Background: ORF45 of Kaposi sarcoma-associated herpesvirus (KSHV) causes sustained activation of p90 ribosomal S6 kinases (RSKs).
Results: ORF45 increases phosphorylation of eIF4B through p90 RSKs.
Conclusion: The ORF45/RSK axis promotes protein translation during lytic replication.
Significance: This mechanism is crucial for understanding of translational regulation during KSHV lytic replication.
Keywords: MAP Kinases (MAPKs), Phosphorylation, Translation Initiation Factors, Translation Regulation, Virus, Kaposi Sarcoma Associated Herpesvirus (KSHV), ORF45, eIF4B, p90 Ribosomal S6 Kinase (RSK), Tegument
Abstract
Open reading frame 45 (ORF45) of Kaposi sarcoma-associated herpesvirus (KSHV) causes sustained activation of p90 ribosomal S6 kinase (RSK), which is crucial for KSHV lytic replication, but the exact functional roles remain to be determined. To characterize the biological consequence of persistent RSK activation by ORF45, we screened known cellular substrates of RSK. We found that ORF45 induced phosphorylation of eukaryotic translation initiation factor 4B (eIF4B), increased its assembly into translation initiation complex, and subsequently facilitated protein translation. The ORF45/RSK-mediated eIF4B phosphorylation was distinguishable from that caused by the canonical AKT/mammalian target of rapamycin/ribosomal S6 kinase and MEK/ERK/RSK pathways because it was resistant to both rapamycin (an mammalian target of rapamycin inhibitor) and U1026 (an MEK inhibitor). The rapamycin and U1026 doubly insensitive eIF4B phosphorylation was induced during KSHV reactivation but was abolished if either ORF45 or RSK1/2 were ablated by siRNA, a pattern that is correlated with reduced lytic gene expression as we observed previously. Ectopic expression of eIF4B but not its phosphorylation-deficient mutant form increased KSHV lytic gene expression and production of progeny viruses. Together, these results indicated that ORF45/RSK axis-induced eIF4B phosphorylation is involved in translational regulation and is required for optimal KSHV lytic replication.
Introduction
Protein translation is primarily regulated at the step of initiation at which the small ribosome subunit is recruited to the 5′-end of mRNA and scans toward the start codon where the initiation complex joins the large subunit to form a complete ribosome (80 S) and begin polypeptide formation (1–4). Many eukaryotic translation initiation factors (eIFs) are involved in the initiation process. Most cellular mRNAs initiate translation through assembly of the cap-binding complex eIF4F on the 5′-cap structure (m7GpppN). The heterotrimeric eIF4F consists of eIF4E, the cap-binding protein; eIF4G, a scaffolding protein; and eIF4A, an RNA helicase. The 43 S preinitiation complex, which consists of the 40 S small ribosome subunit, eIF2-GTP-Met-tRNA ternary complex, eIF3, and eIF1A, is recruited by the cap-binding complex to the 5′-proximal region of mRNA and scans along the 5′-untranslated region (5′-UTR) to the start codon.
The eIF4B is an auxiliary factor that potentiates ribosome recruitment to the mRNA and stimulates translation of both capped and uncapped mRNAs (1). eIF4B is an avid RNA-binding protein that binds to both mRNA and 18 S rRNA simultaneously (2, 3). It interacts with the ribosome-bound eIF3 through protein-protein interactions (4). eIF4B is also known to stimulate eIF4A helicase activity, which unwinds the inhibitory secondary structure in the 5′-UTR of some mRNA species (5–7).
Phosphorylation of eIF4B by p70 ribosomal S6 kinase (S6K)3 or p90 ribosomal S6 kinase (RSK) increases its association with the initiation complex where it interacts with the eIF4A and eIF3. S6K is activated through the AKT/mTOR/S6K pathway, whereas RSK is activated through the MEK/ERK/RSK pathway (8–11). RSK and S6K are also known as kinases that phosphorylate the ribosomal S6 protein (rpS6). The S6K phosphorylates rpS6 at all four serine residues, Ser-235, Ser-236, Ser-240, and Ser-244, in an mTOR-dependent manner, whereas RSK exclusively phosphorylates rpS6 at Ser-235 and Ser-236 independently of mTOR (12, 13). Phosphorylation of rpS6 promotes the recruitment of the 40 S ribosome to the mRNA and therefore has a paramount effect on translation (13, 14).
Viruses rely on the cellular translational machinery for synthesis of their own proteins; therefore, they have evolved a variety of strategies to control that machinery (15, 16). Some viruses encode products that directly modify various eIFs or recruit them to viral mRNAs. Other viruses instead modulate cellular signaling pathways that regulate translation factors and ultimately hijack the host translational machinery for production of viral proteins. The MEK/ERK/RSK and AKT/mTOR/S6K cascades are commonly activated and exploited by a variety of DNA and RNA viruses (17–22). Kaposi sarcoma-associated herpesvirus (KSHV) is the etiological agent of endothelial neoplasm Kaposi sarcoma and two lymphoproliferative diseases, primary effusion lymphoma and multicentric Castleman disease. KSHV activates both MEK/ERK/RSK and AKT/mTOR/S6K signaling pathways by multiple mechanisms (23–32). We recently found that the immediate early and tegument protein open reading frame 45 (ORF45) of KSHV interacts with RSK1 and RSK2 and causes sustained activation of RSK and ERK during lytic replication (17). We further revealed the underlying mechanism and demonstrated that ORF45 forms complexes with RSK and ERK, retains active pRSK and pERK in the complexes, and protects them from dephosphorylation, thus promoting sustained activation of these kinases (33). Moreover, we showed that depletion of RSK expression by siRNA or inhibition of RSK activity by specific inhibitors reduces KSHV lytic gene expression, suggesting that the RSK signaling is required for optimal KSHV lytic replication, but its exact roles remain to be determined (17, 33). In the work reported here, we showed that ORF45-mediated sustained activation of ERK/RSK signaling contributes to the phosphorylation of eIF4B and plays a critical role in translation regulation during KSHV lytic infection.
EXPERIMENTAL PROCEDURES
Cells and Reagents
Body cavity-based lymphoma 1 (BCBL1) cells, a line derived from primary effusion lymphoma and latently infected with KSHV, were cultured in Roswell Park Memorial Institute (RPMI) 1640 medium supplemented with 10% heat-inactivated FBS. Human embryonic kidney (HEK) 293T cells and 293 cells were cultured in DMEM supplemented with 10% FBS. iSLK, a cell line of endothelial origin, was engineered to express doxycycline-inducible RTA (a key KSHV gene that is essential and sufficient for initiating the lytic replication program from latency). iSLK.219 (clone 10), derived from iSLK, is latently infected with a recombinant rKSHV.219 virus (34). The resultant cells maintain strict latency but support robust lytic reactivation upon induction with doxycycline. Both iSLK cell lines were cultured and maintained as described previously (35). BAC36, a bacterial artificial chromosome clone that carries the entire KSHV genome (36), and BAC-stop45, an ORF45-null mutant that carries a premature stop codon in the ORF45 coding region, have been described previously (37). 12-O-tetradecanoylphorbol-13-acetate (TPA), sodium butyrate, and anti-FLAG and anti-HA antibodies were purchased from Sigma. Antibodies detecting ERK1/2, eIF4B, rpS6, and the phosphorylated forms pERK1/2 (Thr-202/Tyr-204), peIF4B (Ser-422), pS6 (Ser-235/Ser-236), pS6 (Ser-240/Ser-244), pRSK1 (Thr-359/Ser-363), pRSK1 (Ser-380), and pRSK1 (Thr-573) along with rapamycin and U0126 were purchased from Cell Signaling Technology (Beverly, MA). An RSK inhibitor, BI-D1870, was purchased from the Medical Research Council Protein Phosphorylation Unit, University of Dundee (Dundee, Scotland, UK). Rat anti-latency-associated nuclear antigen antibody was purchased from Advanced Biotechnologies (Columbia, MD). Rabbit anti-RSK1 and -RSK2 antibodies were purchased from Upstate (now Millipore, Billerica, MA). Mouse monoclonal anti-RTA was a generous gift from Dr. Ueda (Hamamatsu University School of Medicine, Hamamatsu, Japan). Anti-ORF45 and anti-K8 antibodies have been described previously (38, 39). 7-Methyl-GTP (m7GTP)-Sepharose 4B beads were from GE Healthcare. Recombinant vaccinia virus vTF7-3 and lentivirus vector pHIV7 were gifts from Dr. Hengli Tang at The Florida State University.
