Abstract
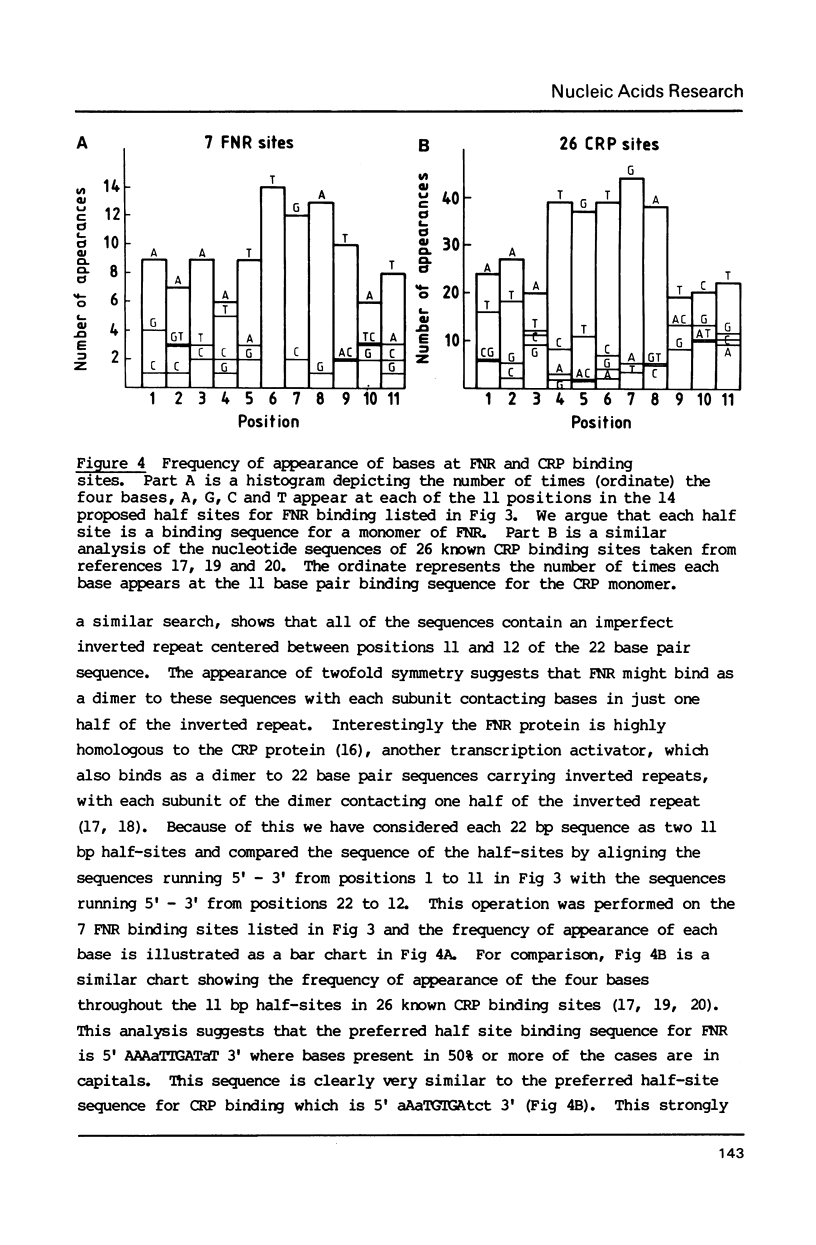
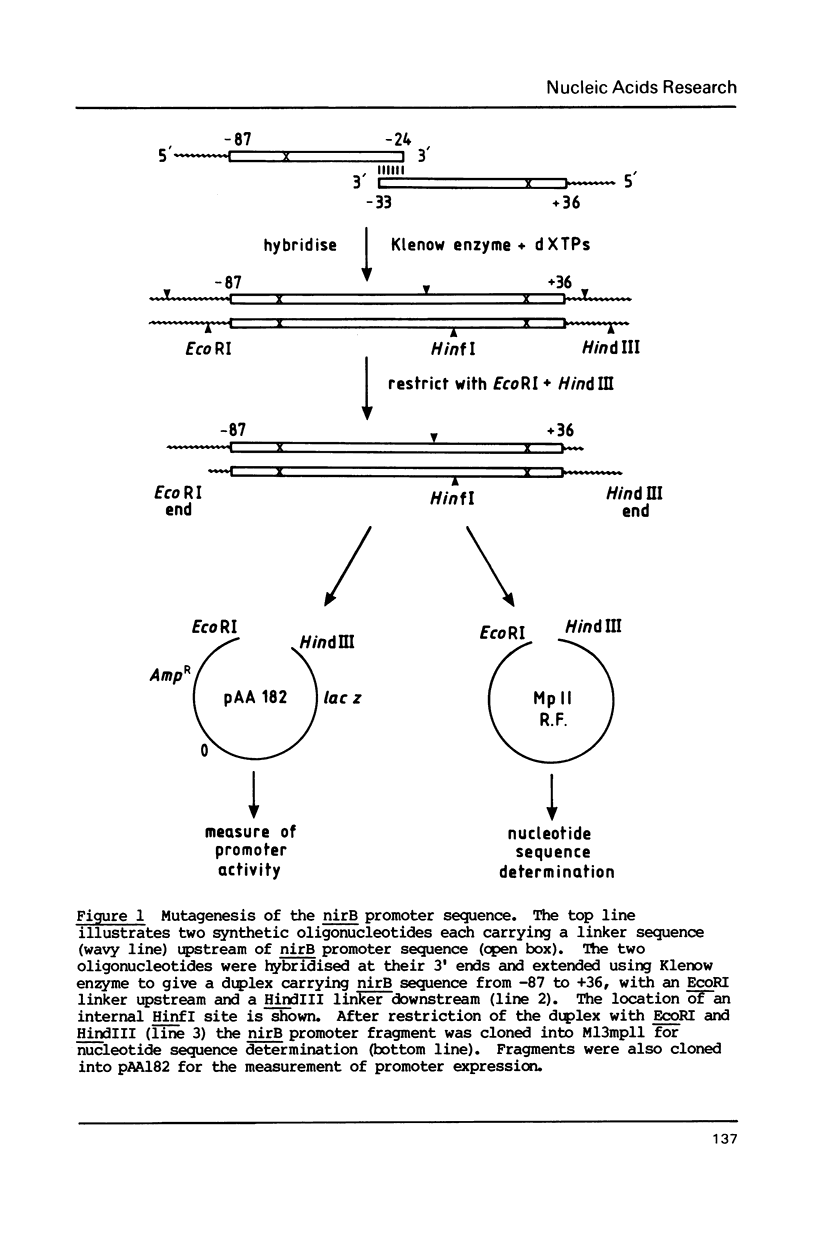
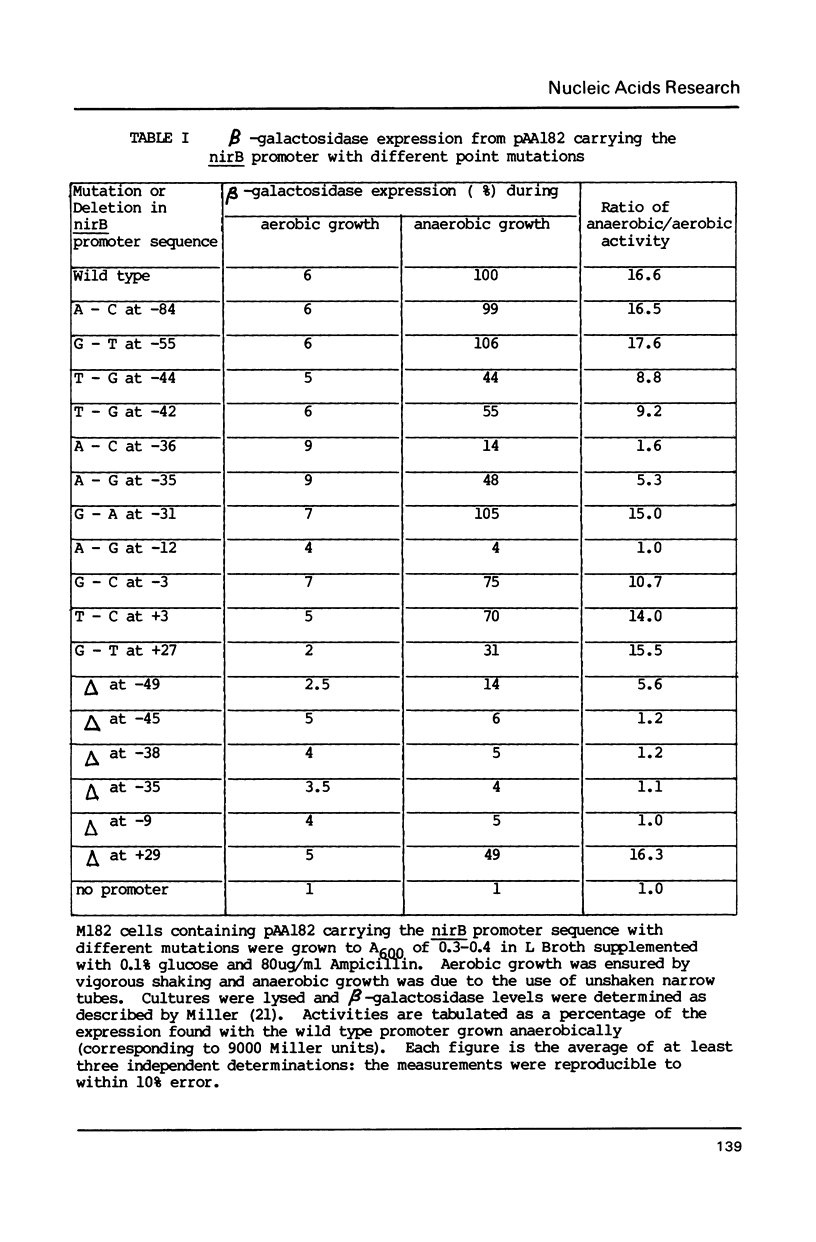
During anaerobic growth of E. coli, the FNR protein activates transcription initiation at the nirB promoter. After chemical synthesis using deliberately contaminated nucleotides, we isolated a series of recombinant plasmids with single point mutations or one base pair deletions in the nirB promoter. The effects of these alterations on the anaerobic induction of promoter activity were measured. Mutations that abolish anaerobic induction identify the -10 hexamer sequence whilst changes that allow reduced induction suggest positions involved in FNR binding. Comparison of the nucleotide sequence of the nirB promoter with other promoters that are regulated by FNR show clear homologies, suggesting consensus sequences for FNR binding sites, and confirming that some of the point mutations described here do indeed act by weakening FNR binding.
Full text
PDF








Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Berg O. G., von Hippel P. H. Selection of DNA binding sites by regulatory proteins. II. The binding specificity of cyclic AMP receptor protein to recognition sites. J Mol Biol. 1988 Apr 20;200(4):709–723. doi: 10.1016/0022-2836(88)90482-2. [DOI] [PubMed] [Google Scholar]
- Busby S., Dreyfus M. Segment-specific mutagenesis of the regulatory region in the Escherichia coli galactose operon: isolation of mutations reducing the initiation of transcription and translation. Gene. 1983 Jan-Feb;21(1-2):121–131. doi: 10.1016/0378-1119(83)90154-3. [DOI] [PubMed] [Google Scholar]
