Abstract
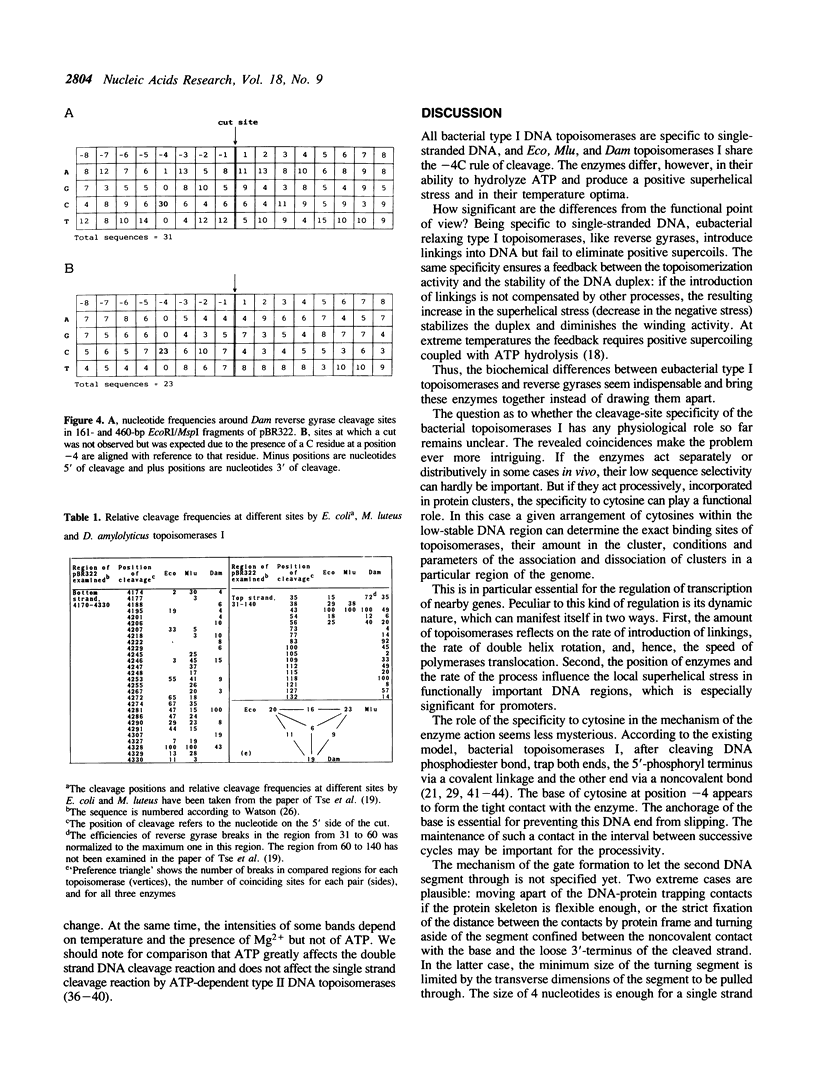
ATP-dependent type I topoisomerases from extremely thermophilic archaebacteria--reverse gyrases--drive positive supercoiling of DNA. We showed that reverse gyrase from Desulfurococcus amylolyticus breaks the DNA at specific sites and covalently binds to the 5' end. In 30 out of 31 sites located in pBR322 DNA fragments, cleavage occurs at the sequence 5'---CNNN/---(N is any base). The same rule was previously shown to hold for single-stranded DNA breakage by eubacterial topoisomerases I. The relative cleavage frequencies at different sites depend on Mg2+ and temperature. We discuss the possible physiological and mechanistic role of the above specificity of the bacterial topoisomerases I.
