Abstract
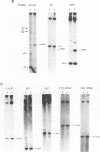
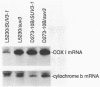
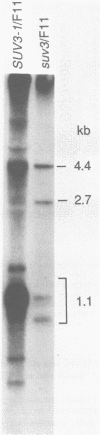
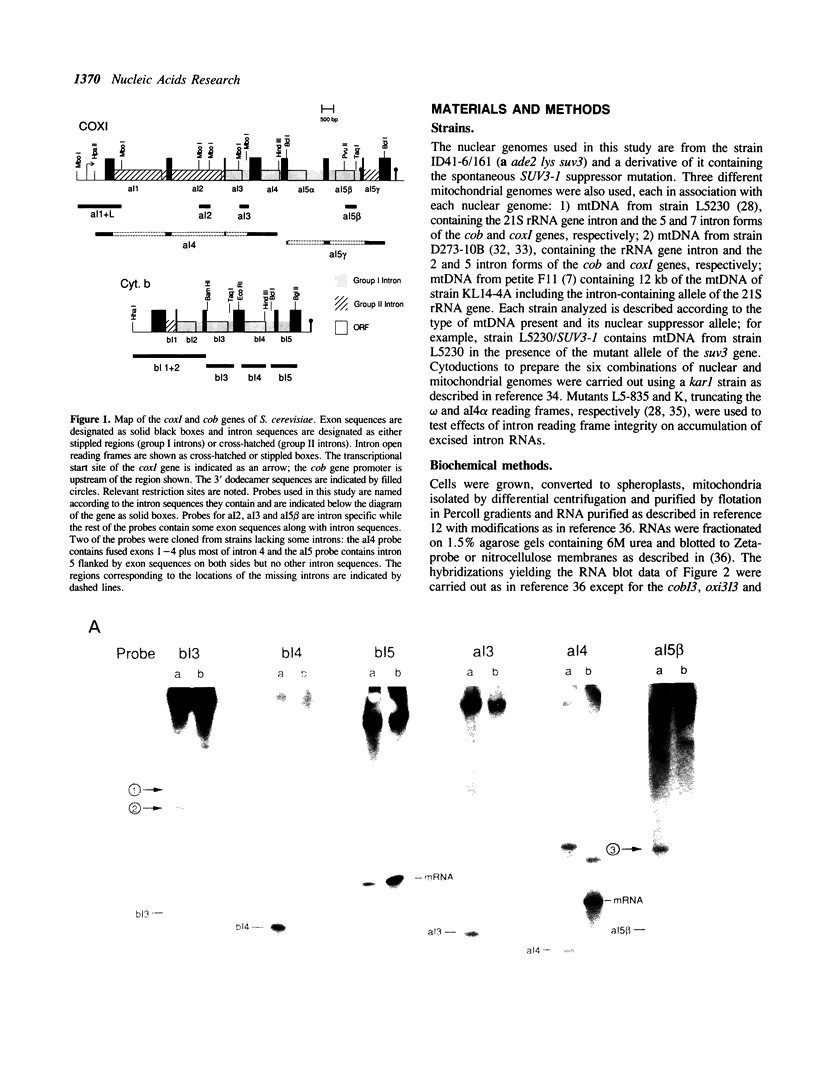
The SUV3-1 mutation was isolated earlier as a suppressor of a deletion of a conserved RNA processing site (dodecamer) near the 3' end of the var1 gene. Previous studies indicate that the suppressor enhances translation of mutant var1 messages; unexpectedly, it also causes over-accumulation of excised intron RNA of the large rRNA gene intron and blocks cleavage at the dodecamer site within that intron. In this study most mitochondrial genes in SUV3-1 and suv3 nuclear contexts are surveyed for changes in levels of mRNA, for interference with dodecamer cleavage and splicing and for levels of excised intron RNAs. SUV3-1 has little or no effect on the size or abundance of unspliced RNAs tested. It results, however, in a marked increase in the abundance of seven of eight excised group I intron RNAs tested, most of which are not detectable in wild-type (suv3) strains. The suppressor lowers levels of the cob and coxl mRNAs about 2-5 and 20-fold, respectively. The effect on coxl mRNA results from a decrease in the splicing of its intron 5 beta. Despite the reduction in these mRNA levels, the amounts of coxl and cyt b polypeptides were close to wild-type levels in SUV3-1 cells. These data show that the suv3 gene plays a prominent role in post-transcriptional and translation events in yeast mitochondria.
