Abstract
The advent of powerful genomics technologies has uncovered many fundamental aspects of biology, including the mechanisms of cancer; however, it has not been appropriately matched by the development of global approaches to discover new medicines against human diseases. Here we describe a unique high-throughput screening strategy by high-throughput sequencing, referred to as HTS2, to meet this challenge. This technology enables large-scale and quantitative analysis of gene matrices associated with specific disease phenotypes, therefore allowing screening for small molecules that can specifically intervene with disease-linked gene-expression events. By initially applying this multitarget strategy to the pressing problem of hormone-refractory prostate cancer, which tends to be accelerated by the current antiandrogen therapy, we identify Peruvoside, a cardiac glycoside, which can potently inhibit both androgen-sensitive and -resistant prostate cancer cells without triggering severe cytotoxicity. We further show that, despite transcriptional reprogramming in prostate cancer cells at different disease stages, the compound can effectively block androgen receptor-dependent gene expression by inducing rapid androgen receptor degradation via the proteasome pathway. These findings establish a genomics-based phenotypic screening approach capable of quickly connecting pathways of phenotypic response to the molecular mechanism of drug action, thus offering a unique pathway-centric strategy for drug discovery.
Keywords: chemical screening, gene signature
It is of utmost importance to match the power of functional genomics in interrogating diseased cells/tissues with potent drug-discovery approaches. Although target-centric approaches have been favored in the past decade, phenotypic screening appears to have out-paced such mechanism-based screening strategies in discovering “first-in-class” drugs, thus igniting recent debate on the merit of target-based strategies (1). This debate is important because analysis of US Food and Drug Administration-approved drugs in recent decades have revealed low productivity in drug research and development, despite staggering investment in the pharmaceutical industry (2).
Although phenotypic approaches score the final functional outcomes, it is challenging to optimize candidate drugs without knowing their mechanism of action and many procedures have limited capacity in implementing high-throughput screening. In contrast, target-centric approaches have their own problems because specific molecular hypotheses based on the existing knowledge may or may not be related to disease phenotype. A proposed solution to these problems is to monitor the collective response of all relevant genes to a specific disease phenotype (3), but this has been a major challenge with any existing technologies.
A “quick-win/fast-fail” strategy has been proposed to streamline initial candidate hits in early phases to offset high attrition rates in drug discovery (4). This strategy begs the question of how to retain the advantages of both phenotypic and target-centric screening approaches to quickly lead to promising drug candidates. One idea is to take molecular approaches to conduct “phenotypic screenings” by using a set of genes to report a disease phenotype, thus allowing screening for small molecules that can effectively block disease-associated gene-expression events. Several proof-of-concept experiments for this gene-signature approach have been performed on a number of disease paradigms by PCR (5) or hybridization on Luminex beads (6, 7). However, none of these methods has yet reached the throughput or cost-effectiveness that is required for large-scale applications in drug discovery.
Here, we report the development of a pathway-centric high-throughput screening strategy by taking advantage of the ever-increasing power of high-throughput sequencing (this strategy is hereafter referred to as HTS2). The technology permits quantitative analysis of a gene matrix directly in cell lysates. We have initially applied this technology on a prostate cancer cell (LNCaP) model to identifying small molecules that can block the androgen receptor (AR)-mediated gene expression, because AR overexpression has been linked to prostate cancer progression to androgen-refractory, incurable tumors (8, 9). By following the AR pathway, we identified several classes of compounds, one of which belongs to cardiac glycosides currently used for treating congestive heart failure and arrhythmias, but also known for their broad anticancer activities on cancer-cell models (10, 11). Interestingly, a recent epidemiological study revealed that long-term use of cardiac glycosides has significant benefit in preventing prostate cancer (12). We now showed that this activity is likely a result of the ability of cardiac glycosides to cause AR destabilization, thereby effectively blocking AR-dependent gene expression and cell proliferation on both androgen-sensitive and -resistant prostate cancer cells. These findings validate the potential of this recently developed HTS2 technology in pathway-centric chemical screenings, matching the advances in functional genomics to the development of new anticancer therapies.
Results
HTS Platform Based on Next-Generation Sequencing.