Plasmids
Plasmids pKH3-RSK1, pKH3-RSK2 wild type, pKH3-RSK2 active (Y707A), and pKH3-RSK2 kinase-dead (K100A/Y707A) constructs were obtained from Dr. Deborah Lannigan (University of Virginia). Plasmid pCR3.1-ORF45 full length and its derivatives have been described previously (17, 33). Plasmid pRK7-HA-S6K1 (8984) was purchased from Addgene (Cambridge, MA) (40). The T7 promoter-driven luciferase reporter was constructed by insertion of REN-HCV IRES-LUC fragment from the bicistronic Renilla-HCV IRES-firefly luciferase plasmid (9) between EcoRV and XbaI into pGEM-T Easy at EcoRV sites. The coding sequence of human eIF4B was amplified from BCBL1 cDNA (41) and inserted into lentivirus vector pHIV7 (42) at BamHI and NotI sites to yield pHIV7-eIF4B plasmid. A point mutation was introduced with the QuikChange mutagenesis kit (Stratagene, La Jolla, CA) to yield pHIV7-eIF4B-S422A plasmid.
In Vitro Kinase Assay
RSK and S6K kinase activities were assayed in vitro with GST-S6 peptide (218KEAKEKRQEQIAKRRRLSSLRASTSKSESSQK249) as described previously (17). Briefly, HEK293 cells seeded in 100-mm dishes were transfected with 10 μg of HA-tagged RSK1 or S6K1 expression vector. After serum starvation for 24 h, the transfected cells were treated with TPA for 10 min or left untreated as controls. Whole-cell lysates were made, and the HA-RSK1 or HA-S6K1 proteins were immunoprecipitated with 50 μl of EZview Red anti-HA affinity beads. After two washes with the lysis buffer and three washes with TBS buffer (50 mm Tris-HCl, pH 7.4, 150 mm NaCl), the immunoprecipitated beads were resuspended in 100 μl of TBS plus 1 mm PMSF, 1 mm Na3VO4, 1× protease inhibitor mixture (Roche Applied Science). The kinase reaction was performed by incubation of 5 μl of the precipitated complexes with 2.5 μg of GST-S6 substrate in 25 μl of 1× kinase assay buffer (25 mm HEPES, pH 7.5, 50 mm NaCl, 20 mm β-glycerophosphate, 1 mm DTT, 20 mm MgCl2, 1 mm Na3VO4, 1 μg/ml BSA, 20 μm ATP, 5 μCi of [γ-32P]ATP). The reactions were kept at 30 °C for 30 min and stopped by addition of SDS-PAGE loading buffer. After fractionation of samples by 10% SDS-PAGE, the gel was dried and exposed to x-ray film or analyzed by phosphorimaging.
Retrovirus-based Transduction
The knockdown of gene expression by a retrovirus-based siRNA vector including target sequences for RSK1, RSK2, ERKs, and ORF45 has been described previously (17, 33). The BCBL1 cells that stably express eIF4B wild type or S422A mutant were established by standard lentivirus vector protocols (43).
Pulldown Assay of m7GTP-Sepharose
m7GTP-Sepharose 4B beads were incubated with an equal amount of whole-cell extract (∼1 mg) at 4 °C for 2–4 h and then washed four times with whole-cell lysis buffer. The proteins bound to the Sepharose beads were subjected to SDS-PAGE followed by Western blot analysis.
Polysomal Fractionation
Polysomal ribosome fractions were separated by sucrose density gradient centrifugation as described by Roux et al. (13). Briefly, 10 min before harvest, 100 μg/ml cycloheximide was added to the culture medium. Cells were washed in PBS supplemented with 100 μg/ml cycloheximide and harvested in polysome lysis buffer (50 mm Tris-HCl, pH 7.4, 250 mm KCl, 5 mm MgCl2, 250 mm sucrose, 1% Triton X-100, 1.3% sodium deoxycholate, 100 μg/ml cycloheximide, and 1× EDTA-free protease inhibitor mixture from Roche Applied Science). Samples were incubated on ice for 15 min and then centrifuged at 10,000 × g for 10 min at 4 °C. The supernatant was layered on a 15–50% linear sucrose gradient (w/v) and centrifuged in an SW41 rotor at 35,000 rpm (150,000 × g) for 150 min at 4 °C. After centrifugation, 1-ml fractions were collected from the gradient. RNAs were isolated from each fraction with TRIzol reagent (Invitrogen) and quantified by absorbance at 260 nm.
Infection of Vaccinia Virus vTF7-3 and Translation Assay
Vaccinia virus vTF7-3 was amplified on Vero cells. Briefly, vTF7-3 virus stock was mixed with an equal volume of 10-fold diluted trypsin-EDTA solution (Invitrogen; final trypsin concentration, 0.025%) for 30 min at 37 °C, and then 1 ml of DMEM with 2.5% FBS was added to the solution to neutralize the trypsin. One milliliter of trypsin-digested vTF7-3 virus was added to a monolayer of Vero cells (>50% confluence) in a 100-mm dish and incubated for 2 h at 37 °C. Then the viral inoculum was removed, and 10 ml of fresh medium with 2.5% FBS per dish was added. After 3–4 days of incubation, cells were collected when most cells became round. The cells were washed with PBS once, resuspended in 1 ml of PBS per dish, subjected to three rounds of freeze/thaw at −80 and 37 °C, and then centrifuged at 10,000 × g for 10 min for removal of cell debris. The supernatant was collected, titrated, and stored at −80 °C. For translation reporter assays, HEK293 or RSK siRNA-transduced HEK293 cells (80–90% confluence) were infected with trypsin-digested vTF7-3 with a multiplicity of infection of 10 for 1 h at 37 °C. During the incubation, DNA-Lipofectamine complexes were prepared by mixing Lipofectamine 2000 with the reporter plasmid pGEM-REN-HCV IRES-LUC and ORF45-expressing vectors. The vTF7-3-infected cells were rinsed once and provided with fresh medium with 2.5% FBS; DNA-Lipofectamine complexes were then added. After 36 h, cell extracts were analyzed for luciferase activity with a Dual-Luciferase assay reporter system (Promega, Madison, WI).
Amplification and Infection of Adenovirus Ad-RTA
The recombinant Ad-RTA virus was a gift from Dr. Britt Glaunsinger at the University of California, Berkeley and was amplified in HEK293 cells as described previously (44). BCBL1 cells (1 × 106 cells) were infected with a serial dilution (multiplicity of infection ranging from 1 to 1000) of Ad-RTA, and expressions of KSHV lytic genes were analyzed by Western blots. An optimal multiplicity of infection of 10 was used in most experiments.
Statistical Analysis
Data are shown as average values with S.D. The p value was determined by Student's t test.
RESULTS
ORF45 Increases Phosphorylation of eIF4B and rpS6
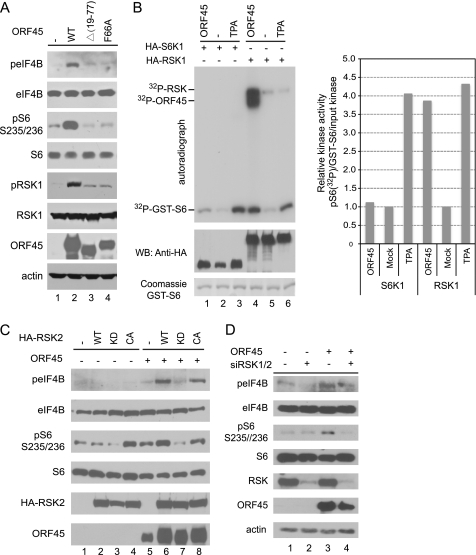
We recently found that KSHV ORF45 causes sustained activation of RSK1 and RSK2. We further demonstrated that depletion of RSK expression or inhibition of RSK activity reduces KSHV lytic gene expression, indicating that RSK signaling is required for optimal KSHV lytic replication (17, 33), but the exact functional roles of the persistently active RSK remain to be determined. To characterize the biological consequence of RSK activation by ORF45, we screened some of the known cellular substrates of RSK by Western blot using phosphorylation-specific antibodies (45, 46). Because ORF45 of KSHV is the only one among its homologues of γ2 (Rhadinovirus) herpesviruses that is localized predominantly in the cytoplasm (47), we were particularly interested in the cytoplasmic substrates of RSK. We found two cytoplasmic substrates, eIF4B and ribosomal protein S6, whose phosphorylations were consistently increased in cells transfected with wild-type pCR3.1-ORF45 but not in cells transfected with the mutant pCR3.1-ORF45-Δ(19–77) or pCR3.1-ORF45-F66A, both of which failed to activate RSK (Fig. 1A). Recent studies have shown that eIF4B and rpS6 can be phosphorylated by both RSK and S6K (9, 13). To determine whether S6K contributes to the phosphorylation of eIF4B and rpS6 induced by ORF45, we cotransfected HA-tagged kinase expression vectors with or without pCR3.1-ORF45 plasmid into HEK293 cells and immunoprecipitated the kinases with anti-HA affinity beads to perform in vitro kinase assays. As expected, the RSK activity was dramatically induced by ORF45 and comparable with that caused by TPA stimulation (Fig. 1B, left panel, lanes 4 and 6, compare the 32P-GST-S6 signals). As reported previously (17), ORF45 was coimmunoprecipitated very efficiently with RSK and phosphorylated by the associated active kinases, producing strong 32P-ORF45 and 32P-RSK signals (Fig. 1B, lane 4). In contrast, the S6K activity was induced marginally by ORF45 in comparison with that induced by TPA (Fig. 1B, left panel, lanes 1 and 3, compare the 32P-GST-S6 signals). The effects of ORF45 on RSK and S6K kinase activities are more quantitatively illustrated in Fig. 1B (right bar graph) where the relative kinases activities were normalized to the amounts of input GST-S6 substrate and HA-tagged kinases. The above results suggested that RSK rather than S6K is the principal kinase activated by ORF45 and is responsible for the increased eIF4B and rpS6 phosphorylation induced by ORF45.
