Background: MHC-peptide multimers are used to analyze and sort antigen-specific CD8+ T cells.
Results: NTA-His tag containing multimers can be switched from stable binding to rapid dissociation.
Conclusion: These reagents allow improved analysis and sorting of bona fide T cells.
Significance: New tools for improved CD8+ T cells analysis.
Keywords: Annexin, Antigen, Cysteine-mediated Cross-linking, Immunology, Lymphocyte, Major Histocompatibility Complex (MHC), T Cell, T Cell Receptor, Multimers, Tetramers
Abstract
MHC-peptide multimers containing biotinylated MHC-peptide complexes bound to phycoerythrin (PE) streptavidin (SA) are widely used for analyzing and sorting antigen-specific T cells. Here we describe alternative T cell-staining reagents that are superior to conventional reagents. They are built on reversible chelate complexes of Ni2+-nitrilotriacetic acid (NTA) with oligohistidines. We synthesized biotinylated linear mono-, di-, and tetra-NTA compounds using conventional solid phase peptide chemistry and studied their interaction with HLA-A*0201-peptide complexes containing a His6, His12, or 2×His6 tag by surface plasmon resonance on SA-coated sensor chips and equilibrium dialysis. The binding avidity increased in the order His6 < His12 < 2×His6 and NTA1 < NTA2 < NTA4, respectively, depending on the configuration of the NTA moieties and increased to picomolar KD for the combination of a 2×His6 tag and a 2×Ni2+-NTA2. We demonstrate that HLA-A2–2×His6-peptide multimers containing either Ni2+-NTA4-biotin and PE-SA- or PE-NTA4-stained influenza and Melan A-specific CD8+ T cells equal or better than conventional multimers. Although these complexes were highly stable, they very rapidly dissociated in the presence of imidazole, which allowed sorting of bona fide antigen-specific CD8+ T cells without inducing T cell death as well as assessment of HLA-A2-peptide monomer dissociation kinetics on CD8+ T cells.
Introduction
Conjugation of proteins, DNA/RNA, fluorescent dyes to Qdots, microspheres, sensor or microarray chips, or ELISA plates is crucial for many applications in basic and applied sciences. The extremely strong binding of biotin to streptavidin (SA)2 (KD = ∼1 × 10−14 m) has been extensively used for such applications (1). Although for some applications, this practically irreversible binding is desirable, for others it is not, e.g. (i) introduction of macromolecules into cells by means of a delivery peptide reversibly conjugated to the molecules of interest (2); (ii) purification of recombinant proteins via reversible adsorption on an affinity matrix, e.g. Ni2+-NTA column (3); or (iii) reversible staining and sorting of antigen-specific T cells (4, 5). MHC-peptide tetramers are reagents that are widely used to enumerate, analyze, and isolate antigen-specific T cells (6, 7). CD8+ T cells express clonotypic T cell antigen receptors (TCR) that bind cognate MHC class I-peptide complexes. Although the binding of monomers to T cells is unstable, the binding of tetramers is stable and allows detection, enumeration, and isolation of antigen specific CD8+ T cells by flow cytometry. MHC class I-peptide monomers are produced by refolding of MHC heavy and light chain in the presence of a peptide of interest and subsequent biotinylation at a C-terminal biotinylation sequence peptide (BSP) by means of the biotin-transferase BirA; they are subsequently tetramerized by incubation with phycoerythrin (PE) or allophycocyanine-conjugated SA (6, 7).
A major shortcoming of these tetramers is that they avidly bind to and cross-link cell surface TCR and CD8, thereby inducing strong T cell activation, which frequently provokes T cell death (4, 5, 8, 9). Thus, isolation of antigen-specific CD8+ T cells by tetramers harbors the risk of substantial T cell loss. To circumvent this, “reversible” tetramers have been developed that contain low affinity biotin analogues and therefore dissociate upon addition of free biotin (4, 5, 9). However, although these reagents improved sorting and cloning of live antigen-specific CD8+ T cells, they are costly to produce and of limited stability, even at room temperature, commonly used for FACS sorting. Furthermore, there is strong evidence that the dissociation kinetic of TCR-MHC-peptide complexes is a key determinant for antigen-specific CD8+ T cell activation (10, 11). Dissociation kinetics have been assessed by measuring tetramer dissociation on CD8+ T cells. However, results from such experiments are error prone, e.g. because of rebinding of dissociated tetramers and internalization that rapidly occurs at elevated temperatures. Based on our previous demonstration that dissociation kinetics can be accurately assessed on living CD8+ T cells by using MHC-peptide monomers and TCR photoaffinity labeling (12), we reasoned that reversible tetramers might provide an alternative, generally applicable means to conclusive measure of MHC-peptide monomers dissociation kinetics on living cells.
Here we describe the preparation of MHC-peptide multimers that are built on Ni2+-NTA moieties and oligohistidine-tagged HLA-A*0201-influenza matrix peptide58–66 (A2/Flu) complexes. Mono-Ni2+-NTA compounds form reversible coordination complexes with oligohistidines, which for a His6 have a KD of ∼10−6 m (13–17). Although this is sufficient for purification of His-tagged recombinant proteins from culture supernatants (3), it is not sufficient for the preparation of staining reagents, which must be stable for months. Previous studies have shown that two His6 tags joined by a flexible linker greatly strengthens their binding to mono-Ni2+-NTA groups (18, 19). Moreover, it has been demonstrated that the affinity of linear or branched di-, tri-, and tetra-Ni2+-NTA compounds for His6- and His10-tagged molecules increases substantially with the number of Ni2+-NTA entities up to subnanomolar KD (2, 20–24). However, the synthesis of such NTA compounds requires advanced organic chemistry, which limits their application to specialized laboratories. Here we describe the synthesis of biotinylated linear mono-, di-, and tetra-NTA compounds based on conventional, i.e. widely accessible, solid phase peptide synthesis. We prepared A2/Flu monomers containing a C-terminal His6 or His12 or a double hexahistidine (2×His6) tag and studied their interactions with the different NTA compounds by surface plasmon resonance (SPR) and equilibrium dialysis. We report that the KD decreased in the order NTA1 > NTA2 > NTA4 and His6 > His12 > 2×His6, respectively, reaching 1.9 × 10−11 m for the combination of NTA4 and 2×His6 A2/Flu complexes. Although these complexes are remarkably stable, they dissociate rapidly upon the addition of imidazole, which we demonstrate permits FACS sorting of bona fide antigen-specific CD8+ T cells and conclusive assessment of MHC-peptide monomer dissociation kinetics on CD8+ T cells.
EXPERIMENTAL PROCEDURES
Chemical Synthesis of NTA Compounds
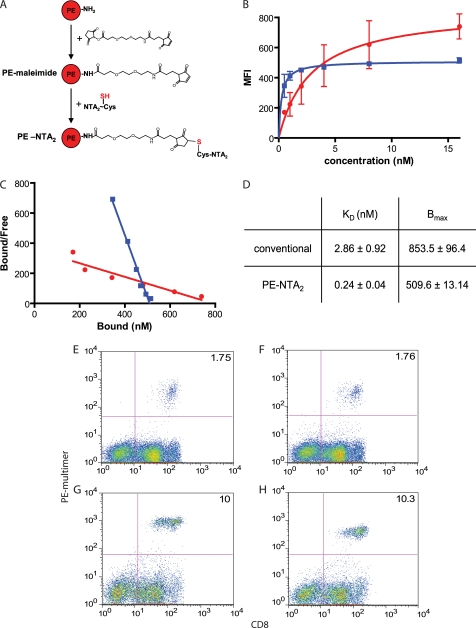
Detailed descriptions of the synthesis and characterization of the NTA compounds 1–4 (see Fig. 1A) are provided in the supplemental materials. In brief, the compounds 1, 2, and 4 were obtained by alkylation of the corresponding thio-peptide precursors with N-(5-(3-maleimidopropionylamino)-1-carboxy-pentyl)iminodiacetic acid (Dojindo Laboratories, Kumamoto, Japan) in phosphate buffer (0.1 m; pH 7.2). The precursors were obtained by conventional solid phase peptide chemistry using chlorotrityl resins. Compound 3 was obtained by reacting the precursor peptide, which was synthesized on a rink resin and upon deprotection contained two orthogonal carboxyl groups, with H2N-NTA(but)3 (23) in dimethylformamide. The thiopeptide used for alkylation of PE-maleimide was synthesized like compound 2, except that Fmoc-Cys(Acm)-OH was used instead of lysine-ϵ-amino-caproyl-biotin. The Acm protected group resisted TFA treatment and was removed with HgII in acetic acid (20%) for 1 h and then purified by HPLC (Rt = 13.9 min). The yield was 37% (ESI-MS calculated for C45H72N9O16S2 [M+H]+ was 1384.5; found was 1385.3).
