Abstract
Rapid synthesis of the polyamine catabolic enzyme spermidine/spermine-N1-acetyltransferase (SSAT) in response to increased polyamines is an important polyamine homeostatic mechanism. Indirect evidence has suggested that there is an important control mechanism involving the release of a translational repressor protein that allows the immediate initiation of SSAT protein synthesis without RNA transcription, maturation, or translocation. To identify a repressor protein, we used a mass spectroscopy-based RNA-protein interaction system and found six proteins that bind to the coding region of SSAT mRNA. Individual small interfering RNA (siRNA) experiments showed that nucleolin knockdown enhances SSAT translation. Nucleolin exists in several isoforms, and we report that the isoform that binds to SSAT mRNA undergoes autocatalysis in the presence of polyamines, a result suggesting that there is a negative feedback system that helps control the cellular content of polyamines. Preliminary molecular interaction data show that a nucleolin isoform binds to a 5′ stem-loop of the coding region of SSAT mRNA. The glycine/arginine-rich C terminus of nucleolin is required for binding, and the four RNA recognition motif domains are included in the isoform that blocks SSAT translation. Understanding SSAT translational control mechanisms has the potential for the development of therapeutic strategies against cancer and obesity.
INTRODUCTION
Polyamines are small positively charged molecules present in all cells. The common polyamines putrescine, spermidine, and spermine are essential for cell growth. The rate of synthesis and content both increase with increased cell proliferation (47). Polyamine functions include stabilization of polynucleotides, regulation of transcription and translation, control of enzyme activities, modulation of ion channels, and response to oxidative stress (42, 61). Because so many processes are affected, levels are maintained within a relatively narrow range by shifts in anabolism/catabolism and import/export (1, 46). Manipulation of polyamine metabolism has been an anticancer strategy, with pool depletion in tumor cells used as a surrogate marker of efficacy (21, 34, 38, 48).
Many mechanisms contribute to control of eukaryotic polyamine metabolic enzymes (15, 41, 43, 50). Ornithine decarboxylase (ODC), the rate-limiting anabolic enzyme, is regulated allosterically, at transcription, at translation, and by “ODC antizyme,” a protein that binds to ODC monomers, thereby blocking homodimerization required for activity and accelerating monomer degradation (41). Antizyme itself is regulated by a translation control mechanism involving polyamine-induced ribosomal frame-shifting (15, 35). Spermidine/spermine-N1-acetyltransferase (SSAT) is the principal catabolic regulator. SSAT acetylates spermidine and spermine using acetyl-coenzyme A (CoA), thereby altering their charge and facilitating excretion. SSAT basal activity is very low but increases quickly when polyamines are in excess (11). There is evidence for transcription, translation, and posttranslation control (11), but transcription seems not to play a major role. For example, transgenic mice with more than 50 copies of the SSAT gene have large quantities of mRNA but only moderately increased enzymatic activity (44, 56). Furthermore, SSAT protein and enzymatic activity can increase markedly in response to elevated polyamines with little change in mRNA (19, 40, 44, 56). Despite the widely held postulation that SSAT is controlled mostly at translation, little is known about the mechanism. Earlier work suggested the 5′ and 3′ untranslated regions (UTRs) are not involved, the 5′ terminus of the coding region is involved, and an unknown repressor is involved (8, 40). Control at translation has advantages for rapid initiation of protein synthesis. When mRNA is presynthesized but translation is repressed, initiation can be achieved quickly by repression release without transcription, mRNA maturation, or translocation (20, 29). This allows proteins to be translated at the correct time, in the correct place, and at a high rate (3). Translation could be controlled at initiation, elongation, or termination, but repression is most often associated with control of initiation (55). Several translational control mechanisms that perturb these processes have been described, including heavily structured 5′ UTRs of mRNA, translational repressor proteins that interact with mRNA, and microRNAs (20, 45).
Our interest in SSAT translation control was driven by previous efforts to control polyamine metabolism for therapeutic purposes and work with transgenic mice that revealed a strong connection between SSAT activity and control of lipid metabolism. The original goals of the work we report here were to identify the SSAT translation repressor protein and to begin characterizing the mechanism involved. We identified an isoform of the abundant cellular protein nucleolin as the repressor, and our data show this isoform binds to a stem-loop at the extreme 5′ end of the mRNA coding region, likely stabilizing the loop and thus blocking translation. We also found that high polyamines selectively induce isoform degradation, which suggests operation of a negative feedback system: high polyamine concentrations induce isoform degradation, thereby releasing SSAT translation suppression, and increased SSAT activity promotes polyamine catabolism, thereby correcting polyamine concentrations. Conversely, low polyamines allow the nucleolin isoform to remain, thus inhibiting polyamine loss.
MATERIALS AND METHODS
Gene templates used for PCRs.
A plasmid containing human SSAT cDNA was obtained from Openbiosystems (accession no. BC002503). The cDNA of nucleolin was prepared using the Superscript III one-step reverse transcriptase PCR (RT-PCR) system (Invitrogen) and 500 ng of HEK293T cell total RNA. The following primers were used: forward, 5′ ATG GTG AAG CTC GCG AAG GC 3′; reverse, 5′ GGG AAA GCA GAG GGA CAG AAG C 3′. The PCR product was gel purified and cloned in the Pgem T vector (Promega) and was verified by sequencing.
Cell culture and recombinant constructs for protein expression.
HEK293T cells (ATCC) were grown in Dulbecco's modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum. The PLEX-MCS plasmid (Openbiosystems) was used for the expression of recombinant proteins. The plasmid was modified to include a C-terminal purification tag containing a Strep tag sequence and a 6-His sequence (CCPGCCASWSHPQFEKGGSNWSHPQFEKGGTGSENLYFQGRGGSENLYFQGEGTGSHHHHHH) (22). The following 12 SSAT constructs were created: ORF SSAT, Δ4-30, Δ4-45, Δ4-75, Δ52-117, Δ31-75, 5UTR+SSAT, 5UTR+Δ4-30SSAT, 5UTR+Δ49-114SSAT, 5UTR(ΔuORFs)+SSAT, 5UTR(ΔuORFs)+Δ4-30SSAT, and 5UTR(ΔuORFs)+Δ49-114SSAT. The first 6 were generated by performing PCR with a common reverse primer (5′ TCC CAC CGG TCT CCT CTG TTG CCA TTT TTA GC 3′) containing a restriction site for AgeI (Italicized) and 6 forward primers containing a BamHI (bold and italicized) restriction site and the Kozak sequence (lowercase): ORFSSAT (5′ CGG GAT CCg ccg cca cca tgG CTA AAT TCG TGA TCC G 3′), Δ 4-30 (5′ CGG GAT CCg ccg cca cca tgG CCG CCG ACT GCA GTG AC 3′), Δ 4-45 (5′ CGG GAT CCg ccg cca cca tgG ACA TAC TGC GGC TGA TCA AG 3′), Δ 4-75 (5′ CGG GAT CCg ccg cca cca tgA AAT ATG AAT ACA TGG AAG AAC AAG T 3′), Δ 31-75 (5′ CGG GAT CCg ccg cca cca tgG CTA AAT TCG TGA TCC GCC CAG CCA CTA AAT ATG AAT ACA TGG AAG AAC AAG 3”), and Δ 52-117 (5′ CGG GAT CCg ccg cca cca tgG CTA AAT TCG TGA TCC GCC CAG CCA CTG CCG CCG ACT GCA GTG ACA TAG ATC TGC TAG AAG ATG GTT TTG G 3′). The PCR products obtained with these oligonucleotides were purified with a PCR cleanup kit (Promega), digested with BamHI and AgeI (Fermentas), and ligated in the plasmid mentioned above. All the clones were confirmed by sequencing.
Another three different versions of SSAT were produced as follows. The construct “5′UTR+SSAT” was created with a forward oligonucleotide (5′ CGG GAT CCC TGG TGT TTA TCC GTC ACT CG 3′) containing the BamHI restriction site (bold and italicized) and the common reverse primer for the other 6 constructs of SSAT.
The construct “5′UTR+Δ 4-30 SSAT” (data not shown) was created by ligating two PCR products using a restriction site for BglII that is present inside the open reading frame of SSAT at position 117 to 122. The first PCR product, “segment 1A,” was created with a forward primer (5′ CGG GAT CCC TGG TGT TTA TCC GTC ACT CG 3′) containing a BamHI restriction site (italicized) and a reverse primer (5′ TAG CAG ATC TTT TTC AGT TAA GAT TAC TTG TTC TTC CAT GTA TTC ATA TTT AGC CAG CTC CTT GAT CAG CCG CAG TAT GTC ACT GCA GTC GGC GGC CAT TTT CGT CTT TTG CTT TTC TT 3′). The second PCR product, “segment 2,” was created with a forward primer (5′ GAA AAA GAT CTG CTA GAA GAT GGT T 3′) and a reverse primer (5′ TCC CAC CGG TCT CCT CTG TTG CCA TTT TTA GC 3′) containing a restriction site for AgeI (bold and italicized). Segment 1A and segment 2 PCR products were cleaned up, digested with BglII, and ligated with T4 DNA ligase (Invitrogen). A third PCR was set up using this ligation product as the template, the forward primer of segment 1A, and the reverse primer of segment 2. This PCR product was digested with BamHI and AgeI and ligated in the expression plasmid.
The construct “5′UTR+Δ49-114 SSAT” was generated using the same technique (data not shown). A PCR product, “segment 1B,” was created with the same forward primer for segment 1A and the reverse primer (5′ TAG CAG ATC TTT GTC ACT GCA GTC GGC GGC 3′). Again, this product was cleaned up, digested with BglII, and ligated with segment 2. A third PCR was setup using the forward primer of segment 1A, the reverse primer of segment 2, and the ligation reaction between segment 1B and segment 2 as the template. This PCR product was digested with BamHI and AgeI and ligated in the expression plasmid.
Three additional variants of SSAT, with a 5′ UTR where the AUG codons of the two upstream open reading frames (uORFs) were eliminated by replacing the G with A, were created by using a forward primer (CGG GAT CCC TGG TGT TTA TCC GTC ACT CGC CGA GGT TCC TTG GGT CAT AGT GCC AGC CTG ACT GAG AAG AGG ACG CTC CCG GGA GAC GAA TAA GGA ACC ACC TCC TCC TAC TGT TCA AGT A 3′) containing BamHI (bold and italicized) and a reverse primer (5′ TCC CAC CGG TCT CCT CTG TTG CCA TTT TTA GC 3′) containing AgeI (bold and italicized). The previously described constructs “5′UTR+SSAT,” “5′UTR+Δ4-30 SSAT,” and “5′UTR+Δ49-114 SSAT” were used as templates to generate the PCR products“5′UTR(Δ uORFs)+SSAT,” “5′UTR(ΔuORFs)+Δ4-30 SSAT,” and “5′UTR(ΔuORFs)+Δ49-114 SSAT,” respectively.