FIGURE 1.
ORF45/RSK axis induces phosphorylation of eIF4B and rpS6. A, ORF45 induces eIF4B and rpS6 phosphorylation. HEK293 cells were cotransfected with plasmid pKH3-RSK1 and plasmids expressing FLAG-tagged wild-type ORF45 or mutant constructs Δ(19–77) and F66A, both of which lost most of their ability to activate RSK. After serum starvation for 24 h, whole-cell lysates were immunoblotted with antibodies as indicated. B, ORF45 preferentially stimulates RSK rather than S6K kinase activity. The HA-tagged expression vector pKH3-RSK1 or pRK7-HA-S6K1 plasmids were cotransfected with pCR3.1-ORF45 or empty vector into HEK293 cells. After serum starvation for 24 h, the transfected cells were treated with TPA or vehicle (DMSO) for 10 min, and whole-cell lysates were then prepared. The HA-RSK1 and HA-S6K1 kinases were immunoprecipitated with anti-HA affinity beads and used in in vitro kinase assays with GST-S6 as substrate. Signals on the autoradiograph are phosphorylated GST-S6, autophosphorylated HA-RSK1, and ORF45 that was coimmunoprecipitated with RSK and phosphorylated by the associated kinases (17). The immunoprecipitated complexes were also analyzed by Western blot (WB) with antibodies as indicated. The right panel represents the relative kinase activities that were normalized to the input GST-S6 and HA-tagged kinases. Quantitative analysis of the band intensities in the left panel was performed with ImageJ software from NIH. C, RSK activity is required for the ORF45-induced eIF4B and rpS6 phosphorylation. HEK293 cells were cotransfected with ORF45 expression vector plus HA-tagged wild-type RSK2 (WT), RSK2-KD mutant (K100A/Y707A), or RSK2-CA mutant (Y707A) constructs in combinations as indicated. Beginning 48 h after transfection, the cells were serum-starved for an additional 24 h. Whole-cell lysates were analyzed by Western blot with antibodies as indicated. D, knockdown of RSK1 and RSK2 expression by siRNAs inhibits ORF45-induced eIF4B phosphorylation. RSK1/2 siRNA- or control siRNA-transduced HEK293 cells (17) were transfected with ORF45-expressing vectors or mock-transfected with control empty vectors. After serum starvation for 24 h, the cell lysates were analyzed by Western blots with antibodies as indicated.
To confirm the principal role of RSK, we next used an RSK2 kinase-dead (KD) mutant (RSK2-K100A/Y707A) and a constitutively active (CA) mutant (RSK2-Y707A) to determine whether RSK activity was needed for the ORF45-induced phosphorylation of eIF4B and rpS6. As shown in Fig. 1C, in the presence of ORF45, overexpression of the RSK2-WT and the RSK2-CA increased the phosphorylation of eIF4B and rpS6, but overexpression of the RSK2-KD did not increase or actually reduced the phosphorylation (Fig. 1C). In the absence of ORF45, overexpression of the RSK2-WT or the RSK2-CA did not induce noticeable eIF4B phosphorylation, but overexpression of the RSK2-CA increased the basal level of rpS6 phosphorylation, whereas overexpression of the RSK2-KD reduced the basal level of rpS6 phosphorylation (Fig. 1C). This experiment suggests that RSK activity is required for the ORF45-induced eIF4B phosphorylation and that ORF45 specifically increases eIF4B phosphorylation by RSK. Furthermore, we used HEK293-derived stable cell lines in which both RSK1 and RSK2 have been knocked down by retrovirus-delivered siRNAs (17) and found that depletion of both RSK1 and RSK2 reduced the levels of ORF45-induced eIF4B and rpS6 phosphorylation (Fig. 1D). Together, these results supported induction of eIF4B and rpS6 phosphorylation by the ORF45/RSK axis.
ORF45 Promotes Translation
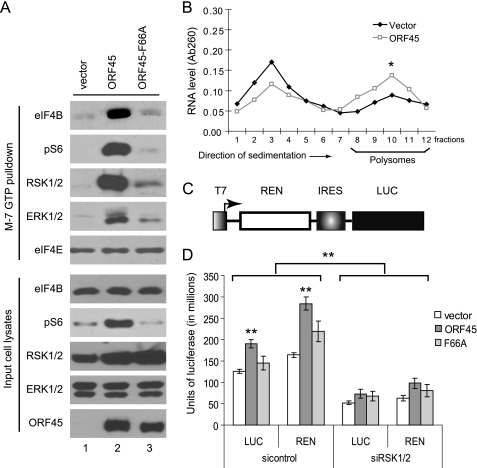
Phosphorylation of eIF4B and rpS6 increases their associations with the translation initiation complex and subsequently promotes translation (9, 13). We next determined whether ORF45 affects the recruitment of eIF4B and rpS6 into the initiation complex using m7GTP (which mimics the m7GpppN mRNA cap)-Sepharose beads. As shown in Fig. 2A, ORF45 increased the recruitment of eIF4B and pS6 into the mRNA cap-associated complex, but the ORF45-F66A mutant had little effect (Fig. 2A). In contrast, ORF45 did not affect binding of eIF4E to m7GTP-Sepharose beads. Interestingly, ORF45 also increased association of RSK and ERK with the complex (Fig. 2A, top) in agreement with the previous study showing that RSK and ERK are recruited to polysomes during active translation (48).
FIGURE 2.
ORF45 facilitates translation. A, ORF45 promotes assembly of the translation initiation complex. HEK293 cells were transfected with pCR3.1-ORF45, pCR3.1-ORF45-F66A, or empty vector. Forty-eight hours after transfection, cell lysates were prepared and incubated with m7GTP-Sepharose beads at 4 °C. After washes, the bead-bound proteins and the input cell lysates were analyzed by Western blots with antibodies as indicated. B, ORF45 promotes association of mRNAs with polysomes. Lysates from pCR3.1-ORF45 or empty vector-transfected HEK293 cells were loaded on 15–50% (w/v) sucrose gradients. After centrifugation, 1-ml fractions were collected (fraction 1, 15% sucrose; fraction 12, 50% sucrose) and used to purify RNA. The concentration of RNA was measured by absorbance at 260 nm (Ab260). The experiments were performed in duplicate (*, p < 0.05). C, schematic presentation of the bicistronic reporter plasmid (pGEM-REN-HCV IRES-LUC). T7, T7 promoter; REN, Renilla luciferase; IRES, internal ribosomal entry site of HCV; LUC, firefly luciferase. D, ORF45 promotes protein translation. HEK293 cells stably transduced with control siRNA or siRNAs against RSK1 and RSK2 were infected with a recombinant vaccinia virus, vTF7-3 (multiplicity of infection = 10), that expresses bacteriophage T7 RNA polymerase. The infected cells were then transfected with the reporter plasmid pGEM-REN-HCV IRES-LUC plus pCR3.1-ORF45, pCR3.1-ORF45-F66A, or empty vector pCR3.1. Luciferase activities were measured 36 h after transfection with the Dual-Luciferase assay kit (Promega). The experiments were performed three times, each in duplicate (**, p < 0.01). Error bar represents standard deviation.
To determine the effect of ORF45 on the recruitment of mRNA to active translational machinery, we performed polysomal fractionation over a sucrose gradient. Measurement of the absorbance of each fraction (from top to bottom) at 260 nm revealed two major peaks representing ribosomal and polysomal RNAs. As shown in Fig. 2B, expression of ORF45 caused a shift from the ribosomal fractions to the polysomal fractions, indicating that the percentage of RNA in the polysomal fraction was increased (Fig. 2B). The magnitude of effect was comparable with that caused by perturbations of the RSK or related pathways reported in the literature (13, 49). These results suggested that ORF45 promotes the assembly of the translation initiation complex and recruitment of mRNA into active translation machinery, therefore promoting protein translation.
