Abstract
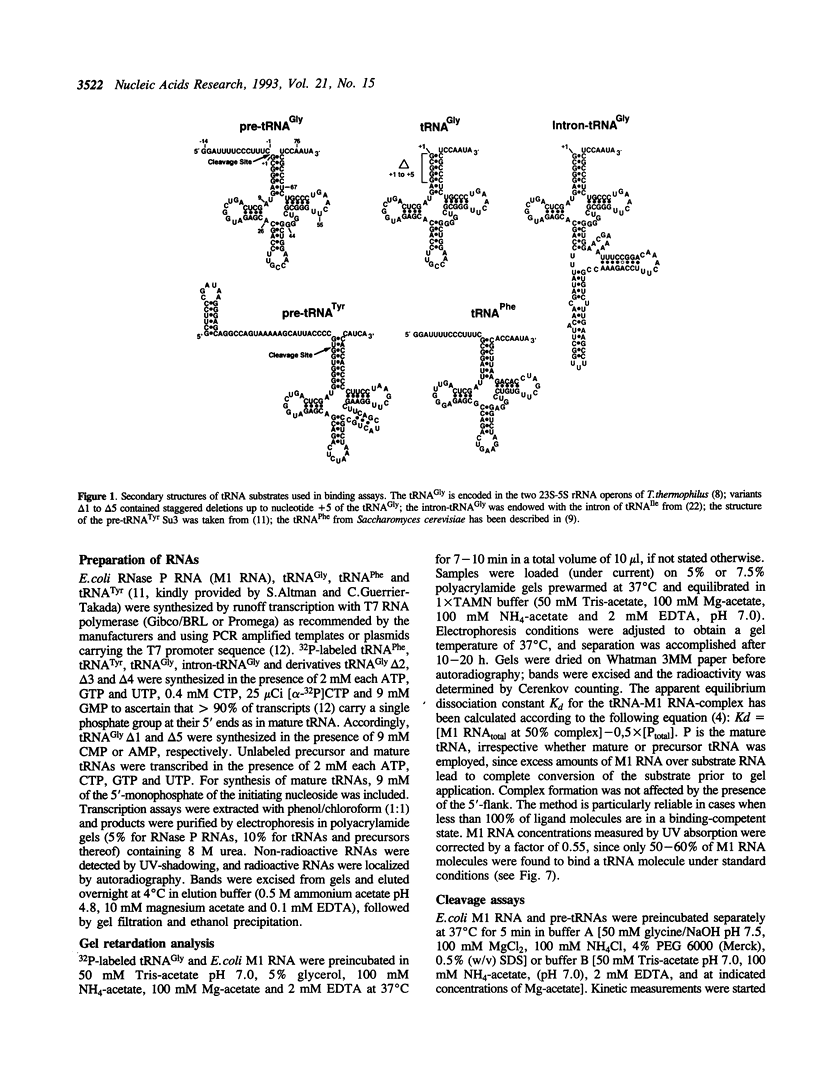
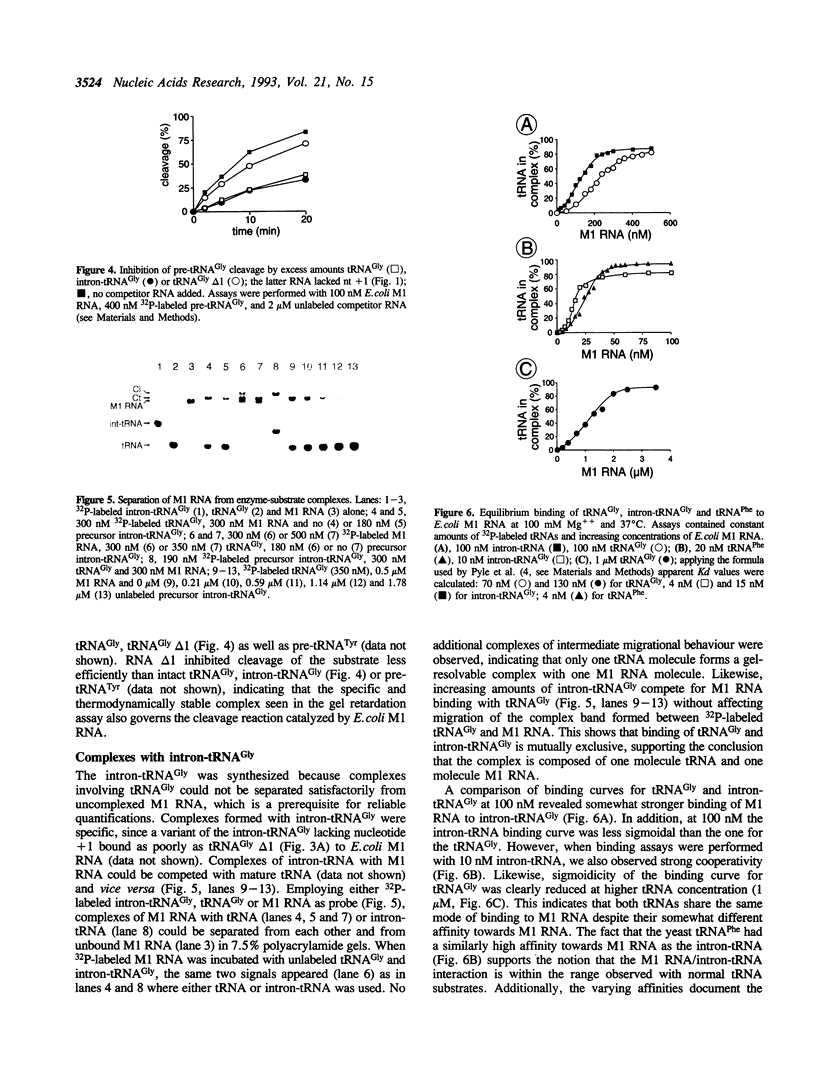
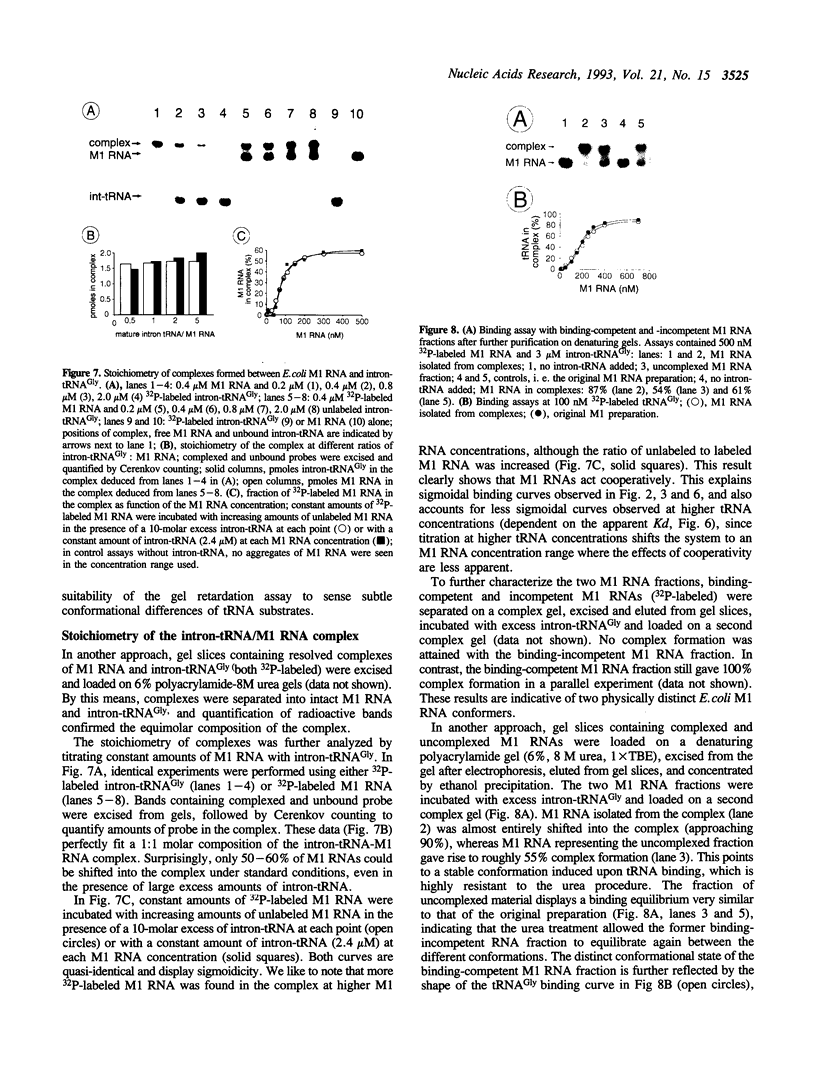
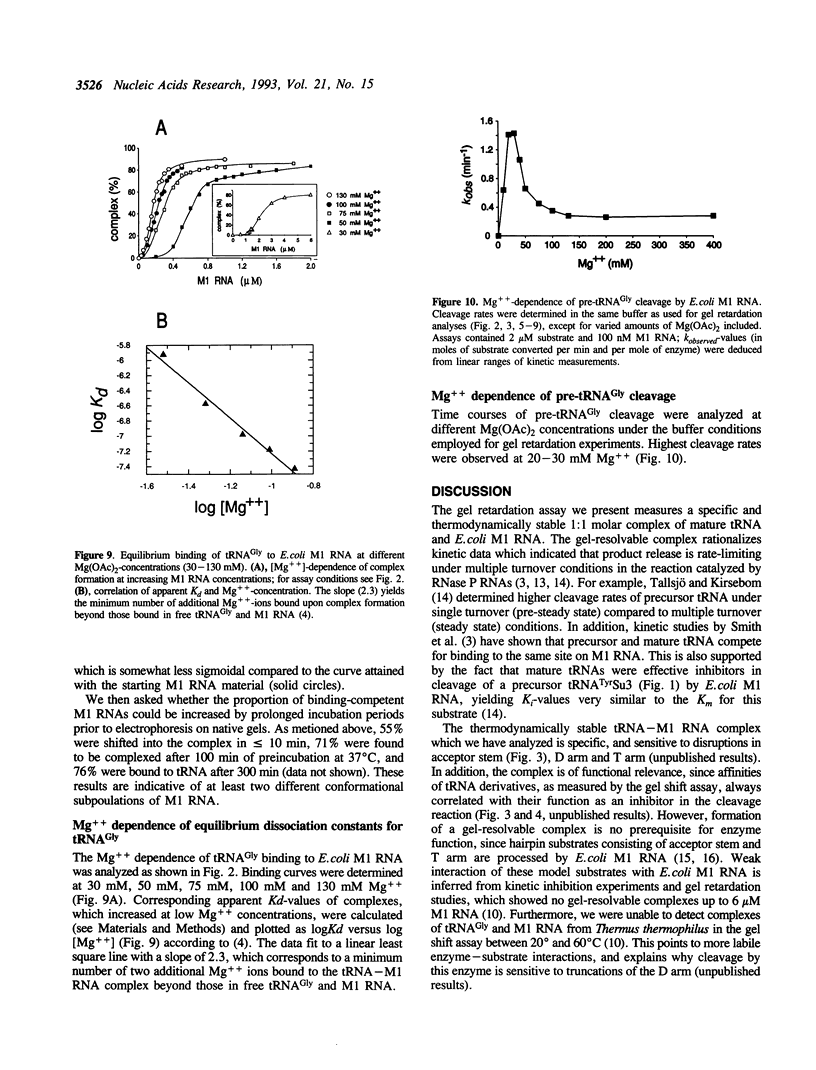
We have analyzed complexes between tRNA and E. coli M1 RNA by electrophoresis in non-denaturing polyacrylamide gels. The RNA subunit of E. coli RNase P formed a specific complex with mature tRNA molecules. A derivative of the tRNA(Gly), endowed with the intron of yeast tRNA(ile) (60 nt), was employed to improve separation of complexed and unbound M1 RNA. Binding assays with tRNA(Gly) and intron-tRNA(Gly) as well as analysis of intron-tRNA/M1 RNA complexes on denaturing gels showed that one tRNA is bound per molecule of M1 RNA. A tRNA carrying a truncation as small as the 5'-nucleotide had a strongly reduced affinity to M1 RNA and was also a weak competitor in the cleavage reaction, suggesting that nucleotide +1 is a major determinant of tRNA recognition and that the thermodynamically stable tRNA-M1 RNA complex is relevant for enzyme function. Binding was shown to be dependent on the M1 RNA concentration in a cooperative fashion. Only a fraction of M1 RNAs (50-60%) readily formed a complex with intron-tRNA(Gly), indicating that distinct conformational subpopulations of M1 RNA may exist. Formation of the M1 RNA-tRNA(Gly), complex was very similar at 100 mM Mg++ and Ca++, corroborating earlier data that Ca++ is competent in promoting M1 RNA folding and tRNA binding. Determination of apparent equilibrium constants (app Kd) for tRNA(Gly) as a function of the Mg++ concentration supports an uptake of at least two additional Mg++ ions upon complex formation. At 20-30 mM Mg++, highest cleavage rates but strongly reduced complex formation were observed. This indicates that tight binding of the tRNA to the catalytic RNA at higher magnesium concentrations retards product release and therefore substrate turnover.