Five recombinant fragments of nucleolin were expressed: N-term, R1234, R1234GAR, R12, and R34. The segments were amplified by PCR, digested with both BamHI and AgeI, and ligated into the expression plasmid. The oligonucleotides used to amplify each fragment are the following: “N-term,” forward (5′ CGG GAT CCg ccg cca cca tgG TGA AGC TCG CGA AGG 3′) and reverse (5′ TCC CAC CGG TAG TCG GTT CTG TGC CTT CCA 3′); “R1234GAR,” forward (5′ CGG GAT CCg ccg cca cca tgA CGG CTT TCA ATC TCT TTG TTG 3′) and reverse (5′ TCC CAC CGG TTT CAA ACT TCG TCT TCT TTC CTT G 3′); “R1234,” forward (5′ CGG GAT CCg ccg cca cca tgA CGG CTT TCA ATC TCT TTG TTG 3′) and reverse (5′ TCC CAC CGG TTT CAC CCT TAG GTT TGG CCC 3′); “R12,” forward (5′ CGG GAT CCg ccg cca cca tgA CGG CTT TCA ATC TCT TTG TTG 3′) and reverse (5′ TCC CAC CGG TTT GAC CTT TCT CTC CAG TAT AGT ACA G 3′); and “R34,” forward (5′ CGG GAT CCg ccg cca cca tgG AAT CAA AAA CTC TGG TTT TAA GC 3′) and reverse (5′ TCC CAC CGG TTT CAC CCT TAG GTT TGG CCC 3′). In all the forward primers, the restriction site for BamHI is bold and italicized and the Kozak sequence is in lowercase. In all the reverse primers, the restriction site for AgeI is also bold and italicized.
In addition, an enhanced green fluorescent protein (eGFP) construct and a mutant called Loop eGFP were cloned. The plasmid pLVTHM (Addgene.org clone 12247) was used as the template to amplify by PCR the eGFP gene. The PCR products were digested with BamHI and AgeI and cloned in the PLEX-MCS vector described above. The construct called “eGFP” was created with the following primers: forward, 5′ cgG GAT CCg ccg cca cca tgG TGA GCA AGG GCG AG 3′; reverse, 5′ TCC CAC CGG TTC GAG ATC TGA GTC CGG ACT T 3′. The construct called “eGFPloop” was created with the forward primer 5′ cgG GAT CCg ccg cca cca tgG CTA AAT TCG TGA TCC GCC CAG CCA CTG CCG CCG ACT GCA GTG ACA TAC TGC GGC TGA TCA AGG AGC TGG CTA TGG TGA GCA AGG GCG AG 3′ and the same reverse primer described for the construct called “eGFP.” In all the forward primers, the restriction site for BamHI is bold and italicized and the Kozak sequence is in lowercase. In the reverse primer the restriction site for AgeI is also bold and italicized.
The integrity of all the constructs was verified by sequencing. Lipofectamine 2000 (Invitrogen) was used for plasmid transfection, following the manufacturer's recommendations. A HEK293T stable cell line containing ORF SSAT was selected using puromycin, following the manufacturer's recommendations (Openbiosystems). In studies involving spermine supplementation, 1 mM aminoguanidine was added to the culture medium to inhibit amine oxidase activity and consequent toxicity due to the generation of reactive oxygen species. A HEK293T cell viability assay using Alamar blue (Invitrogen) in the presence of incremental concentrations of spermine was evaluated after 48 h of exposure.
Chimeric RNA synthesis.
In order to isolate SSAT RNA-interacting proteins, a chimeric RNA was designed and produced by in vitro transcription (IVT): a DNA template for IVT was created by linking the first 170 and the last 181 nucleotides of the open reading frame of SSAT with the RNA aptamer sequence specific for streptomycin binding (65). This aptamer contained an internal recognition site for the restriction enzyme BanI. A set of primers was designed to amplify the first 170 bp of the SSAT open reading frame (ORF) (section 1): forward (5′TAA TAC GAC TCA CTA TAG GGA TGG CTA AAT TCG TGA TCC G 3′) and reverse (5′ CCG TGG TGC CCT TGC GGG CAG AAG TCC AAA TGC GAT CCT TCG CAA CCA GGC AGT GGT AAA AG 3′). The forward primer contains the T7 promoter region (underlined), and the reverse primer contains part of the sequence of the RNA aptamer, including the BanI recognition site (bold and italicized). The last 181 bp of the ORF of SSAT were amplified using the following primers: forward (5′ CAA GGG CAC CAC GGT CGG ATC CTC TAA GCC AGG TTG CAA TGA G3′) and reverse (5′ TCA CTC CTC TGT TGC CAT TTT T 3′). The forward oligonucleotide contains the rest of the sequence of the streptomycin binding aptamer starting with the BanI sequence (bold and italicized); this amplicon is called section 2. Both section 1 and 2 were digested with BanI, purified, and ligated with T4 DNA ligase (Invitrogen). The ligated product was used as the template in a PCR using the forward primer used to create section 1 with the reverse primer used to create section 2. The product of this reaction was gel purified, cloned in the Pgem T vector (Promega), and verified by sequencing. For the in vitro transcription reaction, the Transcriptaid T7 high-yield transcription kit (Fermentas) was used as directed by the manufacturer. Using the forward primer for section 1 and the reverse primer for section 2, a PCR with the plasmid containing the chimera design as the template was performed. The PCR product was purified using a PCR cleanup kit (Promega), and 200 ng of the product was used per IVT reaction. This RNA molecule is referred to as “1-516.”
Six additional RNA truncated constructs of this molecule were generated using the same procedure with the following oligonucleotides: RNA “1-486,” forward (5′ TAA TAC GAC TCA CTA TAG GGA TGG CTA AAT TCG TGA TCC G 3′) and reverse (5′ CTC CTT GTC GAT CTT GAA CAG TC 3′); RNA “50-486,” Forward (5′ TAA TAC GAC TCA CTA TAG GGT ACT GCG GCT GAT CAA GGA G 3′) and the same reverse primer as that for RNA “1-486”; RNA “118-486,” forward (5′ TAA TAC GAC TCA CTA TAG GGG ATC TGC TAG AAG ATG GTT TTG G 3′) and the same reverse primer as that for RNA “1-486”; RNA “50-417,” forward (5′ TAA TAC GAC TCA CTA TAG GGT ACT GCG GCT GAT CAA GGA G 3′) and reverse (5′ GAA GTT GAT GGA TGG TTC ATT CC 3′); RNA “118-417,” forward (5′ TAA TAC GAC TCA CTA TAG GGG ATC TGC TAG AAG ATG GTT TTG G 3′) and the same reverse primer as that for RNA “50-417.” The T7 promoter sequence is underlined.
As a control, a green fluorescent protein (GFP)-aptamer construct was prepared using the following primers: forward (TAA TAC GAC TCA CTA TAG GGA TGG TGA GCA AGG GCG AG) and reverse (GGG GGG GGG GGG ATC CGA CCG TGG TGC CCT TGC GGG CAG AAG TCC AAA TGC GAT CCC CCC CCC CCC CGC CCT CGA ACT TCA CCT C). The T7 promoter sequence is underlined, and the streptomycin aptamer is included in the reverse primer. The PCR product obtained with eGFP as a template was used for in vitro RNA transcription as described above.
Cytoplasmic extracts.
Cytoplasmic fractions of HEK293T cells were obtained following cellular lysis in a hypotonic buffer (0.1× PBS, 0.001% Triton X-100, and protease inhibitor cocktail [Thermo Fisher]). Briefly, hypotonic lysis buffer (200 μl) was added to the HEK293T cell pellet. The resulting suspension was agitated by pipetting and was placed in ice for 15 min. Following incubation, the suspension was vortexed for 1 min and centrifuged at 16,000 × g for 30 min. The supernatant was collected, and the protein was quantified by the Bradford method (Bio-Rad). This cytoplasmic fraction was used as the source of protein for the RNA-protein interaction studies.
Identification of RNA-interacting proteins.
RNA-interacting proteins were isolated using a recently reported method (63). Briefly, 150 μg of the previously described chimeric molecule was obtained using a high-yield T7 in vitro transcription system, following the manufacturer's instructions (Fermentas). The chimera was purified using Megaclear (Ambion) and eluted with 100 μl of the provided elution buffer. The chimera was renatured in a thermocycler using the following cycle: 5 min at 56°C and 10 min at 37°C. A volume of 900 μl of column buffer (50 mM Tris HCl, pH 7.5, 5 mM MgCl2, and 250 mM NaCl) was added to the RNA chimera and kept on ice. A Sepharose-streptomycin purification column was prepared as previously described (63). To prevent nonspecific binding, the column was blocked with 20 μg yeast tRNA (Sigma) per 1 ml column buffer. The chimeric RNA was added to the column and was left to interact for 10 min. After the column was washed, 1 mg of a cytoplasmic protein fraction resuspended in 1 ml of the column buffer was added to the column. The lysate was left to interact with the RNA chimera for 10 min. The column was washed 8 times with 1 ml of column buffer, and the bound proteins were eluted with 2 ml of 10 μM streptomycin in column buffer. The eluents were concentrated to 50 μl using a Nanosep 3000 centrifugal device (Pall Corp) and separated with SDS-PAGE. The gel was stained with Sypro ruby (Invitrogen).
The isolated proteins were identified by gel electrophoresis-liquid chromatography-mass spectroscopy (gel-LCMS) technology using a Bruker HCTultra ion trap mass spectrometer as previously described (16). To identify the segment of SSAT mRNA recognized by nucleolin, 6 truncated RNA chimera molecules were prepared using the oligonucleotides described above. An RNA-protein interaction assay was performed using each of these truncated molecules as described above. The eluted proteins were concentrated, separated by SDS-PAGE, and transferred to a nitrocellulose membrane for Western blot analysis using antinucleolin monoclonal antibody (Santa Cruz Biotechnology).
siRNA knockdown experiments.
