Abstract
Low concentrations of the dissolved leonardite humic acid HuminFeed® (HF) prolonged the lifespan and enhanced the thermal stress resistance of the model organism Caenorhabditis elegans. However, growth was impaired and reproduction delayed, effects which have also been identified in response to other polyphenolic monomers, including Tannic acid, Rosmarinic acid, and Caffeic acid. Moreover, a chemical modification of HF, which increases its phenolic/quinonoid moieties, magnified the biological impact on C. elegans. To gain a deep insight into the molecular basis of these effects, we performed global transcriptomics on young adult (3 days) and old adult (11 days) nematodes exposed to two different concentrations of HF. We also studied several C. elegans mutant strains in respect to HF derived longevity and compared all results with data obtained for the chemically modified HF. The gene expression pattern of young HF-treated nematodes displayed a significant overlap to other conditions known to provoke longevity, including various plant polyphenol monomers. Besides the regulation of parts of the metabolism, transforming growth factor-beta signaling, and Insulin-like signaling, lysosomal activities seem to contribute most to HF’s and modified HF’s lifespan prolonging action. These results support the notion that the phenolic/quinonoid moieties of humic substances are major building blocks that drive the physiological effects observed in C. elegans.
Keywords: humic substances, hydroxybenzene, gene expression, aging, longevity, stress, TGF-beta, Caenorhabditis elegans
Introduction
Several studies with different model organisms have demonstrated that mild chemical stress trains cellular stress response pathways, e.g., biotransformation and antioxidant systems, which can ultimately result in lifespan extension; for a review see Kourtis and Tavernarakis (2011). Recently, we were able to show that exposure to a specific humic substance preparation, HuminFeed® (HF), significantly extends the lifespan of the nematode Caenorhabditis elegans (Steinberg et al., 2007). HF is weathered leonardite humic material characterized by high functional group content (Meinelt et al., 2007). By analogy it has been concluded that the effective building blocks may be hydroxybenzene groups. To confirm the biological impact of these structures, HF was chemically modified by increasing the concentrations of phenolic and quinonoid functional groups (Menzel et al., 2011). This chemical modification boosted the antioxidant properties of HF both in vitro and in vivo. Moreover, modified HF caused a significantly increased tolerance toward thermal stress in C. elegans and extended its lifespan (Menzel et al., 2011). In contrast, HF and the modified substances delayed the onset of reproduction and caused a reduction in overall body length. The underlying molecular basis of these HF mediated effects is, to date, unknown.
To define the transcriptional responses of HF exposure, we conducted global gene expression analyses using the Affymetrix® whole genome DNA microarray platform. Nematodes were exposed to two different concentrations of HF over a 3- or 11-days incubation period. We also assessed the effect of Huminfeed–Hydroquinone (HF-HQ), a HF derivate chemically enriched with hydroquinone.
Initially, we defined the differently expressed genes (DEGs), many of which displayed concentration dependent changes in expression. Selected results were confirmed by quantitative real-time RT-PCR. Subsequent investigations included gene ontology (GO; Ashburner et al., 2000) and Kyoto encyclopedia of genes and genomes (KEGG; Kanehisa, 2002) pathway analyses. Moreover, over-represented gene expression mountains and gene classes were evaluated according to Kim et al. (2001). In doing so, we were able to identify gene classes and pathways that returned a significant over-representation in HF or HF-HQ treated nematodes. A meta-analysis compared our findings with recently published data specific to either age-related gene expression, the genetic background of longevity mutants, the transcriptional profile of polyphenol treated nematodes or infection/immunity-related gene expression. This allowed us to pinpoint genes and associated pathways predicted to be key players in HF mediated longevity.
To substantiate the importance of these genes and pathways, loss of function mutants were tested for their ability to extend the lifespan in response to HF or HF-HQ exposure. In summary, this study provides new evidence that specific humic substances induce a complex mode of action. Moreover, humic substances are not limited (as previously thought) to act indirectly, e.g., via the unspecific binding to organic and inorganic compounds or the shuttling of electrons in microbial redox reactions, but rather extends the lifespan of C. elegans by means of regulatory and stress response pathways.