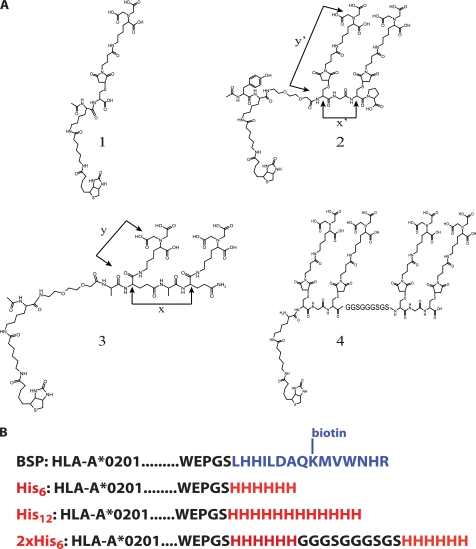
FIGURE 1.
The NTA compounds and His tag under study. A, structures of the four peptidic NTA compounds (compounds 1–4) in this study. The approximate maximal distances are: x = 7 Å, y = 8 Å, x′ = 5 Å, and y′ = 14 Å. B, sequence of the different sequences added C-terminally at the HLA-A2 heavy chain, including the biotinylation sequence peptide BSP, His6, His12, or two His6 tags (2xHis6) separated by a flexible GGGSGGGSGS spacer (as in compound 4).
Surface Plasmon Resonance Experiments
SPR experiments were performed on a BIACore 3000 instrument using SA-coated, CM5 sensor chips (BIACore; GE Healthcare). The chips were loaded by injecting compounds 1–4 in flow cells 2, 3, and 4 at a rate of 10 μl/min in running buffer (10 mm HEPES, 150 mm NaCl, 50 μm EDTA, pH 7.4). The average degree of loading was ∼100 RU (range, 75–130 RU). Free biotin binding sites were blocked with biotin. Cell 1 was used as control for changes in bulk refractive index. All of the experiments were performed in duplicate at 25 °C and repeated two to four times. All of the buffers were filtered and degassed prior to use. For kinetic experiments A2-His6 Flu peptide complexes were diluted in running buffer in 1:2 serial dilutions starting from 22 μm, and A2-His12 or A2–2×His6 peptide complexes in 1:3 serial dilutions, starting from 7.4 μm and passed over the Ni2+-NTA-biotin-SA-coated and control flow cells at 50 μl/min. As described previously (16, 17), the steps were as follows: (i) rinsing the flow cells with running buffer; (ii) activation of peptide-NTA with Ni2+ (500 μm NiCl 2+ in running buffer); (iii) rinsing in running buffer; (iv) injection of His-tagged A2/Flu complexes at different concentrations and measuring changes in RU measured over 60 s and then over an uninterrupted dissociation period of 6 min; (v) His-tagged protein removal by washing with imidazole solution (500 mm in water); (vi) NiCl2+ removal with regeneration buffer (10 mm HEPES, 150 mm NaCl, 0.005% polysorbate 20, 350 mm EDTA, pH 7.4); and (vii) rinse in dispenser buffer (10 mm HEPES, 150 mm NaCl, 0.005% polysorbate 20, 3 mm EDTA, pH 7.4). The kon and koff values were calculated assuming 1:1 Langmuir binding, and data were analyzed using BIAevaluation 4.1 software and a global fit algorithm. For each concentration of His-tagged A2/Flu complexes, the response of the control cell was subtracted.
Multimers
The preparation of the A2-BSP heavy chain has been described previously (4, 25, 26). Those containing the A2-His tags were prepared analogously using phosphorylated forward and reverse primers encoding the His6, His12, and 2×His6 tags with BamHI in 5′ and HindIII in 3′; complementary primers were annealed. A2/Flu monomers were obtained by refolding of the different A2 heavy chains in the presence of β2m and influenza matrix peptide58–66 (GILGFVFTL) and purified on a Superdex S75 column as described (4, 25, 26). BSP A2/Flu58–66 multimers were prepared by mixing biotinylated A2/Flu monomers with SA-PE (Invitrogen). For the preparation of SA-PE-NTA multimers, SA-PE was first incubated with compound 1, 2, 3, or 4 (5-fold molar excess) at 4 °C for 1 h, followed by incubation for 30 min with NiSO4 (10 mm). Excess of reagents were removed using spin columns (ZebaTM Spin Desalting Columns; Thermo Scientific). His-tagged A2/Flu monomers were incubated at 4 °C with SA-PE-biotin-Ni2+-NTA at a 10-fold molar excess for ≥16 h. Melan A and Epstein-Barr virus (EBV) monomers were prepared likewise using the Melan A peptide 26–35 (ELAGIGILTV) or EBV BMFl1peptide 280–288 (GLCTLVAML). A2/Flu58–66 PE-streptamer was obtained from IBA (Lucerna Ltd, Lucerne, Switzerland). PE-NTA2 conjugates were prepared by first reacting PE (Sigma) (50 nm) in phosphate buffer (0.1 m, pH 7.2 with SM(PEG)2 (succinimidyl-[(N-maleimidopropionamido)-diethyleneglycol] ester (Pierce) (10 mm) at room temperature for 2 h. Excess reagents were removed by spin columns. The resulting PE-maleimide was incubated under argon in phosphate buffer (100 mm, pH 7.0) with cysteine-NTA2 (50 mm) at room temperature for 2 h. After incubation for 30 min with NiSO4 (100 mm), excess reagents were removed by spin columns; the concentration of Ni2+-NTA2-PE was determined by Abs at 565 nm.
Cells under Study
PBMC were stimulated with peptides as described (25). The Flu-, Melan A-, and EBV-specific CD8+ T cell clones or lines were obtained by limiting dilution cloning of peptide-stimulated PBMC from healthy donors or a melanoma patient Lau 1164 and cultured in RPMI 1640 medium supplemented with 8% human serum and rIL-2 (150 units/ml) (CTL medium).
Multimer Binding Assays
For binding studies, peptide-stimulated PBMC (5 × 104) were incubated for 30 min at room temperature with graded concentrations of A2/Flu multimersin 50 μl of FACS buffer (PBS supplemented with 0.5% BSA, 15 mm HEPES, and 0.02% NaN3). After washing, cell-associated fluorescence was measured on a LSR II flow cytometer (BD, Allschwil, Switzerland) using gating on live cells. The data were processed using the FlowJo software (Tree Star, Inc., Ashland, OR). For dissociation experiments, PBMC were incubated for 60 min at 4 °C with 8 nm multimer and washed, and after different periods of incubation in FACS buffer containing or not anti-HLA2 mAb BB7.2 mAb (100 μg/ml; Serotec) at the indicated temperatures, cell-associated fluorescence was determined by flow cytometry. In some experiments washed cells were resuspended in FACS buffer supplemented with 100 mm imidazole or 10 mm biotin and reincubated for different periods of time. For cell viability experiments CD8+ T cells (105 cells) were incubated at 4 °C for 1 h and stained with 8 nm of multimers for 15 min at 4 °C. After staining with conventional BSP multimers, annexin V-FITC, and DAPI (Molecular Probes, Basel, Switzerland), the cells were analyzed by flow cytometry. Live cells were enumerated as a percentage of annexin V− and DAPI− cells in the live lymphocyte gate, using untreated cells as reference (100%).
CD8+ T Cell Sorting
Flu peptide-stimulated PBMC were incubated for 1 h at 4 °C, stained for 15 min at 4 °C with 4 nm of conventional BSP, SA-PE-NTA4-A2–2×His6 multimers, or PE-streptamer. After washing, the cells were FACS sorted on a FACS Aria sorter (BD, Allschwil, Switzerland). A2/Flu multimer+ cells (5 × 105 cells) were collected in medium containing imidazole (100 mm) or 10 mm biotin, washed, and plated in a 96-well plate. After incubation in CTL medium for 1, 4, 24, and 48 h, the number of viable, cells was enumerated by flow cytometry upon staining with A2/Flu BSP multimer, gating on live lymphocytes. The number of tetramer+ live cells at time 0 was referred to as 100%.
