Abstract
Comparison of the deduced amino acid sequences of DNA-[N6-adenine]-methyltransferases has revealed several conserved regions. All of these enzymes contain a DPPY-motif, or a variant of it. By site-directed mutagenesis of a cloned T4 dam gene, we have altered the first proline residue in this motif (located in conserved region IV of the T4 Dam-MTase) to alanine or threonine. The mutant enzymic forms, P172A and P172T, were overproduced and purified. Kinetic studies showed that compared to the wild-type (wt) the two mutant enzymic forms had: (i) an increased (6 and 23-fold, respectively) Km for substrate, S-adenosyl-methionine (AdoMet) and an increased (6 and 23-fold) Ki for product, S-adenosyl-homocysteine (AdoHcy); (ii) a slightly reduced (1.5 and 3-fold lower) kcat; (iii) a strongly reduced kcat/KmAdoMet (10 and 80-fold); and (iv) the same Km for substrate DNA. Equilibrium dialysis studies showed that the mutant enzymes had a reduced (3 and 7-fold lower) Ka for AdoMet; all forms bound two molecules of AdoMet. Taken together these data indicate that the P172A and P172T alterations resulted primarily in a reduced affinity for AdoMet. This suggests that the DPPY-motif is important for AdoMet-binding, and that region IV contains an AdoMet-binding site.
Full text
PDF
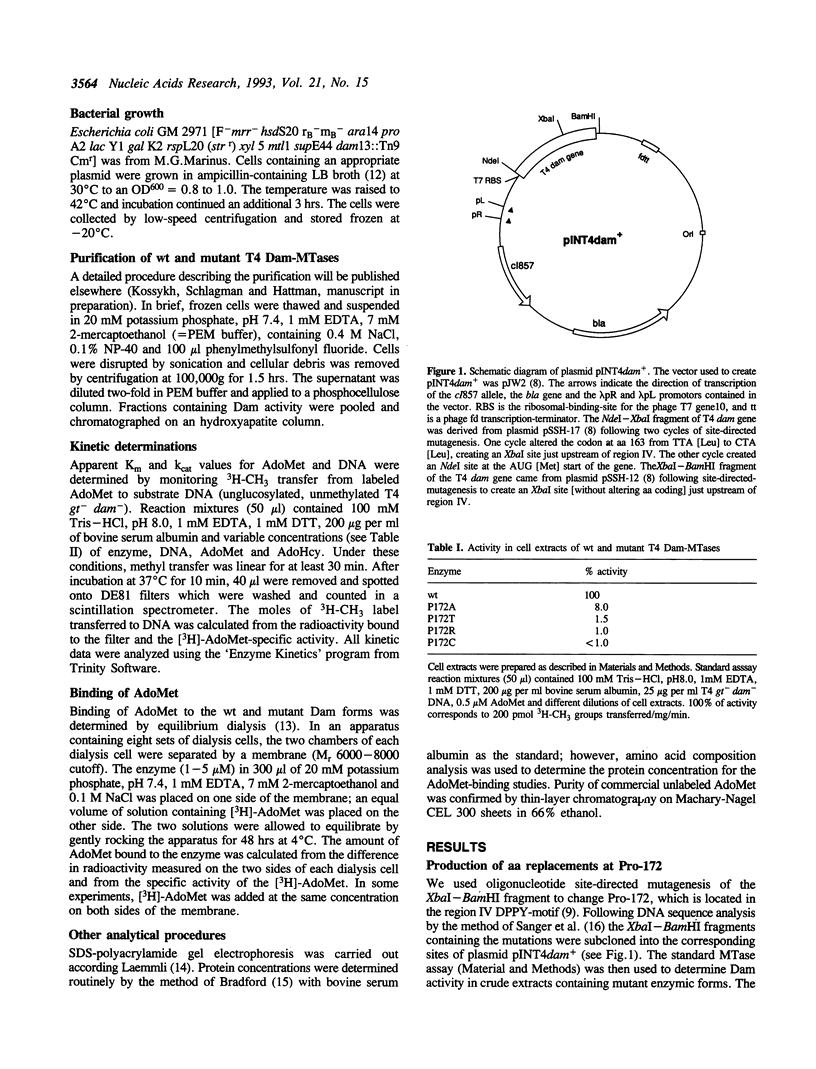

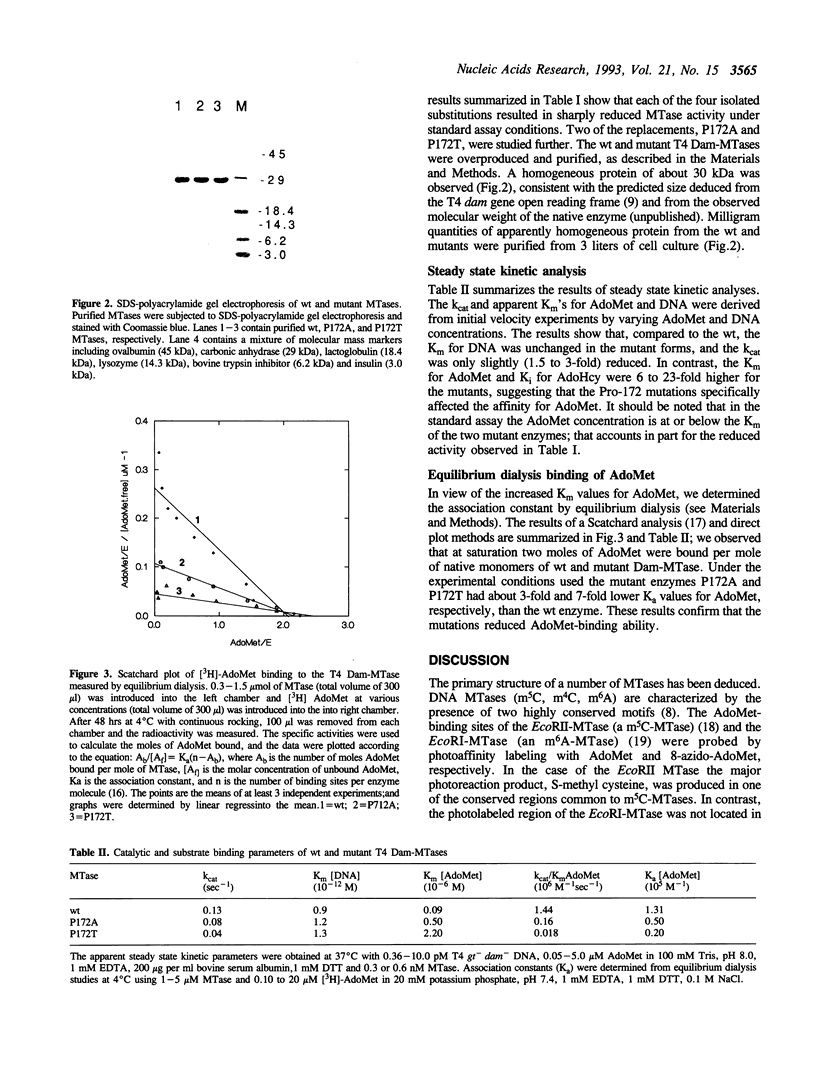

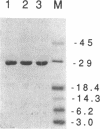
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bergerat A., Guschlbauer W. The double role of methyl donor and allosteric effector of S-adenosyl-methionine for Dam methylase of E. coli. Nucleic Acids Res. 1990 Aug 11;18(15):4369–4375. doi: 10.1093/nar/18.15.4369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Guschlbauer W. The DNA and S-adenosylmethionine-binding regions of EcoDam and related methyltransferases. Gene. 1988 Dec 25;74(1):211–214. doi: 10.1016/0378-1119(88)90289-2. [DOI] [PubMed] [Google Scholar]
- Hattman S., Wilkinson J., Swinton D., Schlagman S., Macdonald P. M., Mosig G. Common evolutionary origin of the phage T4 dam and host Escherichia coli dam DNA-adenine methyltransferase genes. J Bacteriol. 1985 Nov;164(2):932–937. doi: 10.1128/jb.164.2.932-937.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klimasauskas S., Timinskas A., Menkevicius S., Butkienè D., Butkus V., Janulaitis A. Sequence motifs characteristic of DNA[cytosine-N4]methyltransferases: similarity to adenine and cytosine-C5 DNA-methylases. Nucleic Acids Res. 1989 Dec 11;17(23):9823–9832. doi: 10.1093/nar/17.23.9823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkel T. A., Bebenek K., McClary J. Efficient site-directed mutagenesis using uracil-containing DNA. Methods Enzymol. 