Materials and Methods
Nematodes
Maintenance of large synchronous cultures of old nematodes is challenging due to the offspring generated during the onset of reproductive output. Rather than using fluorodeoxyuridine to inhibit embryonic development, which has recently been shown to affect the worm (Aitlhadj and Stürzenbaum, 2010; Davies et al., 2012), this study utilized the C. elegans mutant strain GE24, pha-1(e2123), a putative transcriptional regulator of the pharyngeal precursor cells (Granato et al., 1994a). The mutant allele pha-1(e2123) is temperature sensitive; reproduction resembles wild type at 15°C, but is 100% embryonic lethal at 25°C. At the restrictive temperature, pharyngeal tissues of mutant embryos fail to undergo terminal differentiation and morphogenesis. After passing embryogenesis at the permissive temperature, however, a temperature shift does not affect pharyngeal functionality. Previously introduced as a selectable genetic marker (Granato et al., 1994b), we used pha-1(e2123) to maintain and follow a bulk preparation of synchronized nematodes. For reasons of comparison, we used the wild type strain Bristol N2 for all qRT-PCR experiments.
The lifespan assay included, besides N2 and pha-1(e2123), the following mutant strains: asah-1(tm495); RB1855, cyp-34A9(ok2401); DA465, eat-2(ad465); TK22, mev-1(kn1); AM1, osr-1(rm1); AU1, sek-1(ag1); VC199, sir-2.1(ok434); and MT2605, unc-43(n498n1186). All nematode strains were maintained on nematode growth medium (NGM) plates using Escherichia coli OP50 as food source according to standard procedures (Brenner, 1974; Sulston and Hodgkin, 1988).
Humic materials
HuminFeed® (HF; Humintech GmbH, Düsseldorf, Germany) was made by an alkaline extraction process of highly oxidized lignite (for a detailed and comparative physicochemical analysis see Meinelt et al., 2007). Our experiments used the same HF batch as previously chemically characterized. Moreover, we utilized a formaldehyde polycondensation product between HF and hydroquinone, namely HF-HQ, as described in Menzel et al. (2011). HF was used solely for practical reasons; it does not constitute an advertisement for this product.
Cultivation for the gene expression screen
Untreated nematodes were chunked onto control plates (no HF) and treatment plates [0.2 and 2.0 mM dissolved organic carbon (DOC) of HF and HF-HQ, respectively] and incubated at 15°C for 5 days. Then, a synchronous culture was generated by filtering worms through a 10-μm membrane (SM 16510/11, Sartorius, Germany), a pore size that retains all but first stage juveniles (L1). For each individual experiment, 15,000 larvae were distributed to three freshly prepared plates (Ø = 94 mm) and cultivated at 25°C to the young adult stage (3 days) or an older adult stage (11 days). Nematodes were fed every third day by replenishing the bacterial suspension and supplemented with a fresh preparation of humic substances. Following the respective exposures, nematodes were harvested by rinsing off with M9 buffer, rewashed twice, frozen in liquid nitrogen, and stored at −80°C until use. Each condition was cultivated in triplicate.
RNA preparation
Total RNA of each individual condition (n = 3) was isolated using an innuSPEED Tissue RNA Kit (AnalytikJena, Jena, Germany), which included an improved homogenization step with a SpeedMill (AnalytikJena, Jena, Germany) and the removal of genomic DNA through an initial spin filter column step. The RNA quality and quantity was analyzed both spectroscopically (NanoDrop 1000, ThermoScientific, UK) and by means of Agilent’s Bioanalyzer 2100 equipped with a RNA 6000 Nano kit (Agilent Technologies, Santa Clara, CA, USA). All RNA-samples showed no signs of degradation as indicated by Agilent’s RNA integrity numbers of 9 or 10.
DNA microarrays
Procedure
The processing of each RNA sample, the first- and second-strand cDNA synthesis as well as cRNA synthesis, labeling, and fragmentation was performed with a MessageAmp™ Premier RNA Amplification Kit (Ambion, Austin, TX, USA). We used the GeneChip® C. elegans genome platform (Affymetrix, Santa Clara, CA, USA), representing 22,548 different transcripts. To conduct the microarray hybridization experiments, we followed the specifications from Affymetrix’s GeneChip® hybridization, wash, and stain kit. The automated washing steps were performed in a GeneChip® fluidics station 450 (Affymetrix), and scans conducted by means of a GeneChip® scanner 3000 7G (Affymetrix). Triplicate GeneChips® were run for each condition.
