bartonella spp. recovered from E. helvum bats were typical for this species of bats only. This pattern of cospeciation resembles the Bartonella spp.–host relations observed in cotton rats (18). Similarly, the gltA sequences from all strains obtained from Miniopterus spp. bats have not been found in bats of other bat genera. More investigations of Bartonella spp. in diverse bat species are required to test this hypothesis.
No evidence is available to suggest whether novel strains of Bartonella spp. found in bats from Kenya cause human illness. However, relevant surveillance in Kenya and other African countries has not been implemented. The significance of African bats in public and veterinary health is not understood because of a lack of surveillance. Bats are known as principal reservoir hosts of lyssaviruses. In Africa, these include Lagos bat virus, which circulates in pteropid bats (including E. helvum and R. aegyptiacus), and Duvenhage virus, which circulates in insectivorous bats (although specific reservoirs have not been established). In addition, Shimoni bat virus was identified recently in an insectivorous bat Hipposideros commersoni (32). Furthermore, multiple species of African bats have been shown to harbor coronaviruses (33). Nipah virus was identified in straw-colored fruit bats E. helvum (34), and Marburg virus was identified in tomb bats R. aegyptiacus (35). Circulation patterns of these agents in bat populations have not been sufficiently studied.
Characterization of the isolates obtained from bats and comparison with those obtained from human cases associated with Bartonella spp. of unknown origin (e.g., B. tamiae) can be helpful in the search for potential reservoirs. The reagents prepared from bat isolates of Bartonella spp. can be used for serologic surveys conducted in high-risk areas of the world. Application of antigens produced from bat-derived strains is especially relevant to serologic investigations of cases of infectious disease of unknown origin among persons occupationally exposed to bats. In addition, cocirculation of Bartonella spp. with other pathogens in a bat population can affect the bat’s ecology and pathobiology. These aspects need further investigation.
Biography
Dr Kosoy is the chief of the Bartonella Laboratory, Bacterial Diseases Branch, Division of Vector Borne Diseases, Centers for Disease Control and Prevention. His research interests include ecology of zoonotic infectious diseases and population biology of bacterial pathogens.
Footnotes
Suggested citation for this article: Kosoy M, Bai Y, Lynch T, Kuzmin IV, Niezgoda M, Franka R, et al. Bartonella spp. in bats, Kenya. Emerg Infect Dis [serial on the Internet]. 2010 Dec [date cited]. Available from http://dx.doi.org/10.3201/eid1612.100601
References
- 1.Halpin K, Young PL, Field HE, Mackenzie JS. Isolation of Hendra virus from pteropid bats: a natural reservoir of Hendra virus. J Gen Virol. 2000;81:1927–32. [DOI] [PubMed] [Google Scholar]
- 2.Li W, Shi Z, Yu M, Ren W, Smith C, Epstein JH, et al. Bats are natural reservoirs of SARS-like coronaviruses. Science. 2005;310:676–9. 10.1126/science.1118391 [DOI] [PubMed] [Google Scholar]
- 3.Wang LF, Shi Z, Zhang S, Field H, Daszak P, Eaton BT. Review of bats and SARS. Emerg Infect Dis. 2006;12:1834–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Williams CJ. Bats as the reservoir for outbreaks of emerging infectious diseases. Euro Surveill. 2005;10(11):E051110.4. PMID: 16794279 [DOI] [PubMed]