To demonstrate directly that ORF45 affects translation, we adapted a luciferase reporter assay similar to one used previously by Sonenberg and co-workers (50) to measure translation efficiency in mammalian cells. Briefly, HEK293 cells were first infected with a recombinant vaccinia virus, vTF7-3, that expresses bacteriophage T7 RNA polymerase (51). The infected cells were then cotransfected with plasmid pCR3.1-ORF45 and a bicistronic luciferase reporter expressed from the T7 promoter (Fig. 2C). In this system, the mRNA of luciferase is transcribed by T7 RNA polymerase instead of eukaryotic RNA polymerase and translated by cellular translation machinery. Among the T7-transcribed RNAs in cells, about 5–10% are capped by the vaccinia capping enzyme and are translated, whereas the uncapped products are largely untranslated (50, 52, 53). Because the design minimizes possible effects of the complex transcriptional regulation in mammalian cells, the system is an ideal surrogate assay for measuring translation efficiency (50). As shown in Fig. 2C, both firefly luciferase activity (which represents cap-independent translation) and Renilla luciferase activity (which represents cap-dependent translation) were increased about 50% in ORF45-expressing cells over those in the control cells. The promotion of translation by ORF45 apparently depends on RSK activation because an ORF45-F66A mutant was less able to increase luciferase activity. Furthermore, knockdown of both RSK1 and RSK2 expression by siRNA decreased luciferase activity. These results suggested that ORF45 promotes cellular translation through activation of RSK.
RSK Is Required for eIF4B and rpS6 Phosphorylation during KSHV Lytic Replication
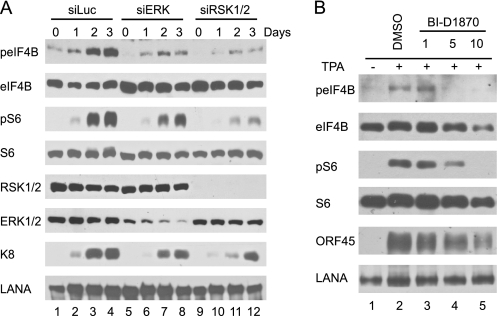
Having demonstrated that ORF45-induced RSK signaling promotes translation, we next determined the role of RSK signaling in translational control during KSHV lytic replication. We first examined whether depletion of RSK affects eIF4B and rpS6 phosphorylation during KSHV reactivation. We used BCBL1-derived cell lines in which RSK1/2 or ERK1/2 have been knocked down by retrovirus-delivered siRNAs as described previously (17, 33). As shown in Fig. 3A, sustained phosphorylations of eIF4B and rpS6 were observed in the TPA-induced control cells transduced with siRNA against luciferase, but the levels of phosphorylation were dramatically lower in cells transduced with siRNAs against RSK1/2 or ERK (Fig. 3A). To determine the role of RSK activity in eIF4B and rpS6 phosphorylation, we induced BCBL1 cells in the presence of a specific RSK inhibitor, BI-D1870 (17, 54). As shown in Fig. 3B, BI-D1870 reduced eIF4B and rpS6 phosphorylation in a dose-dependent manner. These results suggested that RSK-dependent signaling is essential for eIF4B and rpS6 phosphorylation during KSHV replication.
FIGURE 3.
RSK is required for eIF4B and rpS6 phosphorylation during KSHV lytic replication. A, depletion of RSK and ERK expression by siRNAs decreases eIF4B and rpS6 phosphorylation. RSK1/2-, ERK-, and luciferase (Luc) siRNA-transduced BCBL1 cells were induced with TPA (17, 33). Cells were collected at different times, and cell lysates were analyzed by Western blot with antibodies as indicated. B, inhibition of RSK activity blocks eIF4B and rpS6 phosphorylation. BCBL1 cells were induced with TPA in the absence and presence of increasing concentrations of RSK inhibitor BI-D1870 for 48 h. Whole-cell lysates were then prepared and analyzed by Western blot with antibodies as indicated. LANA, latency-associated nuclear antigen.
ORF45/RSK Axis Contributes Preferentially to eIF4B Phosphorylation during KSHV Lytic Infection
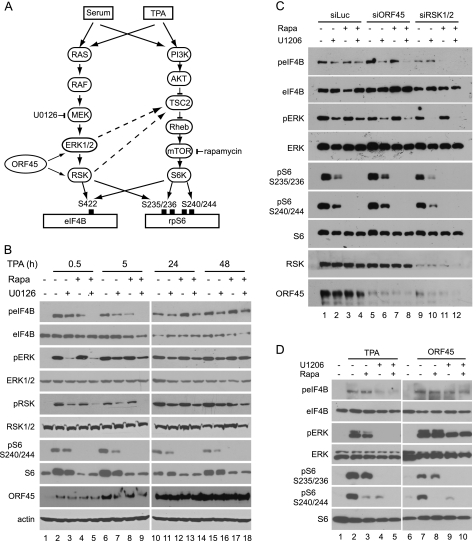
eIF4B and rpS6 can be phosphorylated through two signaling pathways, MEK/ERK/RSK and AKT/mTOR/S6K. The AKT/mTOR/S6K signaling was inhibited by rapamycin, and the MEK/ERK/RSK signaling was inhibited by U0126 (Fig. 4A). We next used these two inhibitors to determine the contribution of each pathway to the phosphorylation of eIF4B and rpS6 during KSHV lytic reactivation. S6K phosphorylates rpS6 at all four serine residues (Ser-235, Ser-236, Ser-240, and Ser-244), whereas RSK does so only at Ser-235 and Ser-236; therefore, the antibody against rpS6 at Ser-240/Ser-244 detects S6K-specific phosphorylation of rpS6 protein (12, 13). In the early stages of KSHV reactivation (0.5 and 5 h after induction), rapamycin blocked rpS6 phosphorylation at Ser-240/Ser-244, and U0126 inhibited ERK and RSK phosphorylation efficiently, but neither of them blocked eIF4B phosphorylation completely, whereas a combination of the two did (Fig. 4B, left panel). This result indicated that both the ERK/RSK and mTOR/S6K pathways contribute to the phosphorylation of eIF4B in the early stage of KSHV reactivation. In the late stages of KSHV lytic replication (24 and 48 h after induction), rapamycin blocked rpS6 phosphorylation at Ser-240/Ser-244 almost completely as expected, whereas U1026 only slightly reduced ERK, RSK, and rpS6 phosphorylation (Fig. 4B, right panel). The minor inhibition of rpS6 phosphorylation by U1026 probably reflects the cross-talk between ERK/RSK and mTOR/S6K signaling cascades (55–57). Interestingly, even a combination of rapamycin and U0126 did not inhibit eIF4B phosphorylation completely (Fig. 4B, right panel), suggesting that a virus-encoded or -induced factor(s) is involved in the induction of the rapamycin and U1026 doubly insensitive eIF4B phosphorylation in the late stage of KSHV lytic replication. Because ORF45-mediated activation of RSK is resistant to U1026 (17) and independent of mTOR signaling and thus resistant to rapamycin too (Fig. 4, A and D), we speculated that the ORF45/RSK axis is likely to be the major candidate signaling that contributes preferentially to the eIF4B phosphorylation during the late stage of KSHV lytic replication.
FIGURE 4.
ORF45/RSK axis contributes to eIF4B phosphorylation in KSHV late lytic infection. A, schematic representation of two converging signaling pathways, RAS/RAF/MEK/ERK/RSK and PI3K/AKT/mTOR/S6K, that are involved in eIF4B phosphorylation. B, rapamycin (Rapa) and U0126 dually insensitive eIF4B phosphorylation during KSHV lytic replication. BCBL1 cells were induced with TPA and then treated with U0126 (25 μm) and/or rapamycin (50 nm) for 2 h at the time points indicated. Cell lysates were analyzed by Western blotting with the antibodies indicated. C, ORF45 and RSK contribute to the rapamycin and U0126 doubly insensitive eIF4B phosphorylation. BCBL1 cells transduced by siRNAs against RSK1 and RSK2, ORF45, or control luciferase (Luc) were treated with TPA for 24 h; U0126 and/or rapamycin was added to the medium and incubated for 2 h. The whole-cell lysates were then analyzed by Western blot with antibodies as indicated. D, ORF45 induces rapamycin and U0126 doubly insensitive eIF4B phosphorylation. HEK293 cells were transfected with pCR3.1-ORF45 or empty vector and grown in the presence of 10% FBS for 48 h. U0126 and/or rapamycin was added to the medium and incubated for 2 h; then whole-cell extracts were analyzed by Western blot with antibodies as indicated. TSC2, tuberous sclerosis protein 2.
To confirm that the ORF45/RSK axis indeed contributes to phosphorylation of eIF4B during the late stage of KSHV lytic replication, we examined eIF4B phosphorylation in TPA-induced BCBL1-derived stable cells in which ORF45 or RSK1/2 had been knocked down by siRNAs (17, 33). In the control siRNA-transduced cells, neither rapamycin nor U0126 reduced eIF4B phosphorylation significantly, but in RSK1/2 or ORF45 siRNA-transduced cells, the combined treatment blocked eIF4B phosphorylation, supporting the idea that the ORF45/RSK signaling axis contributes to rapamycin and U0126 doubly insensitive eIF4B phosphorylation during KSHV replication (Fig. 4C). Interestingly, although eIF4B phosphorylation was resistant to rapamycin and U0126 in KSHV lytically infected cells, rpS6 phosphorylation remained sensitive to both rapamycin and U0126 during KSHV lytic replication (Fig. 4C). As expected, rapamycin inhibited rpS6 phosphorylation at Ser-240/Ser-244 completely because these sites were phosphorylated only by S6K and not by RSK (Fig. 4C). Although Ser-235/Ser-236 can be phosphorylated by both RSK and S6K, rpS6 phosphorylation at these sites remained sensitive to either rapamycin or U0126 inhibition (Fig. 4C), suggesting that the ORF45/RSK axis contributes little to direct rpS6 phosphorylation or contributes indirectly presumably through phosphorylation of tuberous sclerosis protein 2 (Fig. 4A) (55, 58). Together, these results indicated that the ORF45/RSK signaling axis contributes preferentially to eIF4B phosphorylation during KSHV lytic replication.