Full text
PDF

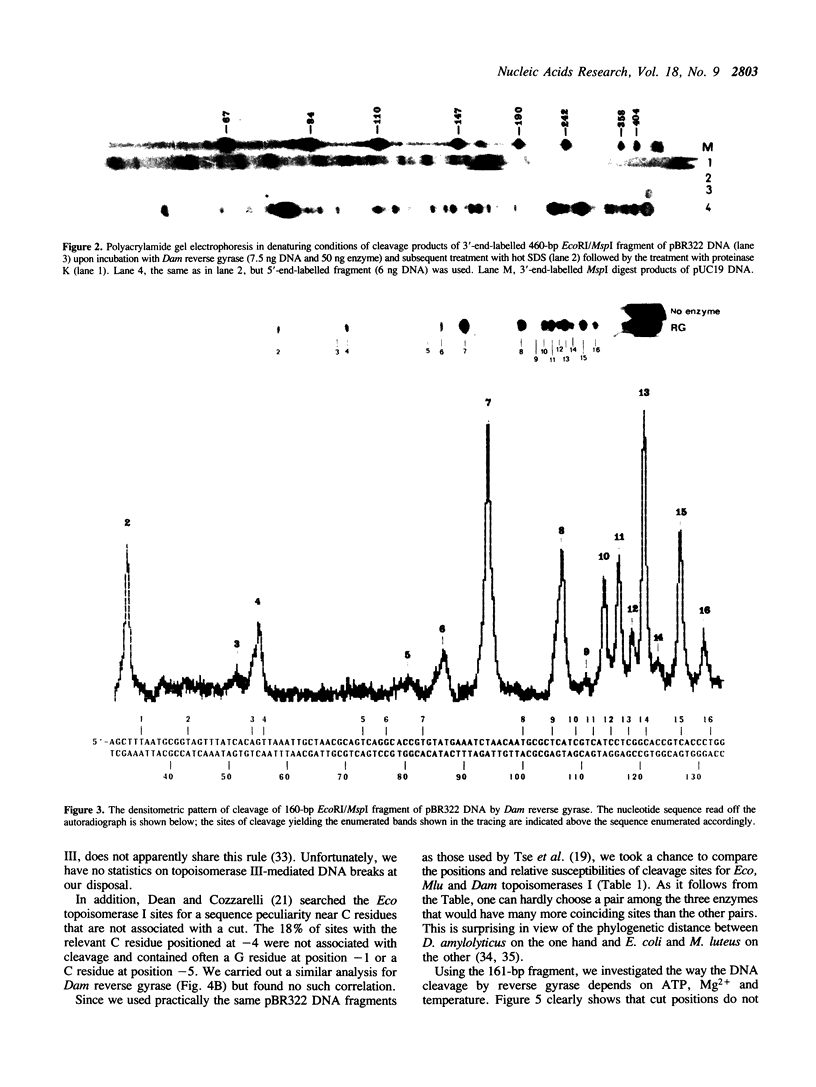

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Brown P. O., Cozzarelli N. R. Catenation and knotting of duplex DNA by type 1 topoisomerases: a mechanistic parallel with type 2 topoisomerases. Proc Natl Acad Sci U S A. 1981 Feb;78(2):843–847. doi: 10.1073/pnas.78.2.843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caruthers M. H., Beaucage S. L., Efcavitch J. W., Fisher E. F., Goldman R. A., deHaseth P. L., Mandecki W., Matteucci M. D., Rosendahl M. S., Stabinsky Y. Chemical synthesis and biological studies on mutated gene-control regions. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 1):411–418. doi: 10.1101/sqb.1983.047.01.048. [DOI] [PubMed] [Google Scholar]
- Champoux J. J. Strand breakage by the DNA untwisting enzyme results in covalent attachment of the enzyme to DNA. Proc Natl Acad Sci U S A. 1977 Sep;74(9):3800–3804. doi: 10.1073/pnas.74.9.3800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dean F. B., Cozzarelli N. R. Mechanism of strand passage by Escherichia coli topoisomerase I. The role of the required nick in catenation and knotting of duplex DNA. J Biol Chem. 1985 Apr 25;260(8):4984–4994. [PubMed] [Google Scholar]
- Depew R. E., Liu L. F., Wang J. C. Interaction between DNA and Escherichia coli protein omega. Formation of a complex between single-stranded DNA and omega protein. J Biol Chem. 1978 Jan 25;253(2):511–518. [PubMed] [Google Scholar]
- Domanico P. L., Tse-Dinh Y. C. Cleavage of dT8 and dT8 phosphorothioyl analogues by Escherichia coli DNA topoisomerase I: product and rate analysis. Biochemistry. 1988 Aug 23;27(17):6365–6371. doi: 10.1021/bi00417a026. [DOI] [PubMed] [Google Scholar]
- Drlica K. Biology of bacterial deoxyribonucleic acid topoisomerases. Microbiol Rev. 1984 Dec;48(4):273–289. doi: 10.1128/mr.48.4.273-289.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gellert M. DNA topoisomerases. Annu Rev Biochem. 1981;50:879–910. doi: 10.1146/annurev.bi.50.070181.004311. [DOI] [PubMed] [Google Scholar]
- Honigberg S. M., Radding C. M. The mechanics of winding and unwinding helices in recombination: torsional stress associated with strand transfer promoted by RecA protein. Cell. 1988 Aug 12;54(4):525–532. doi: 10.1016/0092-8674(88)90074-8. [DOI] [PubMed] [Google Scholar]
- Jaxel C., Nadal M., Mirambeau G., Forterre P., Takahashi M., Duguet M. Reverse gyrase binding to DNA alters the double helix structure and produces single-strand cleavage in the absence of ATP. EMBO J. 1989 Oct;8(10):3135–3139. doi: 10.1002/j.1460-2075.1989.tb08466.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirkegaard K., Pflugfelder G., Wang J. C. The cleavage of DNA by type-I DNA topoisomerases. Cold Spring Harb Symp Quant Biol. 1984;49:411–419. doi: 10.1101/sqb.1984.049.01.047. [DOI] [PubMed] [Google Scholar]