Data interpretation and statistical analysis
Pre-processing of DNA microarray raw data included probe-specific background correction, summarization of probe set values, and normalization using the GCRMA algorithm with CARMAweb 1.4, an R- and Bioconductor-based web service for microarray data analysis (Rainer et al., 2006)1. Then, the data were initially filtered for missing values and subjected to a CLEAR-test that combines differential expression and variability using the GEPAS software (Herrero et al., 2003)2. For selection of DEGs, an unpaired t-test was performed followed by a significance analysis of microarray (SAM) test including a calculation that estimates the false discovery rate (FDR). FDR, reducing on the one hand type I errors for null associations, was set to a non-stringent level of <12.5%, mainly to guard from an increase of type II error (Swain et al., 2010) and also based on findings by Levine et al. (2011), which described 12.5% as most acceptable optimum level of FDR, representing the 90th percentile of the normal distribution curve. DEGs exceeding a fold change of 1.25 were further analyzed with respect to their functional clustering. We chose this fold-cut-off to allow an interpretation that is biologically meaningful, akin to the notion that data of sound technical and experimental quality which returns strong, statistically significant, absolute signal intensities is sufficiently robust to justify a fold-cut-off of >1.2 (Grigoryev et al., 2004; McCarthy and Smyth, 2009). This analysis was conducted using the functional annotation clustering tool of the Database for Annotation, Visualization, and Integrated Discovery (DAVID; Huang et al., 2007)3. This tool identified annotation categories including, e.g., GO terms and bio-pathways that are significantly enriched within the gene list, followed by a multiple sample correction (Benjamini and Hochberg, 1995). The resultant annotation clusters were ranked according to the statistical significance of cluster enrichment.
Representation factor
To assess the level of overlap between different conditions we calculated the representation factor (RF) in order to explore the fold enrichment. The RF identifies the level of enrichment (of individual transcripts) between gene lists (Kim et al., 2001; Evans et al., 2008). The choice of N(genome) was based on the values recommended by the authors. Intersection P-values were calculated from the hypergeometric distribution. RF were considered significant when RF > 1.
Validation of DNA microarray data by qRT-PCR
qRT-PCR analyses were conducted with samples from control and 2.0 mM DOC HF as well as HF-HQ exposed N2 wild type nematodes. The cultivation conditions of N2 wild type and pha-1(e2123) were identical. β-Actin (act-1) was used as reference gene, which did not change significantly in the DNA microarray data. A total of 1 μg RNA was reverse transcribed into cDNA (Menzel et al., 2005). Quantitative real-time amplification was performed in a MyiQ single color qPCR detection system (BIO-RAD, Germany) using the double-stranded DNA intercalating fluorescent agent EvaGreen for amplicon detection. Each reaction consisted of the qPCR Green Core Kit (Jena Bioscience, Germany), 200 nM of each primer pair, and cDNA template equivalent to 5 ng RNA starting material. The relative expression of the target genes was calculated by means of the comparative 2−ΔΔCt method (Livak and Schmittgen, 2001). All experiments for each selected gene were performed in duplicate; RT-negatives were also run for each sample and gene to confirm the absence of DNA contamination. The list of primers with their corresponding PCR-efficiencies (91–100%) is given in Table S1 in Supplementary Material; at least one primer of each pair spanned an intron to avoid amplification of genomic DNA.
Lifespan assay and statistical evaluation
The lifespan of C. elegans was investigated as previously described (Pietsch et al., 2009) using synchronized L4 larvae and a growth temperature of 20°C. However, pha-1(e2123) was pre-cultured at 15°C until the L4 state, and then maintained at 20°C. The concentrations of HF and HF-HQ were 0 and 0.4 mM DOC, respectively, mixed both into the agar and to the bacterial lawn. The first day of adulthood was defined as day 1. We performed three independent trials, each comprising 10 small agar plates (Ø = 35 mm) and 150 nematodes per trial. Animals were scored daily for survival until all worms had died. Median and mean lifespan and percentage changes (compared to controls) were determined. The statistical significance of alterations in the mean lifespan was calculated using the log-rank test (Azen et al., 1977), available online from the Bioinformatics group at the Walter and Eliza Hall Institute of Medical Research (Melbourne, Australia)4. Blinding of studies was not possible due to the color of humic material, which also stains the NGM agar.