- Cole S. T. Nucleotide sequence and comparative analysis of the frd operon encoding the fumarate reductase of Proteus vulgaris. Extensive sequence divergence of the membrane anchors and absence of an frd-linked ampC cephalosporinase gene. Eur J Biochem. 1987 Sep 15;167(3):481–488. doi: 10.1111/j.1432-1033.1987.tb13362.x. [DOI] [PubMed] [Google Scholar]
- Cole S. T. Nucleotide sequence coding for the flavoprotein subunit of the fumarate reductase of Escherichia coli. Eur J Biochem. 1982 Mar 1;122(3):479–484. doi: 10.1111/j.1432-1033.1982.tb06462.x. [DOI] [PubMed] [Google Scholar]
- Gaston K., Chan B., Kolb A., Fox J., Busby S. Alterations in the binding site of the cyclic AMP receptor protein at the Escherichia coli galactose operon regulatory region. Biochem J. 1988 Aug 1;253(3):809–818. doi: 10.1042/bj2530809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griffiths L., Cole J. A. Lack of redox control of the anaerobically-induced nirB+ gene of Escherichia coli K-12. Arch Microbiol. 1987 May;147(4):364–369. doi: 10.1007/BF00406134. [DOI] [PubMed] [Google Scholar]
- Hawley D. K., McClure W. R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983 Apr 25;11(8):2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jayaraman P. S., Peakman T. C., Busby S. J., Quincey R. V., Cole J. A. Location and sequence of the promoter of the gene for the NADH-dependent nitrite reductase of Escherichia coli and its regulation by oxygen, the Fnr protein and nitrite. J Mol Biol. 1987 Aug 20;196(4):781–788. doi: 10.1016/0022-2836(87)90404-9. [DOI] [PubMed] [Google Scholar]