- Kirkegaard K., Wang J. C. Bacterial DNA topoisomerase I can relax positively supercoiled DNA containing a single-stranded loop. J Mol Biol. 1985 Oct 5;185(3):625–637. doi: 10.1016/0022-2836(85)90075-0. [DOI] [PubMed] [Google Scholar]
- Kreuzer K. N., Alberts B. M. Site-specific recognition of bacteriophage T4 DNA by T4 type II DNA topoisomerase and Escherichia coli DNA gyrase. J Biol Chem. 1984 Apr 25;259(8):5339–5346. [PubMed] [Google Scholar]
- Lee M. P., Sander M., Hsieh T. S. Single strand DNA cleavage reaction of duplex DNA by Drosophila topoisomerase II. J Biol Chem. 1989 Aug 15;264(23):13510–13518. [PubMed] [Google Scholar]
- Liu L. F. DNA topoisomerases--enzymes that catalyse the breaking and rejoining of DNA. CRC Crit Rev Biochem. 1983;15(1):1–24. doi: 10.3109/10409238309102799. [DOI] [PubMed] [Google Scholar]
- Liu L. F., Wang J. C. Supercoiling of the DNA template during transcription. Proc Natl Acad Sci U S A. 1987 Oct;84(20):7024–7027. doi: 10.1073/pnas.84.20.7024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Maxwell A., Gellert M. Mechanistic aspects of DNA topoisomerases. Adv Protein Chem. 1986;38:69–107. doi: 10.1016/s0065-3233(08)60526-4. [DOI] [PubMed] [Google Scholar]
- Nakasu S., Kikuchi A. Reverse gyrase; ATP-dependent type I topoisomerase from Sulfolobus. EMBO J. 1985 Oct;4(10):2705–2710. doi: 10.1002/j.1460-2075.1985.tb03990.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osheroff N. Biochemical basis for the interactions of type I and type II topoisomerases with DNA. Pharmacol Ther. 1989;41(1-2):223–241. doi: 10.1016/0163-7258(89)90108-3. [DOI] [PubMed] [Google Scholar]
- Sander M., Hsieh T. Double strand DNA cleavage by type II DNA topoisomerase from Drosophila melanogaster. J Biol Chem. 1983 Jul 10;258(13):8421–8428. [PubMed] [Google Scholar]
- Shibata T., Nakasu S., Yasui K., Kikuchi A. Intrinsic DNA-dependent ATPase activity of reverse gyrase. J Biol Chem. 1987 Aug 5;262(22):10419–10421. [PubMed] [Google Scholar]
- Slesarev A. I., Kozyavkin S. A. DNA substrate specificity of reverse gyrase from extremely thermophilic archaebacteria. J Biomol Struct Dyn. 1990 Feb;7(4):935–942. doi: 10.1080/07391102.1990.10508533. [DOI] [PubMed] [Google Scholar]
- Slesarev A. I. Positive supercoiling catalysed in vitro by ATP-dependent topoisomerase from Desulfurococcus amylolyticus. Eur J Biochem. 1988 Apr 15;173(2):395–399. doi: 10.1111/j.1432-1033.1988.tb14012.x. [DOI] [PubMed] [Google Scholar]
- Sugino A., Higgins N. P., Brown P. O., Peebles C. L., Cozzarelli N. R. Energy coupling in DNA gyrase and the mechanism of action of novobiocin. Proc Natl Acad Sci U S A. 1978 Oct;75(10):4838–4842. doi: 10.1073/pnas.75.10.4838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsao Y. P., Wu H. Y., Liu L. F. Transcription-driven supercoiling of DNA: direct biochemical evidence from in vitro studies. Cell. 1989 Jan 13;56(1):111–118. doi: 10.1016/0092-8674(89)90989-6. [DOI] [PubMed] [Google Scholar]