Cytolytic Assays
Cytolytic activity was assessed in a 4-h chromium release experiment using as targets T2 cells (5 × 105 cells in 100 μl of FCS) previously incubated with 100 μCi of 51Cr for 1 h at 37 °C, washed, and then incubated with serial dilutions of specific peptide for an additional 1 h at room temperature. As negative control, 51Cr-labeled T2 cells were loaded with an irrelevant peptide. An E:T ratio 10/1 was used. After 4 h of coincubation at 37 °C, the released 51Cr was assessed in 100 μl of supernatant, and the specific lysis was calculated as a percentage of specific lysis as follows: [(sample release − spontaneous release)/(maximal release − spontaneous release)] × 100.
A2/Flu58–66 Monomer Dissociation Kinetics
Cloned Flu-specific BC74 CTL were incubated with PE-Ni2+-NTA2-A2/Flu-2×His6 multimers (6 nm) containing Cy5-labeled β2m, prepared by alkylation of β2m containing the S88C mutation with Cy5-maleimide (GE Healthcare) at 4 °C for 45 min, washed, and incubated for variable periods at 4 °C in FACS buffer. PE and Cy5 fluorescence was measured before and after the addition of imidazole (100 mm, dilution factor: 60-fold).
RESULTS
Synthesis of Mono-, Di-, and Tetra-NTA Compounds
To indentify suitable NTA moieties, we synthesized linear mono-, di-, and tetra-NTA peptides using conventional peptide synthesis. Based on the observation that the affinity of His6-tagged molecules for Ni2+-NTA increases with their valence (20–24), we initially synthesized the NTA2 compound 3 (Fig. 1A and supplemental Fig. S1A). The coupling of the precursor (P3) with tri-but NTA-lysine in solution was inefficient. The sequence Ala-Glu-Ala-Glu- (where Glu indicates inverted Glu) was used, because with Ala-Glu-Ala-Glu or Glu-Glu sequences, very poor coupling yields were obtained, presumably because of steric hindrance of adjacent bulky tri-but-lysines. Moreover, tri-but NTA-lysine has to be prepared (23), and the carrier peptide cannot contain groups reactive to activated carboxyl residues (i.e. OH, SH, or NH2), which imposes constraints on their synthesis. We therefore established another strategy in which peptides containing one, two, or four cysteines were first synthesized (supplemental Fig. S1B) and then alkylated in aqueous solution with commercially available maleimido-NTA. The corresponding NTA thioethers (compounds 1, 2, and 4) were obtained in high yields and purity. The NTA4 compound 4 contains two NTA2 moieties (as in compound 2) linked by a flexible GGGSGGGSGS spacer.
Preparation of His-tagged A2/Flu Monomers
To find a highly stable Ni2+-NTA-His tag pair, we prepared A2/Flu monomers containing the C terminus of the α3 domain of the heavy chain His6, His12, or 2×His6 tag (Fig. 1B). These heavy chains were refolded in the presence of β2m and Flu matrix peptide58–66. The refolding efficiency of the 2×His6-tagged complex was nearly as high (98%) as the one of the BSP complex (data not shown). For the His6-tagged complex, the refolding efficiency was ∼85%, but for the His12-tagged complex it was only 60%. Essentially the same results were obtained when using other peptides or other HLA alleles (data not shown).
SPR Binding Studies on Ni2+-NTA Complexes and His-tagged HLA-A2/Flu Complexes
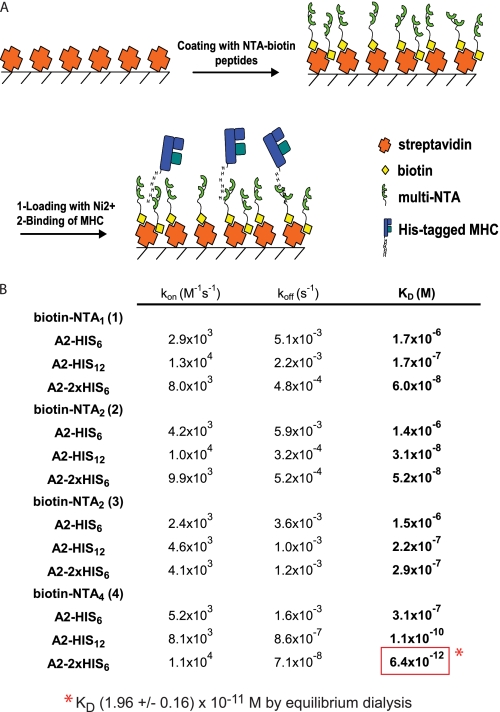
To study the interaction of the different A2/Flu monomers and biotinylated NTA molecules (Fig. 1), the latter were adsorbed on SA-coated sensor chips, using an equal or a very low degree of loading (∼100 RU). After saturation with biotin and NiCl2, graded concentrations of the different His-tagged A2/Flu complexes were passed over the chips, and the changes in RU were measured for 1 min; thereafter the decrease of RU was recorded upon flowing wash buffer (Fig. 2A and supplemental Fig. S3). Because the binding of biotin to SA is exceedingly stable (1), we disregarded dissociation of biotin from SA and calculated the binding on (kon) and off rates (koff), assuming a 1:1 Langmuir binding and from these the KD. A nearly five log difference between the binding of the His6-tagged A2/Flu complexes to immobilized Ni2+-NTA1 and the binding of 2×His6-tagged complexes to Ni2+-NTA4 was observed (Fig. 2B and supplemental Table S1). Because the latter value is beyond the range of accurate SPR measurements, we verified this KD value by equilibrium dialysis using 125I-labeled NTA4 as radioactive ligand. This analysis gave a KD value of 1.96 × 10−11 m, i.e. ∼3-fold higher than the one determined by SPR (supplemental Fig. S4), most likely explained by an overestimate of the koff by SPR, because of the rebinding of analyte.
FIGURE 2.
Binding of His-tagged A2/Flu monomers to immobilized Ni2+-NTA moieties assessed by SPR. A, scheme of SPR binding assay. Streptavidin-coated sensor chips were loaded with biotinylated NTA compounds (100 RU each) and loaded with NiCl2, and changes in RU were measured upon incubation with His-tagged A2/Flu complexes. B, summary of SPR binding parameters for the indicated Ni2+-NTA compounds A2/Flu complexes. The lowest KD is boxed in red. The mean values were calculated from at least three experiments. Standard deviations are shown in supplemental Table S1. The KD indicated as footnote was determined by equilibrium dialysis (see supplemental Fig. S4).
For the His6-tagged A2/Flu complex, the KD decreased modestly on the order NTA1 > NTA2 > NTA4. However, for His12-tagged complexes, the affinity increased more substantially, notably by 3 orders of magnitude in the case of NTA4. Consistent with previous reports, we observed that the KD was over 100-fold lower for A2/Flu complexes carrying a 2×His6 tag compared with those carrying a single His6 tag (18, 19).
Binding Isotherms of NTA1, NTA2, and NTA4-biotin-SA-PE Multimers on Flu-specific CD8+ T Cell Populations
To examine the usefulness of the different Ni2+-NTA-His tag combinations for the preparation of CD8+ T cell-staining reagents, SA-PE was loaded with the different biotin-NTA compounds and conjugated with His-tagged A2/Flu58–66 monomers. The room temperature binding isotherms of A2/Flu multimers containing the 2×His6 tag exhibited striking differences depending on the NTA moiety (Fig. 3, A and D). The NTA4 containing multimers displayed the most avid binding, followed by the NTA2 (compound 2) multimers. The KD values calculated from Scatchard analysis were 1.5 and 2.7 nm, respectively. Nonspecific staining was essentially background (not shown). The multimers containing NTA2 (compound 3) bound markedly less avidly and for the NTA1 (compound 1)-containing multimers, no KD could be conclusively determined. For comparison, we included commercial A2/Flu58–66 streptamers, which exhibited less avid binding than the NTA4 and NTA2 (compound 2)-containing multimers. We repeated this experiment using multimers containing His12-tagged A2/Flu complexes. The binding of all NTA-containing multimers were reduced compared with the 2×His6 containing counterparts (Fig. 3, B and D). The difference was particularly pronounced for multimers containing NTA2 compound 2. The results from the staining experiments correlated with those from the SPR experiments, i.e. the apparent KD observed for the multimer binding on cells showed the same hierarchy as those determined by SPR (Figs. 2B and 3D) and demonstrate that multimers containing Ni2+-NTA2 (compound 2) and Ni2+-NTA4 (compound 4) and 2×His6-tagged A2/peptide complexes are highly stable and efficient CD8+ T cell-staining reagents.