Six genes were targeted by small interfering RNA (siRNA) using one molecule per target of Silencer Select RNAi (Applied Biosystems). The following identification (ID) numbers were provided: ENO1 (enolase 1) (siRNA ID s-4682), SSB (siRNA ID s-13468), CSDA (DNA-binding protein A) (siRNA ID s224989), YBX1 (Y box protein 1) (siRNA ID s-9732), DXH9 (AtP-dependent RNA helicase A) (siRNA ID s-4019), and NCL (nucleolin) (siRNA ID s-9312). Silencer select (Applied Biosystems) nontargeting siRNA was used as a negative control. The transfection was performed using the manufacturer's instructions. Briefly, 600,000 HEK293T cells stably expressing the wild-type SSAT ORF were seeded in 6-well plates and transfected with Lipofectamine 2000 (Invitrogen) in Opti-MEM medium with a siRNA (10 nM). The cells were incubated for 60 h, and the cytoplasmic fraction was collected as described above. The proteins were quantified and analyzed by Western blotting.
Nucleolin-SSAT RNA interaction in situ.
The procedure described by Niranjanakumari et al. (39) was used to demonstrate protein-RNA interactions within cells. Briefly, cells were exposed to formaldehyde to cross-link proteins and nucleic acids and then sonicated to lyse and randomly fragment aggregates to allow subsequent immunoprecipitation. Monoclonal antibodies bound to protein G-coated magnetic beads (Dynabeads, catalog number 100-03D; Invitrogen, Carlsbad, CA) were used to immunoprecipitate selected proteins with their cross-linked molecules, including RNA. Cross-links were reversed with heat, and RNA was isolated with Trizol reagent (Invitrogen); DNA was removed with DNase I that had been inactivated prior to either one-step RT-PCR or PCR without the reverse transcriptase step. These oligonucleotides were used: forward (5′ GCT AAA TTC GTG ATC CGC C 3′) and reverse (5′ CTG CCG CCC TTC TCG AAC T 3′). The reverse oligonucleotide annealed on the purification tag present only in the recombinant SSAT transcript. The following cycle was used: 30 min at 60°C for reverse transcription, followed by 35 cycles of 94°C for 20 s, 60°C for 30 s, and 68°C for 60 s, and then 5 min at 68°C.
RNA-protein interaction with nucleolin fragments.
Five different fragments of nucleolin, “N-term,” “R1234GAR,” “R1234,” “R12,” and “R34,” were overexpressed by transient transfection as follows: The plasmids of the constructs were obtained using the high-yield plasmid Maxiprep system (Promega) and were transiently transfected into 5 × 106 HEK293T cells each using Lipofectamine 2000. The cells were allowed to express the recombinant proteins for 48 h and were then washed 3 times in cold PBS. A cytoplasmic fraction was obtained as described above and quantified by the Bradford method (Bio-Rad). To determine the nucleolin segment interacting with the SSAT mRNA, the following RNA-protein interaction assay was performed. Five columns of Sepharose-streptomycin were prepared as described above, and the RNA bait molecule “1-516” (150 μg) was allowed to interact with each of the columns for 10 min. After the column was washed, 1 mg of a cytoplasmic protein fraction from cells overexpressing each of the recombinant fragments of nucleolin was resuspended in 1 ml of the column buffer and was added to the column. The lysate was left to interact for 10 min with the RNA bait. The column was washed 8 times with 1 ml of column buffer, and the bound proteins were eluted with 2 ml of 10 μM streptomycin in column buffer. The eluents were concentrated to 50 μl using a Nanosep 3000 instrument (Pall Corp), separated with SDS-PAGE, and transferred to a nitrocellulose membrane for Western blotting using the anti-His-C-term-horseradish peroxidase (HRP) monoclonal antibody.
2DE and 2D Western blotting.
Cytoplasmic lysates (40 μg) from HEK293T cells and lysates from cells treated for 12 h with 2 mM spermine were subjected to two-dimensional (2D) gel electrophoresis (2DE) separation (pI, 3 to 6) as previously described (4, 5). In addition, a sample of RNA-interacting proteins obtained from the Sepharose-streptomycin-chimeric RNA column described above was subjected to the same 2DE separation. The sample was initially concentrated to 30 μl using a Nanosep 3000 instrument (Pall Corp) and dialyzed with 0.1× PBS for 12 h using a Slide-A-Lyzer 7K Mini unit (Pierce Biotech). After separation by 2DE, the proteins were transferred to a nitrocellulose membrane using a Mini Trans-Blot system (Bio-Rad) and were subjected to Western blotting using an antinucleolin monoclonal antibody.
Antibodies and chemicals.
For Western blots, the monoclonal antibody Anti-His(C-term)-HRP (Invitrogen) was used to detect all recombinant proteins. The following antibodies from Santa Cruz Biotechnologies were used for other proteins: nucleolin monoclonal (SC-8031), actin monoclonal (SC-47778), SSB (SC-80655), CSDA (SC-21318), YBX1 (SC-18057), and ENO1 (SC-15343). Spermine and aminoguanidine were from Sigma, and N1,N11-diethylnorspermine (DENSPM) was kindly provided by Carl Potter (Roswell Park Cancer Institute, Buffalo, NY).
RESULTS
Cell line overexpressing SSAT mRNA.
It is difficult to detect changes in SSAT expression because the amount of protein in cells is very low; therefore, we generated a stable HEK293T cell line that overexpresses the open reading frame of SSAT (HEK293T-SSAT), as shown in Fig. 1. The HEK293T-SSAT cells have markedly increased SSAT mRNA, but that does not produce a detectable increase in the SSAT protein without stimulation by the polyamine analogue N1,N13-diethylnorspermine (DENSPM), a well-characterized inducer of SSAT translation (8).
Fig 1.
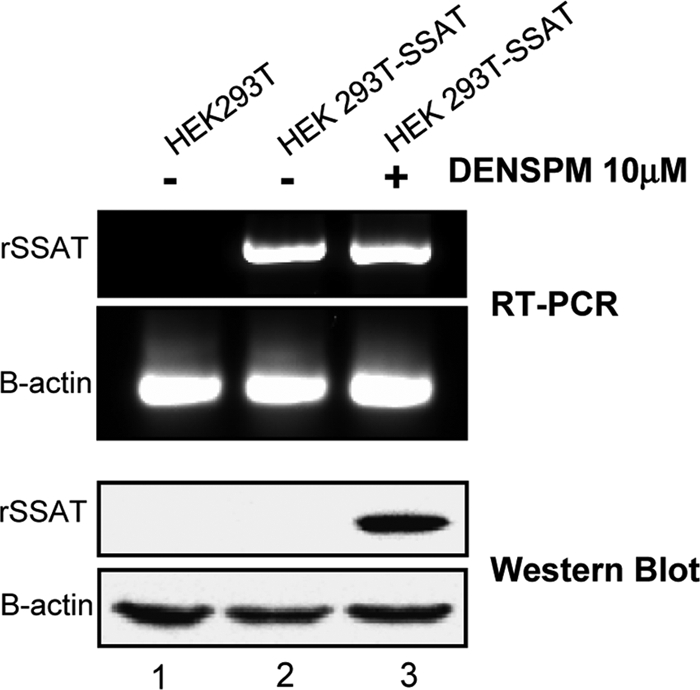
DENSPM-induced translation of recombinant His-tagged SSAT. The top panels show semiquantitative RT-PCR comparing SSAT RNA of control HEK293T cells and that of cells stably transfected with a plasmid containing the SSAT ORF with a His tag (HEK 293T-SSAT). The bottom panels show anti-His tag Western blots of the same cells. Actin served as the target and loading controls. Transfection alone markedly increased SSAT RNA, but the SSAT protein remained undetected. A 12-h exposure to 10 μM DENSPM, a polyamine analogue and potent inductor of SSAT translation, strongly induced the His-tagged SSAT protein with little or no further increase in SSAT RNA.
Proteins that bind to SSAT RNA.
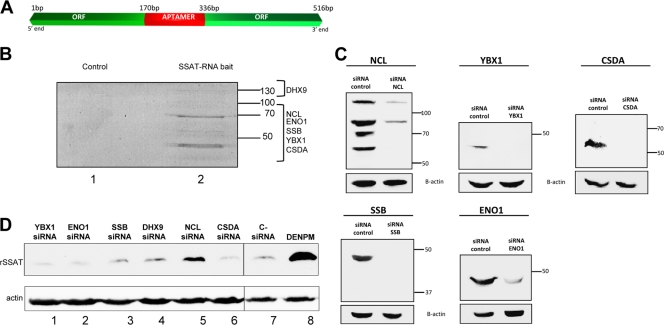
Because prior publications suggested that an unidentified protein represses translation by interacting with the coding region of the SSAT transcript (8), we sought repressor candidates based on the ability to bind to SSAT mRNA. Based on previous data indicating involvement of the 5′ coding region (8), we created a chimeric RNA bait containing the first 170 bp and last 181 bp of the human SSAT open reading frame linked by a streptomycin-binding RNA aptamer (63) (Fig. 2A). The aptamer allowed the bait to bind to a Sepharose-streptomycin column which we used to capture proteins that bind to SSAT RNA. HEK293T cell lysate was incubated in the column, unbound proteins were washed from the column, and bound proteins were eluted. Analysis by gel electrophoresis-liquid chromatography-mass spectroscopy (Gel-LCMS) identified six proteins in the eluate (Fig. 2B and Table 1), all of which had previously been described as having RNA-binding properties. To test for nonspecific binding, a control column with aptamer-linked GFP RNA was constructed; no proteins were found in the control eluate (Fig. 2B).
Fig 2.
SSAT RNA-interacting proteins and knockdown effects. (A) Chimeric RNA. Bait RNA was created using the first 170 bases of the 5′ end of the SSAT ORF and the last 181 bases of the 3′ end linked by a streptomycin-binding aptamer (red) that allowed the bait to bind to a Sepharose-streptomycin column. (B) Proteins interacting with SSAT RNA. HEK293T cell cytoplasmic proteins were incubated in the column, unbound proteins were washed away, and bound proteins were eluted with streptomycin-containing buffer. Eluted proteins were separated by SDS-PAGE and stained with Sypro ruby. No proteins were detected in the control column eluate, but multiple proteins were eluted from the SSAT-RNA column and then identified by mass spectroscopy (Table 1). Proteins and approximate gel positions are labeled: ENO1 (enolase 1), YBX1 (Y box protein 1), CSDA (DNA-binding protein A), SSB (La protein), DHX9 (ATP-dependent RNA helicase A), and NCL (nucleolin). (C) siRNA knockdowns. Western blots demonstrate individual siRNA knockdown of RNA-interacting proteins. (D) Knockdown effects on SSAT translation. Expression of recombinant SSAT was assayed by anti-His tag Western blotting with actin as the loading control. Lanes 1 to 6 are from cells treated with the indicated siRNA. Lane 7 is a negative control from cells treated with nontargeting siRNA. Lane 8 is a positive control with proteins from cells exposed to DENSPM to induce SSAT translation. Only nucleolin knockdown significantly increased expression of recombinant SSAT (lane 5).