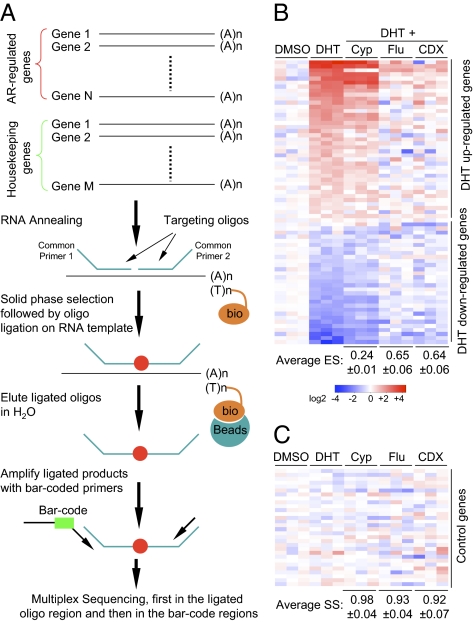
As diagrammed in Fig. 1A, we first profile gene expression in a chosen cell type to define a panel of genes associated with a disease phenotype. To quantify these genes in a high-throughput manner, we use the RNA annealing, selection, ligation (RASL) strategy, originally designed to profile mRNA isoforms using pooled pairs of oligonucleotides, each flanked by a universal primer to target specific splice junctions in spliced mRNAs (13, 14). Upon annealing to total RNA followed by solid-phase selection via oligo-dT or biotinylated total RNA, paired DNA probes templated by specific RNA sequences can be ligated by T4 DNA ligase, thus converting singleton probes to PCR amplicons. Minimal bias is introduced in this process because of the uniform length and relatively balanced GC content of designed probes (15). Instead of quantifying the amplified products on microarrays, as before, we can now use deep-sequencing to directly count the correctly ligated products. By using bar-coded primers (each contains a unique 7-nt sequence), multiple samples can be pooled for parallel quantification by first reading target sequences followed by sequencing the bar-coded region to decode different samples in the pool.
Fig. 1.
The scheme of the HTS2 technology. (A) The flow of the HTS2 technology. (B and C) Representative responses to androgen (DHT) and the effects of known antiandrogen compounds on a panel of DHT-responsive genes (B) and a set of housekeeping and cytotoxicity-related genes (C). Three vehicle-treated samples are averaged to serve as the baseline and variations from the baseline are color-coded: red for DHT-induced genes and blue for DHT-suppressed genes. The ES, SS, and SD are shown on the bottom of each panel for each compound.
This experimental scheme is fully amenable to automation and direct transcript analysis in the cell lysate, two critical parameters for high-throughput applications. As hybridization can take place in the presence of detergent and high salt, the annealing step is fully compatible to standard hybridization conditions. Biotinylated oligo-dT included in the annealing step captures spliced mRNA from the cell lysate along with annealed probes on them via streptavidin selection. RNA capture can be alternatively performed on oligo-dT–coated plates, which produced similar results. All subsequent washing and ligation steps are carried out on the solid phase. Ligated products are released in H2O, converted to bar-coded amplicons by PCR, and pooled for deep sequencing. We have fully implemented this HTS2 strategy on a Biomek FX robot.
Robust Performance of HTS2 on Androgen-Regulated Genes.
To demonstrate this unique technology, we initially selected two dozen androgen-responsive genes for comparison between fold-differences detected by HTS2 and RT-qPCR on mock-treated and androgen (dihydrotestosterone, DHT)-treated LNCaP cells, which validated the ability of the HTS2 technology in detecting quantitative differences (Fig. S1A). We noticed that different probe sets against the same genes often showed differences in annealing/ligation, thus resulting in different tag counts, but fold-differences were relatively consistent (Fig. S1B). This finding actually allowed us to select low-efficiency probe sets for abundant transcripts and high-efficiency probe sets for low-copy transcripts to balance the sequence space for transcripts of varying abundance.
By requiring at least 200 counts per transcript in a panel consisting of ∼100 targets, we initially tested pooling all samples in 384 wells for sequencing in a single lane on the Illumina GAII system, which normally generates ∼20 million reads, thus sufficient for the required count capacity (200 × 100 × 384 = ∼8 million reads). More recently, we further successfully tested an even higher multiplexicity by pooling samples from four such 384-well plates for sequencing on HiSeq2000, which has the capacity to produce up to 200 million counts per lane. Thus, on a pair of eight-lane flowcells in a single HiSeq2000 run, the system permits screening >20 thousand samples (16 × 4 × 384 = 24,576).
Application of HTS2 to Analysis of Antiandrogen Activities.