- Tse-Dinh Y. C., McCarron B. G., Arentzen R., Chowdhry V. Mechanistic study of E. coli DNA topoisomerase I: cleavage of oligonucleotides. Nucleic Acids Res. 1983 Dec 20;11(24):8691–8701. doi: 10.1093/nar/11.24.8691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tse-Dinh Y. C. Uncoupling of the DNA breaking and rejoining steps of Escherichia coli type I DNA topoisomerase. Demonstration of an active covalent protein-DNA complex. J Biol Chem. 1986 Aug 15;261(23):10931–10935. [PubMed] [Google Scholar]
- Tse Y. C., Kirkegaard K., Wang J. C. Covalent bonds between protein and DNA. Formation of phosphotyrosine linkage between certain DNA topoisomerases and DNA. J Biol Chem. 1980 Jun 25;255(12):5560–5565. [PubMed] [Google Scholar]
- Vosberg H. P. DNA topoisomerases: enzymes that control DNA conformation. Curr Top Microbiol Immunol. 1985;114:19–102. doi: 10.1007/978-3-642-70227-3_2. [DOI] [PubMed] [Google Scholar]
- Wang J. C. DNA topoisomerases. Annu Rev Biochem. 1985;54:665–697. doi: 10.1146/annurev.bi.54.070185.003313. [DOI] [PubMed] [Google Scholar]
- Wang J. C. DNA topoisomerases: nature's solution to the topological ramifications of the double-helix structure of DNA. Harvey Lect. 1985 1986;81:93–110. [PubMed] [Google Scholar]
- Wang J. C. Interaction between DNA and an Escherichia coli protein omega. J Mol Biol. 1971 Feb 14;55(3):523–533. doi: 10.1016/0022-2836(71)90334-2. [DOI] [PubMed] [Google Scholar]
- Wang J. C. Recent studies of DNA topoisomerases. Biochim Biophys Acta. 1987 Jun 6;909(1):1–9. doi: 10.1016/0167-4781(87)90040-6. [DOI] [PubMed] [Google Scholar]
- Watson N. A new revision of the sequence of plasmid pBR322. Gene. 1988 Oct 30;70(2):399–403. doi: 10.1016/0378-1119(88)90212-0. [DOI] [PubMed] [Google Scholar]
- Woese C. R. Bacterial evolution. Microbiol Rev. 1987 Jun;51(2):221–271. doi: 10.1128/mr.51.2.221-271.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R., Fox G. E. Phylogenetic structure of the prokaryotic domain: the primary kingdoms. Proc Natl Acad Sci U S A. 1977 Nov;74(11):5088–5090. doi: 10.1073/pnas.74.11.5088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu H. Y., Shyy S. H., Wang J. C., Liu L. F. Transcription generates positively and negatively supercoiled domains in the template. Cell. 1988 May 6;53(3):433–440. doi: 10.1016/0092-8674(88)90163-8. [DOI] [PubMed] [Google Scholar]
- Yang L., Jessee C. B., Lau K., Zhang H., Liu L. F. Template supercoiling during ATP-dependent DNA helix tracking: studies with simian virus 40 large tumor antigen. Proc Natl Acad Sci U S A. 1989 Aug;86(16):6121–6125. doi: 10.1073/pnas.86.16.6121. [DOI] [PMC free article] [PubMed] [Google Scholar]