Table 1.
SSAT RNA binding proteins identified by Gel-LCMSa
| UniProt accession no. | Description (gene name) | Theoretical mass (Da) | Identified peptide(s) | Protein score |
|---|---|---|---|---|
| Q08211 | ATP-dependent RNA helicase A (DHX9) | 140,869 | AAECNIVVTQPR | 135 |
| YQILPLHSQIPR | ||||
| LQISHEAAACITGLR | ||||
| ELDALDANDELTPLGR | ||||
| AENNSEVGASGYGVPGPTWDR | ||||
| LSMSQLNEK | ||||
| DVVQAYPEVR | ||||
| SSVNCPFSSQDMK | ||||
| QPAIISQLDPVNER | ||||
| P67809 | Nuclease-sensitive element binding protein 1 (YBX1) | 35,903 | GAEAANVTGPGGVPVQGSK | 248 |
| SVGDGETVEFDVVEGEK | ||||
| AADPPAENSSAPEAEQGGAE | ||||
| EDGNEEDKENQGDETQGQQPPQR | ||||
| P19338 | Nucleolin (NCL) | 76,568 | NDLAVVDVR | 231 |
| EVFEDAAEIR | ||||
| GLSEDTTEETLK | ||||
| EAMEDGEIDGNK | ||||
| TGISDVFAK | ||||
| SISLYYTGEK | ||||
| P05455 | Lupus La protein (SSB) | 46,808 | QKLEEDAEMK | 170 |
| FSGDLDDQTCR | ||||
| IIEDQQESLNK | ||||
| LTTDFNVIVEALSK | ||||
| P16989 | DNA binding protein A (CSDA) | 40,066 | AGEAPTENPAPPTQQSSAE | 49 |
| GAEAANVTGPDGVPVEGSR | ||||
| P06733 | Alpha enolase (ENO1) | 47,139 | VNQIGSVTESLQACK | 110 |
Mass spectrum processing was performed using the Mascot Distiller software program (version 2.3.0.0) with search and quantitation toolbox options. The generated deisotoped peak list was submitted to an in-house Mascot 2.2 server for searching against the Swiss-Prot database (version 56.6). Mascot search parameters were set as follows: species, Homo sapiens; enzyme, trypsin with maximal 1 missed cleavage; fixed modification, cysteine carbamidomethylation; variable modification, methionine oxidation; 0.50-Da mass tolerance for precursor peptide ions; and 0.6 Da for MS/MS fragment ions. All peptides matches were filtered using an ion score cutoff of 30. The following two criteria were used to evaluate protein identification: one peptide with ion score of ≥50, two or more peptides with at least one ion score of ≥32 (P < 0.05 threshold) and cumulative Mascot scores of ≥50; for all the proteins with cumulative MOWSE scores of ≥50 and ≤80, the theoretical and experimental gel molecular weights had to be consistent. When these criteria were used to search against a reversed decoy Swiss-Prot database, there was no false-positive match (false discovery < 0.5%).
Nucleolin knockdown and increased SSAT translation.
To assess which of the SSAT mRNA-binding proteins might be responsible for repression of SSAT translation, we knocked them down individually in HEK293T-SSAT cells using RNA interference (siRNA). Knockdown efficacy was demonstrated by Western blotting (Fig. 2C) for all but DHX9 (antibodies available for DHX9 did not produce useful blots). Nucleolin knockdown enhanced SSAT expression (Fig. 2D), suggesting this as the elusive SSAT translational repressor.
Interaction of nucleolin and SSAT RNA in situ.
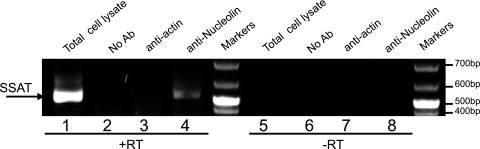
To demonstrate that nucleolin-SSAT RNA interaction is not an in vitro artifact, we used an established method to demonstrate association in vivo (39). The method involves treating intact cells with the cross-linking agent formaldehyde, followed by immunoprecipitation to isolate selected proteins and the macromolecules associated with them. After cross-linking and cell lysis, nucleolin was immunoprecipitated and the presence of SSAT RNA in the precipitate was detected by RT-PCR (Fig. 3). Lanes 1 to 4 contain products of SSAT RT-PCRs; lanes 5 to 8 contain products of control reactions that omitted reverse transcriptase. The arrow shows the position of the SSAT cDNA product when total cell lysate is used as the target. The band in lane 4 shows the presence of SSAT RNA in the antinucleolin immunoprecipitate; lane 8 shows that this product is dependent on RNA, since no cDNA was produced when reverse transcriptase was omitted. Controls in lanes 2 and 6 show that SSAT RNA is not present when protein G-coated beads do not contain bound antibody. Controls in lanes 3 and 7 show that SSAT RNA is not present when the beads bind antibody against the highly prevalent protein actin, thus indicating that SSAT-RNA is not trapped by nonspecific precipitation. Sequencing the PCR product in lane 4 confirmed the in vivo association of nucleolin and SSAT RNA (data not shown).
Fig 3.
Nucleolin-SSAT RNA interaction in situ. Protein-RNA coprecipitation was detected using RT-PCR. Lanes 1 to 4 are products of RT-PCRs. Lanes 5 to 8 are negative controls omitting reverse transcriptase. Lanes 1 and 5 are positive controls with the PCR target being whole-cell lysate. Lanes 2 and 6 are negative controls with the PCR target being sham immunoprecipitate from protein G-coated beads without bound antibody (No Ab). Lanes 3 and 7 are negative controls with the PCR target being immunoprecipitate using antiactin monoclonal antibody. Lanes 4 and 8 used immunoprecipitate from beads with antinucleolin monoclonal antibody as the PCR target. The data show SSAT RNA coprecipitates with nucleolin.
Nucleolin regions necessary for SSAT RNA binding.
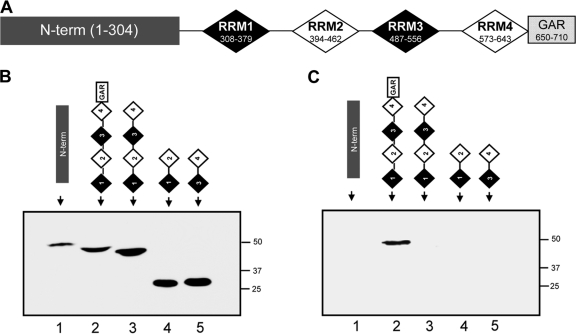
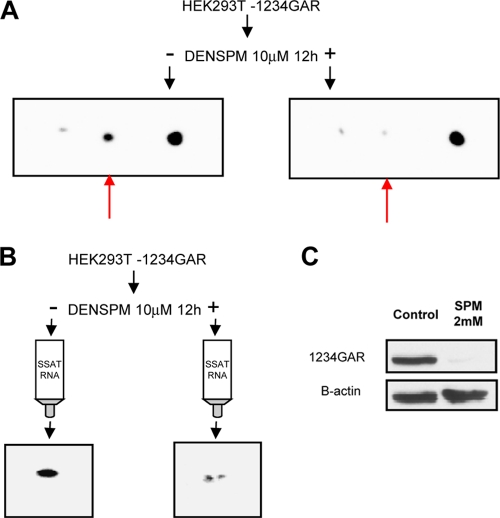
Nucleolin contains 6 domains (Fig. 4A): an N-terminal (N-term) domain, 4 “RNA recognition motif” domains (RRM1, RRM2, RRM3, and RRM4), and a C-terminal glycine/arginine-rich (GAR) domain (60). To determine which are required for binding to SSAT mRNA, HEK293T cells were transiently transfected to produce recombinant nucleolin fragments. For efficiency and to increase binding stringency, lysates rather than isolated protein fragments were incubated with individual chimeric SSAT RNA columns that were processed as described above (Fig. 4B). The composite Western blot of eluted proteins shown in Fig. 4C reveals that the GAR domain is necessary for binding. Because we were not successful in creating recombinant fragments that included the GAR region unless RRM1, RRM2, RRM3, and RRM4 were also included, subsequent studies were conducted with peptides containing RRM1, RRM2, RRM3, RRM4, and GAR (1234GAR). The ability of DENSPM, a polyamine analogue, to increase SSAT translation strongly is well known (8, 40), so we tested whether DENSPM would affect the His-tagged 1234GAR segment. Two-dimensional Western blot analysis of HEK293T cells overexpressing His-tagged 1234GAR showed that the segment exists as three different isoforms, and exposure to DENSPM resulted in degradation of one (Fig. 5A, red arrow). DENSPM exposure also reduced the amount of recombinant segment that binds to the SSAT-RNA column (Fig. 5B). Spermine also caused degradation of the 1234GAR segment (Fig. 5C). Together these suggest that the SSAT-RNA binding site and the autocatalytic site both lie within the 1234GAR region.
Fig 4.
Glycine/arginine-rich nucleolin domain is required for SSAT mRNA binding. (A) Nucleolin schematic with amino acid numbers: N-term, N-terminal region; RRM1 to RRM4, “RNA recognition motif” domains 1 to 4; GAR, glycine/arginine-rich domain. (B) Western blots using an anti-His tag monoclonal antibody confirmed expression of nucleolin fragments by cells transiently transfected with plasmids containing the indicated constructs (native nucleolin or fragments not detected). (C) Anti-His tag Western blots of SSAT-RNA column eluates from transiently transfected cell lysates show that the binding does not occur without the GAR domain.
Fig 5.
DENSPM and spermine effects on recombinant nucleolin peptide 1234GAR. HEK293T cells were transiently transfected to overexpress His-tagged 1234GAR (HEK293T-1234GAR); anti-His antibody was used to detect the peptide. (A) 2D blots were prepared as for Fig. 2: control HEK293T NCL-1234GAR cells are on the left, and cells exposed to 10 mM DENSPM for 12 h are on the right. Red arrows indicate a peptide isoform present in controls but absent in cells treated with DENSPM. (B) 2D blots of peptides that bound to SSAT RNA columns; peptides from controls are on the left, and those from cells exposed to 10 mM DENSPM for 12 h are on the right. 1234GAR peptides bind to the column and are markedly reduced in cells exposed to DENSPM. (C) One-dimensional (1D) Western blots show 1234GAR peptide autocatalysis in cells exposed to 2 mM spermine for 12 h.