To confirm further the critical role of ORF45 in eIF4B phosphorylation, we next determined whether expression of ORF45 is sufficient for induction of rapamycin and U1026 doubly insensitive eIF4B phosphorylation. We transiently transfected the ORF45 expression vector into HEK293 cells and then examined eIF4B phosphorylation after rapamycin and/or U0126 treatment. As shown in Fig. 4D, the TPA-induced eIF4B phosphorylation was dramatically reduced by U0126 but not by rapamycin, indicating that ERK and RSK act as the principal kinases for eIF4B phosphorylation upon TPA stimulation. In contrast, the ORF45-induced eIF4B phosphorylation was not inhibited completely by rapamycin or U0126, supporting the conclusion that ORF45 is sufficient to induce rapamycin and U0126 doubly insensitive eIF4B phosphorylation.
ORF45/RSK Axis Is Involved in Translation Regulation during KSHV Lytic Infection
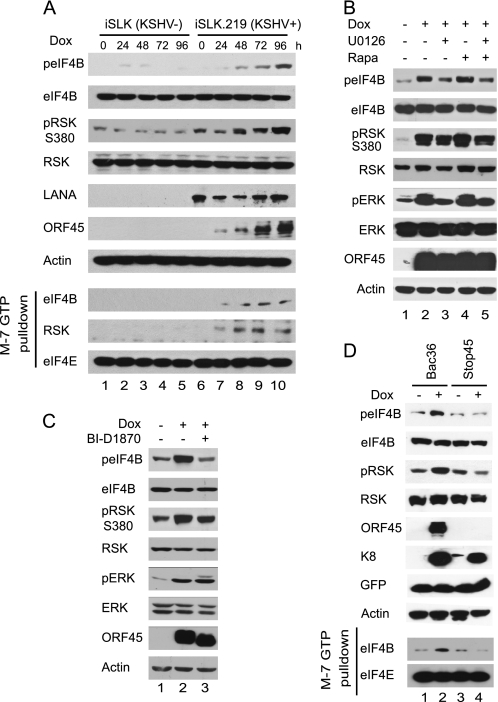
We have demonstrated that depletion of RSK by siRNA or inhibition of RSK activity by inhibitor BI-D1870 reduces the expression of KSHV lytic genes and production of infectious progeny viruses (17). We wished to determine whether the ORF45/RSK axis is involved in the regulation of the translation program during KSHV lytic replication. Recently, a doxycycline-inducible KSHV-producing cell line, iSLK.219, has been established (35). The cells maintain strict latency but support robust viral reactivation upon induction with doxycycline (Dox), which triggers expression of RTA (a key KSHV gene that is sufficient for initiating the lytic replication program from latency), routinely resulting in the expression of lytic genes in more than 95% cells. Dox induction also obviates the need for chemical stimuli, such as TPA, which has pleiotropic effects on cell signaling. These advantages make this cell/virus system an ideal one in which to analyze eIF4B phosphorylation and to determine the effect of the ORF45/RSK axis during the KSHV lytic cycle. When the iSLK.219 cells were treated with Dox, phosphorylation of eIF4B and RSK increased from 24 to 96 h after treatment (Fig. 5A, lanes 11–14), whereas phosphorylation in the uninfected parental iSLK cells remained largely unchanged under the same conditions (Fig. 5A, top panel, lanes 4–7). Phosphorylation of eIF4B consequently resulted in increased association with mRNA cap complexes in the KSHV-infected cells (Fig. 5A, lower panel). This experiment demonstrated unambiguously that KSHV lytic replication induces eIF4B phosphorylation and increases its association of eIF4b with translation initiation complexes.
FIGURE 5.
ORF45/RSK axis is required for efficient assembly of translation initiation complex during KSHV lytic replication. A, KSHV lytic replication induces eIF4B phosphorylation and its association with the translation initiation complex. The iSLK.219 cells maintain strict latency but support robust lytic reactivation upon induction with doxycycline (35). The iSLK.219 and its parental uninfected iSLK cells were treated with Dox. At the indicated time after treatment, cell lysates were prepared and analyzed by Western blot with the specified antibodies (top panel). The lysates were also used in m7GTP pulldown assays (lower panel). B, KSHV lytic replication induces phosphorylation of eIF4B through a distinct pathway. The iSLK cells were treated with Dox for 70 h; U0126 (25 μm) and/or rapamycin (Rapa) (50 nm) was added to the medium and incubated for 2 h. The cell lysates were then analyzed by Western blot with antibodies as indicated. C, phosphorylation of eIF4B during KSHV lytic replication requires RSK activity. The iSLK.219 cells were induced with Dox for 70 h and then treated with 10 μm BI-D1870 for 2 h. Cell lysates were analyzed by Western blotting with the antibodies indicated. D, KSHV-induced phosphorylation of eIF4B depends on ORF45. The iSLK cells were transfected with BAC36 or BAC-stop45 and then subjected to hygromycin selection, becoming cells latently infected with wild-type or ORF45-null recombinant KSHV viruses. The cells were treated with doxycycline for 3 days. Cell lysates were then prepared and analyzed by Western blots with antibodies as indicated. LANA, latency-associated nuclear antigen.
When the Dox-induced cells (70 h after induction) were treated with rapamycin and/or U1206, the phosphorylation of eIF4B was not blocked completely by either agent alone or by a combination of the two (Fig. 5B). The results were reminiscent of what we observed during KSHV lytic replication in the TPA-induced BCBL1 cells (Fig. 4B), confirming that eIF4B is phosphorylated through a distinct pathway caused by viral lytic replication. The phosphorylation of eIF4B was abolished by treatment with BI-D1870 (Fig. 5C), suggesting that RSK activity is required for the phosphorylation during KSHV lytic replication. Because ORF45 is phosphorylated efficiently by RSKs (17), a shift in electrophoretic mobility of ORF45 attested to effective inhibition of RSK activity by BI-D1870, although no apparent change in phosphorylation of RSKs was observed as reported previously (54). These experiments confirmed that KSHV lytic replication induces rapamycin and U0126 doubly insensitive eIF4B phosphorylation that requires RSK activity.
To determine whether the loss of ORF45 affects the phosphorylation of eIF4B and its association with the initiation complex during KSHV lytic replication, we next transduced the iSLK cells with the bacterial artificial chromosome (BAC) clone of KSHV wild-type BAC36 (36) and ORF45-null mutant BAC-stop45 as described previously (37) and then treated the stably transduced cells with Dox to initiate lytic replication. As shown in Fig. 5D, the Dox treatment increased the levels of phosphorylated RSK and eIF4B in the BAC36-iSLK cells but did not do so in the BAC-stop45-transduced cells under the same conditions (Fig. 5D, top panel). Consequently, the Dox-induced increase of the association of eIF4B with the initiation complex was abolished by null mutation of ORF45 (Fig. 5D, lower panel), suggesting that RSK activity is required for recruitment of eIF4B to the translation initiation complex during KSHV lytic replication. Together, these results supported the conclusion that ORF45-mediated RSK signaling plays a role in translational regulation by facilitating recruitment of eIF4B into the active initiation complex during KSHV lytic replication.