Full text
PDF


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Forster A. C., Altman S. External guide sequences for an RNA enzyme. Science. 1990 Aug 17;249(4970):783–786. doi: 10.1126/science.1697102. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Gardiner K., Marsh T., Pace N., Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983 Dec;35(3 Pt 2):849–857. doi: 10.1016/0092-8674(83)90117-4. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Haydock K., Allen L., Altman S. Metal ion requirements and other aspects of the reaction catalyzed by M1 RNA, the RNA subunit of ribonuclease P from Escherichia coli. Biochemistry. 1986 Apr 8;25(7):1509–1515. doi: 10.1021/bi00355a006. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Lumelsky N., Altman S. Specific interactions in RNA enzyme-substrate complexes. Science. 1989 Dec 22;246(4937):1578–1584. doi: 10.1126/science.2480641. [DOI] [PubMed] [Google Scholar]
- Hartmann R. K., Toschka H. Y., Erdmann V. A. Processing and termination of 23S rRNA-5S rRNA-tRNA(Gly) primary transcripts in Thermus thermophilus HB8. J Bacteriol. 1991 Apr;173(8):2681–2690. doi: 10.1128/jb.173.8.2681-2690.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartmann R. K., Ulbrich N., Erdmann V. A. An unusual rRNA operon constellation: in Thermus thermophilus HB8 the 23S/5S rRNA operon is a separate entity from the 16S rRNA operon. Biochimie. 1987 Oct;69(10):1097–1104. doi: 10.1016/0300-9084(87)90009-5. [DOI] [PubMed] [Google Scholar]
- Herschlag D., Cech T. R. Catalysis of RNA cleavage by the Tetrahymena thermophila ribozyme. 1. Kinetic description of the reaction of an RNA substrate complementary to the active site. Biochemistry. 1990 Nov 6;29(44):10159–10171. doi: 10.1021/bi00496a003. [DOI] [PubMed] [Google Scholar]
- Holm P. S., Krupp G. The acceptor stem in pre-tRNAs determines the cleavage specificity of RNase P. Nucleic Acids Res. 1992 Feb 11;20(3):421–423. doi: 10.1093/nar/20.3.421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kahle D., Wehmeyer U., Krupp G. Substrate recognition by RNase P and by the catalytic M1 RNA: identification of possible contact points in pre-tRNAs. EMBO J. 1990 Jun;9(6):1929–1937. doi: 10.1002/j.1460-2075.1990.tb08320.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirsebom L. A., Svärd S. G. The kinetics and specificity of cleavage by RNase P is mainly dependent on the structure of the amino acid acceptor stem. Nucleic Acids Res. 1992 Feb 11;20(3):425–432. doi: 10.1093/nar/20.3.425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClain W. H., Guerrier-Takada C., Altman S. Model substrates for an RNA enzyme. Science. 1987 Oct 23;238(4826):527–530. doi: 10.1126/science.2443980. [DOI] [PubMed] [Google Scholar]
- Milligan J. F., Uhlenbeck O. C. Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol. 1989;180:51–62. doi: 10.1016/0076-6879(89)80091-6. [DOI] [PubMed] [Google Scholar]
- Pyle A. M., McSwiggen J. A., Cech T. R. Direct measurement of oligonucleotide substrate binding to wild-type and mutant ribozymes from Tetrahymena. Proc Natl Acad Sci U S A. 1990 Nov;87(21):8187–8191. doi: 10.1073/pnas.87.21.8187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reich C., Olsen G. J., Pace B., Pace N. R. Role of the protein moiety of ribonuclease P, a ribonucleoprotein enzyme. Science. 1988 Jan 8;239(4836):178–181. doi: 10.1126/science.3122322. [DOI] [PubMed] [Google Scholar]
- Sampson J. R., Uhlenbeck O. C. Biochemical and physical characterization of an unmodified yeast phenylalanine transfer RNA transcribed in vitro. Proc Natl Acad Sci U S A. 1988 Feb;85(4):1033–1037. doi: 10.1073/pnas.85.4.1033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schlegl J., Fürste J. P., Bald R., Erdmann V. A., Hartmann R. K. Cleavage efficiencies of model substrates for ribonuclease P from Escherichia coli and Thermus thermophilus. Nucleic Acids Res. 1992 Nov 25;20(22):5963–5970. doi: 10.1093/nar/20.22.5963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith D., Burgin A. B., Haas E. S., Pace N. R. Influence of metal ions on the ribonuclease P reaction. Distinguishing substrate binding from catalysis. J Biol Chem. 1992 Feb 5;267(4):2429–2436. [PubMed] [Google Scholar]
- Tallsjö A., Kirsebom L. A. Product release is a rate-limiting step during cleavage by the catalytic RNA subunit of Escherichia coli RNase P. Nucleic Acids Res. 1993 Jan 11;21(1):51–57. doi: 10.1093/nar/21.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thurlow D. L., Shilowski D., Marsh T. L. Nucleotides in precursor tRNAs that are required intact for catalysis by RNase P RNAs. Nucleic Acids Res. 1991 Feb 25;19(4):885–891. doi: 10.1093/nar/19.4.885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vioque A., Arnez J., Altman S. Protein-RNA interactions in the RNase P holoenzyme from Escherichia coli. J Mol Biol. 1988 Aug 20;202(4):835–848. doi: 10.1016/0022-2836(88)90562-1. [DOI] [PubMed] [Google Scholar]