Abstract
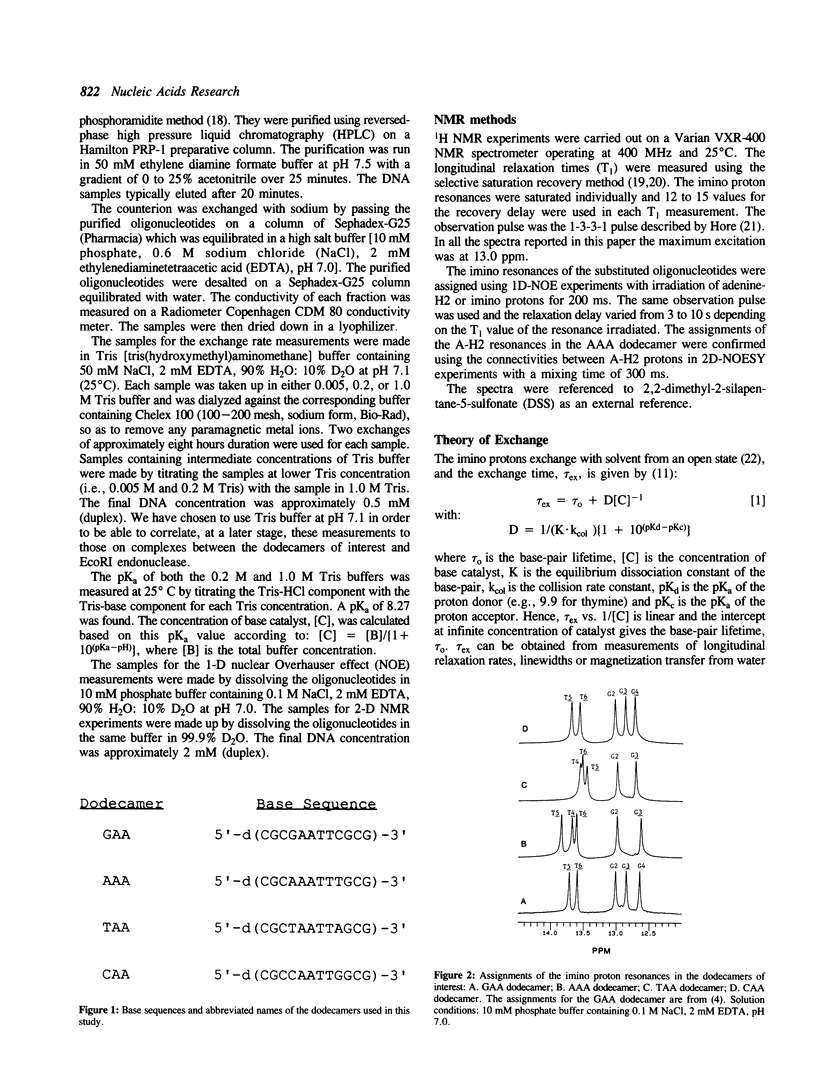
We have used nuclear magnetic resonance (NMR) spectroscopy to measure the lifetimes of individual base-pairs in the palindromic DNA oligonucleotide 5'-d(CGCGAATTCGCG)-3' and in three other dodecamers with symmetrical base substitutions in the sites underlined. The resonances of the hydrogen-bonded imino protons in each of the substituted oligomers in the duplex form have been assigned using one dimensional nuclear Overhauser effect (1-D NOE) experiments. The lifetimes have been obtained from the dependence of selective longitudinal relaxation times and linewidths of the imino proton resonances on the concentration of base catalyst (Tris) at 25 degrees C and in the presence of 50 mM NaCl. The lifetimes of the central A.T base-pairs have been found to depend on base sequence. They are greatly increased in the dodecamer 5'-d(CGCAAATTTGCG)-3' which contains an A3T3 tract. The lifetimes of the central A.T base-pairs in 5'-d(CGCGAATTCGCG)-3', 5'-d(CGCTAATTAGCG)-3' and 5'-d(CGCCAATTGGCG)-3' are comparable. In all dodecamers, the lifetime of the A.T base-pair at the 5'-end of the AnTn tract is the shortest. The anomalous opening kinetics of the A.T base-pairs can be correlated to the bending properties of the corresponding sequences.