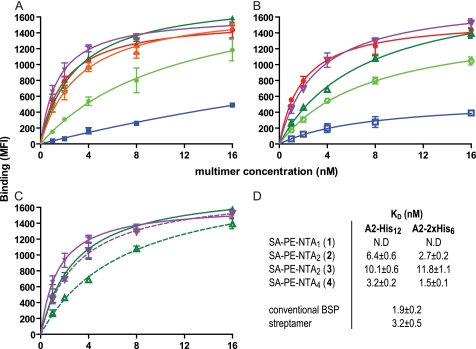
FIGURE 3.
Binding isotherms of SA-PE-NTA A2/Flu multimers on Flu-specific CD8+ T cell population. A, Flu58–66 peptide-stimulated PBMC were incubated at ambient temperature for 30 min with graded concentrations of SA-PE A2/Flu multimers containing a 2×His6 tag and biotinylated NTA1 (compound 1) (blue squares), NTA2 (compound 2) (dark green triangles), NTA2 (compound 3) (light green circles), or NTA4 (compound 4) (purple triangles). For comparison, conventional BSP multimers (red circles) and streptactin multimers (orange diamonds) are included. After washing, cell-associated fluorescence was assessed by flow cytometry. B, the same experiment was performed with SA-PE A2/Flu multimers containing a His12 tag. C, comparison of the binding of SA-PE A2/Flu multimers containing either a 2×His6 tag (solid lines, full symbols) or a His12 tag (dashed lines, empty symbols) and NTA2 (compound 2) (green triangles) or NTA4 (compound 4) (purple triangles), respectively. D, dissociation constants (KD) and maximal binding values (Bmax) observed for the indicated SA-PE A2/Flu multimers. The mean values and S.D. were calculated from at least three experiments.
NTA-His Tag Built Multimers Are Rapidly Reversible
To examine the stability of the staining of Flu-specific CD8+ T cells by different A2/Flu multimers, polyclonal Flu-specific CD8+ T cells were incubated in the cold with equal concentrations of multimers, washed, and incubated for different periods of time at ambient temperature. The stability of 2×His6 tag containing multimers increased in the order NTA1 (compound 1) < NTA2 (compound 2) < NTA4 (compound 4) (Fig. 4A). The dissociation of the NTA4 and BSP multimers were remarkably slow, reaching ∼92% of initial binding after 2 h. The sluggish dissociation of BSP multimers has been shown to be accounted for in part by rebinding (27); indeed in the presence of unlabeled tetramers or anti-A2 antibody, considerably faster dissociation kinetics were observed (half-lives of 11.9 and 5.4 min, respectively; supplemental Fig. S5). For the NTA2 (compound 2)-containing multimer, the dissociation was slightly faster, reaching ∼82% after 2 h. However, for A2/Flu streptamer and NTA1-containing multimers, the dissociation was considerably faster, reaching after 2 h ∼60 and 27%, respectively. For multimers containing the His12 tag, the dissociations were slightly slower, notably in the case of NTA1, with half-maximal dissociations (t½) of 34.5 versus 56.4 min (Fig. 4, A and E).
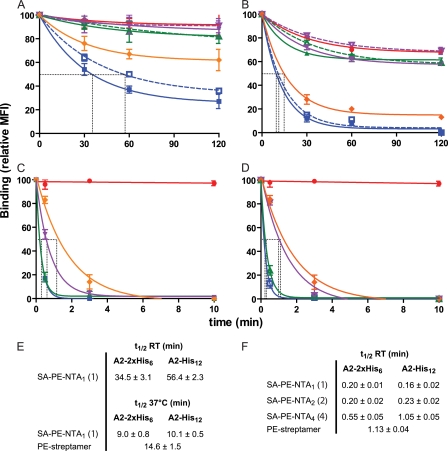
FIGURE 4.
Dissociation kinetics of SA-PE A2/Flu complexes. A and B, Flu58–66 peptide-stimulated PBMC were incubated at 4 °C for 30 min with 8 nm of SA-PE A2/Flu complexes containing NTA1 (blue squares), NTA2 (compound 2, green triangles), or NTA4 (purple triangles) and either a 2×His6 (solid lines) or His12 tag (dashed lines). For comparison, BSP multimers (red circles) and streptamers (orange diamonds) are included. After washing, the cells were incubated at room temperature (A) or 37 °C (B), and after the indicated period cell-associated fluorescence was assessed by flow cytometry. C and D, the experiment shown in A was repeated for 2×His6- (C) or His12-containing complexes (D) in the presence of imidazole (100 mm) or 10 mm biotin (for streptamer). The dashed gray lines indicate half-maximal dissociation. E and F, compilations of half-lives (t½) observed in the absence (E) or presence (F) of imidazole or biotin. The mean values and S.D. were calculated from at least three experiments. MFI, mean fluorescence intensity; RT, room temperature.
When repeating this dissociation experiment at 37 °C, significantly accelerated kinetics were observed, especially for the streptamer (t½ = 14.6 min), and the multimers containing NTA1 (t½ = 9.0 and 10.1 min for 2×His6 and His12 complexes, respectively) (Fig. 4, B and E). The dissociations of the His12-containing multimers were slower compared with the 2×His6-containing ones, which seems at odd with the SPR experiments, according to which the koff were lower for 2×His6 than for His12 containing A2/Flu complexes (Fig. 2B). However, in the latter experiments, monomeric complexes were examined, whereas in the former experiments, multimeric ones and their dissociation from cells may depend on additional parameters.
We then investigated the multimer dissociation in the presence of imidazole, which disrupts Ni2+-NTA-histidine complexes (11–18). To find out the maximal imidazole concentration tolerated by our polyclonal CD8+ T cells, we incubated these with graded concentrations of imidazole for 1 h at 37 °C and assessed their viability by trypan blue exclusion counting. Upon incubation with 100 mm imidazole, the cell viability was ≥99%; however, it significantly decreased at >200 mm (data not shown). In the presence of 100 mm imidazole, the dissociations of the 2×His6 tag-containing multimers at room temperature were remarkably rapid. For the NTA1- and NTA2 (compound 2)-containing multimers, dissociations with t½ of ∼0.2 min were barely measurable by flow cytometry (Fig. 4, C, D, and F). For the NTA4-containing multimers, the dissociation was slower with t½ of 0.5 and 1 min for 2×His6- and His12-containing multimers, respectively. It is noteworthy that in the presence of EDTA (20 mm) the Ni2+-NTA4–2×His6 A2/Flu complexes were stable, arguing that NTA4 (compound 4) more avidly chelated Ni2+ than EDTA (data not shown). For comparison, we assessed the dissociation kinetics of A2/Flu streptamers. Upon addition of 10 mm biotin, the staining of the polyclonal CD8+ T cells decreased at room temperature with a t½ of 1.1 min, i.e. slower than the NTA-containing multimers (Fig. 4, C, D, and F).
Collectively these results demonstrate that MHC I-peptide multimers built on NTA2 (compound 2)- or NTA4 (compound 4)-2×His6 tag chelate complexes stain Flu-specific CD8+ T cells as efficiently as or better than conventional BSP multimers. Importantly, they rapidly dissociated in the presence of imidazole, while being highly stable in its absence.
Reversible NTA Multimers Minimally Affect CD8+ T Cell Viability
Numerous immunological studies require sorting of antigen-specific CD8+ T cells by FACS (or magnetic-activated cell sorting). Because conventional BSP multimers affect CD8+ T cell viability (4–7), we examined what impact reversible staining with SA-PE-NTA4-containing multimers has on cell viability. Randomly chosen cloned CTL specific for Flu matrix peptide 58–66, Epstein-Barr peptide EBV 280–288, or a CTL line specific for Melan A 26–35 were incubated at 4 °C with the corresponding multimers containing BSP or SA-PE-NTA4, and cell-associated fluorescence analyzed by flow cytometry. The staining with SA-PE-NTA4 multimers was slightly higher as compared with conventional multimers (Fig. 5A). The stained cells after washing with 100 mm imidazole were incubated at 37 °C for different periods of time, and viable cells were enumerated by flow cytometry. The percentages of live, i.e. DAPI− and annexin V− cells were markedly lower on cells treated with conventional as compared with reversible multimers (Fig. 5B). To assess the functional integrity of thus treated CTL, we examined their cytotoxicity. As shown in Fig. 5C the CTL treated with BSP, but not SA-PE-NTA4 multimers exhibited reduced lysis, in terms of both maximal lysis and EC50. Remarkably, BSP multimer-treated CTL displayed increased nonspecific lysis, which is probably explained by transfer of peptide from BSP multimers to cell-associated MHC I molecules, giving rise to aberrant CTL-CTL and CTL-target cell interactions (28, 29).