Full text
PDF
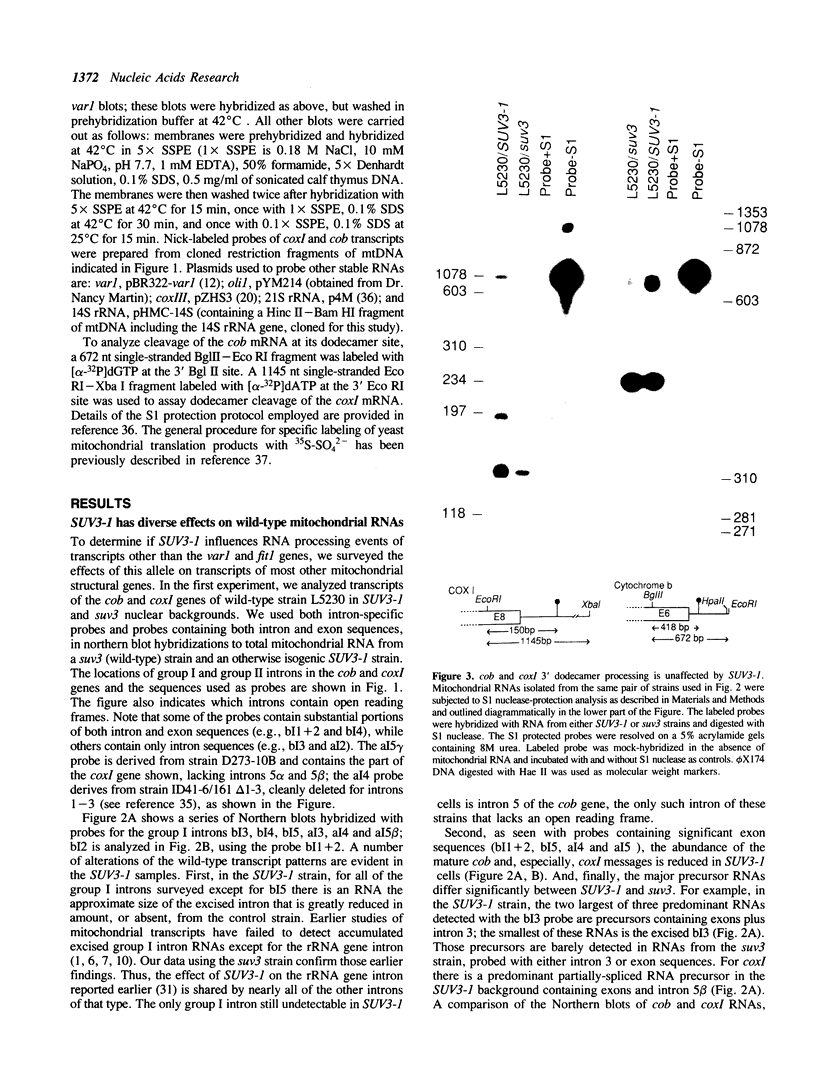
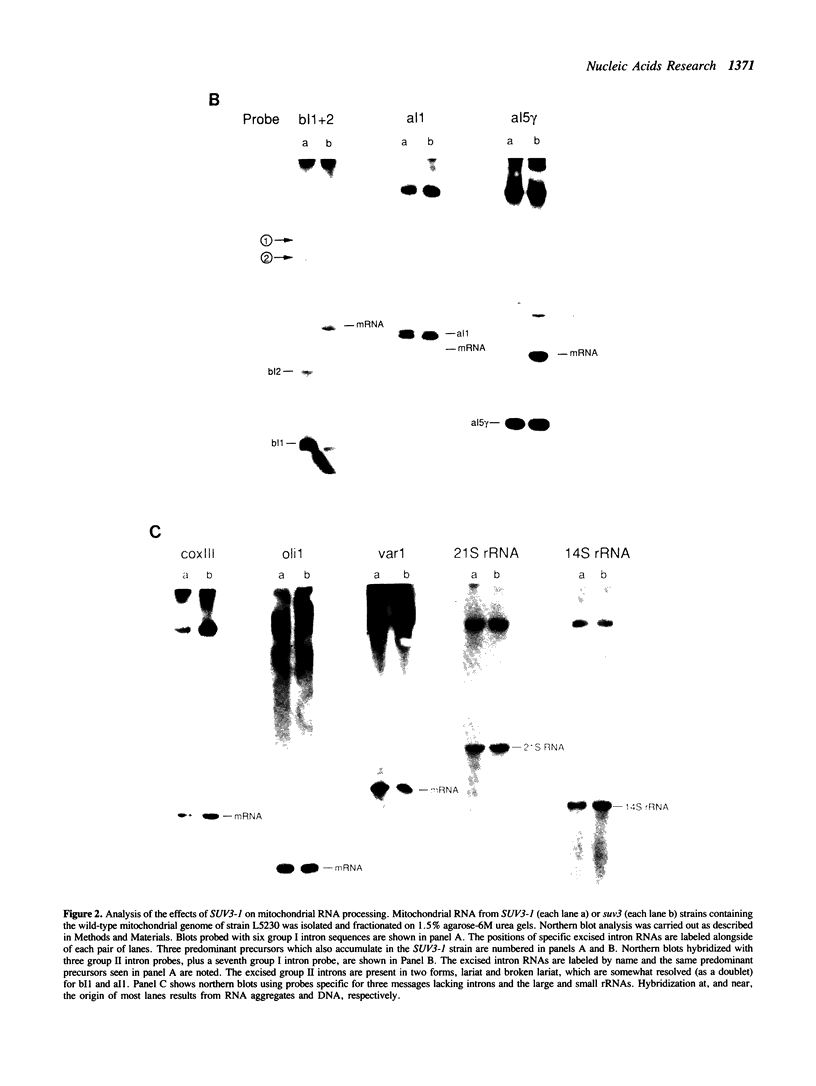





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arnberg A. C., Van Ommen G. J., Grivell L. A., Van Bruggen E. F., Borst P. Some yeast mitochondrial RNAs are circular. Cell. 1980 Feb;19(2):313–319. doi: 10.1016/0092-8674(80)90505-x. [DOI] [PubMed] [Google Scholar]
- Bonitz S. G., Coruzzi G., Thalenfeld B. E., Tzagoloff A., Macino G. Assembly of the mitochondrial membrane system. Structure and nucleotide sequence of the gene coding for subunit 1 of yeast cytochrme oxidase. J Biol Chem. 1980 Dec 25;255(24):11927–11941. [PubMed] [Google Scholar]
- Bonitz S. G., Homison G., Thalenfeld B. E., Tzagoloff A., Nobrega F. G. Assembly of the mitochondrial membrane system. Processing of the apocytochrome b precursor RNAs in Saccharomyces cerevisiae D273-10B. J Biol Chem. 1982 Jun 10;257(11):6268–6274. [PubMed] [Google Scholar]
- Cheng M. Y., Hartl F. U., Martin J., Pollock R. A., Kalousek F., Neupert W., Hallberg E. M., Hallberg R. L., Horwich A. L. Mitochondrial heat-shock protein hsp60 is essential for assembly of proteins imported into yeast mitochondria. Nature. 1989 Feb 16;337(6208):620–625. doi: 10.1038/337620a0. [DOI] [PubMed] [Google Scholar]
- Christianson T., Rabinowitz M. Identification of multiple transcriptional initiation sites on the yeast mitochondrial genome by in vitro capping with guanylyltransferase. J Biol Chem. 1983 Nov 25;258(22):14025–14033. [PubMed] [Google Scholar]
- Colleaux L., d'Auriol L., Betermier M., Cottarel G., Jacquier A., Galibert F., Dujon B. Universal code equivalent of a yeast mitochondrial intron reading frame is expressed into E. coli as a specific double strand endonuclease. Cell. 1986 Feb 28;44(4):521–533. doi: 10.1016/0092-8674(86)90262-x. [DOI] [PubMed] [Google Scholar]
- Costanzo M. C., Fox T. D. Product of Saccharomyces cerevisiae nuclear gene PET494 activates translation of a specific mitochondrial mRNA. Mol Cell Biol. 1986 Nov;6(11):3694–3703. doi: 10.1128/mcb.6.11.3694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Costanzo M. C., Fox T. D. Specific translational activation by nuclear gene products occurs in the 5' untranslated leader of a yeast mitochondrial mRNA. Proc Natl Acad Sci U S A. 1988 Apr;85(8):2677–2681. doi: 10.1073/pnas.85.8.2677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delahodde A., Goguel V., Becam A. M., Creusot F., Perea J., Banroques J., Jacq C. Site-specific DNA endonuclease and RNA maturase activities of two homologous intron-encoded proteins from yeast mitochondria. Cell. 1989 Feb 10;56(3):431–441. doi: 10.1016/0092-8674(89)90246-8. [DOI] [PubMed] [Google Scholar]