Full text
PDF




Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Benevides J. M., Wang A. H., van der Marel G. A., van Boom J. H., Thomas G. J., Jr Crystal and solution structures of the B-DNA dodecamer d(CGCAAATTTGCG) probed by Raman spectroscopy: heterogeneity in the crystal structure does not persist in the solution structure. Biochemistry. 1988 Feb 9;27(3):931–938. doi: 10.1021/bi00403a014. [DOI] [PubMed] [Google Scholar]
- Braunlin W. H., Bloomfield V. A. 1H NMR study of the base-pairing reactions of d(GGAATTCC): salt and polyamine effects on the imino proton exchange. Biochemistry. 1988 Feb 23;27(4):1184–1191. doi: 10.1021/bi00404a018. [DOI] [PubMed] [Google Scholar]
- Caruthers M. H., Beaucage S. L., Becker C., Efcavitch J. W., Fisher E. F., Galluppi G., Goldman R., deHaseth P., Matteucci M., McBride L. Deoxyoligonucleotide synthesis via the phosphoramidite method. Gene Amplif Anal. 1983;3:1–26. [PubMed] [Google Scholar]
- Coll M., Frederick C. A., Wang A. H., Rich A. A bifurcated hydrogen-bonded conformation in the d(A.T) base pairs of the DNA dodecamer d(CGCAAATTTGCG) and its complex with distamycin. Proc Natl Acad Sci U S A. 1987 Dec;84(23):8385–8389. doi: 10.1073/pnas.84.23.8385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickerson R. E., Drew H. R. Structure of a B-DNA dodecamer. II. Influence of base sequence on helix structure. J Mol Biol. 1981 Jul 15;149(4):761–786. doi: 10.1016/0022-2836(81)90357-0. [DOI] [PubMed] [Google Scholar]
- Diekmann S., McLaughlin L. W. DNA curvature in native and modified EcoRI recognition sites and possible influence upon the endonuclease cleavage reaction. J Mol Biol. 1988 Aug 20;202(4):823–834. doi: 10.1016/0022-2836(88)90561-x. [DOI] [PubMed] [Google Scholar]
- Drew H. R., Wing R. M., Takano T., Broka C., Tanaka S., Itakura K., Dickerson R. E. Structure of a B-DNA dodecamer: conformation and dynamics. Proc Natl Acad Sci U S A. 1981 Apr;78(4):2179–2183. doi: 10.1073/pnas.78.4.2179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Englander S. W., Downer N. W., Teitelbaum H. Hydrogen exchange. Annu Rev Biochem. 1972;41:903–924. doi: 10.1146/annurev.bi.41.070172.004351. [DOI] [PubMed] [Google Scholar]
- Fratini A. V., Kopka M. L., Drew H. R., Dickerson R. E. Reversible bending and helix geometry in a B-DNA dodecamer: CGCGAATTBrCGCG. J Biol Chem. 1982 Dec 25;257(24):14686–14707. [PubMed] [Google Scholar]
- Frederick C. A., Grable J., Melia M., Samudzi C., Jen-Jacobson L., Wang B. C., Greene P., Boyer H. W., Rosenberg J. M. Kinked DNA in crystalline complex with EcoRI endonuclease. Nature. 1984 May 24;309(5966):327–331. doi: 10.1038/309327a0. [DOI] [PubMed] [Google Scholar]
- Guéron M., Kochoyan M., Leroy J. L. A single mode of DNA base-pair opening drives imino proton exchange. Nature. 1987 Jul 2;328(6125):89–92. doi: 10.1038/328089a0. [DOI] [PubMed] [Google Scholar]
- Hare D. R., Wemmer D. E., Chou S. H., Drobny G., Reid B. R. Assignment of the non-exchangeable proton resonances of d(C-G-C-G-A-A-T-T-C-G-C-G) using two-dimensional nuclear magnetic resonance methods. J Mol Biol. 1983 Dec 15;171(3):319–336. doi: 10.1016/0022-2836(83)90096-7. [DOI] [PubMed] [Google Scholar]
- Johnston P. D., Redfield A. G. An NMR study of the exchange rates for protons involved in the secondary and tertiary structure of yeast tRNA Phe. Nucleic Acids Res. 1977 Oct;4(10):3599–3615. doi: 10.1093/nar/4.10.3599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kochoyan M., Lancelot G., Leroy J. L. Study of structure, base-pair opening kinetics and proton exchange mechanism of the d-(AATTGCAATT) self-complementary oligodeoxynucleotide in solution. Nucleic Acids Res. 1988 Aug 11;16(15):7685–7702. doi: 10.1093/nar/16.15.7685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kochoyan M., Leroy J. L., Guéron M. Proton exchange and base-pair lifetimes in a deoxy-duplex containing a purine-pyrimidine step and in the duplex of inverse sequence. J Mol Biol. 1987 Aug 5;196(3):599–609. doi: 10.1016/0022-2836(87)90036-2. [DOI] [PubMed] [Google Scholar]