- 5.Kuzmin IV, Niezgoda M, Franka R, Agwanda B, Markotter W, Beagley JC, et al. Lagos bat virus in Kenya. J Clin Microbiol. 2008;46:1451–61. 10.1128/JCM.00016-08 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Calisher CH, Childs JE, Field HE, Holmes KV, Schountz T. Bats: important reservoir hosts of emerging viruses. Clin Microbiol Rev. 2006;19:531–45. 10.1128/CMR.00017-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bennett M. Bats and human emerging diseases. Epidemiol Infect. 2006;134:905–7. 10.1017/S0950268806006674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.van der Poel WH, Lina PH, Kramps JA. Public health awareness of emerging zoonotic viruses of bats: a European perspective. Vector Borne Zoonotic Dis. 2006;6:315–24. 10.1089/vbz.2006.6.315 [DOI] [PubMed] [Google Scholar]
- 9.Ghatak S, Banerjee R, Agarwal RK, Kapoor KN. Zoonoses and bats: a look from human health viewpoint. J Commun Dis. 2000;32:40–8. [PubMed] [Google Scholar]
- 10.Evans NJ, Bown K, Timofte D, Simpson VR, Birtles RJ. Fatal borreliosis in bat caused by relapsing fever spirochete, United Kingdom. Emerg Infect Dis. 2009;15:1331–3. 10.3201/eid1508.090475 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Concannon R, Wynn-Owen K, Simpson VR, Birtles RJ. Molecular characterization of haemoparasites infecting bats (Microchiroptera) in Cornwall, UK. Parasitology. 2005;131:489–96. 10.1017/S0031182005008097 [DOI] [PubMed] [Google Scholar]
- 12.Loftis AD, Gill JS, Schriefer ME, Levin ML, Eremeeva ME, Gilchrist MJ, et al. Detection of Rickettsia, Borrelia, and Bartonella in Carios kelleyi (Acari: Argasidae). J Med Entomol. 2005;42:473–80. 10.1603/0022-2585(2005)042[0473:DORBAB]2.0.CO;2 [DOI] [PubMed] [Google Scholar]
- 13.Reeves WK, Loftis AD, Gore JA, Dasch GA. Molecular evidence for novel Bartonella species in Trichobius major (Diptera: Streblidae) and Cimex adjunctus (Hemiptera: Cimicidae) from two southeastern bat caves. U.S.A. J Vector Ecol. 2005;30:339–41. [PubMed] [Google Scholar]
- 14.Reeves WK, Rogers TE, Durden LA, Dasch GA. Association of Bartonella with the fleas (Siphonaptera) of rodents and bats using molecular techniques. J Vector Ecol. 2007;32:118–22. 10.3376/1081-1710(2007)32[118:AOBWTF]2.0.CO;2 [DOI] [PubMed] [Google Scholar]
- 15.Birtles RJ. Bartonellae as elegant hemotropic parasites. Ann N Y Acad Sci. 2005;1063:270–9. 10.1196/annals.1355.044 [DOI] [PubMed] [Google Scholar]
- 16.Dehio C. Bartonella interactions with endothelial cells and erythrocytes. Trends Microbiol. 2001;9:279–85. 10.1016/S0966-842X(01)02047-9 [DOI] [PubMed] [Google Scholar]
- 17.Chomel BB, Boulouis HJ, Breitschwerdt EB, Kasten RW, Vayssier-Taussat M, Birtles RJ, et al. Ecological fitness and strategies of adaptation of Bartonella species to their hosts and vectors. Vet Res. 2009;40:29. 10.1051/vetres/2009011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kosoy M, Regnery R, Tzianabos T, Marston E, Jones D, Green D, et al. Distribution, diversity and host specificity of Bartonella in rodents from the southeastern United States. Am J Trop Med Hyg. 1997;57:578–88. [DOI] [PubMed] [Google Scholar]
- 19.Anderson B, Neuman MA. Bartonella spp. as emerging human pathogens. Clin Microbiol Rev. 1997;10:203–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Daly JS, Worthington MG, Brenner DJ, Moss CW, Hollis DG, Weyant RS, et al. Rochalimaea elizabethae sp. nov. isolated from a patient with endocarditis. J Clin Microbiol. 1993;31:872–81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Welch DF, Carroll KC, Hofmeister EK, Persing DH, Robison DA, Steigerwalt AG, et al. Isolation of a new subspecies, Bartonella vinsonii subsp. arupensis, from a cattle rancher: identity with isolates found in conjunction with Borrelia burgdorferi and Babesia microti among naturally infected mice. J Clin Microbiol. 