We next applied the HTS2 platform to identify small molecules that can effectively block androgen-induced gene expression in LNCaP cells. Unlike early efforts based on a single expression reporter (16), we selected a panel of 70 androgen-induced (both up- and down-regulated) genes to represent the AR pathway, determined by RNA-seq (17). We also included 30 housekeeping and cytotoxicity-related genes (18) as specificity controls (Table S1). The robotic RASL assay demonstrated high reproducibility among biological repeats of mock-treated and DHT-treated samples (Fig. S2 A and B) and the ability to detect anticipated androgen responses (Fig. S2C). We then tested several known androgen antagonists, including Cyproterone (Cyp), Flutamide (Flu), and Bicalutamide (CDX). Cyp showed a modest effect but both Flu and CDX were more potent than Cyp in blocking DHT-induced gene expression (Fig. 1B), all of which had minimal effect on the built-in controls (Fig. 1C).
Previous works used summed gene expression, k-nearest neighbors, or naïve Bayes classification to quantify the antiandrogen effect (5). These methods generate an index to each compound based on alteration in gene expression toward a desired direction. Using a panel of controls to simultaneously evaluate the specificity, we generated both the efficacy and specificity scores for each compound by using summed gene expression (Methods). The Efficacy score (ES) measures independent contribution of each gene to antagonizing the DHT effect, which is summed to produce a score ranging from −1 (for full effect in enhancing the DHT effect) to +1 (for full effect in suppressing the DHT effect). The Specificity score (SS) determines the number of control genes that remain unaltered (twofold), ranging from 0 (for nonspecific effect) to +1 (for lack of nonspecific effect). Therefore, we are able to generate a pair of numerical indexes for each of the known antiandrogen compounds (Fig. 1 B and C). The data fully concur with different potencies of these drugs in treating prostate cancer patients in the clinic (19), demonstrating the ability of the HTS2 platform in characterizing androgen antagonists based on the collective response of androgen-responsive genes.
Identification of Unique Antiandrogen Compounds.
On the established HTS2 platform, we next screened a collection of compounds (∼4,000), consisting of human-experienced drugs, a set of natural products, and various synthetic inhibitors of known enzymes. We included CDX as a benchmark for antiandrogen activity and full-effect mimics (LNCaP cells not treated with DHT or compound, thus mimicking complete suppression of the DHT effect). As expected, most full-effect mimics (blue dots) showed high ES and SS scores; DHT-treated samples (pink dots) produced low ES and high SS scores; and treatments with CDX (red dots) generated relatively high ES and SS scores (Fig. 2A and Fig. S3). Interestingly, although most test compounds (green dots) lacked any antiandrogen effects (low ES scores), we identified a number of compounds that exhibited high ES and SS scores, indicating that these candidate hits may function similarly or better than CDX in inhibiting DHT-responsive genes in LNCaP cells.
Fig. 2.
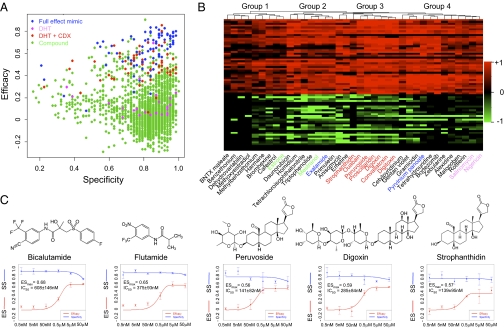
Identification of unique anti-AR compounds. (A) Two-dimensional plot of screened compounds (plots of individual plates are shown in Fig. S3). Blue, full-effect mimics (no DHT treatment and no compound; pink, DHT treatment alone; red, CDX on DHT-treated cells; and green, compound on DHT-treated cells. (B) Clustering analysis of top candidate hits. Red represents the effect on suppressing DHT-induced genes; green shows the effect on restoring DHT-repressed genes. The compounds identified from published screenings as described in the text are highlighted in green, blue, and pink; cardiac glycosides are labeled in red. (C) Titration and deduced ESmax and IC50 for individual compounds. SDs are based on triplicate measurements. Compound structure is shown on top of each titration curve.
The anti-DHT effects of top candidates are displayed by hierarchical clustering based on their impact on individual genes in the panel where red represents suppression of DHT-induced gene expression ranging from 0 (no effect) to +1 (full suppression), whereas green indicates restoration of DHT-repressed gene expression, which ranges from 0 (no effect) to −1 (full restoration) (Fig. 2B). This analysis revealed four groups: groups 1 and 4 showed effective suppression of DHT-induced genes, but little effect on restoring DHT down-regulated genes, indicating that these compounds may impair gene expression in some general ways. In comparison, groups 2 and 3 antagonized DHT-responsive genes in both directions.
