Abstract
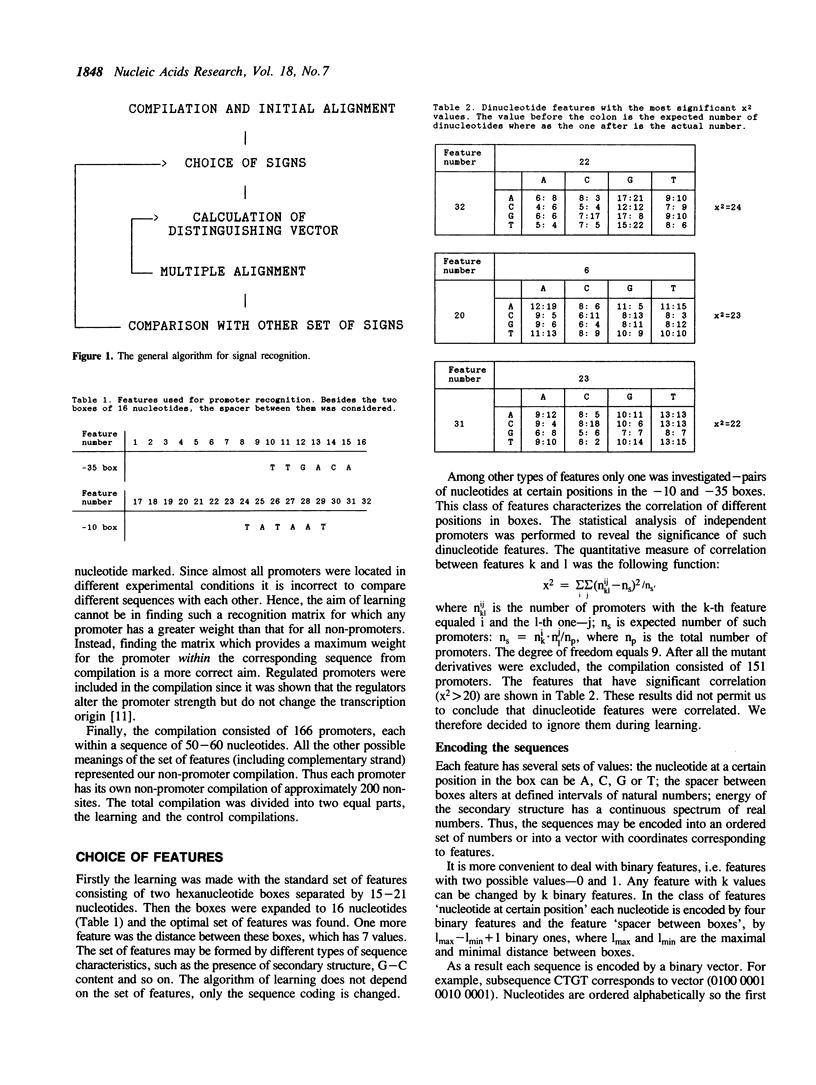
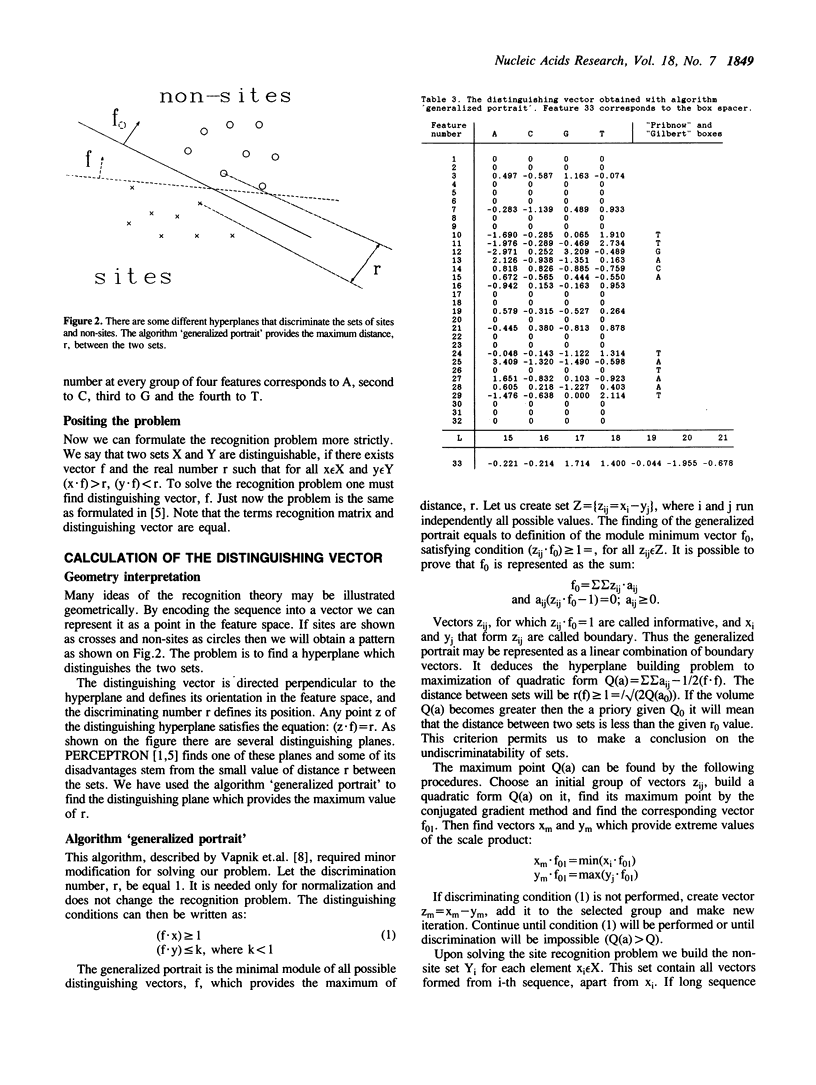
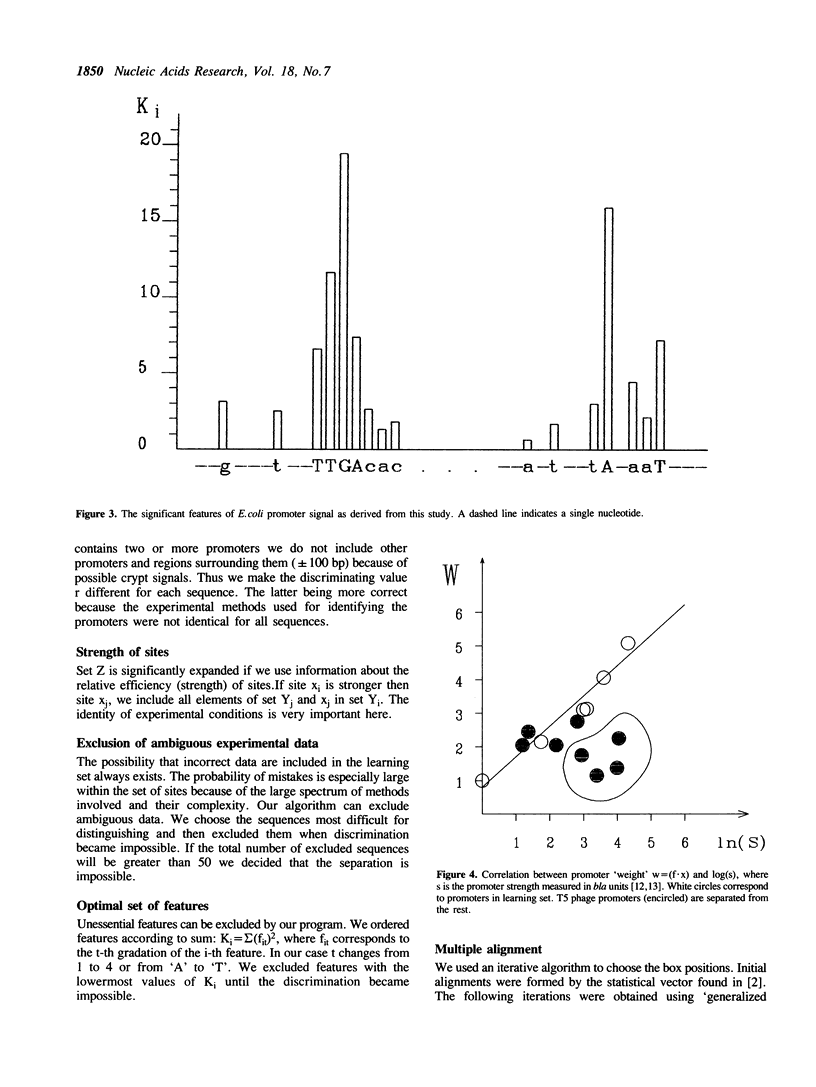
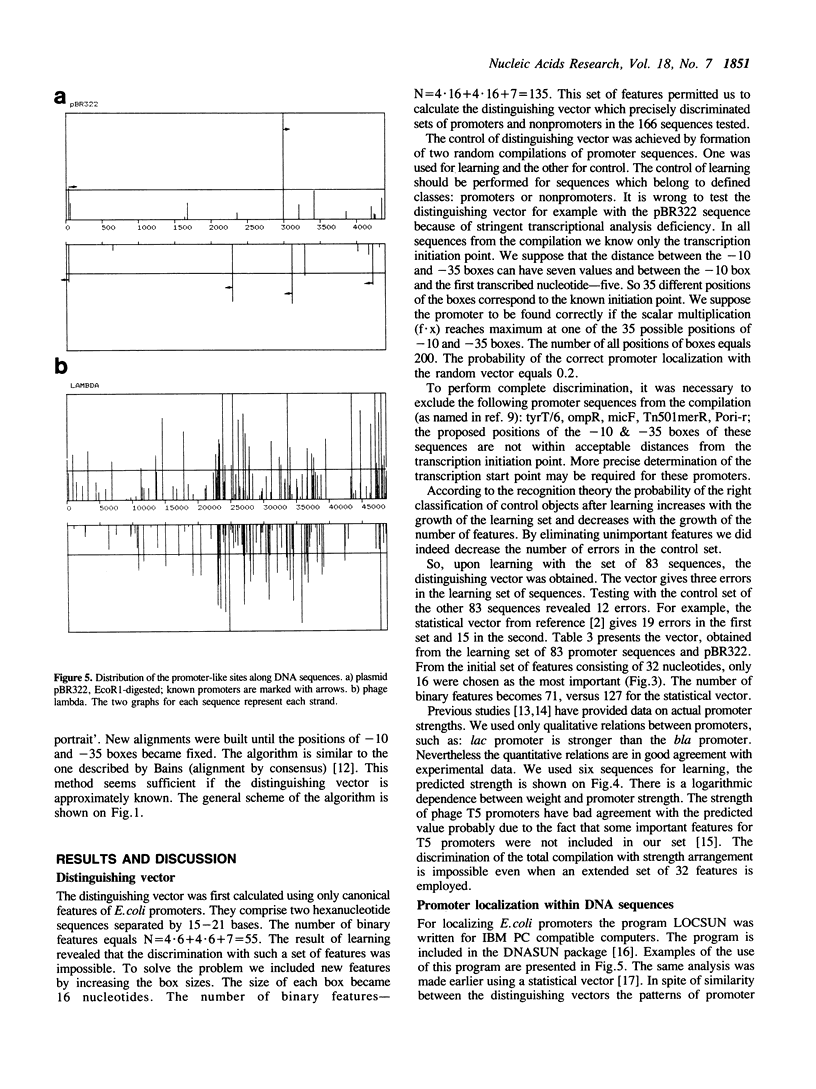
An algorithm from the pattern recognition theory 'generalized portrait' was used to find a distinguishing vector (scoring matrix) for E. coli promoters. We have attempted to solve three closely linked problems: (i) the selection of significant features of the signal; (ii) subsequent multiple alignment and (iii) calculation of the vector coordinates. Promoters with known strength have been successfully ranked in the correct order using this vector. We demonstrate the use of this method in predicting the location of promoters. A revised consensus promoter sequence is also presented.
Full text
PDF


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aleksandrov N. N., Mironov A. A. Raspoznavanie promotorov Escherichia coli po pervichnoi strukture DNK. Mol Biol (Mosk) 1987 Jan-Feb;21(1):242–249. [PubMed] [Google Scholar]
- Bains W. MULTAN: a program to align multiple DNA sequences. Nucleic Acids Res. 1986 Jan 10;14(1):159–177. doi: 10.1093/nar/14.1.159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brendel V., Trifonov E. N. A computer algorithm for testing potential