eIF4B Phosphorylation Promotes KSHV Lytic Replication
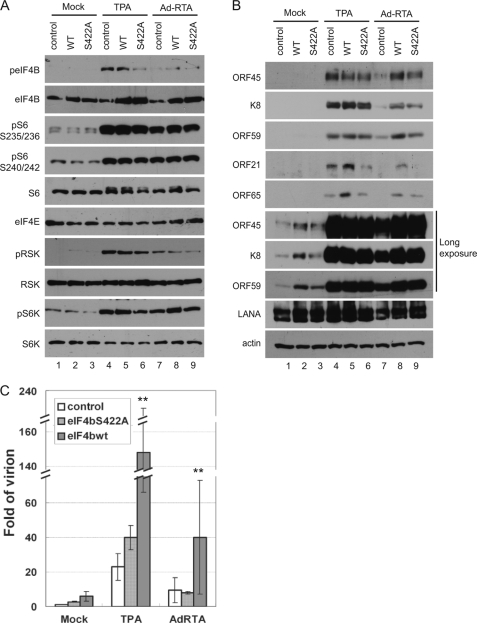
To determine the role of eIF4B phosphorylation in KSHV lytic infection, we ectopically expressed the eIF4B wild type or the phosphorylation-deficient S422A mutant in BCBL1 cells with lentivirus-based vectors and then treated the cells with TPA or Ad-RTA, an adenovirus-based vector expressing RTA, to induce KSHV lytic replication (44). As expected, the expression and phosphorylation of RSK, rpS6, S6K, and other proteins involved in translational regulation were not affected by overexpression of either eIF4B or eIF4B-S422A (Fig. 6A), but the expression of wild-type eIF4B increased the expressions of both early (ORF45, ORF59, and K8) and late KSHV lytic genes (ORF21 and ORF65), whereas expression of the phosphorylation-deficient eIF4B-S422A mutant showed no or a less dramatic effect (Fig. 6B). In contrast, overexpression of either one did not affect the latent expression of latency-associated nuclear antigen gene. Interestingly, the difference in the late lytic gene expression (ORF21 and ORF65) was much clearer in both TPA- and Ad-RTA-induced cells, whereas the difference in early lytic gene expression (ORF45, K8, and ORF59) was less so in TPA-induced cells but was more obvious in cells undergoing spontaneous or Ad-RTA-induced lytic replication presumably because the TPA-mediated cellular signaling pathways, including the ERK/RSK and mTOR/S6K pathways (Fig. 4A), induced phosphorylation of the endogenous eIF4B and thus blurred the effect. The effect of overexpression of eIF4B on viral lytic gene expression seems to occur mostly at the translational level because real time RT-PCR analysis revealed no significant difference in the levels of mRNAs (data not shown). These results suggested that eIF4B phosphorylation has a more dramatic effect on the translation of KSHV late lytic genes. Furthermore, overexpression of eIF4B increased the yield of KSHV virions significantly (about 5-fold) in cells induced by either TPA or Ad-RTA and in cells undergoing spontaneous lytic replication, whereas eIF4B-S422A did not (Fig. 6C). These data suggested that eIF4B phosphorylation promotes KSHV lytic gene expression and virion production. In summary, the ORF45/RSK axis-mediated eIF4B phosphorylation contributes to optimal KSHV lytic replication.
FIGURE 6.
eIF4B phosphorylation promotes KSHV lytic replication. A and B, ectopic expression of wild-type eIF4B but not its phosphorylation-deficient mutant increases expression of KSHV lytic genes. BCBL1 cells were stably transduced with lentivirus-based vectors that express eIF4B wild type or S422A mutant. The transduced cells were induced by TPA or Ad-RTA (a recombinant adenovirus expressing RTA) for 3 days. Phosphorylation of upstream kinases RSK and S6K and components of translation initiation complexes (A) and the expression of KSHV lytic genes (B) in cell lysates were determined by Western blot with antibodies as indicated. C, eIF4B phosphorylation promotes KSHV virion production. The yields of KSHV virion were estimated by real time PCR for determination of viral genomic copy numbers (**, p < 0.01). Error bar represents standard deviation. LANA, latency-associated nuclear antigen.
DISCUSSION
In the studies reported here, we demonstrated that ORF45-mediated activation of RSK results in phosphorylation of eIF4B and subsequently promotes its assembly into the translation initiation complex and recruitment of mRNA to the active translation machinery. We further demonstrated that depletion of RSK expression by siRNA, inhibition of RSK activity, or knock-out of ORF45 causes reduction of eIF4B phosphorylation, which is correlated with a lower level of viral lytic gene expression as we observed previously (17, 33). In addition, we found that ectopic expression of eIF4B in lytically KSHV-infected cells increases both viral lytic gene expression and progeny virus yield. Together, our data support the conclusion that the ORF45/RSK axis-mediated eIF4B phosphorylation is involved in translational control during KSHV lytic replication and is required for optimal expression of viral lytic genes.
Viruses rely on the cellular translational machinery for production of viral proteins; therefore, they have evolved a variety of strategies for superseding cellular mRNA translation while maintaining the synthesis of their own gene products. Many viruses encode products that directly modify various eIFs or recruit these factors to viral mRNAs (15, 16). For example, some RNA viruses, such as picornavirus, cleave the eIF4G with viral proteases to disrupt cap-dependent translation in favor of viral internal ribosomal entry site (IRES)-dependent protein synthesis. More commonly, viruses subvert the reversible phosphorylation process of eIF4E or its repressor 4E-BP to facilitate translational selectivity for viral mRNA during infection (15, 16). Both α (such as herpes simplex virus-1) and β (such as human cytomegalovirus) herpesviruses are known to modify eIF4F during lytic infection (16, 59–62). Translational control during γ herpesvirus lytic infection has not been well studied. A recent study by Arias et al. (63) demonstrated that KSHV reactivation causes profound changes in the protein synthesis profile that occur in parallel with phosphorylation of eIF4E by MNK1 (mitogen-activated protein kinase-interacting serine/threonine kinase) and phosphorylation of 4E-BP1 possibly by mTOR Complex 1. Interestingly, the authors observed that rapamycin could reduce 4E-BP phosphorylation efficiently but had no significant impact on KSHV lytic gene expression. They also noticed that eIF4E phosphorylation increased association of eIF4G but not itself with the m7GTP beads. These observations imply that a rapamycin-insensitive mechanism other than phosphorylation of 4E-BP and eIF4E is involved in translational regulation during KSHV lytic replication. We found that ORF45-mediated ERK/RSK signaling induces phosphorylation not only of MNK1 but also of eIF4B and rpS6 (Fig. 1).4 We demonstrated that the ORF45/RSK axis induces rapamycin-insensitive eIF4B phosphorylation, potentiates assembly of initiation complex, and subsequently promotes translation. Ectopic expression of eIF4B but not the phosphorylation-deficient S422A mutant in KSHV-infected primary effusion lymphoma cells increased viral lytic gene expression and progeny virus production. On the basis of these data, we believe that the ORF45/RSK-mediated eIF4B phosphorylation plays a critical role in translation regulation during KSHV lytic replication.
We have shown previously that the loss of RSK function has a profound impact on KSHV lytic gene expression and progeny virion production (17). The impact can be attributed partially to the roles of RSKs in translational regulation, but RSKs do not seem to be absolutely essential for global translation because Rsk1−/Rsk2−/Rsk3− triple knock-out mice are viable (45, 64). We noticed that growth of HEK293 and BCBL1 cells in which both RSK1 and RSK2 were significantly knocked down by siRNAs was comparable with that of control cells under normal culture conditions. Redundant pathways therefore seem to be involved in phosphorylation of eIF4B (8). eIF4B and rpS6 phosphorylation are known to be cell type- and stimulus-dependent. For example, the serum-induced eIF4B phosphorylation at Ser-422 requires RSK signaling, but the insulin-induced eIF4B phosphorylation does not rely on RSK but requires mTOR signaling (9). Currently, three AGC kinase family members, namely S6K, RSK, and AKT, have been shown to phosphorylate eIF4B at Ser-422 (8). These kinases seem to be modulated by other cellular factors in cell type- and stimulus-dependent manners. In lytically KSHV-infected cells, both RSK1 and RSK2 are associated with and activated by ORF45. Consequently, the ORF45-induced RSK signaling becomes a crucial pathway involved in eIF4B phosphorylation during the viral lytic cycle (17). Interestingly, although ORF45 increased phosphorylation of both eIF4B and rpS6, we found that phosphorylation of eIF4B was not sensitive to rapamycin but that phosphorylation of rpS6 was, suggesting that eIF4B is phosphorylated by ORF45-activated RSK directly, whereas rpS6 is phosphorylated by mTOR/S6K-dependent signaling that could be activated by RSK indirectly through phosphorylation of tuberous sclerosis protein 2 (55, 58) or other KSHV gene products, such as viral G protein-coupled receptor or K1 (27–29, 31). In any case, our data support the conclusion that the ORF45/RSK axis plays a crucial role in translational control during KSHV lytic replication through phosphorylation of eIF4B. The redundancy of the eIF4B phosphorylation pathways in general and the strict requirement of RSK for eIF4B phosphorylation in lytically KSHV-infected cells suggest that RSK is an ideal target for anti-KSHV intervention. Regulation of cellular signaling is expected to be cell type-specific; further studies are needed to determine whether the ORF45/RSK axis is universally required for KSHV lytic gene expression in other cell types, such as primary endothelial cells. Examination of the activation status of RSK and eIF4B in Kaposi sarcoma lesions and in specimens of other KSHV-associated diseases, such as primary effusion lymphoma and multicentric Castleman disease, will also be important.