- Dieckmann C. L., Koerner T. J., Tzagoloff A. Assembly of the mitochondrial membrane system. CBP1, a yeast nuclear gene involved in 5' end processing of cytochrome b pre-mRNA. J Biol Chem. 1984 Apr 25;259(8):4722–4731. [PubMed] [Google Scholar]
- Dieckmann C. L., Pape L. K., Tzagoloff A. Identification and cloning of a yeast nuclear gene (CBP1) involved in expression of mitochondrial cytochrome b. Proc Natl Acad Sci U S A. 1982 Mar;79(6):1805–1809. doi: 10.1073/pnas.79.6.1805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Douglas M., Finkelstein D., Butow R. A. Analysis of products of mitochondrial protein synthesis in yeast: genetic and biochemical aspects. Methods Enzymol. 1979;56:58–66. doi: 10.1016/0076-6879(79)56009-1. [DOI] [PubMed] [Google Scholar]
- Farrelly F., Zassenhaus H. P., Butow R. A. Characterization of transcripts from the Var1 region on mitochondrial DNA of Saccharomyces cerevisiae. J Biol Chem. 1982 Jun 10;257(11):6581–6587. [PubMed] [Google Scholar]
- Frontali L., Palleschi C., Francisci S. Transcripts of mitochondrial tRNA genes in Saccharomyces cerevisiae. Nucleic Acids Res. 1982 Nov 25;10(22):7283–7293. doi: 10.1093/nar/10.22.7283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halbreich A., Pajot P., Foucher M., Grandchamp C., Slonimski P. A pathway of cytochrome b mRNA processing in yeast mitochondria: specific splicing steps and an intron-derived circular DNA. Cell. 1980 Feb;19(2):321–329. doi: 10.1016/0092-8674(80)90506-1. [DOI] [PubMed] [Google Scholar]
- Hensgens L. A., Arnberg A. C., Roosendaal E., van der Horst G., van der Veen R., van Ommen G. J., Grivell L. A. Variation, transcription and circular RNAs of the mitochondrial gene for subunit I of cytochrome c oxidase. J Mol Biol. 1983 Feb 15;164(1):35–58. doi: 10.1016/0022-2836(83)90086-4. [DOI] [PubMed] [Google Scholar]
- Hensgens L. A., Bonen L., de Haan M., van der Horst G., Grivell L. A. Two intron sequences in yeast mitochondrial COX1 gene: homology among URF-containing introns and strain-dependent variation in flanking exons. Cell. 1983 Feb;32(2):379–389. doi: 10.1016/0092-8674(83)90457-9. [DOI] [PubMed] [Google Scholar]
- Jacquier A., Dujon B. An intron-encoded protein is active in a gene conversion process that spreads an intron into a mitochondrial gene. Cell. 1985 Jun;41(2):383–394. doi: 10.1016/s0092-8674(85)80011-8. [DOI] [PubMed] [Google Scholar]