- Jones H. M., Gunsalus R. P. Transcription of the Escherichia coli fumarate reductase genes (frdABCD) and their coordinate regulation by oxygen, nitrate, and fumarate. J Bacteriol. 1985 Dec;164(3):1100–1109. doi: 10.1128/jb.164.3.1100-1109.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lambden P. R., Guest J. R. Mutants of Escherichia coli K12 unable to use fumarate as an anaerobic electron acceptor. J Gen Microbiol. 1976 Dec;97(2):145–160. doi: 10.1099/00221287-97-2-145. [DOI] [PubMed] [Google Scholar]
- Li S. F., DeMoss J. A. Promoter region of the nar operon of Escherichia coli: nucleotide sequence and transcription initiation signals. J Bacteriol. 1987 Oct;169(10):4614–4620. doi: 10.1128/jb.169.10.4614-4620.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Macdonald H., Cole J. Molecular cloning and functional analysis of the cysG and nirB genes of Escherichia coli K12, two closely-linked genes required for NADH-dependent nitrite reductase activity. Mol Gen Genet. 1985;200(2):328–334. doi: 10.1007/BF00425444. [DOI] [PubMed] [Google Scholar]
- Newman B. M., Cole J. A. The chromosomal location and pleiotropic effects of mutations of the nirA+ gene of Escherichia coli K12: the essential role of nirA+ in nitrite reduction and in other anaerobic redox reactions. J Gen Microbiol. 1978 May;106(1):1–12. doi: 10.1099/00221287-106-1-1. [DOI] [PubMed] [Google Scholar]
- Oliphant A. R., Nussbaum A. L., Struhl K. Cloning of random-sequence oligodeoxynucleotides. Gene. 1986;44(2-3):177–183. doi: 10.1016/0378-1119(86)90180-0. [DOI] [PubMed] [Google Scholar]
- Raibaud O., Schwartz M. Positive control of transcription initiation in bacteria. Annu Rev Genet. 1984;18:173–206. doi: 10.1146/annurev.ge.18.120184.001133. [DOI] [PubMed] [Google Scholar]