We note two estrogens, Estrone and Mestranol (green in Fig. 2B), in the top hits, suggesting that the activation of the estrogen pathway might interfere with the status of the androgen pathway, which is consistent with the reported effect of estrogen in inhibiting prostate cancer cells through the estrogen receptor β (ERβ) (20). Two compounds Pyrvinium Panoate and Exalamide (blue in Fig. 2B) were previously identified from an AR conformational screen (21, 22). We also identified several ionophors (pink in Fig. 2B), including Salinomycin, a compound recently shown to selectively inhibit breast cancer stem cells (23), and Nigericin, which was previously identified from a screen based on suppressing the prostate-specific antigen (PSA) (16).
Interestingly, a major fraction of candidate hits belongs to the cardiac glycoside family, including Strophanthidin, Ouabain, Proscillaridin, Peruvoside, Digoxin, Concallatoxin, and Digitoxin (red in Fig. 2B), all in group 3. We selected several of these cardiac glycosides to confirm their anti-androgen effects and derive the maximum effect based on the ES (ESmax) and the half-maximum inhibitory concentration (IC50) (Fig. 2C). The results reveal that cardiac glycosides, particularly Peruvoside and Strophanthidin, are more potent than CDX and Flu in blocking DHT-induced gene expression in LNCaP cells (Fig. 2C).
Peruvodise Potently Inhibits Cell Proliferation Without Inducing Cytotoxicity.
We next focused on understanding how cardiac glycosides suppress AR-mediated gene expression for two reasons. First, various cardiac glycosides exhibit broad anticancer effects, including prostate cancer, both in vitro (24, 25) and on castration-resistant prostate tumors in animal models (26). Second, a recent epidemiological study revealed that long-term use of Digoxin, a cardiac glycoside widely used to treat congestive heart diseases, significantly reduces the risk of prostate cancers (12). Our results suggest that cardiac glycosides may directly intervene with the AR pathway in prostate cancer cells.
To determine if the inhibitory effect on prostate cancer cells depends on a functional AR, we tested several cardiac glycosides at 5 μM (the same concentration we used in the primary screen) on two pairs of isogenic prostate cancer cells (Fig. 3A). One pair is androgen-sensitive LNCaP cells and their androgen-resistant derivative LNCaP-abl cells; the other pair is androgen-resistant, AR-negative PC3 cells and AR re-expressed PC3-AR cells. All three cardiac glycosides tested began to inhibit proliferation of LNCaP cells within the first 24 h of treatment, with little effect on LNCaP-abl cells and no effect on PC3 and PC3-AR cells. By day 2, the cardiac glycosides were quite effective in inhibiting LNCaP-abl cells, but still lacked effect on PC3 cells. Interestingly, PC3-AR cells appear to have gained a degree of sensitivity to cardiac glycosides (Fig. 3A). By day 3, proliferation was inhibited in all cell types. These observations suggest that a functional AR may render PC3 cells sensitive to inhibition by cardiac glycosides.
Fig. 3.
Effects of lead compounds on cell proliferation and apoptosis. (A) Effect of each compound (5 μM) on DHT-treated LNCaP cells and on androgen-independent LNCaP-abl, PC3, and PC3-AR cells. (B) Effect of each compound at different concentrations on androgen-resistant LNCaP-abl cells. (C) Apoptosis induced on LNCaP (Left) and LNCaP-abl (Right) cells by individual compounds with Nigericin as a positive control. (D) Competitive binding determined with 3H-labeled DHT in the presence of increasing concentrations of individual cold DHT, CDX, and Peruvoside.
To further characterize these cardiac glycosides, we treated LNCaP-abl cells for 3 d with each compound at different concentrations (Fig. 3B). Peruvoside blocked cell growth at 50 nM; both Digoxin and Strophanthidin required 500 nM to achieve the same effect on the androgen-resistant LNCaP-abl cells, although Strophanthidin effectively inhibited androgen-induced gene expression as Peruvoside did on androgen-sensitive LNCaP cells. Early studies suggest that cardiac glycosides inhibit cell proliferation through the induction of apoptosis (24–26). We asked whether induced apoptosis was sufficient to account for the strong effect of Peruvoside on inhibiting cell proliferation. By monitoring activated Caspases 3 and 7, we found that Peruvoside has a detectable degree of induced apoptosis on LNCaP cells, but no effect on LNCaP-abl cells (Fig. 3C). Thus, Peruvoside can effectively block cell proliferation without triggering general cytotoxic response on androgen-resistant prostate cancer cells. In contrast, Nigercin induced apoptosis on both LNCaP and LNCaP-abl cells.