- Kreike J., Schulze M., Pillar T., Körte A., Rödel G. Cloning of a nuclear gene MRS1 involved in the excision of a single group I intron (bI3) from the mitochondrial COB transcript in S. cerevisiae. Curr Genet. 1986;11(3):185–191. doi: 10.1007/BF00420605. [DOI] [PubMed] [Google Scholar]
- Labouesse M., Herbert C. J., Dujardin G., Slonimski P. P. Three suppressor mutations which cure a mitochondrial RNA maturase deficiency occur at the same codon in the open reading frame of the nuclear NAM2 gene. EMBO J. 1987 Mar;6(3):713–721. doi: 10.1002/j.1460-2075.1987.tb04812.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Locker J., Rabinowitz M. Transcription in yeast mitochondria: analysis of the 21 S rRNA region and its transcripts. Plasmid. 1981 Nov;6(3):302–314. doi: 10.1016/0147-619x(81)90038-x. [DOI] [PubMed] [Google Scholar]
- Macreadie I. G., Scott R. M., Zinn A. R., Butow R. A. Transposition of an intron in yeast mitochondria requires a protein encoded by that intron. Cell. 1985 Jun;41(2):395–402. doi: 10.1016/s0092-8674(85)80012-x. [DOI] [PubMed] [Google Scholar]
- McGraw P., Tzagoloff A. Assembly of the mitochondrial membrane system. Characterization of a yeast nuclear gene involved in the processing of the cytochrome b pre-mRNA. J Biol Chem. 1983 Aug 10;258(15):9459–9468. [PubMed] [Google Scholar]
- Miller D. L., Martin N. C. Characterization of the yeast mitochondrial locus necessary for tRNA biosynthesis: DNA sequence analysis and identification of a new transcript. Cell. 1983 Oct;34(3):911–917. doi: 10.1016/0092-8674(83)90548-2. [DOI] [PubMed] [Google Scholar]
- Müller P. P., Reif M. K., Zonghou S., Sengstag C., Mason T. L., Fox T. D. A nuclear mutation that post-transcriptionally blocks accumulation of a yeast mitochondrial gene product can be suppressed by a mitochondrial gene rearrangement. J Mol Biol. 1984 Jun 5;175(4):431–452. doi: 10.1016/0022-2836(84)90178-5. [DOI] [PubMed] [Google Scholar]
- Nagley P., Linnane A. W. Expression of mitochondrial DNA in Saccharomyces cerevisiae: the construction of sets of isonuclear haploid strains containing different specified mitochondrial genomes. Biochem Biophys Res Commun. 1978 Nov 29;85(2):585–592. doi: 10.1016/0006-291x(78)91203-2. [DOI] [PubMed] [Google Scholar]
- Nobrega F. G., Tzagoloff A. Assembly of the mitochondrial membrane system. DNA sequence and organization of the cytochrome b gene in Saccharomyces cerevisiae D273-10B. J Biol Chem. 1980 Oct 25;255(20):9828–9837. [PubMed] [Google Scholar]