Polyamine-induced nucleolin degradation.
Polyamines had previously been reported to accelerate nucleolin degradation (57, 58) by autocatalysis (12, 18); therefore, we considered the possibility of a negative feedback control system in which excess polyamines cause nucleolin degradation and thus increased SSAT, which leads to polyamine reduction. However, polyamine-induced degradation had been demonstrated only in vitro using partially purified nucleolin, and therefore a necessary test of this hypothesis was determining whether this occurs within cells. We exposed cells to 2 mM spermine for 12 or 48 h, a concentration chosen to create a reasonable gradient across the cell membrane and thus increase intracellular concentrations. Intracellular spermine in control HEK293T cells was 612 ± 32 μM, and that in cells exposed to 2 mM spermine for 12 h was 792 ± 49 μM; both of these concentrations are within range of reported normal values, and exposure for 48 h was not toxic (Fig. 6F).
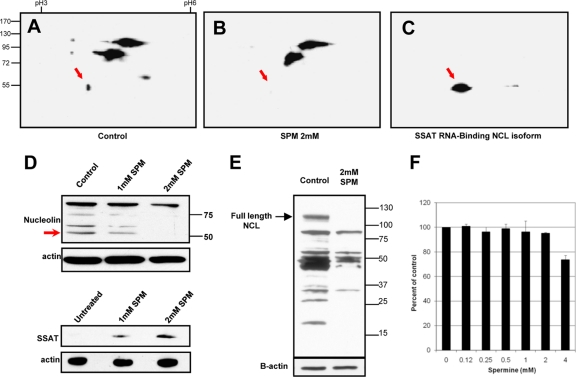
Fig 6.
Exogenous spermine reduces 55-kDa nucleolin isoform. 2D Western blots in panels A, B, and C used IPG strips (pI 3 to 6) for the first separation, SDS-PAGE for the second, and monoclonal antinucleolin antibody for the blots. Red arrows point to the position of the 55-kDa isoform that binds to the SSAT-RNA column and is missing or reduced in cells exposed to spermine. (A) HEK293T cells grown under control conditions. (B) HEK293T cells exposed to 2 mM spermine for 12 h. (C) Proteins eluted from a SSAT-RNA interaction column (high intensity of the 55-kDa spot results from column enrichment). (D) Western blots of cells exposed to 0, 1, and 2 mM spermine for 12 h. Upper and lower blots are from the same 3 cultures: upper panels show dose-dependent spermine-induced degradation of the 55-kDa nucleolin isoform, and lower panels show corresponding spermine-induced increases in the SSAT protein. (E) Polyamine-mediated nucleolin degradation by 48 h of spermine exposure. (F) An Alamar blue assay was used to test viability of HEK293T cells grown for 48 h in the indicated spermine concentrations. Aminoguanidine (1 mM) was included in all cultures to inhibit generation of reactive oxygen species by amine oxidase activity of serum in the media.
Nucleolin is known to exist in cells as multiple isoforms due to posttranslational modification and/or degradation. Western blots in Fig. 6 using a monoclonal antinucleolin antibody demonstrate the presence of multiple isoforms; bands at 105 kDa are considered intact protein (18). Red arrows indicate the position of an isoform present in control cells (Fig. 6A) but absent in cells exposed to spermine (Fig. 6B and D). This same isoform is eluted from a SSAT RNA-binding column (Fig. 6C). Blots in the upper 2 panels of Fig. 6D show dose-dependent spermine-induced degradation of this isoform (red arrow), as well as an overall reduction of material reacting with the monoclonal antibody. The lower 2 panels of Fig. 6D show dose-dependent increases in the SSAT protein, which otherwise is not detectable. Longer exposure to 2 mM spermine results in a loss of full-length nucleolin, as well as lighter isoforms, suggesting the possibility of dynamic isoform conversion.
Characterization of nucleolin isoform that represses SSAT translation.
To determine the regions of the 55-KDa nucleolin isoform that interact with SSAT RNA, ion trap mass spectroscopy analysis of the proteins eluted from the SSAT RNA column was performed. The analysis produced partial sequences for four nucleolin peptides, all of which are in the C-terminal half of nucleolin and span the four RRM domains (Fig. 7). Thus, the repressor contains not only the GAR region of nucleolin but also the RNA-binding regions.
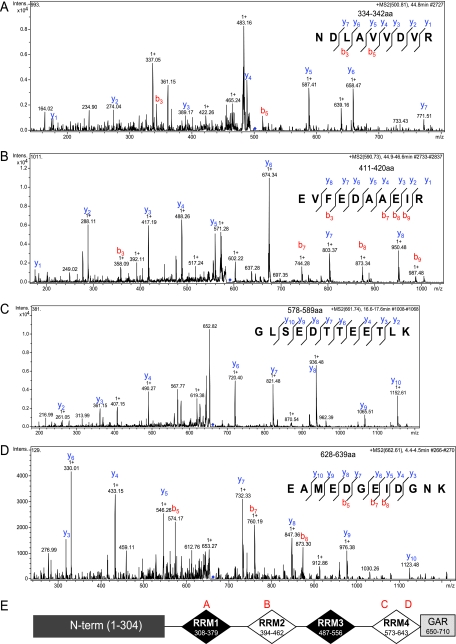
Fig 7.
Characterization of 55-kDa nucleolin isoform by mass spectroscopy. The nucleolin isoform interacting with SSAT RNA was separated by 2DE, trypsin digested, and analyzed by mass spectroscopy. The tandem mass spectrometry (MS/MS) profile of each the four identified peptides (panels A to D) and their representative sequence is shown on the right of each panel. The location of each peptide is shown in panel E.
Nucleolin binds to the 5′ end of the coding region of SSAT mRNA.
To identify the nucleolin binding region of SSAT mRNA, we created six additional Sepharose-streptomycin-chimeric SSAT RNA columns with truncated constructs of the chimeric RNA used for the column that bound nucleolin originally (Fig. 8A). HEK293T cell lysate was allowed to bind to these 6 columns and to a column with the original chimeric construct. Antinucleolin Western blots show that the entire chimera columns bound nucleolin, except those lacking nucleotides 50 to 118 (Fig. 8B, lanes 7 and 9). Thus, nucleolin binding depends on nucleotides within the 50–118 region.
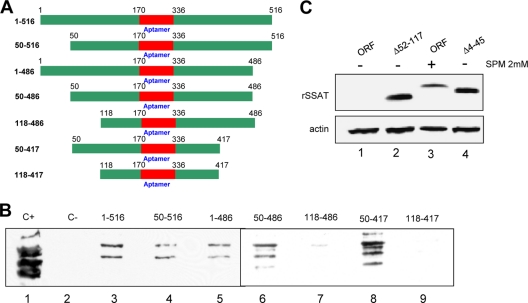
Fig 8.
SSAT RNA bases essential for nucleolin binding and translation repression. (A) Chimeric RNA-bait molecules. Top schematic, labeled 1-516, is the original chimera used to isolate nucleolin RNA binding proteins. Graphics and labels of others indicate nucleotides omitted in other bait molecules. (B) Nucleotides essential for nucleolin binding. Individual columns were prepared using the seven bait molecules plus a control with nonspecific RNA. HEK293T cell cytoplasmic lysate was incubated with the columns, and bound proteins were eluted and then concentrated for Western blot analysis using an antinucleolin monoclonal antibody. “C+” is a positive control using whole HEK293T lysate. “C-” is a negative control from the nonspecific RNA column. Lanes 3 to 9 are from columns prepared with the indicated bait molecules. All the columns bound nucleolin except those with baits lacking nucleotides 1 to 117. Since the 50–486 and 50–417 columns did bind nucleolin, binding depends on nucleotides within the 50-to-118 range. (C) Nucleotides essential for SSAT translational repression. HEK293T cells were transiently transfected with plasmids containing the complete SSAT ORF (ORF) and mutants using constructs lacking ORF nucleotides 52 to 117 (Δ52-117) or 4 to 45 (Δ4-45). Anti-His antibody was used for the blots, with antiactin for the loading control. Lanes 1 and 3 show that translation of the full SSAT ORF occurs only when the cells are exposed to spermine. Lanes 2 and 4 show that the Δ52-117 and Δ4-45 mutants are translated without spermine and thus both regions are required for translational control even if nucleolin binds to only one.
Earlier work showed that elimination of nucleotides 4 to 45 released translational repression, suggesting the presence of a control element in this region (8). Our data show nucleolin is involved in repression and binds to SSAT RNA, and the binding site is within nucleotides 50 to 118 of the SSAT coding sequence, suggesting the control element lies within this region. We considered explanations for this conflict: an error in the earlier work, existence of multiple control elements, and a single element incorporating both the 4–45 and 50–118 regions. To test these possibilities, we transiently transfected HEK293T cells using plasmids containing the complete SSAT ORF, a reconstructed Δ4-45 mutant, and a Δ52-117 mutant. Figure 8C shows that both the Δ4–45 and the Δ52–117 mutations eliminated translational repression, a finding explained in the next section.
SSAT RNA 5′ stem-loop.
Mfold analysis of the RNA secondary structure (65) revealed a 75-base-pair stem-loop at the extreme 5′ end of the ORF (nucleotides 2 to 76) with a free energy of −30.8 kcal/mol (Fig. 9A). This is not an artifact created by analysis of the ORF without the UTRs, since the same loop is predicted when the full transcript sequence is used. A stem-loop at this position could act as a translational repressor, a possibility supported by secondary structure analysis showing that both the Δ4-45 and Δ52-117 mutations impair stem-loop formation and thus both regions could be parts of the same control element.
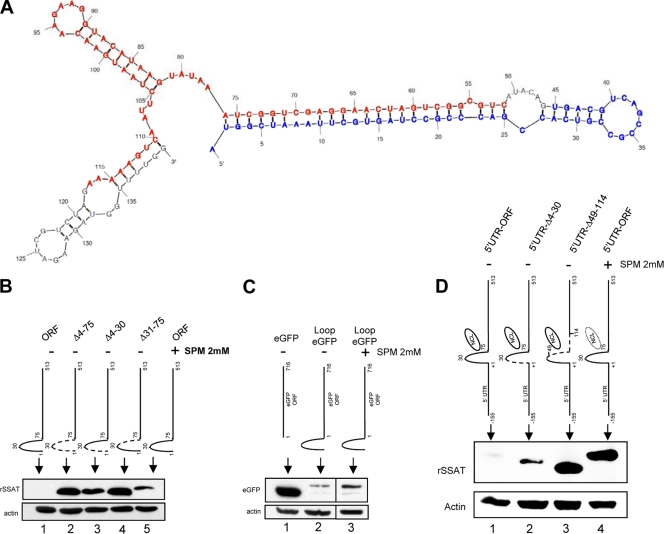
Fig 9.