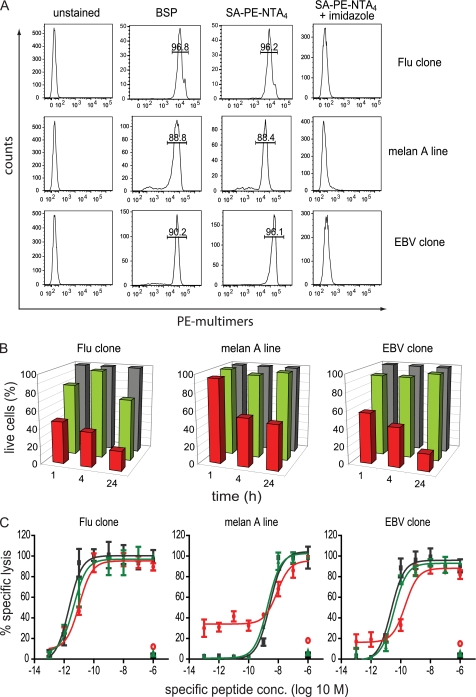
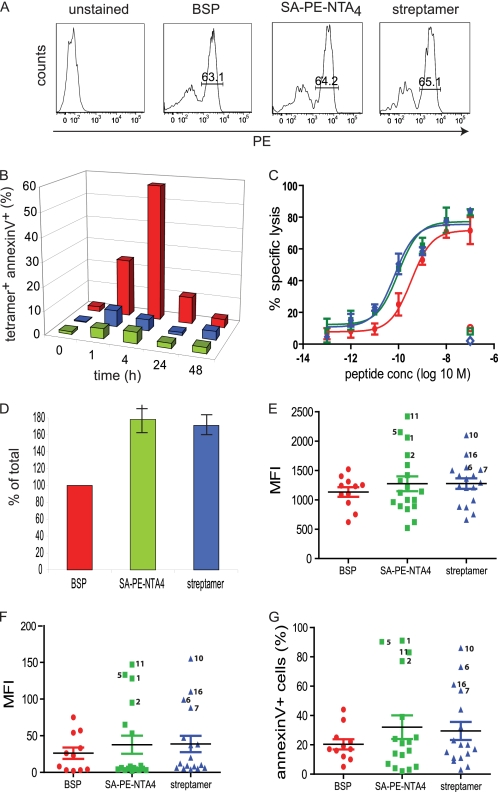
FIGURE 5.
Impact of multimers on CD8+ T cell viability. A, HLA-A2-restricted CD8+ T cells specific for influenza matrix 58–66 (clone BC74), Melan A 26–35 (melan A), or EBV BMLF1 280–288 (clone BC25) were stained at 4 °C with 5 nm BSP or reversible SA-PE-NTA4 multimers containing the respective 2×His6-tagged A2/peptide complexes. After washing with or without imidazole (100 mm) cell-associated fluorescence was measured by flow cytometry. The inserted values represent the cell percentages for the indicated gates. B, the same cells were treated likewise with BSP (red) or NTA4 multimers (green) or left untreated (gray), incubated at 37 °C for the indicate periods of time, stained with annexin V-FITC and DAPI, and analyzed by flow cytometry. Live, annexin V, and DAPI cells are represented as percentages, with 100% referring to untreated cells. One of three experiments is shown. C, alternatively, cells treated likewise with BSP (red) or NTA4 multimers (green) or left untreated (black) were tested in a 51Cr release assay using as targets T2 cells pulsed with graded concentrations of the respective specific or as negative control unspecific peptide. The mean values and S.D. of specific lysis were calculated from two experiments.
Because these experiments showed considerable variations between clones, we next used polyclonal, Flu58–66 peptide-stimulated PBMC. The cells were stained by A2/Flu58–66 BSP, SA-PE-NTA4 multimers, or streptamer to comparable extents (Fig. 6A). A constant number of Flu-specific cells were FACS sorted at 4 °C, and after different periods of incubation at 37 °C, the percentage of dying cells assessed by annexin V staining. Cells sorted with BSP exhibited extensive apoptosis, cumulating after 4 h with 57% of annexin V+ cells (Fig. 6B). By contrast, cells sorted with the reversible NTA4 multimer displayed only ∼6% of apoptotic cells, marginally less than streptamer-sorted cells. To assess the functional integrity of the sorted cells they were tested in a 51Cr release assay. For sake of fair comparison, we preincubated the BSP multimer-sorted cells with anti-A2 mAb BB7.2 to achieve maximal multimer dissociation (supplemental Fig. S5 and Ref. 27). As shown in Fig. 6C, BSP multimer-sorted cells exhibited significantly reduced target cell killing compared with SA-PE-NTA4 multimer of streptamer-sorted cells and increased nonspecific lysis, as observed on clones (Fig. 5C). For further analysis, we cloned sorted cells by limiting dilution. Compared with BSP multimer-sorted cells ∼1.8- and 1.7-fold more clones were obtained from NTA4 multimer- and streptamer-sorted cells, respectively (Fig. 6D). A2/Flu58–66 BSP multimer staining of clones showed that those derived from NTA4 multimer-sorted and to a lesser degree streptamer-sorted cells contained a fraction of high avidity specificities (Fig. 6E). This was also observed when CD8 binding-deficient A2227/228/Flu58–66 multimers were used (Fig. 6F). Upon incubation with A2/Flu58–66 BSP multimer at 37 °C, the brightly stained clones became annexin V+ (Fig. 6G). These results indicate that sorting of Flu-specific CTL with BSP multimers results in selective loss of high avidity, cell death-prone CD8+ T cells.
FIGURE 6.
FACS sorting of Flu-specific CD8+ T cells using A2/Flu multimers. A, Flu58–66 peptide stimulated PBMC were incubated for 15 min at 4 °C with 5 nm of A2/Flu58–66 multimers containing BSP, SA-PE-NTA4, and 2×His6 or A2/Flu58–66 streptamer, washed, and analyzed by flow cytometry. The inserted values represent cell percentages for the indicated gates. B, 100,000 BSP multimer+ cells (red), NTA4 multimer+ cells (green), or streptamer+ cells (blue) were FACS-sorted, washed with imidazole or biotin-containing medium, and after different periods of incubation at 37 °C analyzed by flow cytometry upon staining with annexin V and BSP multimers. Percentages of tetramer+ and annexin V+ cells are shown, with 100% being the input cell number. One of two experiments is shown. C, alternatively the cells sorted with BSP multimers were incubated with anti-A2 mAb BB7.2 (100 μg/ml) at room temperature for 20 min (red) or sorted with SA-PE-NTA4 (green) multimers or streptamer (blue) and washed with imidazole- or biotin-containing medium were incubated with 51Cr-labeled T2 cells in the presence of graded concentrations of Flu58–66 peptide for 4 h at 37 °C. The mean values and S.D. of specific lysis were calculated from two experiments. D, sorted cells (300 cells each) treated as in C were cloned by limiting dilution (0.5 cells/well), and after 14 days the growing clones were enumerated. The mean values and S.D. were calculated from two experiments. E and F, randomly chosen clones from BSP-sorted (red), SA-PE-NTA4 (green) multimer-sorted or streptamer-sorted (blue) cells were stained at room temperature for 30 min with conventional A2/Flu BSP (E) or A2227/228/Flu BSP multimer (F) (5 mm) and analyzed by flow cytometer. G, alternatively the cloned cells were stained with A2/Flu BSP multimer (5 nm) and annexin V and analyzed by flow cytometry after incubation at 37 °C for 30 min. The numbers indicate individual clones. MFI, mean fluorescence intensity.