As cardiac glycosides are themselves cardiotonic steroids (see their core steroid structure in Fig. 2C), and in light of a recent finding that Digoxin can directly bind to a specific nuclear receptor (RORγt) in T cells (27), we examined whether Peruvoside might compete for androgen binding. We found no evidence for Peruvoside to bind to the ligand binding pocket in AR, because it could not compete with binding of 3H-labeled DHT to endogenous AR on LNCaP cells. In contrast, unlabeled DHT and CDX could completely or partially compete, respectively (Fig. 3D). These results ruled out the possibility that Peruvoside acts as a competitive androgen antagonist.
Cardiac Glycosides Block the Entire AR Pathway.
Our primary screen was conducted on androgen-sensitive LNCaP cells, yet these compounds also exhibited significant effects on androgen-resistant LNCaP-abl cells that have a largely distinct AR-dependent gene-expression program as a result of transcriptional reprogramming (28). We therefore asked how they might affect AR-mediated gene expression, even after transcriptional reprogramming, by performing genome-wide analysis by RNA-seq (29).
We first identified AR-dependent genes on LNCaP-abl cells by AR RNAi, and then compared them to altered genes in cardiac glycoside-treated cells (Fig. 4). Based on a stringent cutoff (>twofold, P < 0.01), we identified 2,056 genes that responded to at least one treatment (AR knockdown or treatment with individual cardiac glycosides). When ranked by averaged responses to cardiac glycoside treatments, we found that the induced gene expression in AR knockdown cells largely matched those in cardiac glycoside-treated cells and most of the overlapped genes changed in the same direction. Taken together, these data demonstrated that cardiac glycosides are able to selectively and effectively block AR-dependent gene expression in LNCaP-abl cells, even though the AR program has been dramatically altered compared with androgen-sensitive LNCaP cells to support their androgen independent growth.
Fig. 4.
Global analysis of cardiac glycosides in comparison with AR RNAi on LNCaP-abl cells. (A) LNCaP-abl cells treated with 5 μM of individual cardiac glycosides were compared with the effect of AR RNAi (the knockdown efficiency is shown next to the heatmap). Significant changes (>twofold; P < 0.01) were identified, which added up to a total of 2,056 genes that were either up-regulated (red) or down-regulated (blue) on at least one treatment condition. (B) Venn diagrams of overlapped changes between AR RNAi and individual cardiac glycosides. (C) Overlapped genes showed changes largely in the same directions.
Peruvoside Acts as the Potent Inducer of AR Degradation.
To further understand how cardiac glycosides were able to specifically block AR-dependent gene expression, we asked how various cardiac glycosides might modulate AR expression. By RT-qPCR, we found that these compounds had little effect on AR expression, even though they effectively blocked the induction of the AR-regulated KLK3 gene (Fig. 5 A and B). However, by Western blot analysis, we found that the AR protein was rapidly degraded in cardiac glycoside-treated cells (Fig. 5C). Peruvoside again emerged as the most potent inducer of AR degradation with an estimated IC50 of 10–20 nM, similar to its IC50 value based on cell proliferation (Fig. 3B). AR degradation induced by cardiac glycosides took place 3–6 h posttreatment (Fig. 5D). Peruvoside was also most potent in inducing AR degradation in PC3-AR cells (Fig. 5E).
Fig. 5.
Cardiac glycosides induce rapid AR degradation. (A and B) Effect on the expression of KLK3 (A) and AR (B) in LNCaP-abl cells. SDs are based on triplicate experiments. (C and D) Western blot of AR in LNCaP-abl cells treated with different concentrations of compound for 24 h (C) or with Digoxin for different periods of time (D). (E) Western blot of AR in PC3-AR cells treated with different concentrations of compound for 24 h. (F) Knockdown of the catalytic subunit (ATP1α1) of the Na+/K+ ATPase, showing no effect on AR protein in LNCaP-abl cells. (G) Prevention of induced AR degradation by the proteasome inhibitor MG132. Note that a truncated AR because of calpain-mediated AR cleavage (31) or alternative splicing (32) became detectable in MG132-treated LNCaP-abl cells.