1999;37:2598–601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kerkhoff FT, Bergmans AM, van Der Zee A, Rothova A. Demonstration of Bartonella grahamii DNA in ocular fluids of a patient with neuroretinitis. J Clin Microbiol. 1999;37:4034–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Kosoy M, Murray M, Gilmore RD, Bai Y, Gage KL. Bartonella strains from ground squirrels are identical to Bartonella washoensis isolated from a human patient. J Clin Microbiol. 2003;41:645–50. 10.1128/JCM.41.2.645-650.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Breitschwerdt EB, Kordick DL. Bartonella infection in animals: carriership, reservoir potential, pathogenicity, and zoonotic potential for human infection. Clin Microbiol Rev. 2000;13:428–38. 10.1128/CMR.13.3.428-438.2000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Roux V, Eykyn SJ, Wyllie S, Raoult D. Bartonella vinsonii subsp. berkhoffii as an agent of afebrile blood culture–negative endocarditis in a human. J Clin Microbiol. 2000;38:1698–700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Eremeeva ME, Gerns HL, Lydy SL, Goo JS, Ryan ET, Mathew SS, et al. Bacteremia, fever, and splenomegaly caused by a newly recognized Bartonella species. N Engl J Med. 2007;356:2381–7. 10.1056/NEJMoa065987 [DOI] [PubMed] [Google Scholar]
- 27.Kosoy M, Morway C, Sheff K, Bai Y, Colborn J, Chalcraft L, et al. Bartonella tamiae sp. nov., a newly recognized pathogen isolated from three human patients from Thailand. J Clin Microbiol. 2008;46:772–5. 10.1128/JCM.02120-07 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.La Scola B, Zeaiter Z, Khamis A, Raoult D. Gene-sequence-based criteria for species definition in bacteriology: the Bartonella paradigm. Trends Microbiol. 2003;11:318–21. 10.1016/S0966-842X(03)00143-4 [DOI] [PubMed] [Google Scholar]
- 29.Kosoy M, Mandel E, Green D, Marston E, Jones D, Childs J. Prospective studies of Bartonella of rodents. Part II. Diverse infections in a single rodent community. Vector Borne Zoonotic Dis. 2004;4:296–305. 10.1089/vbz.2004.4.296 [DOI] [PubMed] [Google Scholar]
- 30.Billeter SA, Levy MG, Chomel BB, Breitschwerdt EB. Vector transmission of Bartonella species with emphasis on the potential for tick transmission. Med Vet Entomol. 2008;22:1–15. 10.1111/j.1365-2915.2008.00713.x [DOI] [PubMed] [Google Scholar]
- 31.Dick CW, Patterson BD. Bat flies: obligate ectoparasites of bats. Micromammals and macroparasites: from evolutionary ecology to management. Morand S, et al. (Eds). Springer-Verlag, Tokyo, Japan, 2006;179–94. [Google Scholar]
- 32.Kuzmin IV, Mayer AE, Niezgoda M, Markotter W, Agwanda B, Breiman RF, et al. Shimoni bat virus, a new representative of the Lyssavirus genus. Virus Res. 2010;149:197–210. 10.1016/j.virusres.2010.01.018 [DOI] [PubMed] [Google Scholar]
- 33.Tong S, Conrardy C, Ruone S, Kuzmin IV, Guo X, Tao Y, et al. Detection of novel SARS-like and other coronaviruses in bats from Kenya. Emerg Infect Dis. 2009;15:482–5. 10.3201/eid1503.081013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Drexler JF, Corman VM, Gloza-Rausch F, Seebens A, Annan A, Ipsen A, et al. Henipavirus RNA in African bats. PLoS ONE. 2009;4:e6367. 10.1371/journal.pone.0006367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Towner JS, Amman BR, Sealy TK, Carroll SA, Comer JA, Kemp A, et al. Isolation of genetically diverse Marburg viruses from Egyptian fruit bats. PLoS Pathog. 2009;5:e1000536. 10.1371/journal.ppat.1000536 [DOI] [PMC free article] [PubMed] [Google Scholar]