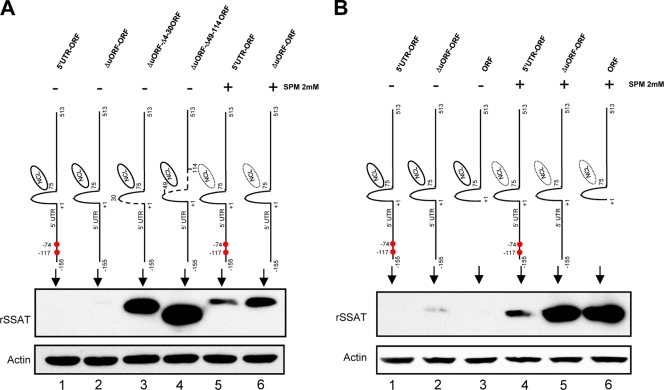
SSAT RNA stem-loop and translational repression. (A) Mfold-calculated RNA secondary structure of first 140 bp of the SSAT ORF. Bases deleted in the Δ52-117 construct are marked in red; blue indicates the Δ4-45 construct. (B) Transiently transfected HEK293T cells. Transfecting plasmids contained the full SSAT ORF (ORF) or mutants with deletions that impair formation of a stem-loop by bases 2 to 76. The Δ4-75 mutant lacks the entire stem-loop, Δ4-30 lacks the first half, and Δ31-75 lacks the second half. Anti-His Western blots show that protein expression by the wild type is dependent on spermine exposure (lanes 1 and 5), but all mutants express protein without spermine (lanes 2, 3, and 4). (C) Fusion of the SSAT ORF stem-loop with a reporter. HEK293T cells were transiently transfected with plasmids producing RNA for the His-tagged reporter protein eGFP or RNA with the SSAT 1–75 stem-loop appended in frame to the 5′ end of eGFP (Loop eGFP). Anti-His Western blots show that Loop eGFP expression is repressed (lane 2) compared to that of eGFP (lane 1), and repression is partially relieved by exposure to 2 mM spermine (lane 3). (D) 5′ UTR and SSAT ORF stem-loop repression. To determine whether the 5′ORF stem-loop represses translation when the full 5′ UTR is present, HEK293T cells were transiently transfected with plasmids containing the 5′ UTR of SSAT RNA and the full coding region (5′UTR-ORF), the 5′ UTR with Δ4-30 ORF (5′UTR-Δ4-30), and the 5′UTR with Δ49-114 ORF (5′UTR-Δ49-114). Expression of 5′UTR-ORF required spermine exposure (lanes 1 and 4) while the mutants did not. Expression of 5′UTR-Δ49-114 (lane 3), which doesn't form the stem-loop and lacks the nucleolin binding region, was greater than that of 5′UTR-Δ4-30 (lane 2), which also doesn't form the stem-loop but does have the nucleolin binding region; the same relative relationship can be seen in panel B, although it is not as marked.
To test this further, we constructed three additional mutants calculated to be deficient in stem-loop formation: Δ4-75, Δ4-30, and Δ31-75. Cells were transiently transfected, and expression was compared to those of cells transfected with the wild type ORF. All 3 mutants were more strongly expressed than the wild type ORF, even when the wild type was treated with spermine (Fig. 9B). We then asked whether this loop could repress translation of another protein. The first 75 bp of the SSAT ORF, which contain the stem-loop, were spliced onto the 5′ end of a reporter gene ORF (eGFP). The stem-loop did repress GFP expression, an effect partially reversed by exogenous spermine (Fig. 9C).
Ordinarily, the ability of an mRNA stem-loop to repress translation relates to 5′ cap proximity; loops more than 50 bp downstream of the cap generally do not affect translation (32). Because the SSAT mRNA of humans contains a long 5′ UTR, placing this stem-loop 155 bp downstream of the 5′ cap, it was necessary to determine whether the stem-loop has an effect within the context of the full-length SSAT mRNA. To do this, we created two mutants containing the complete 5′ UTR but with deletions that interfere with the 5′ ORF stem-loop: “5′ UTR+Δ4-30 SSAT” lacks the first half of the loop but can bind nucleolin, while “5′ UTR+Δ49-114 SSAT” lacks both the stem-loop and the nucleolin binding region. Expression of both mutants was greater than that of the wild type, but expression was greatest in the mutant unable to form the stem-loop and also unable to bind nucleolin (Fig. 9D). These results indicate the 5′ ORF stem-loop does affect translation despite being far downstream of the 5′ cap. Although the greatest nucleolin effect is likely loop stabilization, binding does affect SSAT translation even if the loop cannot form.
Translational control by uORFs.
SSAT contains 2 upstream ORFs (uORFs) (27), features reported capable of modulating translation (9). One begins at −117 and codes for 4 amino acids (aa). The other begins at −74, overlaps the main ORF, and codes for 29 aa (27). The ribosomal scanning model of translation initiation holds that translation starts once the 43S ribosomal subunit finds the first AUG codon, where it is joined by the 60S unit to form a functional ribosome. The presence of uORFs in certain transcripts seems to delude the ribosome machinery so that translation starts upstream of the actual initiation codon, dramatically reducing translation of the intended protein (29, 33, 51). To test the prediction that these uORFs resist translation, we constructed mutants similar to the ones described in Fig. 9D, but with the uORF initiation codons eliminated by replacing codon G's with A's. The result of uORF elimination was increased expression, as predicted (Fig. 10A). Several points indicate that repression by the uORFs and the nucleolin-binding stem-loop are independent systems: the mutant lacking both the uORFs and the nucleolin binding region of the stem-loop had the highest level of expression (Fig. 10A); mutants lacking the uORFs but having the stem-loop expressed SSAT at a low level even in the absence of spermine (Fig. 10B), and spermine induced expression was increased by elimination of either the 5′ UTR or the uORFs within the 5′ UTR (Fig. 10B). Collectively, these results indicate that the uORFs act as spermine-independent barriers for ribosome read-through that could be constitutive or could respond to as yet unknown factors.
Fig 10.
SSAT expression is elevated in mutants lacking uORFs. Red dots show locations of uORF starting codons. HEK293T cells were transiently transfected with plasmids having SSAT RNA 5′uORF plus the full coding region (5′UTR-ORF), mutations eliminating uORF starting codons plus the full coding region (ΔuORF-ORF), mutations eliminating the uORFs plus coding region deletions preventing 5′ORF stem-loop formation (ΔuORF-Δ4-3 ORF), or mutations eliminating the uORFs plus coding region deletions preventing 5′ ORF stem-loop formation and nucleolin binding (ΔuORF-Δ49-114ORF). (A) uORFs affect translation. Lanes 1 and 2 show that elimination of uORFs does not allow SSAT expression without spermine, but lanes 5 and 6 show that uORF elimination enhances spermine-induced expression. Lanes 3 and 4 show that elimination of both the uORFs and stem-loop allows greater expression, and further elimination of the nucleolin binding site enhances expression further. (B) uORF translational control is not affected by spermine. SSAT expression was compared among three constructs that require spermine for translation: 5′UTR-ORF, ΔuORF-ORF, and ORF. Since spermine-induced SSAT translation of ORF was markedly less than that for 5′UTR ORF but the difference was eliminated when the start codons of the uORFs were eliminated, that indicates that the translation repression by uORFs is independent of the presence of spermine. The slight translation leakage of ΔuORF-ORF without spermine supports this interpretation.
DISCUSSION
Previous results showing accumulation of SSAT mRNA without a significant increase in protein, as well as increases in protein without corresponding increases in mRNA, suggested control at the translation level (40). Later data supported this with indications that the 5′ end of the mRNA ORF and a repressor protein were involved, but neither the protein nor the mechanism of repression was identified (8). Here, we report the repressor protein to be an isoform of nucleolin which acts by binding to a 5′ stem-loop of SSAT mRNA and blocks translation.
Using column-immobilized RNA, we isolated 6 proteins that bind to SSAT RNA; nucleolin was one of the 6 (Fig. 2). Nucleolin knockdown by siRNA caused a strong increase in SSAT translation, indicating a repressor function (Fig. 2). Knockdown of SSB (the La protein) and DHX9 (ATP-dependent RNA helicase A) had lesser effects, but we have not yet pursued those. Association of nucleolin with SSAT RNA within cells was demonstrated by coprecipitation after cross-linking (Fig. 3).
Using 2D Western blots, we found SSAT RNA binding to be restricted to an isoform of nucleolin. The data show that the same isoform degrades quickly in cells exposed to exogenous spermine, a treatment known to induce SSAT (Fig. 6). This is consistent with earlier reports that spermine accelerates autocatalysis of semipurified nucleolin in vitro (57, 58), but the mechanism involved is unknown. Tests of recombinant nucleolin fragments show the C-terminal GAR region of nucleolin to be necessary for binding (Fig. 4). Those data and mass spectroscopy analysis of the isoform isolated from cells (Fig. 7) indicate that the isoform encompasses all 4 RNA recognition motif (RRM) domains and the C-terminal GAR region, a domain also known to have nucleic acid binding properties (60). For cells expressing a recombinant fragment of nucleolin containing the 4 RRM domains and GAR region, there was a marked loss of the fragment upon treatment with the polyamine analogue DENSPM, a treatment known to induce SSAT (Fig. 5). We hypothesize that this nucleolin isoform serves as the sensor in a negative feedback control system operating as follows: when polyamines are low, the isoform is present and suppresses SSAT translation; when polyamines rise, the isoform degrades and thus SSAT translation is derepressed, SSAT activity increases, and polyamines decrease. Figure 11 is a graphical representation of our hypothesis.
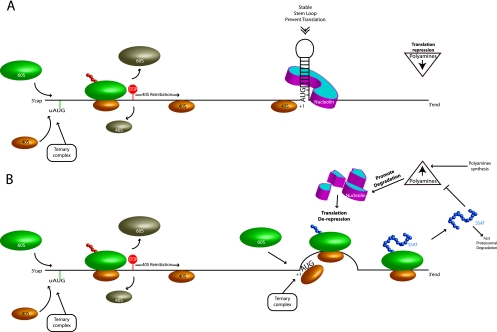
Fig 11.