Cardiac glycosides are well-known inhibitors of the Na+/K+ ATPase via their direct binding to the sodium pump (11), which was recently shown to be sufficient to impair the induction of the IFN-β pathway (30). However, cardiac glycoside-induced hypoxia inducible factor (HIF)-1α degradation in prostate cancer cells appears independent of this mechanism because RNAi knock down of the catalytic subunit ATP1a1 of the sodium pump had little effect on the stability of HIF-1α (26). We observed little effect of ATP1α1 RNAi in AR (Fig. 5F). These observations suggest that the broad anticancer effect of cardiac glycosides may result from enhanced degradation of key cancer gene products. Finally, we provided evidence that the 26S proteasome pathway is likely responsible for AR degradation, because the effect could be fully suppressed by MG132 in cardiac glycoside-treated cells (Fig. 5G). Interestingly, we found that the MG132 treatment not only prevented degradation of full-length AR, but also stabilized a truncated AR, which has been attributed to alternative splicing of AR transcripts and posttranslational cleavage of AR by the protease Calpaine and this truncated AR has been shown to contribute to androgen resistance in prostate cancer cells (31, 32). The induced proteolytical degradation of AR by cardiac glycosides may thus prove to be an effective therapeutic strategy against advanced prostate tumors rising from diverse mechanisms.
Discussion
Our present study elaborates a powerful pathway-centric HTS by using the latest deep-sequencing technology. This approach offers a number of advantages over conventional chemical screening strategies. The approach does not require prior identification of specific drug targets, thus equally applicable to both “druggable” and “nondruggable” disease paradigms. This multitarget, pathway-centric approach relies on the behavior of endogenous genes (instead of engineered reporter) and permits identification of hits that intervene with any potential attack points in the pathway. The approach also overcomes the central shortcoming of phenotypic screening because specific gene-expression responses provide critical clues to potential molecular mechanisms. Thus, the approach described herein may help implement a recommended quick-win/fast-fail strategy in early phases of drug discovery to improve the drug research and development productivity (4).
The present screening reidentified a number of compounds previously scored from a PSA reporter system (16) or from an AR conformation change-based screen (21, 22). Interestingly, a group of compounds identified from our screen belongs to the family of cardiac glycosides, with Peruvoside showing the most potent effect. Cardiac glycosides have been previously shown for their broad anticancer activities (11). Our genome-wide analysis demonstrated that they could largely mimic AR RNAi, explaining their antiproliferation effects on both androgen-sensitive and refractory prostate cancer cells that still depend on AR for growth.
Cardiac glycosides have been best characterized as inhibitors of the Na+/K+ ATPase in the cell, but a long list of other potential mechanisms has also been documented in the literature (11). We tentatively rule out the mechanism for induced AR degradation because RNAi against the major subunit of the enzyme had little effect on AR integrity. By whatever mechanism, the induction of AR degradation provides a plausible mechanism for the observed effect of cardiac glycosides in preventing prostate cancer among congestive heart disease patients treated with a widely prescribed cardiac glycoside (Dixogin) compared with untreated groups (12). Specific cardiac glycosides may therefore be further developed as therapeutic modalities against androgen-resistant prostate cancer.
Finally, we wish to emphasize the broad utility of the HTS2 technology in both basic and translational research. Analysis of a pathway-specific gene signature coupled with perturbation of the pathway by RNAi has been used to deduce gene networks and crosstalk among Toll-like receptors in response to diverse pathogens (33). The HTS2 technology would permit more comprehensive studies in combination with genome-wide RNAi to systematically deduce regulatory networks underlying diverse biological pathways. The HTS2 technology may also be used to link SNPs to causal mutations in human diseases because the underlying RASL assay has the single nucleotide resolution in monitoring gene expression and mRNA isoforms (13, 14). Thus, the HTS2 technology offers a general platform for large-scale genetics and chemical genetics studies.
Methods
Culture conditions for various prostate cancer cells, AR RNAi, and Western blot analysis were as described (34, 35). The AR binding assay was as previously described (21). Methods for chemical screening and for scoring compound efficacy and specificity are detailed in SI Methods.
Supplementary Material
Acknowledgments
The authors thank H.-J. Kung of the University of California at Davis for providing isogenic PC3 and PC3-AR cells, and H. Klocker of Innsbruck Medical University, Austria for sending us LNCaP-abl cells. This work was supported by the Challenge Award from the Prostate Cancer Foundation (to S.D., X.-D.F, and M.G.R.), and National Human Genome Research Institute Grant HG004659 (to X.-D.F.). M.G.R. is an Investigator of the Howard Hughes Medical Institute.