Results
Transcript profiling by whole genome microarray following HF and HF-HQ treatment
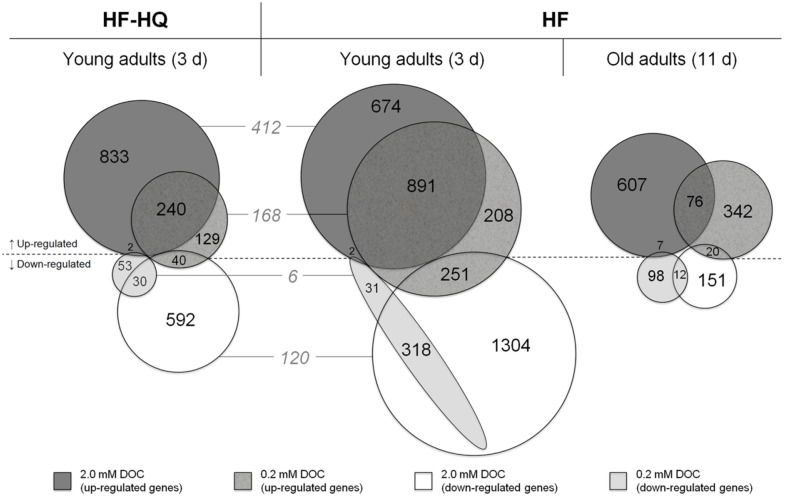
The Venn diagrams in Figure 1 present an overview of the number of genes that were significantly up- or down-regulated in response to the humic substances and the respective overlap between the two concentrations, 0.2 and 2.0 mM DOC. Due to the low threshold (a minimum fold change in gene expression of 1.25), many genes were classed as DEGs, especially in young adults exposed to HF. However, the extended incubation time of 11 days was characterized by a substantial decline in the number of DEGs, in particular down-regulated genes. The HF-HQ derived data resemble the results from young adults exposed to HF, albeit overall less DEGs were identified. The intersection between HF-HQ and HF at 0.2 and 2.0 mM DOC comprised of 174 and 532 DEGs, respectively. An extensive overlap was observed between the two concentrations per HF condition (Figure 1). The complete data can be viewed in the National Center for Biotechnology Information’s (NCBI) Gene Expression Omnibus (GEO) database (accession number GSE35360) and are also given in Table S2 in Supplementary Material, including expression values, statistics, and gene annotations.
Figure 1.
Over-represented genes. The Venn diagrams show the overlap of significantly up- or down-regulated genes (fold change >1.25 or <0.8) per concentration in nematodes exposed for 3 days to HF (center), HF-HQ (left), and for 11 days to HF (right). The gray colored numbers given in italics represent the overlap between HF and HF-HQ derived DEG lists.
Validation of transcript profiles of selected genes and conditions by qRT-PCR
A validation of the microarray experiment was deemed to be important to (i) allow a comparison between the pha-1(e2123) strain (used for microarray experiments) and the N2 wild type (used for qRT-PCR), and (ii) evaluate the expression levels at reduced exposure times (24 and 48 h as well as 72 h). Samples generated for qRT-PCR originated from worms exposed to 2.0 mM DOC of HF or HF-HQ, respectively. Overall, both methods (microarray and qPCR) and genotypes [pha-1(e2123) and wild type] returned analogous expression profiles at 72 h exposure in 9 of 10 genes tested; only F15E11.13 could not be confirmed (Table 1). This suggests that pha-1(e2123) and wild type are essentially interchangeable. The inclusion of further time points revealed that the majority of the selected genes did not respond rapidly to the exposure, exceptions to this were F15E11.13 and, to some extent, cyp-34A9 and skn-1.
Table 1.
Quantitative PCR of 10 HF-responsive genes identified by DNA microarray.
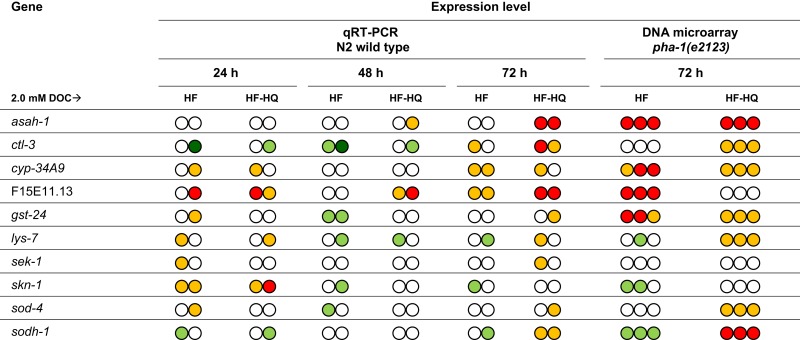 |
Fold change:  <0.3,
<0.3,  0.3–0.7,
0.3–0.7,  0.7–1.5,
0.7–1.5,  1.5–3.0,
1.5–3.0,  >3.0.