Model of the SSAT translational control mechanism. (A) SSAT translational repression in the presence of low levels of polyamines. Success of ribosome scanning to the correct initiation codon is reduced in the presence of uORFs. Translation therefore depends on either leaky scanning or reinitiation of ribosome scanning 3′ to the uORFs. Those ribosome subunits escaping uORF control are then blocked by the strong stem-loop at the initiation site, which is stabilized by nucleolin. (B) Polyamine-regulated translational derepression of SSAT. An isoform of nucleolin blocks translation by binding to and stabilizing the stem-loop at the extreme 5′ end of the ORF. Increased polyamines cause autocatalysis of this nucleolin isoform, thereby reducing the stability of the stem-loop and allowing ribosomal subunits that escaped the uORF control to initiate translation. This increase in expression acetylates the polyamines and lowers their concentration, which reduces nucleolin degradation and thereby reestablishes translation repression. Together, these provide a negative feedback mechanism to regulate cellular polyamine concentrations.
Nucleolin is an abundant protein present in both the nucleus and cytoplasm, as well as the cell surface (7). Among the many known nucleolin functions are roles in ribosome biogenesis (13), stabilization of cytoplasmic RNA (64), assistance of pre-mRNA maturation (26), formation of cell surface receptors (54), and repression of p53 translation, possibly by promoting formation of a 5′ UTR stem-loop (23, 59). Nucleolin is known to exist as the complete 710-aa polypeptide and as isoforms that differ according to phosphorylation, glycosylation, and protein cleavage (12, 37). Although the existence of isoforms is well known, to our knowledge this is the first description of a function being isoform specific; it may be that such specificity contributes to control of other nucleolin functions.
Bioinformatic and mutation analyses together provided information about the region of SSAT RNA where nucleolin binds. Our computation results agreed with a previously reported analysis (8, 40) in predicting a high-energy stem-loop at the extreme 5′ end of the ORF, composed of bases 1 to 76 (Fig. 9). We find the same loop predicted regardless of whether the SSAT mRNA ORF is analyzed alone or within the context of the complete message. Mutation analysis showed the site of nucleolin binding to be within nucleotides 50 to 118, which includes 26 of the 76 stem-loop bases (Fig. 8). A role for the stem-loop in translation repression had been suggested (8, 40), and data here confirm that, since mutations interfering with stem-loop formation reduce translation repression (Fig. 8 and 9). Our data also show deletion of the nucleolin binding site independently reduces repression (Fig. 8 and 9). Further evidence for the function of this control element was obtained by splicing the stem-loop onto the 5′ end of a reporter gene; the loop conferred translation repression that was partially relieved by exogenous polyamines (Fig. 9). Data here do not address the previously predicted roles of a second stem-loop at the 5′ end and one at the 3′ end (8, 40).
This system of SSAT translational control has many unusual features. mRNA secondary structure is commonly involved, but the ability of a stem-loop to repress translation is usually lost if it is located more than 50 bases downstream of the 5′ UTR cap; this SSAT mRNA stem-loop element is 155 bases downstream. Repression by UTR elements is common; but while not unknown, ORF involvement is rare. One example of ORF involvement is human thymidylate synthase and dihydrofolate reductase, where translation is controlled by enzyme end products that interact with the mRNA coding region (14, 17). Control of human protein expression by mRNA secondary structure is common, but involvement of either the initiation or termination codons is not (53); the SSAT mRNA stem-loop element incorporates 2 of the 3 initiation codon bases. A somewhat analogous situation does exist in yeast, where some transcripts have a secondary structure involving the initiation codon, and ribosome profiling data suggest they reduce translational efficiency (31).
It had been suggested the 5′ UTR of SSAT mRNA plays no role in translational repression (8), but a recent publication identified uORFs in SSAT RNA and suggested these do play a regulatory role by responding to polyamine concentrations, although no supporting experimental data were provided (27). These are the same uORFs that we found do suppress translation, but spermine had no effect. Polyamine-responsive uORF elements have been found for other polyamine metabolic enzymes (24, 28), and our data do not exclude the possibility of responses to other polyamines or polyamine-related compounds. Alternatively, these uORFs could act as constitutive repressors.
The work here is focused on control of SSAT at the translation level, but other influences are known. Transcription control is less dramatic but can be influential, and translation is known to respond to factors other than polyamines, such as oxidative stress, heat shock, hypoxia, cytokines, growth factors, etc. (2, 10, 62). In addition, nucleolin may not be the only protein that affects SSAT translation. RNA helicase A (DHX9), one of other proteins that bound to the SSAT RNA column, may provide an alternative translational control mechanism. DHX9 is known to promote translation of several mRNAs possessing strong secondary structure (6, 25, 49), a speculation supported by our data showing DHX9 knockdown marginally increased SSAT translation (Fig. 2D). Perhaps it provides additional stabilization of the 5′ ORF stem-loop or increases the effect of the 5′ uORFs. We did not study DHX9 further, partially because we could not find antibodies to supported Western blot expression analysis, but that problem could be overcome, and the relationship to SSAT expression should be examined further. The effect of La protein knockdown was even less than that for DHX9, but a similar case can be made for this protein.
Improved knowledge of SSAT translation control will help direct efforts to develop pharmacological agents to manipulate this enzyme. Control of SSAT is most often associated with cancer drug development, but SSAT control relates to other human diseases and conditions as well. Animal knockout models show that a loss of SSAT dramatically increases diet-induced obesity, which suggests a role in metabolic regulation (30). Low brain SSAT activity is associated with major depression and suicide (52). Compared to healthy controls, SSAT is decreased and polyamines increased in an area of the brain of Parkinson's patients known to be affected by the disease; the suggested mechanism is polyamine induction of α-synuclein polymerization and toxicity (36).
Our data answer a longstanding question regarding control of SSAT translation, reveal an unusual control element within the RNA coding region, and uncover a new function for nucleolin, presenting the novel finding that nucleolin has at least one isoform with a distinct role, but many open questions remain. We know that a recombinant fragment of nucleolin that includes the 4 RNA recognition motif domains and the GAR C terminus (406 of the 710 total amino acids) binds to SSAT RNA, is degraded when expressed by cells exposed to either spermine or DENSPM (Fig. 5 and 6), and has gel motility similar to that of the isoform that binds to nucleolin RNA. These are important clues, but nucleolin is known to undergo glycosylation and phosphorylation that are likely involved in autocatalysis and will alter function. Polyamine-induced SSAT autocatalysis has been known for 20 years, but neither the region responsible for cleaving the protein nor the site of cleavage is known.
In conclusion, we answered an outstanding question by identifying the SSAT translation repressor protein as an isoform of nucleolin that stabilizes a stem-loop at the 5′ end of the mRNA coding region, and we present the hypothesis that this isoform functions as the sensor in a negative feedback regulatory system contributing to control of cellular polyamines. However, as is always the case when new phenomena are discovered, we end with more outstanding questions than we began with.
ACKNOWLEDGMENTS
This work was partially supported by National Institutes of Health (NIH) grant RO1AI064017.
We have no conflict of interest to declare.
Footnotes
Published ahead of print 21 February 2012
REFERENCES
- 1. Alhonen-Hongisto L, Seppanen P, Janne J. 1980. Intracellular putrescine and spermidine deprivation induces increased uptake of the natural polyamines and methylglyoxal bis(guanylhydrazone). Biochem. J. 192:941–945 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Babbar N, Gerner EW, Casero RA. 2006. Induction of spermidine/spermine N-1-acetyltransferase (SSAT) by aspirin in Caco-2 colon cancer cells. Biochem. J. 394:317–324 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Besse F, Ephrussi A. 2008. Translational control of localized mRNAs: restricting protein synthesis in space and time. Nat. Rev. Mol. Cell Biol. 9:971–980 [DOI] [PubMed] [Google Scholar]
- 4. Boden G, et al. 2008. Increase in endoplasmic reticulum stress-related proteins and genes in adipose tissue of obese, insulin-resistant individuals. Diabetes 57:2438–2444 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Boden G, Merali S. 2011. Measurement of the increase in endoplasmic reticulum stress-related proteins and genes in adipose tissue of obese, insulin-resistant individuals. Methods Enzymol. 489:67–82 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Bolinger C, Sharma A, Singh D, Yu LB, Boris-Lawrie K. 2010. RNA helicase A modulates translation of HIV-1 and infectivity of progeny virions. Nucleic Acids Res. 38:1686–1696 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Borer RA, Lehner CF, Eppenberger HM, Nigg EA. 1989. Major nucleolar proteins shuttle between nucleus and cytoplasm. Cell 56:379–390 [DOI] [PubMed] [Google Scholar]
- 8. Butcher NJ, Broadhurst GM, Minchin RF. 2007. Polyamine-dependent regulation of spermidine-spermine N-1-acetyltransferase mRNA translation. J. Biol. Chem. 282:28530–28539 [DOI] [PubMed] [Google Scholar]
- 9. Calvo SE, Pagliarini DJ, Mootha VK. 2009. Upstream open reading frames cause widespread reduction of protein expression and are polymorphic among humans. Proc. Natl. Acad. Sci. U. S. A. 106:7507–7512 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Casero RA, Jr, Pegg AE. 1993. Spermidine/spermine N1-acetyltrans ferase—the turning point in polyamine metabolism. FASEB J. 7:653–661 [PubMed] [Google Scholar]