- Osinga K. A., De Vries E., Van der Horst G., Tabak H. F. Processing of yeast mitochondrial messenger RNAs at a conserved dodecamer sequence. EMBO J. 1984 Apr;3(4):829–834. doi: 10.1002/j.1460-2075.1984.tb01892.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perlman P. S., Butow R. A. Mobile introns and intron-encoded proteins. Science. 1989 Dec 1;246(4934):1106–1109. doi: 10.1126/science.2479980. [DOI] [PubMed] [Google Scholar]
- Séraphin B., Boulet A., Simon M., Faye G. Construction of a yeast strain devoid of mitochondrial introns and its use to screen nuclear genes involved in mitochondrial splicing. Proc Natl Acad Sci U S A. 1987 Oct;84(19):6810–6814. doi: 10.1073/pnas.84.19.6810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabak H. F., Van der Horst G., Osinga K. A., Arnberg A. C. Splicing of large ribosomal precursor RNA and processing of intron RNA in yeast mitochondria. Cell. 1984 Dec;39(3 Pt 2):623–629. doi: 10.1016/0092-8674(84)90469-0. [DOI] [PubMed] [Google Scholar]
- Thalenfeld B. E., Bonitz S. G., Nobrega F. G., Macino G., Tzagoloff A. oli1 Transcripts in wild type and in a cytoplasmic "petite" mutant of yeast. J Biol Chem. 1983 Dec 10;258(23):14065–14068. [PubMed] [Google Scholar]
- Tzagoloff A., Myers A. M. Genetics of mitochondrial biogenesis. Annu Rev Biochem. 1986;55:249–285. doi: 10.1146/annurev.bi.55.070186.001341. [DOI] [PubMed] [Google Scholar]
- Zassenhaus H. P., Martin N. C., Butow R. A. Origins of transcripts of the yeast mitochondrial var 1 gene. J Biol Chem. 1984 May 10;259(9):6019–6027. [PubMed] [Google Scholar]
- Zhu H., Conrad-Webb H., Liao X. S., Perlman P. S., Butow R. A. Functional expression of a yeast mitochondrial intron-encoded protein requires RNA processing at a conserved dodecamer sequence at the 3' end of the gene. Mol Cell Biol. 1989 Apr;9(4):1507–1512. doi: 10.1128/mcb.9.4.1507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu H., Macreadie I. G., Butow R. A. RNA processing and expression of an intron-encoded protein in yeast mitochondria: role of a conserved dodecamer sequence. Mol Cell Biol. 1987 Jul;7(7):2530–2537. doi: 10.1128/mcb.7.7.2530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zinn A. R., Butow R. A. Nonreciprocal exchange between alleles of the yeast mitochondrial 21S rRNA gene: kinetics and the involvement of a double-strand break. Cell. 1985 Apr;40(4):887–895. doi: 10.1016/0092-8674(85)90348-4. [DOI] [PubMed] [Google Scholar]