prokaryotic terminators. Nucleic Acids Res. 1984 May 25;12(10):4411–4427. doi: 10.1093/nar/12.10.4411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deuschle U., Kammerer W., Gentz R., Bujard H. Promoters of Escherichia coli: a hierarchy of in vivo strength indicates alternate structures. EMBO J. 1986 Nov;5(11):2987–2994. doi: 10.1002/j.1460-2075.1986.tb04596.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gentz R., Bujard H. Promoters recognized by Escherichia coli RNA polymerase selected by function: highly efficient promoters from bacteriophage T5. J Bacteriol. 1985 Oct;164(1):70–77. doi: 10.1128/jb.164.1.70-77.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harley C. B., Reynolds R. P. Analysis of E. coli promoter sequences. Nucleic Acids Res. 1987 Mar 11;15(5):2343–2361. doi: 10.1093/nar/15.5.2343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawley D. K., McClure W. R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983 Apr 25;11(8):2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lida Y. DNA sequences and multivariate statistical analysis. Categorical discrimination approach to 5' splice site signals of mRNA precursors in higher eukaryotes' genes. Comput Appl Biosci. 1987 Jun;3(2):93–98. doi: 10.1093/bioinformatics/3.2.93. [DOI] [PubMed] [Google Scholar]
- Mulligan M. E., Hawley D. K., Entriken R., McClure W. R. Escherichia coli promoter sequences predict in vitro RNA polymerase selectivity. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):789–800. doi: 10.1093/nar/12.1part2.789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mulligan M. E., McClure W. R. Analysis of the occurrence of promoter-sites in DNA. Nucleic Acids Res. 1986 Jan 10;14(1):109–126. doi: 10.1093/nar/14.1.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Neill M. C. Escherichia coli promoters. I. Consensus as it relates to spacing class, specificity, repeat substructure, and three-dimensional organization. J Biol Chem. 1989 Apr 5;264(10):5522–5530. [PubMed] [Google Scholar]
- Staden R. Computer methods to locate signals in nucleic acid sequences. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):505–519. doi: 10.1093/nar/12.1part2.505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stormo G. D., Schneider T. D., Gold L., Ehrenfeucht A. Use of the 'Perceptron' algorithm to distinguish translational initiation sites in E. coli. Nucleic Acids Res. 1982 May 11;10(9):2997–3011. doi: 10.1093/nar/10.9.2997. [DOI] [PMC free article] [PubMed] [Google Scholar]