1991;204:125–139. doi: 10.1016/0076-6879(91)04008-c. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lauster R., Kriebardis A., Guschlbauer W. The GATATC-modification enzyme EcoRV is closely related to the GATC-recognizing methyltransferases DpnII and dam from E. coli and phage T4. FEBS Lett. 1987 Aug 10;220(1):167–176. doi: 10.1016/0014-5793(87)80897-9. [DOI] [PubMed] [Google Scholar]
- Lauster R., Trautner T. A., Noyer-Weidner M. Cytosine-specific type II DNA methyltransferases. A conserved enzyme core with variable target-recognizing domains. J Mol Biol. 1989 Mar 20;206(2):305–312. doi: 10.1016/0022-2836(89)90480-4. [DOI] [PubMed] [Google Scholar]
- Pósfai J., Bhagwat A. S., Pósfai G., Roberts R. J. Predictive motifs derived from cytosine methyltransferases. Nucleic Acids Res. 1989 Apr 11;17(7):2421–2435. doi: 10.1093/nar/17.7.2421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reich N. O., Everett E. A. Identification of peptides involved in S-adenosylmethionine binding in the EcoRI DNA methylase. Photoaffinity laveling with 8-azido-S-adenosylmethionine. J Biol Chem. 1990 May 25;265(15):8929–8934. [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schlagman S. L., Miner Z., Fehér Z., Hattman S. The DNA [adenine-N6]methyltransferase (Dam) of bacteriophage T4. Gene. 1988 Dec 20;73(2):517–530. doi: 10.1016/0378-1119(88)90516-1. [DOI] [PubMed] [Google Scholar]
- Som S., Friedman S. Identification of a highly conserved domain in the EcoRII methyltransferase which can be photolabeled with S-adenosyl-L-[methyl-3H]methionine. Evidence for UV-induced transmethylation of cysteine 186. J Biol Chem. 1991 Feb 15;266(5):2937–2945. [PubMed] [Google Scholar]
- Sugisaki H., Kita K., Takanami M. The FokI restriction-modification system. II. Presence of two domains in FokI methylase responsible for modification of different DNA strands. J Biol Chem. 1989 Apr 5;264(10):5757–5761. [PubMed] [Google Scholar]
- Teraoka H., Nierhaus K. H. Measurement of the binding of antibiotics to ribosomal particles by means of equilibrium dialysis. Methods Enzymol. 1979;59:862–866. doi: 10.1016/0076-6879(79)59131-9. [DOI] [PubMed] [Google Scholar]
- Wilson G. G. Organization of restriction-modification systems. Nucleic Acids Res. 1991 May 25;19(10):2539–2566. doi: 10.1093/nar/19.10.2539. [DOI] [PMC free article] [PubMed] [Google Scholar]