In addition to regulating translation, RSKs regulate transcription because they are known to phosphorylate several transcriptional factors, such as cAMP-responsive element-binding protein, c-Fos, CCAAT/enhancer-binding protein, ATF-4, serum response factor, and histone H3 (45, 46). The case for their role in transcriptional regulation is also strengthened by recent discoveries that the ERK and RSK kinases are associated with chromosomes (65–67). We have shown that RSK1 and RSK2 are colocalized with ORF45 in both the cytoplasm and nucleus (17). The ORF45/RSK axis might contribute to optimal lytic gene expression through both translational and transcriptional regulation. Interestingly, although activation of RSK seems to be a conserved function among ORF45 homologues of γ2 herpesviruses,4 KSHV ORF45 is the only one that is predominantly localized in the cytoplasm; the other homologues are mostly nuclear (47). Determining whether the homologues in other γ2 herpesviruses play a role in translational regulation will be interesting.
Because eIF4B stimulates the helicase activity of eIF4A, eIF4B is postulated to be critical for translation of mRNAs with inhibitory secondary structures in their 5′-UTRs. Some cellular mRNAs with extensive 5′-UTRs encode proteins, such as CDC25, BCL-2, c-MYC, and VEGF, that play important roles in the regulation of the cell cycle, cell growth, survival, and apoptosis (68) and may have important implications in KSHV viral pathogenesis. Knowing which genes are preferentially regulated by ORF45/RSK-induced eIF4B phosphorylation will be important. Besides cellular genes, the viral genes might also be differentially regulated. Expressions of herpesviral genes are regulated temporally in a cascade fashion and are often accompanied by down-regulation of cellular gene translations as a result of a virally encoded “shutoff” function that induces accelerated cellular mRNA turnover (69, 70). Some herpesviral genes are encoded on polycistronic mRNA whose translation relies on the IRES. eIF4B is known to interact with the IRESs of several RNA viruses (71–75) and the virion host shutoff protein of herpesvirus simplex 1 (76). How eIF4B is involved in the translation regulation of herpesviral mRNAs remains to be explored. Our preliminary data suggest that the late lytic genes are more dependent on the ORF45/RSK signaling than the early genes, but only further studies will determine the selectivity and specificity of eIF4B phosphorylation-dependent translational regulation.
In most cells, phosphorylation of eIF4B is regulated by both the MEK/ERK/RSK and AKT/mTOR/S6K pathways. Both play critical roles in many important cell processes and have been shown to be deregulated in a plethora of neoplasias. Components of both pathways have been actively explored as targets of antitumor therapeutic strategies (77, 78). Rapamycin has proved very promising in treating KSHV-related diseases. It reduces KSHV-induced primary effusion lymphoma cell growth in culture and inhibits Kaposi sarcoma development in kidney transplant recipients (79, 80), but some primary effusion lymphoma cell lines were found to be relatively resistant to rapamycin (79, 81), and patient resistance to treatment has also been reported (82). The existence of a rapamycin-insensitive translational regulation mechanism suggests that combining an RSK inhibitor with rapamycin could be a successful strategy for the treatment of KSHV-related diseases (83).
Acknowledgments
We are grateful to Drs. Dario Alessi, John Blenis (through Addgene), Britt Glaunsinger, Shou-Jiang Gao, Deborah A. Lannigan, Nahum Sonenberg, Hengli Tang, and Keiji Ueda for kindly providing reagents. We thank Drs. Betty J. Gaffney, Kenneth H. Roux, and Hengli Tang and members of the Zhu laboratory for reading the manuscript and for helpful discussions. We also thank Anne B. Thistle at The Florida State University for excellent editorial assistance.
This work was supported, in whole or in part, by National Institutes of Health Grant R01DE016680 (to F. Z.). This work was also supported by a Bankhead-Coley bridge grant, a Florida State University setup fund, and a Florida State University planning grant (to F. Z.).
E. Kuang and F. Zhu, unpublished data.
- S6K
- ribosomal S6 kinase
- 4E-BP
- 4E-binding protein
- BAC
- bacterial artificial chromosome
- BCBL1
- body cavity-based lymphoma 1
- HCV
- hepatitis C virus
- IRES
- internal ribosomal entry site
- KSHV
- Kaposi sarcoma-associated herpesvirus
- mTOR
- mammalian target of rapamycin
- RSK
- p90 ribosomal S6 kinase
- TPA
- 12-O-tetradecanoylphorbol-13-acetate
- rpS6
- ribosomal S6 protein
- KD
- kinase-dead
- CA
- constitutively active
- Dox
- doxycycline
- RTA
- replication and transcription activator.
REFERENCES
- 1. Gingras A. C., Raught B., Sonenberg N. (1999) Annu. Rev. Biochem. 68, 913–963 [DOI] [PubMed] [Google Scholar]
- 2. Méthot N., Pause A., Hershey J. W., Sonenberg N. (1994) Mol. Cell. Biol. 14, 2307–2316 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Naranda T., Strong W. B., Menaya J., Fabbri B. J., Hershey J. W. (1994) J. Biol. Chem. 269, 14465–14472 [PubMed] [Google Scholar]
- 4. Méthot N., Song M. S., Sonenberg N. (1996) Mol. Cell. Biol. 16, 5328–5334 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Rozen F., Edery I., Meerovitch K., Dever T. E., Merrick W. C., Sonenberg N. (1990) Mol. Cell. Biol. 10, 1134–1144 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Lawson T. G., Lee K. A., Maimone M. M., Abramson R. D., Dever T. E., Merrick W. C., Thach R. E. (1989) Biochemistry 28, 4729–4734 [DOI] [PubMed] [Google Scholar]
- 7. Rogers G. W., Jr., Richter N. J., Lima W. F., Merrick W. C. (2001) J. Biol. Chem. 276, 30914–30922 [DOI] [PubMed] [Google Scholar]
- 8. van Gorp A. G., van der Vos K. E., Brenkman A. B., Bremer A., van den Broek N., Zwartkruis F., Hershey J. W., Burgering B. M., Calkhoven C. F., Coffer P. J. (2009) Oncogene 28, 95–106 [DOI] [PubMed] [Google Scholar]
- 9. Shahbazian D., Roux P. P., Mieulet V., Cohen M. S., Raught B., Taunton J., Hershey J. W., Blenis J., Pende M., Sonenberg N. (2006) EMBO J. 25, 2781–2791 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Holz M. K., Ballif B. A., Gygi S. P., Blenis J. (2005) Cell 123, 569–580 [DOI] [PubMed] [Google Scholar]
- 11. Raught B., Peiretti F., Gingras A. C., Livingstone M., Shahbazian D., Mayeur G. L., Polakiewicz R. D., Sonenberg N., Hershey J. W. (2004) EMBO J. 23, 1761–1769 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Ferrari S., Bandi H. R., Hofsteenge J., Bussian B. M., Thomas G. (1991) J. Biol. Chem. 266, 22770–22775 [PubMed] [Google Scholar]
- 13. Roux P. P., Shahbazian D., Vu H., Holz M. K., Cohen M. S., Taunton J., Sonenberg N., Blenis J. (2007) J. Biol. Chem. 282, 14056–14064 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Ruvinsky I., Meyuhas O. (2006) Trends Biochem. Sci. 31, 342–348 [DOI] [PubMed] [Google Scholar]
- 15. Gale M., Jr., Tan S. L., Katze M. G. (2000) Microbiol. Mol. Biol. Rev. 64, 239–280 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Mohr I. J., Pe'ery T., Mathews M. B. (2007) in Translational Control in Biology and Medicine (Mathews M. B., Sonenberg N., Hershey J. W. B. eds) pp. 545–599, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY [Google Scholar]
- 17. Kuang E., Tang Q., Maul G. G., Zhu F. (2008) J. Virol. 82, 1838–1850 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Buchkovich N. J., Yu Y., Zampieri C. A., Alwine J. C. (2008) Nat. Rev. Microbiol. 6, 266–275 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Pleschka S. (2008) Biol. Chem. 389, 1273–1282 [DOI] [PubMed] [Google Scholar]
- 20. Lambert P. J., Shahrier A. Z., Whitman A. G., Dyson O. F., Reber A. J., McCubrey J. A., Akula S. M. (2007) Expert Opin. Ther. Targets 11, 589–599 [DOI] [PubMed] [Google Scholar]
- 21. Mizutani T. (2007) Ann. N.Y. Acad. Sci. 1102, 86–95 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Panteva M., Korkaya H., Jameel S. (2003) Virus Res. 92, 131–140 [DOI] [PubMed] [Google Scholar]