PE-NTA2-A2/Peptide Multimers: a New Type of T Cell-staining Reagents
To directly conjugate MHC-peptide complexes with PE, we activated it with maleimido-NHS and reacted the resulting maleimido-PE with NTA2-Cys(SH) (Fig. 7A and S2). After incubation with NiCl2, PE-Ni2+-NTA2 was conjugated with A2/Flu58–66-2×His6 monomers, resulting in multimers containing 7–8 MHC-peptide complexes per PE. The staining of Flu peptide stimulated PBMC by A2/Flu58–66 multimers was strikingly different, whether these were conventional or PE-NTA2-containing ones (Fig. 7B). Although the binding of the conventional multimer increased steadily with the multimer concentration, it reached a plateau at ∼5 nm of PE-NTA2 multimer. As seen from Scatchard analysis, the KD value for the PE-NTA2 multimer was ∼12-fold lower than the one for the conventional multimer; however, the maximal binding (Bmax) was modestly higher (1.6-fold) for conventional multimer. We tested these reagents also on Flu peptide-stimulated PBMC from a different donor as well as on Melan A26–35 peptide-stimulated PBMC from a melanoma patient, using the corresponding A2/Melan A multimers. As shown in Fig. 7 (E–H), PE-NTA2 multimers efficiently stained antigen-specific CD8+ T cells, typically with slightly lower mean fluorescence intensity compared with conventional multimers and similar backgrounds.
FIGURE 7.
PE-NTA-A2/peptide multimers, a new type of T cells staining reagents. A, PE was reacted with maleimido-NHS (SM(PEG)2) and the resulting maleimido-PE subsequently with NTA2-Cys (SH). PE-NTA2, after loading with Ni2+, was conjugated with A2/peptide-2×His6 complexes to make biotin SA free PE-NTA2 multimers. B, Flu58–66-peptide stimulated PBMC were incubated at 4 °C for 1 h with graded concentrations of A2/Flu BSP (red circles) or PE-NTA2 multimers (blue squares), and cell-associated PE fluorescence was measured by flow cytometry. C, the binding data of B were subjected to Scatchard analysis. D, the mean values and S.D. of KD and Bmax values were calculated from three experiments. E–H, flow cytometry analysis of Flu58–66 peptide-stimulated PBMC from a healthy donor (E and F) or Melan A26–35 peptide-stimulated PMBC from melanoma patient Lau 1164 (G and H) were incubated at 4 °C with 8 nm of the corresponding BSP (E and G) or PE-NTA2 (F and H) multimers and anti-CD8 antibody. MFI, mean fluorescence intensity.
Assessment of A2/Flu Monomer Dissociation Kinetics on Cloned, Flu-specific CTL
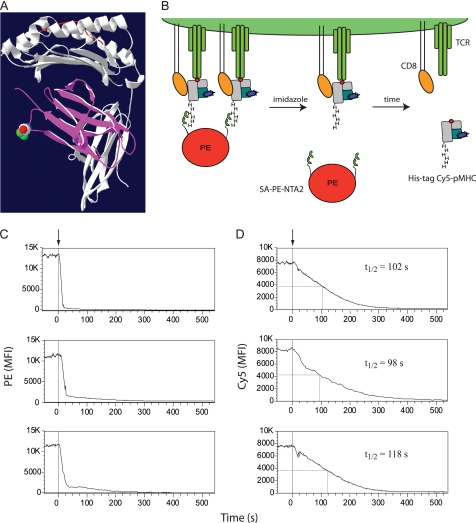
The staining of PE-NTA2 multimers was as very rapidly reversed upon the addition of imidazole (100 mm), as shown for NTA2-biotin-SA-PE at room temperature and at 4 °C (Fig. 4C and data not shown). We therefore examined whether PE-NTA2 MHC-peptide multimers allow assessment of monomer dissociation kinetics as depicted in Figs. 8 (A and B). To this end, we prepared A2/Flu58–66-2×His6 complexes containing Cy5-labeled β2m (Fig. 7A), and we used these PE-NTA2 multimers to stain in the cold cloned Flu-specific BC74 CTL. The washed cells were analyzed by flow cytometry, recording the PE (Fig. 8C) and Cy5 fluorescence before and after the addition of imidazole (Fig. 8D). Although the PE fluorescence reached background within 20–30 s, the Cy5 fluorescence reached background only after 300–350 s. The Cy5 fluorescence decrease exhibited half-maximal dissociation in the range of 98–118 s. Importantly, the monomer dissociation reached background levels, which is in contrast to dissociation experiments with BSP multimers, in which complete dissociation is typically not reached because of multimer rebinding and/or internalization (8, 9). The experiments gave well reproducible results when performed in the cold; at elevated temperatures the monomer dissociation were too rapid to be reliably measured (data not shown). Moreover, for these experiments it is preferable to use 2×His6-NTA2 multimers, because they dissociate faster upon the addition of imidazole than those containing NTA4 and/or the His12 tag (Fig. 4).
FIGURE 8.
Assessment of A2/Flu monomer dissociation kinetics. A, Cy5-labeled A2/Flu58–66 monomers were prepared by refolding with blue β2m obtained by mutating S88C (indicated by spheres) and alkylation with Cy5 maleimide. B, scheme of the dissociation kinetic experiment. CD8+ T cells are stained at 4 °C with A2/Flu58–66 multimers containing PE (red)-NTA2 (two hemi-spheres), 2×His6-tagged MHC (gray)-peptide (red dot) complexes, and Cy5-labeled β2m (blue). Upon the addition of imidazole, the Ni2+-NTA-2×His6 tag complex very rapidly decays in PE-NTA2 and A2/Flu58–66 monomers, which subsequently dissociate from cell-associated TCR (green) and CD8 (orange). C and D, cloned Flu-specific BC74 CD8+ T cells were stained at 4 °C with multimers A2/Flu 58–66 multimers containing Cy5-labeled β2m and either biotin and PE-SA (C) or a 2×His6 tag and PE-NTA2 (D) and analyzed by flow cytometry at 4 °C recording PE (C) or Cy5 (D) fluorescence before and after the addition of imidazole (100 mm) (time 0; arrows and gray lines). The half-maximal dissociations are indicated by gray lines and by their t½ values. Shown are three different experiments. MFI, mean fluorescence intensity.
DISCUSSION
A range of immunological studies require sorted antigen-specific CD8+ (and CD4+) T cells. Whereas FACS is technically highly efficient, a recurrent problem is that MHC-peptide multimers (or antibodies) used for staining persist on sorted cells and provoke cell death and functional alterations (4–9). The present study demonstrates that reversible, highly effective MHC-peptide multimers can be prepared based on Ni2+-NTA-His tag chelate complexes. Previously branched tri-NTA complexes have been described that have up to subnanomolar KD for His6 and His10 tags (22–24). Our results demonstrate that highly stable chelate complexes can be formed with linear peptides carrying orthogonal NTA moieties; the binding of Ni2+-NTA4 peptide (compound 4) to 2×His6-tagged A2/Flu complexes with a KD of 19.6 pm is the highest affinity ever reported for NTA-His tag complexes (Fig. 2 and supplemental Fig. S3). Although previously described tri- and tetra-NTA compounds allow diverse applications, their difficult chemical synthesis make them accessible only to specialized labs (20–24). In only one previous study, a tri-NTA compound was synthesized as a peptide derivative (2). However, as a building block Fmoc-Asp(β-amido-Nϵ-Lys(tri-but-NTA) was used, which is neither commercially available nor facile to synthesize. Moreover, we obtained remarkably poor yields for the coupling of Nϵ-Lys(tri-but-NTA) with peptides containing free orthogonal carboxyl groups, especially when these are adjacent (compound 3; Fig. 1A and supplemental Fig. S1A and unpublished results), and thus argue that coupling of remarkably bulky Nϵ-Lys(tri-but-NTA) or derivatives thereof tend to be inefficient because of steric hindrance. Consistent with this view, we obtained excellent yields for couplings performed in solution on fully deprotected molecules using site-specific alkylation (supplemental Fig. S1B). The strategy to synthesize linear oligocysteine peptides by conventional solid phase peptide synthesis and to alkylate these with commercially available maleimido-NTA renders accessible a wide range oligo-NTA compounds to nonspecialized labs.