- Shaw D. J., Guest J. R. Nucleotide sequence of the fnr gene and primary structure of the Enr protein of Escherichia coli. Nucleic Acids Res. 1982 Oct 11;10(19):6119–6130. doi: 10.1093/nar/10.19.6119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw D. J., Rice D. W., Guest J. R. Homology between CAP and Fnr, a regulator of anaerobic respiration in Escherichia coli. J Mol Biol. 1983 May 15;166(2):241–247. doi: 10.1016/s0022-2836(83)80011-4. [DOI] [PubMed] [Google Scholar]
- Smith M. In vitro mutagenesis. Annu Rev Genet. 1985;19:423–462. doi: 10.1146/annurev.ge.19.120185.002231. [DOI] [PubMed] [Google Scholar]
- Spiro S., Guest J. R. Activation of the lac operon of Escherichia coli by a mutant FNR protein. Mol Microbiol. 1987 Jul;1(1):53–58. doi: 10.1111/j.1365-2958.1987.tb00526.x. [DOI] [PubMed] [Google Scholar]
- Spiro S., Guest J. R. Regulation and over-expression of the fnr gene of Escherichia coli. J Gen Microbiol. 1987 Dec;133(12):3279–3288. doi: 10.1099/00221287-133-12-3279. [DOI] [PubMed] [Google Scholar]
- Webster C., Gaston K., Busby S. Transcription from the Escherichia coli melR promoter is dependent on the cyclic AMP receptor protein. Gene. 1988 Sep 7;68(2):297–305. doi: 10.1016/0378-1119(88)90032-7. [DOI] [PubMed] [Google Scholar]
- Woods S. A., Miles J. S., Roberts R. E., Guest J. R. Structural and functional relationships between fumarase and aspartase. Nucleotide sequences of the fumarase (fumC) and aspartase (aspA) genes of Escherichia coli K12. Biochem J. 1986 Jul 15;237(2):547–557. doi: 10.1042/bj2370547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Crombrugghe B., Busby S., Buc H. Cyclic AMP receptor protein: role in transcription activation. Science. 1984 May 25;224(4651):831–838. doi: 10.1126/science.6372090. [DOI] [PubMed] [Google Scholar]