Footnotes
The authors declare no conflict of interest.
Data deposition: The RNA-seq data reported in this paper have been deposited in the Gene Expression Omnibus (GEO) database, www.ncbi.nlm.nih.gov/geo (accession no. GSE35126).
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1200305109/-/DCSupplemental.
References
- 1.Flordellis CS, Manolis AS, Paris H, Karabinis A. Rethinking target discovery in polygenic diseases. Curr Top Med Chem. 2006;6:1791–1798. doi: 10.2174/156802606778194226. [DOI] [PubMed] [Google Scholar]
- 2.Swinney DC, Anthony J. How were new medicines discovered? Nat Rev Drug Discov. 2011;10:507–519. doi: 10.1038/nrd3480. [DOI] [PubMed] [Google Scholar]
- 3.Imming P, Sinning C, Meyer A. Drugs, their targets and the nature and number of drug targets. Nat Rev Drug Discov. 2006;5:821–834. doi: 10.1038/nrd2132. [DOI] [PubMed] [Google Scholar]
- 4.Paul SM, et al. How to improve R&D productivity: The pharmaceutical industry's grand challenge. Nat Rev Drug Discov. 2010;9:203–214. doi: 10.1038/nrd3078. [DOI] [PubMed] [Google Scholar]
- 5.Stegmaier K, et al. Gene expression-based high-throughput screening (GE-HTS) and application to leukemia differentiation. Nat Genet. 2004;36:257–263. doi: 10.1038/ng1305. [DOI] [PubMed] [Google Scholar]
- 6.Hieronymus H, et al. Gene expression signature-based chemical genomic prediction identifies a novel class of HSP90 pathway modulators. Cancer Cell. 2006;10:321–330. doi: 10.1016/j.ccr.2006.09.005. [DOI] [PubMed] [Google Scholar]
- 7.Corsello SM, et al. Identification of AML1-ETO modulators by chemical genomics. Blood. 2009;113:6193–6205. doi: 10.1182/blood-2008-07-166090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chen CD, et al. Molecular determinants of resistance to antiandrogen therapy. Nat Med. 2004;10:33–39. doi: 10.1038/nm972. [DOI] [PubMed] [Google Scholar]
- 9.Niu Y, et al. Differential androgen receptor signals in different cells explain why androgen-deprivation therapy of prostate cancer fails. Oncogene. 2010;29:3593–3604. doi: 10.1038/onc.2010.121. [DOI] [PubMed] [Google Scholar]
- 10.Goldin AG, Safa AR. Digitalis and cancer. Lancet. 1984;1:1134. doi: 10.1016/s0140-6736(84)92556-x. [DOI] [PubMed] [Google Scholar]
- 11.Prassas I, Diamandis EP. Novel therapeutic applications of cardiac glycosides. Nat Rev Drug Discov. 2008;7:926–935. doi: 10.1038/nrd2682. [DOI] [PubMed] [Google Scholar]
- 12.Platz EA, et al. A novel two-stage, transdisciplinary study identifies digoxin as a possible drug for prostate cancer treatment. Cancer Discov. 2011;2011:68–77. doi: 10.1158/2159-8274.CD-10-0020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Yeakley JM, et al. Profiling alternative splicing on fiber-optic arrays. Nat Biotechnol. 2002;20:353–358. doi: 10.1038/nbt0402-353. [DOI] [PubMed] [Google Scholar]
- 14.Li HR, et al. Two-dimensional transcriptome profiling: Identification of messenger RNA isoform signatures in prostate cancer from archived paraffin-embedded cancer specimens. Cancer Res. 2006;66:4079–4088. doi: 10.1158/0008-5472.CAN-05-4264. [DOI] [PubMed] [Google Scholar]
- 15.Fan JB, et al. A versatile assay for high-throughput gene expression profiling on universal array matrices. Genome Res. 2004;14:878–885. doi: 10.1101/gr.2167504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mashima T, Okabe S, Seimiya H. Pharmacological targeting of constitutively active truncated androgen receptor by nigericin and suppression of hormone-refractory prostate cancer cell growth. Mol Pharmacol. 2010;78:846–854. doi: 10.1124/mol.110.064790. [DOI] [PubMed] [Google Scholar]
- 17.Li H, et al. Determination of tag density required for digital transcriptome analysis: Application to an androgen-sensitive prostate cancer model. Proc Natl Acad Sci USA. 2008;105:20179–20184. doi: 10.1073/pnas.0807121105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Covell DG, et al. Linking tumor cell cytotoxicity to mechanism of drug action: An integrated analysis of gene expression, small-molecule screening and structural databases. Proteins. 2005;59:403–433. doi: 10.1002/prot.20392. [DOI] [PubMed] [Google Scholar]