- 23. Xie J., Ajibade A. O., Ye F., Kuhne K., Gao S. J. (2008) Virology 371, 139–154 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Pan H., Xie J., Ye F., Gao S. J. (2006) J. Virol. 80, 5371–5382 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Ford P. W., Bryan B. A., Dyson O. F., Weidner D. A., Chintalgattu V., Akula S. M. (2006) J. Gen. Virol. 87, 1139–1144 [DOI] [PubMed] [Google Scholar]
- 26. Cohen A., Brodie C., Sarid R. (2006) J. Gen. Virol. 87, 795–802 [DOI] [PubMed] [Google Scholar]
- 27. Sodhi A., Montaner S., Patel V., Gómez-Román J. J., Li Y., Sausville E. A., Sawai E. T., Gutkind J. S. (2004) Proc. Natl. Acad. Sci. U.S.A. 101, 4821–4826 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Sodhi A., Chaisuparat R., Hu J., Ramsdell A. K., Manning B. D., Sausville E. A., Sawai E. T., Molinolo A., Gutkind J. S., Montaner S. (2006) Cancer Cell 10, 133–143 [DOI] [PubMed] [Google Scholar]
- 29. Tomlinson C. C., Damania B. (2004) J. Virol. 78, 1918–1927 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Sharma-Walia N., Krishnan H. H., Naranatt P. P., Zeng L., Smith M. S., Chandran B. (2005) J. Virol. 79, 10308–10329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Bais C., Van Geelen A., Eroles P., Mutlu A., Chiozzini C., Dias S., Silverstein R. L., Rafii S., Mesri E. A. (2003) Cancer Cell 3, 131–143 [DOI] [PubMed] [Google Scholar]
- 32. Chaisuparat R., Hu J., Jham B. C., Knight Z. A., Shokat K. M., Montaner S. (2008) Cancer Res. 68, 8361–8368 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Kuang E., Wu F., Zhu F. (2009) J. Biol. Chem. 284, 13958–13968 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Vieira J., O'Hearn P. M. (2004) Virology 325, 225–240 [DOI] [PubMed] [Google Scholar]
- 35. Myoung J., Ganem D. (2011) J. Virol. Methods 174, 12–21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Zhou F. C., Zhang Y. J., Deng J. H., Wang X. P., Pan H. Y., Hettler E., Gao S. J. (2002) J. Virol. 76, 6185–6196 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Zhu F. X., Li X., Zhou F., Gao S. J., Yuan Y. (2006) J. Virol. 80, 12187–12196 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Zhu F. X., Yuan Y. (2003) J. Virol. 77, 4221–4230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Zhu F. X., Chong J. M., Wu L., Yuan Y. (2005) J. Virol. 79, 800–811 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Schalm S. S., Blenis J. (2002) Curr. Biol. 12, 632–639 [DOI] [PubMed] [Google Scholar]
- 41. Zhu F. X., Cusano T., Yuan Y. (1999) J. Virol. 73, 5556–5567 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Yam P. Y., Li S., Wu J., Hu J., Zaia J. A., Yee J. K. (2002) Mol. Ther. 5, 479–484 [DOI] [PubMed] [Google Scholar]
- 43. Tiscornia G., Singer O., Verma I. M. (2006) Nat. Protoc. 1, 241–245 [DOI] [PubMed] [Google Scholar]
- 44. Bechtel J. T., Liang Y., Hvidding J., Ganem D. (2003) J. Virol. 77, 6474–6481 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Anjum R., Blenis J. (2008) Nat. Rev. Mol. Cell Biol. 9, 747–758 [DOI] [PubMed] [Google Scholar]
- 46. Carriere A., Ray H., Blenis J., Roux P. P. (2008) Front. Biosci. 13, 4258–4275 [DOI] [PubMed] [Google Scholar]
- 47. Li X., Zhu F. (2009) J. Virol. 83, 2531–2539 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Angenstein F., Greenough W. T., Weiler I. J. (1998) Proc. Natl. Acad. Sci. U.S.A. 95, 15078–15083 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Tominaga Y., Tamgüney T., Kolesnichenko M., Bilanges B., Stokoe D. (2005) Mol. Cell. Biol. 25, 8465–8475 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Ferraiuolo M. A., Basak S., Dostie J., Murray E. L., Schoenberg D. R., Sonenberg N. (2005) J. Cell Biol. 170, 913–924 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Fuerst T. R., Niles E. G., Studier F. W., Moss B. (1986) Proc. Natl. Acad. Sci. U.S.A. 83, 8122–8126 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Shatkin A. J. (1976) Cell 9, 645–653 [DOI] [PubMed] [Google Scholar]
- 53. Fuerst T. R., Moss B. (1989) J. Mol. Biol. 206, 333–348 [DOI] [PubMed] [Google Scholar]
- 54. Sapkota G. P., Cummings L., Newell F. S., Armstrong C., Bain J., Frodin M., Grauert M., Hoffmann M., Schnapp G., Steegmaier M., Cohen P., Alessi D. R. (2007) Biochem. J. 401, 29–38 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Roux P. P., Ballif B. A., Anjum R., Gygi S. P., Blenis J. (2004) Proc. Natl. Acad. Sci. U.S.A. 101, 13489–13494 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Iijima Y., Laser M., Shiraishi H., Willey C. D., Sundaravadivel B., Xu L., McDermott P. J., Kuppuswamy D. (2002) J. Biol. Chem. 277, 23065–23075 [DOI] [PubMed] [Google Scholar]
- 57. Wang L., Gout I., Proud C. G. (2001) J. Biol. Chem. 276, 32670–32677 [DOI] [PubMed] [Google Scholar]
- 58. Ma L., Chen Z., Erdjument-Bromage H., Tempst P., Pandolfi P. P. (2005) Cell 121, 179–193 [DOI] [PubMed] [Google Scholar]
- 59. Smith R. W., Graham S. V., Gray N. K. (2008) Biochem. Soc. Trans. 36, 701–707 [DOI] [PubMed] [Google Scholar]
- 60. Walsh D., Mohr I. (2004) Genes Dev. 18, 660–672 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Walsh D., Perez C., Notary J., Mohr I. (2005) J. Virol. 79, 8057–8064 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Walsh D., Mohr I. (2006) Genes Dev. 20, 461–472 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Arias C., Walsh D., Harbell J., Wilson A. C., Mohr I. (2009) PLoS Pathog. 5, e1000334 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Dumont J., Umbhauer M., Rassinier P., Hanauer A., Verlhac M. H. (2005) J. Cell Biol. 169, 227–231 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Dioum E. M., Wauson E. M., Cobb M. H. (2009) Cell 139, 462–463 [DOI] [PubMed] [Google Scholar]
- 66. Lawrence M. C., McGlynn K., Shao C., Duan L., Naziruddin B., Levy M. F., Cobb M. H. (2008) Proc. Natl. Acad. Sci. U.S.A. 105, 13315–13320 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Hu S., Xie Z., Onishi A., Yu X., Jiang L., Lin J., Rho H. S., Woodard C., Wang H., Jeong J. S., Long S., He X., Wade H., Blackshaw S., Qian J., Zhu H. (2009) Cell 139, 610–622 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Shahbazian D., Parsyan A., Petroulakis E., Topisirovic I., Martineau Y., Gibbs B. F., Svitkin Y., Sonenberg N. (2010) Mol. Cell. Biol. 30, 1478–1485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Glaunsinger B., Ganem D. (2004) Mol. Cell 13, 713–723 [DOI] [PubMed] [Google Scholar]
- 70. Kwong A. D., Kruper J. A., Frenkel N. (1988) J. Virol. 62, 912–921 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Ochs K., Saleh L., Bassili G., Sonntag V. H., Zeller A., Niepmann M. (2002) J. Virol. 76, 2113–2122 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Ochs K., Rust R. C., Niepmann M. (1999) J. Virol. 73, 7505–7514 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Rust R. C., Ochs K., Meyer K., Beck E., Niepmann M. (1999) J. Virol. 73, 6111–6113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Meyer K., Petersen A., Niepmann M., Beck E. (1995) J. Virol. 69, 2819–2824 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Baglioni C., Simili M., Shafritz D. A. (1978) Nature 275, 240–243 [DOI] [PubMed] [Google Scholar]
- 76. Doepker R. C., Hsu W. L., Saffran H. A., Smiley J. R. (2004) J. Virol. 78, 4684–4699 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Roberts P. J., Der C. J. (2007) Oncogene 26, 3291–3310 [DOI] [PubMed] [Google Scholar]
- 78. Dowling R. J., Pollak M., Sonenberg N. (2009) BioDrugs 23, 77–91 [DOI] [PubMed] [Google Scholar]
- 79. Sin S. H., Roy D., Wang L., Staudt M. R., Fakhari F. D., Patel D. D., Henry D., Harrington W. J., Jr., Damania B. A., Dittmer D. P. (2007) Blood 109, 2165–2173 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Lee J. S., Li Q., Lee J. Y., Lee S. H., Jeong J. H., Lee H. R., Chang H., Zhou F. C., Gao S. J., Liang C., Jung J. U. (2009) Nat. Cell Biol. 11, 1355–1362 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Nichols L. A., Adang L. A., Kedes D. H. (2011) PLoS One 6, e14535 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Boulanger E., Afonso P. V., Yahiaoui Y., Adle-Biassette H., Gabarre J., Agbalika F. (2008) Am. J. Transplant. 8, 707–710 [DOI] [PubMed] [Google Scholar]
- 83. Nguyen T. L. (2008) Anticancer Agents Med. Chem. 8, 710–716 [DOI] [PubMed] [Google Scholar]