The observation that NTA2 dimer compound 3 exhibited considerably weaker binding than NTA2 dimer compound 2, especially for the His12 and2×His6-tagged HLA-A2/Flu complexes (Figs. 2–5), indicates that the chelate complex stability critically depended on the configuration of NTA compound. The main difference between these two NTA dimers is the length of the side chains carrying the NTA groups (y and y′) and the distance by which they attached to the peptide backbone (x and x′; Fig. 1A). In compound 2, this distance is shorter than in compound 3. A previous study has shown that the avidity of branched tri-Ni2+-NTA molecules for His6-tagged molecules increased with the decreasing length of the spacer linking the NTA moieties to a central nitrogen (24). This argues that the NTA dimer compound 2 exhibited more avid binding than dimer compound 3, because of the shorter spacing distance x′. On the other hand, the longer side chain y′ in compound 2 may permit better interactions of individual Ni2+-NTA moieties with histidine residues on the His tags, namely on the long 2×His6 tag. Although it has been shown that the avidity and stability of Ni2+-NTA-histidine complexes increases with the valence of Ni2+-NTA moieties and the length of the His tags (18–24), our study demonstrates for the first time that the combination of a double His6 tag and a double NTA dimer (compound 4) provides a dramatic gain of avidity and stability. We suggest that the long and flexible GGSGGGSGS linker contained in both entities allows a two-register binding mode, in which two His6 tags and two NTA2 moieties interact in a zipper-like fashion. However, interactions of oligo-Ni2+-NTA moieties with His tags are complex, involving diverse kinetic, entropic, and enthalpic aspects, and more extended studies will be needed to fully understand the underlying structure-binding relations (20–24, 30–32).
It is interesting to note that the conventional A/Flu BSP multimers displayed a modestly lower staining of Flu-specific PBMC than those containing NTA4 and a 2×His6 tag (Fig. 3A). Because the BSP-SA binding is more stable than the Ni2+-NTA4-2×His6 tag complex, this difference may be explained by different configurations of the two types of multimers. Indeed, we have shown previously that soluble MHC I-peptide complexes depending on their configurations can have strikingly different KD and Bmax values (25).
Avid and stable binding of MHC-peptide multimers allows efficient staining of antigen-specific CD8+ T cells and at the same time results in extensive cross-linking of TCR and CD8, which at elevated temperature induces strong T cell activation and rapid T cell death (Figs. 5 and 6 and Refs. 4, 5,8, 9, 26, 33, and 34). It also results in internalization of MHC-peptide complexes and transfer of peptide to cell-associated MHC I molecules, which can cause aberrant CTL functions, e.g. fratricide and bystander killing (26, 28, 29). Because of this, FACS sorting of CD8+ T cells using BSP multimers is prone to infer loss of antigen-specific cells, namely selectively high avidity ones and alteration of the functional integrity of surviving cells (Figs. 5 and 6 and Refs. 4, 5, and 9). This can be avoided by using reversible MHC-peptide multimers, the effectiveness of which depends on: (i) their stability, which determines the staining performance and shelf-life: the here described Ni2+-NTA4-2×His6 tag based multimers are considerably more stable than streptamers and DTB multimers, especially at elevated temperatures, and their shelf-life is the same as for BSP multimers (Fig. 4, Ref. 4, and data not shown); and (ii) their reversibility, which determines how rapidly and quantitatively they can be removed from sorted cells: the induced dissociation of NTA-His tag multimers is clearly faster than the ones of streptamers and DBT multimers (Fig. 4, supplemental Fig. S5, and Ref. 4). Because we have shown that the induced dissociation of DTB multimers from CTL is sufficiently rapid to prevent intracellular Ca2+ mobilization (4), the same is expected for NTA-His tag multimers. Intracellular Ca2+ mobilization is required for most CTL functions, including multimer-induced CD8+ T cell death (8, 26, 33).
The here described NTA-2×His6 chelate complexes allow the preparation of a wide range of conjugates. An attractive application is to couple NTA moieties onto PE (or allophycocyanine), which allows facile and stable conjugation with MHC-peptide-2×His6 complexes (Fig. 7A). CD8+ T cell staining by these conjugates is strikingly different compared with BSP multimers (Figs. 7, B and C). Although BSP multimers are heterogeneous, containing PE-SA conjugates of various stoichiometries and configurations (e.g. orientations of SA to PE), PE-NTA-containing multimers are molecularly better defined, containing one central PE conjugated with seven or eight MHC-peptide complexes (26). This difference is reflected in the binding isotherms, which exhibited a clear saturation in the case of PE-NTA-2×His6 but not BSP multimers and hence better linear regressions in Scatchard analysis (Fig. 7, A–D). This also explains why the PE-NTA-2×His6 multimers bind with lower KD and Bmax than BSP multimers. For these reasons PE-NTA-2×His6 multimers are better suitable for accurate binding studies. Furthermore and importantly, because of the very rapid, imidazole-induced disintegration of NTA-2×His6 multimers, they permit conclusive assessment of MHC-peptide monomer dissociations on living CD8+ T cells (Fig. 8). PE-NTA2-2×His6 multimers are particularly suitable for this, because they provide stable staining and very rapid imidazole-induced disintegration (Figs. 3A and 4A and unpublished data). In contrast to dissociation experiments using BSP multimers, this strategy provides real kinetics of dissociation of MHC-peptide monomers from TCR and CD8 in the absence of artifactual residual rebinding (Fig. 8, C and D, and Refs. 10 and 11). Although this technique is less accurate than SPR measurements, it provides binding data on living cells, which substantially deviate from SPR binding experiments using recombinant TCR and MHC-peptide complexes, notably because of coordinate binding of CD8 to TCR-associated MHC-peptide (7, 12, 27). Recent two-dimensional binding studies (e.g. intercellular) of TCR-MHC-peptide interactions have indicated binding parameters that were strikingly different from those measured in three-dimensional (e.g. SPR) binding studies (35–37). This casted doubt on the value of the latter in terms of predicting T cell activation. This is especially intriguing for CD8+ T cells, because CD8 can bind to TCR-associated cognate as well as to noncognate MHC-peptide complexes and thereby substantially affect TCR-MHC-peptide binding and T cell activation (12, 27, 36). The here described assay (Fig. 8) and TCR photoaffinity labeling (12) constitute interesting alternatives, because they allow MHC-peptide binding studies on living cells in the absence of intercellular adhesion effects.
PE-NTA2 or PE-NTA4 can be equally used for the preparation on MHC class II-peptide and antibody Fab multimers. With the former, the 2×His6 tag is fused C-terminal to the α2 domain, which additionally allows universal and gentle purification of fragile, empty MHC II molecules from culture supernatants; with the latter, a synthetic His tag is conjugated with Fab′ fragments by site-specific alkylation, similarly as described previously (4). The stability and imidazole-induced reversibility of NTA-His tag-based conjugates depends on those of the underlying chelate complexes and not on the type of the conjugate (e.g. PE-NTA or NTA-biotin-SA-PE). In case of NTA4-2×His6-containing multimers, the stability of the underlying chelate complex exceeds by far any of the previously described switchable conjugates, including streptamers and DTB multimers. At the same time their imidazole-induced disintegration is faster than any of the previously reported reversible MHC-peptide multimers, and their production is simpler, which affords a better accessibility (Figs. 2–4 and Refs. 4, 5, 9, and 21–24).
In conclusion, the present work provides a framework of novel types of protein conjugates, namely fluorescent MHC-peptide multimers, built on reversible NTA-His tag chelate complexes. We describe the synthesis of linear NTA peptides that requires no specialized equipment or knowhow and that allows complex formation with 2×His6-tagged proteins of unprecedented stability, while being rapidly switchable upon the addition of imidazole. MHC-peptide multimers built on NTA-His coordinate complexes outperform previously described reversible staining reagents and allow greater diversity of conjugate formation, e.g. PE-NTA-based multimers and hence provide new opportunities to advance T cell analysis.
Acknowledgments
We are indebted to Nicole Montandon for competent technical assistance, Dr. Marek Kosinski (Centre Hospitalier Universitaire Vaudois, Lausanne) for expert assistance with equilibrium dialysis experiments, and Dr. B. Robert (Centre de Recherche en Cancérologie de Montpellier) for critical reading of the manuscript.
This work was supported by grants from the Swiss National Science Foundation (310030_12533/1) and from the Cancer Research Institute.

The on-line version of this article (available at http://www.jbc.org) contains supplemental text, Table S1, and Figs. S1–S5.