- 19.Reid P, Kantoff P, Oh W. Antiandrogens in prostate cancer. Invest New Drugs. 1999;17:271–284. doi: 10.1023/a:1006344807086. [DOI] [PubMed] [Google Scholar]
- 20.McPherson SJ, et al. Essential role for estrogen receptor beta in stromal-epithelial regulation of prostatic hyperplasia. Endocrinology. 2007;148:566–574. doi: 10.1210/en.2006-0906. [DOI] [PubMed] [Google Scholar]
- 21.Jones JO, et al. Non-competitive androgen receptor inhibition in vitro and in vivo. Proc Natl Acad Sci USA. 2009;106:7233–7238. doi: 10.1073/pnas.0807282106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Jones JO, Diamond MI. A cellular conformation-based screen for androgen receptor inhibitors. ACS Chem Biol. 2008;3:412–418. doi: 10.1021/cb800054w. [DOI] [PubMed] [Google Scholar]
- 23.Gupta PB, et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell. 2009;138:645–659. doi: 10.1016/j.cell.2009.06.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yeh JY, Huang WJ, Kan SF, Wang PS. Effects of bufalin and cinobufagin on the proliferation of androgen dependent and independent prostate cancer cells. Prostate. 2003;54:112–124. doi: 10.1002/pros.10172. [DOI] [PubMed] [Google Scholar]
- 25.McConkey DJ, Lin Y, Nutt LK, Ozel HZ, Newman RA. Cardiac glycosides stimulate Ca2+ increases and apoptosis in androgen-independent, metastatic human prostate adenocarcinoma cells. Cancer Res. 2000;60:3807–3812. [PubMed] [Google Scholar]
- 26.Zhang H, et al. Digoxin and other cardiac glycosides inhibit HIF-1alpha synthesis and block tumor growth. Proc Natl Acad Sci USA. 2008;105:19579–19586. doi: 10.1073/pnas.0809763105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Huh JR, Jr, et al. Digoxin and its derivatives suppress TH17 cell differentiation by antagonizing RORγt activity. Nature. 2011;472:486–490. doi: 10.1038/nature09978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wang Q, et al. Androgen receptor regulates a distinct transcription program in androgen-independent prostate cancer. Cell. 2009;138:245–256. doi: 10.1016/j.cell.2009.04.056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Fox-Walsh K, Davis-Turak J, Zhou Y, Li H, Fu XD. A multiplex RNA-seq strategy to profile poly(A+) RNA: Application to analysis of transcription response and 3′ end formation. Genomics. 2011;98:266–271. doi: 10.1016/j.ygeno.2011.04.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ye J, Chen S, Maniatis T. Cardiac glycosides are potent inhibitors of interferon-β gene expression. Nat Chem Biol. 2011;7:25–33. doi: 10.1038/nchembio.476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Libertini SJ, et al. Evidence for calpain-mediated androgen receptor cleavage as a mechanism for androgen independence. Cancer Res. 2007;67:9001–9005. doi: 10.1158/0008-5472.CAN-07-1072. [DOI] [PubMed] [Google Scholar]
- 32.Guo Z, et al. A novel androgen receptor splice variant is up-regulated during prostate cancer progression and promotes androgen depletion-resistant growth. Cancer Res. 2009;69:2305–2313. doi: 10.1158/0008-5472.CAN-08-3795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Amit I, et al. Unbiased reconstruction of a mammalian transcriptional network mediating pathogen responses. Science. 2009;326:257–263. doi: 10.1126/science.1179050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Wang D, et al. Reprogramming transcription by distinct classes of enhancers functionally defined by eRNA. Nature. 2011;474:390–394. doi: 10.1038/nature10006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Li H, Qiu J, Fu X-D. RASL-seq for massive parallel and quantitative analysis of gene expression. Curr Protoc Mol Biol. 2011 doi: 10.1002/0471142727.mb0413s98. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.