- Leroy J. L., Charretier E., Kochoyan M., Guéron M. Evidence from base-pair kinetics for two types of adenine tract structures in solution: their relation to DNA curvature. Biochemistry. 1988 Dec 13;27(25):8894–8898. doi: 10.1021/bi00425a004. [DOI] [PubMed] [Google Scholar]
- Leroy J. L., Kochoyan M., Huynh-Dinh T., Guéron M. Characterization of base-pair opening in deoxynucleotide duplexes using catalyzed exchange of the imino proton. J Mol Biol. 1988 Mar 20;200(2):223–238. doi: 10.1016/0022-2836(88)90236-7. [DOI] [PubMed] [Google Scholar]
- Lycksell P. O., Gräslund A., Claesens F., McLaughlin L. W., Larsson U., Rigler R. Base pair opening dynamics of a 2-aminopurine substituted Eco RI restriction sequence and its unsubstituted counterpart in oligonucleotides. Nucleic Acids Res. 1987 Nov 11;15(21):9011–9025. doi: 10.1093/nar/15.21.9011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClarin J. A., Frederick C. A., Wang B. C., Greene P., Boyer H. W., Grable J., Rosenberg J. M. Structure of the DNA-Eco RI endonuclease recognition complex at 3 A resolution. Science. 1986 Dec 19;234(4783):1526–1541. doi: 10.1126/science.3024321. [DOI] [PubMed] [Google Scholar]
- Nadeau J. G., Crothers D. M. Structural basis for DNA bending. Proc Natl Acad Sci U S A. 1989 Apr;86(8):2622–2626. doi: 10.1073/pnas.86.8.2622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson H. C., Finch J. T., Luisi B. F., Klug A. The structure of an oligo(dA).oligo(dT) tract and its biological implications. Nature. 1987 Nov 19;330(6145):221–226. doi: 10.1038/330221a0. [DOI] [PubMed] [Google Scholar]
- Pardi A., Morden K. M., Patel D. J., Tinoco I., Jr Kinetics for exchange of imino protons in the d(C-G-C-G-A-A-T-T-C-G-C-G) double helix and in two similar helices that contain a G . T base pair, d(C-G-T-G-A-A-T-T-C-G-C-G), and an extra adenine, d(C-G-C-A-G-A-A-T-T-C-G-C-G). Biochemistry. 1982 Dec 7;21(25):6567–6574. doi: 10.1021/bi00268a038. [DOI] [PubMed] [Google Scholar]
- Pardi A., Tinoco I., Jr Kinetics for exchange of imino protons in deoxyribonucleic acid, ribonucleic acid, and hybrid oligonucleotide helices. Biochemistry. 1982 Sep 14;21(19):4686–4693. doi: 10.1021/bi00262a026. [DOI] [PubMed] [Google Scholar]
- Patel D. J., Kozlowski S. A., Marky L. A., Broka C., Rice J. A., Itakura K., Breslauer K. J. Premelting and melting transitions in the d(CGCGAATTCGCG) self-complementary duplex in solution. Biochemistry. 1982 Feb 2;21(3):428–436. doi: 10.1021/bi00532a002. [DOI] [PubMed] [Google Scholar]
- Patel D. J., Shapiro L., Hare D. Nuclear magnetic resonance and distance geometry studies of DNA structures in solution. Annu Rev Biophys Biophys Chem. 1987;16:423–454. doi: 10.1146/annurev.bb.16.060187.002231. [DOI] [PubMed] [Google Scholar]
- Rosenberg J. M., Greene P. Eco RI* specificity and hydrogen bonding. DNA. 1982;1(2):117–124. doi: 10.1089/dna.1.1982.1.117. [DOI] [PubMed] [Google Scholar]
- Sarma M. H., Gupta G., Sarma R. H. Structure of a bent DNA: two-dimensional NMR studies on d(GAAAATTTTC)2. Biochemistry. 1988 May 3;27(9):3423–3432. doi: 10.1021/bi00409a045. [DOI] [PubMed] [Google Scholar]
- Yoon C., Privé G. G., Goodsell D. S., Dickerson R. E. Structure of an alternating-B DNA helix and its relationship to A-tract DNA. Proc Natl Acad Sci U S A. 1988 Sep;85(17):6332–6336. doi: 10.1073/pnas.85.17.6332. [DOI] [PMC free article] [PubMed] [Google Scholar]