>3.0.
Evaluation of transcript profiles based on annotation enrichment analyses
The transcriptional profiles were subjected to detailed analyses to identify pathways linked to the mode of actions of HF and HF-HQ. The principal approach applied an annotation enrichment analysis within the different DEG sets, sub-divided in up- and down-regulated genes.
First, DEGs were assigned to KEGG pathways and mapped to known molecular interaction networks, such as metabolic pathways or environmental information processes (Table S3 in Supplementary Material). The analysis of the two HF derived lists for young adults (3 days) identified 10 KEGG pathways for 0.2 and 16 KEGG pathways for 2.0 mM DOC (Table 2, left part). DEG lists of 3-days-old HF-HQ treated nematodes returned 3 and 12 pathways for the low and the high concentration, respectively (Table 2). Despite some individual differences between both conditions (e.g., the down-regulation of spliceosome specific genes by HF), the overall overlap between concentrations and conditions was significant. Particularly noticeable was the induction of fatty acid metabolism, in particular arachidonic acid (AA) and sphingolipid metabolism, as well as the up-regulation of lysosome related genes. In the 11-days-old HF exposed worms, only four significantly enriched KEGG pathways were modulated, which were restricted to the lower concentration of HF (0.2 mM DOC). Besides the persistent induction of the AA metabolism, HF (0.2 mM DOC) was marked by a distinct up-regulation of the biotransformation machinery, which includes glutathione and cytochrome P450 (CYP) pathways. It should be noted that HF (2 mM DOC) also induced biotransformation associated genes, however, because of the large number of DEGs, they were not found to be significantly enriched.
Table 2.
Overview of over-represented KEGG pathways and gene ontology terms in DEG lists.
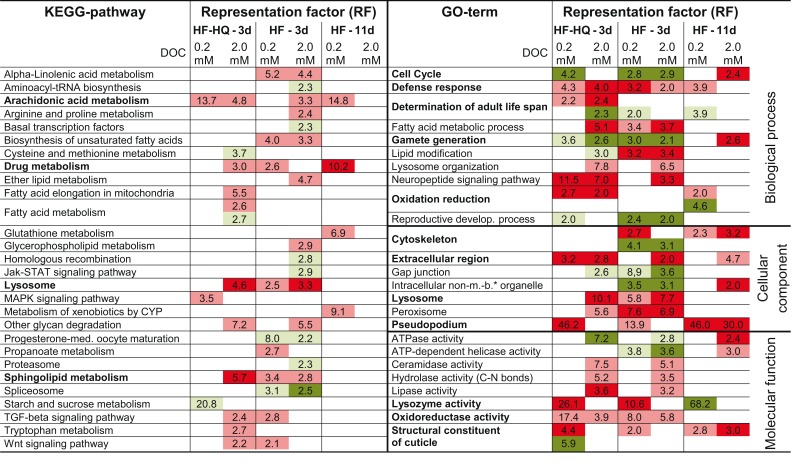 |
Shown is the representation factor for DEGs in the respective list in relation to the overall count in significantly enriched KEGG and GO terms, respectively; red label stands for up-, green for down-regulated genes; respective pale coloring corresponds to P < 0.05, deep colors to P < 0.001; apparently most relevant terms are given in bolt; please find more details in Tables S3 and S4 in Supplementary Material. *Membrane-bound.
Second, DEGs were classified using GO terms to obtain further functional insights into gene expression responses. The GO analysis produced a multitude of significantly enriched terms, many represented by the same genes across and within the three GO domains (biological process, cellular component, and molecular function). Redundancy was removed by applying the “GOTERM_XX_ALL” option in DAVID 3.0, a functional annotation clustering tool. Table S4 in Supplementary Material lists GO terms represented by the largest number of genes within individual functional clusters. A further selection of the 25 most striking terms are given in Table 2 (right part). Again, results from young adult nematodes, exposed either to HF or HF-HQ, were more consistent compared to their older counterparts. As before, lysosomal processes, defense response as well as lipid and fatty acid metabolism were found to be enriched in the group of up-regulated genes. Moreover, humic substances induced the expression of genes coding for constituents of the cuticle and cytoskeleton. The persistent strong induction of the cellular components pseudopodium and extracellular region is caused by a comprehensive up-regulation of various major sperm proteins (MSP). Oxidative/reductive processes, determinants of adult life span, and neuropeptide signaling were found to be more enriched in response to the chemically modified HF-HQ. Both HF preparations seem to slow down the reproductive development of C. elegans (also cell cycle and gamete production) following a short term exposure of 3 days, a process which was seen to be reverted in older nematodes exposed for 11 days.