- SA
- streptavidin
- A2
- HLA-A*0201
- β2m
- β2-microglobulin
- BSP
- biotinylation sequence peptide
- CTL
- cytotoxic T lymphocyte
- MHC
- major histocompatibility complex
- NTA
- nitrilotriacetic acid
- PBMC
- peripheral blood mononuclear cell(s)
- PE
- phycoerythrin
- SPR
- surface plasmon resonance
- TCR
- T cell antigen receptor
- Fmoc
- N-(9-fluorenyl)methoxycarbonyl
- RT
- retention time
- RU
- resonance unit.
REFERENCES
- 1. Stayton P. S., Freitag S., Klumb L. A., Chilkoti A., Chu V., Penzotti J. E., To R., Hyre D., Le Trong I., Lybrand T. P., Stenkamp R. E. (1999) Biomol. Eng. 16, 39–44 [DOI] [PubMed] [Google Scholar]
- 2. June R. K., Gogoi K., Eguchi A., Cui X. S., Dowdy S. F. (2010) J. Am. Chem. Soc. 132, 10680–10682 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Cao H., Lin R. (2009) Biotechnol. Prog. 25, 461–467 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Guillaume P., Baumgaertner P., Angelov G. S., Speiser D., Luescher I. F. (2006) J. Immunol. 177, 3903–3912 [DOI] [PubMed] [Google Scholar]
- 5. Knabel M., Franz T. J., Schiemann M., Wulf A., Villmow B., Schmidt B., Bernhard H., Wagner H., Busch D. H. (2002) Nat. Med. 8, 631–637 [DOI] [PubMed] [Google Scholar]
- 6. Guillaume P., Dojcinovic D., Luescher I. F. (2009) Cancer Immun. 9, 7–13 [PMC free article] [PubMed] [Google Scholar]
- 7. Wooldridge L., Lissina A., Cole D. K., van den Berg H. A., Price D. A., Sewell A. K. (2009) Immunology. 126, 147–164 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Cebecauer M., Guillaume P., Hozák P., Mark S., Everett H., Schneider P., Luescher I. F. (2005) J. Immunol. 174, 6809–6819 [DOI] [PubMed] [Google Scholar]
- 9. Neudorfer J., Schmidt B., Huster K. M., Anderl F., Schiemann M., Holzapfel G., Schmidt T., Germeroth L., Wagner H., Peschel C., Busch D. H., Bernhard H. (2007) J. Immunol. Methods. 320, 119–131 [DOI] [PubMed] [Google Scholar]
- 10. Kalergis A. M., Boucheron N., Doucey M. A., Palmieri E., Goyarts E. C., Vegh Z., Luescher I. F., Nathenson S. G. (2001) Nat. Immunol. 2, 229–234 [DOI] [PubMed] [Google Scholar]
- 11. Dutoit V., Guillaume P., Cerottini J. C., Romero P., Valmori D. (2002) Eur. J. Immunol. 32, 3285–3293 [DOI] [PubMed] [Google Scholar]
- 12. Luescher I. F., Vivier E., Layer A., Mahiou J., Godeau F., Malissen B., Romero P. (1995) Nature 373, 353–356 [DOI] [PubMed] [Google Scholar]
- 13. Schmitt L., Dietrich C., Tampé R. (1994) J. Am. Chem. Soc. 116, 8485–8491 [Google Scholar]
- 14. Dietrich C., Schmitt L., Tampé R. (1995) Proc. Natl. Acad. Sci. U.S.A. 92, 9014–9018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Dorn I. T., Neumaier K. R., Tampé R. (1998) J. Am. Chem. Soc. 120, 2753–2763 [Google Scholar]
- 16. Knecht S., Ricklin D., Eberle A. N., Ernst B. (2009) J. Mol. Recognit. 22, 270–279 [DOI] [PubMed] [Google Scholar]
- 17. Nieba L., Nieba-Axmann S. E., Persson A., Hämäläinen M., Edebratt F., Hansson A., Lidholm J., Magnusson K., Karlsson A. F., Plückthun A. (1997) Anal. Biochem. 252, 217–228 [DOI] [PubMed] [Google Scholar]
- 18. Khan F., He M., Taussig M. J. (2006) Anal Chem. 78, 3072–3079 [DOI] [PubMed] [Google Scholar]
- 19. Steinhauer C., Wingren C., Khan F., He M., Taussig M. J., Borrebaeck C. A. (2006) Proteomics 6, 4227–4234 [DOI] [PubMed] [Google Scholar]
- 20. Tinazli A., Tang J., Valiokas R., Picuric S., Lata S., Piehler J., Liedberg B., Tampé R. (2005) Chemistry 11, 5249–5259 [DOI] [PubMed] [Google Scholar]
- 21. Lata S., Gavutis M., Tampé R., Piehler J. (2006) J. Am. Chem. Soc. 128, 2365–2372 [DOI] [PubMed] [Google Scholar]
- 22. Lata S., Reichel A., Brock R., Tampé R., Piehler J. (2005) J. Am. Chem. Soc. 127, 10205–10215 [DOI] [PubMed] [Google Scholar]
- 23. Huang Z., Park J. I., Watson D. S., Hwang P., Szoka F. C., Jr. (2006) Bioconjug. Chem. 17, 1592–1600 [DOI] [PubMed] [Google Scholar]
- 24. Huang Z., Hwang P., Watson D. S., Cao L., Szoka F. C., Jr. (2009) Bioconjug. Chem. 20, 1667–1672 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Guillaume P., Baumgaertner P., Neff L., Rufer N., Wettstein P., Speiser D. E., Luescher I. F. (2010) Int. J. Cancer 127, 910–923 [DOI] [PubMed] [Google Scholar]
- 26. Guillaume P., Legler D. F., Boucheron N., Doucey M. A., Cerottini J. C., Luescher I. F. (2003) J. Biol. Chem. 278, 4500–4509 [DOI] [PubMed] [Google Scholar]
- 27. Wooldridge L., van den Berg H. A., Glick M., Gostick E., Laugel B., Hutchinson S. L., Milicic A., Brenchley J. M., Douek D. C., Price D. A., Sewell A. K. (2005) J. Biol. Chem. 280, 27491–27501 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Schott E., Bertho N., Ge Q., Maurice M. M., Ploegh H. L. (2002) Proc. Natl. Acad. Sci. U.S.A. 99, 13735–13740 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Ge Q., Stone J. D., Thompson M. T., Cochran J. R., Rushe M., Eisen H. N., Chen J., Stern L. J. (2002). Proc. Natl. Acad. Sci. U.S.A. 99, 13729–13734 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Brellier M., Barlaam B., Mioskowski C., Baati R. (2009) Chem. Eur. J. 15, 12689–12701 [DOI] [PubMed] [Google Scholar]
- 31. Tareste D., Pincet F., Brellier M., Mioskowski C., Perez E. (2005) J. Am. Chem. Soc. 127, 3879–3884 [DOI] [PubMed] [Google Scholar]
- 32. André T., Reichel A., Wiesmüller K. H., Tampé R., Piehler J., Brock R. (2009) Chembiochem. 10, 1878–1887 [DOI] [PubMed] [Google Scholar]
- 33. Angelov G. S., Guillaume P., Cebecauer M., Bosshard G., Dojcinovic D., Baumgaertner P., Luescher I. F. (2006) J. Immunol. 176, 3356–3365 [DOI] [PubMed] [Google Scholar]
- 34. Xu X. N., Purbhoo M. A., Chen N., Mongkolsapaya J., Cox J. H., Meier U. C., Tafuro S., Dunbar P. R., Sewell A. K., Hourigan C. S., Appay V., Cerundolo V., Burrows S. R., McMichael A. J., Screaton G. R. (2001) Immunity 14, 591–602 [DOI] [PubMed] [Google Scholar]
- 35. Huang J., Zarnitsyna V. I., Liu B., Edwards L. J., Jiang N., Evavold B. D., Zhu C. (2010) Nature 464, 932–936 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Jiang N., Huang J., Edwards L. J., Liu B., Zhang Y., Beal C. D., Evavold B. D., Zhu C. (2011) Immunity 34, 13–23 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Huppa J. B., Axmann M., Mörtelmaier M. A., Lillemeier B. F., Newell E. W., Brameshuber M., Klein L. O., Schütz G. J., Davis M. M. (2010) Nature 463, 963–967 [DOI] [PMC free article] [PubMed] [Google Scholar]