- 11. Casero RA, Pegg AE. 2009. Polyamine catabolism and disease. Biochem. J. 421:323–338 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Chen CM, Chiang SY, Yeh NH. 1991. Increased stability of nucleolin in proliferating cells by inhibition of its self-cleaving activity. J. Biol. Chem. 266:7754–7758 [PubMed] [Google Scholar]
- 13. Chen DY, Huang S. 2001. Nucleolar components involved in ribosome biogenesis cycle between the nucleolus and nucleoplasm in interphase cells. J. Cell Biol. 153:169–176 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Chu E, et al. 1993. Identification of an RNA binding site for human thymidylate synthase. Proc. Natl. Acad. Sci. U. S. A. 90:517–521 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Coffino P. 2001. Regulation of cellular polyamines by antizyme. Nat. Rev. Mol. Cell Biol. 2:188–194 [DOI] [PubMed] [Google Scholar]
- 16. Duan X, Kelsen SG, Clarkson AB, Jr, Ji R, Merali S. 2010. SILAC analysis of oxidative stress-mediated proteins in human pneumocytes: new role for treacle. Proteomics 10:2165–2174 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Ercikan-Abali EA, et al. 1997. Dihydrofolate reductase protein inhibits its own translation by binding to dihydrofolate reductase mRNA sequences within the coding region. Biochemistry 36:12317–12322 [DOI] [PubMed] [Google Scholar]
- 18. Fang SH, Yeh NH. 1993. The self-cleaving activity of nucleolin determines its molecular dynamics in relation to cell proliferation. Exp. Cell Res. 208:48–53 [DOI] [PubMed] [Google Scholar]
- 19. Fogel-Petrovic M, Vujcic S, Miller J, Porter CW. 1996. Differential post-transcriptional control of ornithine decarboxylase and spermidine-spermine N-1-acetyltransferase by polyamines. FEBS Lett. 391:89–94 [DOI] [PubMed] [Google Scholar]
- 20. Gebauer F, Hentze MW. 2004. Molecular mechanisms of translational control. Nat. Rev. Mol. Cell Biol. 5:827–835 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Gerner EW, Meyskens FL., Jr 2004. Polyamines and cancer: old molecules, new understanding. Nat. Rev. Cancer 4:781–792 [DOI] [PubMed] [Google Scholar]
- 22. Giannone RJ, et al. 2007. Dual-tagging system for the affinity purification of mammalian protein complexes. Biotechniques 43:296, 298, 300, passim [DOI] [PubMed] [Google Scholar]
- 23. Hanakahi LA, Bu ZM, Maizels N. 2000. The C-terminal domain of nucleolin accelerates nucleic acid annealing. Biochemistry 39:15493–15499 [DOI] [PubMed] [Google Scholar]
- 24. Hanfrey C, et al. 2005. A dual upstream open reading frame-based autoregulatory circuit controlling polyamine-responsive translation. J. Biol. Chem. 280:39229–39237 [DOI] [PubMed] [Google Scholar]
- 25. Hartman TR, et al. 2006. RNA helicase A is necessary for translation of selected messenger RNAs. Nat. Struct. Mol. Biol. 13:509–516 [DOI] [PubMed] [Google Scholar]
- 26. Ishikawa F, Matunis MJ, Dreyfuss G, Cech TR. 1993. Nuclear proteins that bind the pre-mRNA 3′ splice site sequence r(UUAG/G) and the human telomeric DNA sequence d(TTAGGG)n. Mol. Cell. Biol. 13:4301–4310 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Ivanov IP, Atkins JF, Michael AJ. 2010. A profusion of upstream open reading frame mechanisms in polyamine-responsive translational regulation. Nucleic Acids Res. 38:353–359 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Ivanov IP, Loughran G, Atkins JF. 2008. uORFs with unusual translational start codons autoregulate expression of eukaryotic ornithine decarboxylase homologs. Proc. Natl. Acad. Sci. U. S. A. 105:10079–10084 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Jackson RJ, Hellen CUT, Pestova TV. 2010. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 11:113–127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Jell J, et al. 2007. Genetically altered expression of spermidine/spermine N1-acetyltransferase affects fat metabolism in mice via acetyl-CoA. J. Biol. Chem. 282:8404–8413 [DOI] [PubMed] [Google Scholar]
- 31. Kertesz M, et al. 2010. Genome-wide measurement of RNA secondary structure in yeast. Nature 467:103–107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Kozak M. 1989. Circumstances and mechanisms of inhibition of translation by secondary structure in eucaryotic mRNAs. Mol. Cell. Biol. 9:5134–5142 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Kozak M. 2005. Regulation of translation via mRNA structure in prokaryotes and eukaryotes. Gene 361:13–37 [DOI] [PubMed] [Google Scholar]
- 34. Kramer DL, et al. 2001. Polyamine depletion in human melanoma cells leads to G1 arrest associated with induction of p21WAF1/CIP1/SDI1, changes in the expression of p21-regulated genes, and a senescence-like phenotype. Cancer Res. 61:7754–7762 [PubMed] [Google Scholar]
- 35. Kurian L, Palanimurugan R, Godderz D, Dohmen RJ. 2011. Polyamine sensing by nascent ornithine decarboxylase antizyme stimulates decoding of its mRNA. Nature 477:490–494 [DOI] [PubMed] [Google Scholar]
- 36. Lewandowski NM, et al. 2010. Polyamine pathway contributes to the pathogenesis of Parkinson disease. Proc. Natl. Acad. Sci. U. S. A. 107:16970–16975 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Losfeld ME, et al. 2011. N-Glycosylation influences the structure and self-association abilities of recombinant nucleolin. FEBS J. 278:2552–2564 [DOI] [PubMed] [Google Scholar]
- 38. Marton LJ, Pegg AE. 1995. Polyamines as targets for therapeutic intervention. Annu. Rev. Pharmacol. Toxicol. 35:55–91 [DOI] [PubMed] [Google Scholar]
- 39. Niranjanakumari S, Lasda E, Brazas R, Garcia-Blanco MA. 2002. Reversible cross-linking combined with immunoprecipitation to study RNA-protein interactions in vivo. Methods 26:182–190 [DOI] [PubMed] [Google Scholar]
- 40. Parry L, Balana Fouce R, Pegg AE. 1995. Post-transcriptional regulation of the content of spermidine/spermine N1-acetyltransferase by N1N12-bis(ethyl)spermine. Biochem. J. 305(Pt 2):451–458 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Pegg AE. 2006. Regulation of ornithine decarboxylase. J. Biol. Chem. 281:14529–14532 [DOI] [PubMed] [Google Scholar]
- 42. Pegg AE, McCann PP. 1982. Polyamine metabolism and function. Am. J. Physiol. 243:C212–C221 [DOI] [PubMed] [Google Scholar]
- 43. Perez-Leal O, Merali S. 2012. Regulation of polyamine metabolism by translational control. Amino Acids 42:611–617 [DOI] [PubMed] [Google Scholar]
- 44. Pietila M, et al. 1997. Activation of polyamine catabolism profoundly alters tissue polyamine pools and affects hair growth and female fertility in transgenic mice overexpressing spermidine/spermine N1-acetyltransferase. J. Biol. Chem. 272:18746–18751 [DOI] [PubMed] [Google Scholar]
- 45. Pillai RS, Bhattacharyya SN, Filipowicz W. 2007. Repression of protein synthesis by miRNAs: how many mechanisms? Trends Cell Biol. 17:118–126 [DOI] [PubMed] [Google Scholar]
- 46. Porter CW, Bergeron RJ. 1988. Enzyme regulation as an approach to interference with polyamine biosynthesis—an alternative to enzyme inhibition. Adv. Enzyme Regul. 27:57–79 [DOI] [PubMed] [Google Scholar]
- 47. Porter CW, Bergeron RJ. 1983. Spermidine requirement for cell proliferation in eukaryotic cells: structural specificity and quantitation. Science 219:1083–1085 [DOI] [PubMed] [Google Scholar]
- 48. Porter CW, Bernacki RJ, Miller J, Bergeron RJ. 1993. Antitumor activity of N1, N11-bis(ethyl)norspermine against human melanoma xenografts and possible biochemical correlates of drug action. Cancer Res. 53:581–586 [PubMed] [Google Scholar]
- 49. Qiu CH, Ma YH, Wang JQ, Peng SP, Huang YQ. 2010. Lin28-mediated post-transcriptional regulation of Oct4 expression in human embryonic stem cells. Nucleic Acids Res. 38:1240–1248 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Raney A, Law GL, Mize GJ, Morris DR. 2002. Regulated translation termination at the upstream open reading frame in S-adenosylmethionine decarboxylase mRNA. J. Biol. Chem. 277:5988–5994 [DOI] [PubMed] [Google Scholar]
- 51. Sachs MS, Geballe AP. 2006. Downstream control of upstream open reading frames. Genes Dev. 20:915–921 [DOI] [PubMed] [Google Scholar]
- 52. Sequeira A, et al. 2006. Implication of SSAT by gene expression and genetic variation in suicide and major depression. Arch. Gen. Psychiatry 63:35–48 [DOI] [PubMed] [Google Scholar]
- 53. Shabalina SA, Ogurtsov AY, Spiridonov NA. 2006. A periodic pattern of mRNA secondary structure created by the genetic code. Nucleic Acids Res. 34:2428–2437 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Shi HB, et al. 2007. Nucleolin is a receptor that mediates antiangiogenic and antitumor activity of endostatin. Blood 110:2899–2906 [DOI] [PubMed] [Google Scholar]
- 55. Sonenberg N, Hershey JWB, Mathews M. 2000. Translational control of gene expression, 2nd ed Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY [Google Scholar]
- 56. Suppola S, et al. 1999. Overexpression of spermidine/spermine N1-acetyltransferase under the control of mouse metallothionein I promoter in transgenic mice: evidence for a striking post-transcriptional regulation of transgene expression by a polyamine analogue. Biochem. J. 338(Pt 2):311–316 [PMC free article] [PubMed] [Google Scholar]
- 57. Suzuki N, Matsui H, Hosoya T. 1985. Effects of androgen and polyamines on the phosphorylation of nucleolar proteins from rat ventral prostates with particular reference to 110-kDa phosphoprotein. J. Biol. Chem. 260:8050–8055 [PubMed] [Google Scholar]
- 58. Suzuki T, Suzuki N, Hosoya T. 1993. Limited proteolysis of rat liver nucleolin by endogenous proteases: effects of polyamines and histones. Biochem. J. 289(Pt 1):109–115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Takagi M, Absalon MJ, McLure KG, Kastan MB. 2005. Regulation of p53 translation and induction after DNA damage by ribosomal protein L26 and nucleolin. Cell 123:49–63 [DOI] [PubMed] [Google Scholar]
- 60. Tuteja R, Tuteja N. 1998. Nucleolin: a multifunctional major nucleolar phosphoprotein. Crit. Rev. Biochem. Mol. Biol. 33:407–436 [DOI] [PubMed] [Google Scholar]
- 61. Wang JY, Casero RA. 2006. Polyamine cell signaling: physiology, pharmacology, and cancer research. Humana Press, Totowa, NJ [Google Scholar]
- 62. Wang YL, Casero RA. 2006. Mammalian polyamine catabolism: a therapeutic target, a pathological problem, or both? J. Biochem. 139:17–25 [DOI] [PubMed] [Google Scholar]
- 63. Windbichler N, Schroeder R. 2006. Isolation of specific RNA-binding proteins using the streptomycin-binding RNA aptamer. Nat. Protoc. 1:637–640 [DOI] [PubMed] [Google Scholar]
- 64. Zaidi SH, Malter JS. 1995. Nucleolin and heterogeneous nuclear ribonucleoprotein C proteins specifically interact with the 3′-untranslated region of amyloid protein precursor mRNA. J. Biol. Chem. 270:17292–17298 [DOI] [PubMed] [Google Scholar]
- 65. Zuker M. 2003. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31:3406–3415 [DOI] [PMC free article] [PubMed] [Google Scholar]