In a third step, we compared the DEG lists to a library of 59 different gene classes or functionally related groups of genes (for the complete comparison see Table S5 in Supplementary Material), which were assembled into a gene expression map (Kim et al., 2001). Table 3 (left part) shows a selection of the 15 most relevant gene groups in which at least one dataset displays an over-representation. As before, genes coding for determinants of cell structure, lipid metabolism, glutathione transferases, and MSPs were significantly enriched. Lysozyme and protease encoding genes were found to be predominantly up-regulated after 3 days of exposure but down-regulated in 11-days-old HF-treated nematodes.
Table 3.
Overview of over-represented gene classes in DEG lists and comparison to literature data.
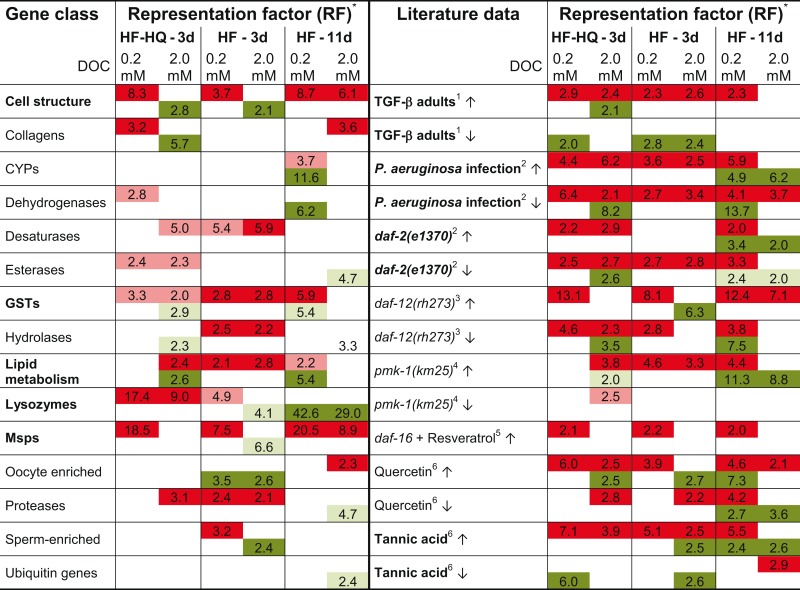 |
*Shown is the representation factor for DEGs in the respective list in relation to the overall count in significantly enriched gene classes or in comparison to literature data; red label and ↑ refer to up-, green label and ↓ to down-regulated genes; respective pale coloring corresponds to P < 0.05, deep colors to P < 0.001; gene glasses and datasets deemed to be most relevant are given in bold; for more details see Tables S5 and S6 in Supplementary Material. 1Shaw et al. (2007), 2Evans et al. (2008), 3Fisher and Lithgow (2006), 4Troemel et al. (2006), 5Viswanathan et al. (2005), 6Pietsch et al. (2012).
Meta-analysis: Comparison of transcript profiles with selected datasets taken from the literature
Datasets from HF and HF-HQ treated nematodes were compared to expression profiles obtained from long-lived mutants, dauer larvae, worms treated with lifespan-extending polyphenols and immunity challenged nematodes. As a control, the analysis included studies addressing the gene expression changes during the C. elegans life-cycle. Table S6 in Supplementary Material summarizes the results of the complete analysis comprising 40 individual data sets. Table 3 (right part) reduces the meta-analysis to the 15 most overlapping data sets. The transcriptional profiles of HF and HF-HQ are closest to the results from Tannic acid treated nematodes and mutants of the transforming growth factor-beta (TGF-β) pathway. In contrast, significant overlaps were limited to up-regulated DEGs in long-lived daf-2(e1370) and daf-12(rh273), as well as nematodes infected with Pseudomonas aeruginosa or exposed to another polyphenol, the flavonoid Quercetin. A significant proportion of genes up-regulated after 3 days were shown to be down-regulated after 11 days. This result demonstrates the level of dynamic transcriptional changes during the HF mediated aging process.
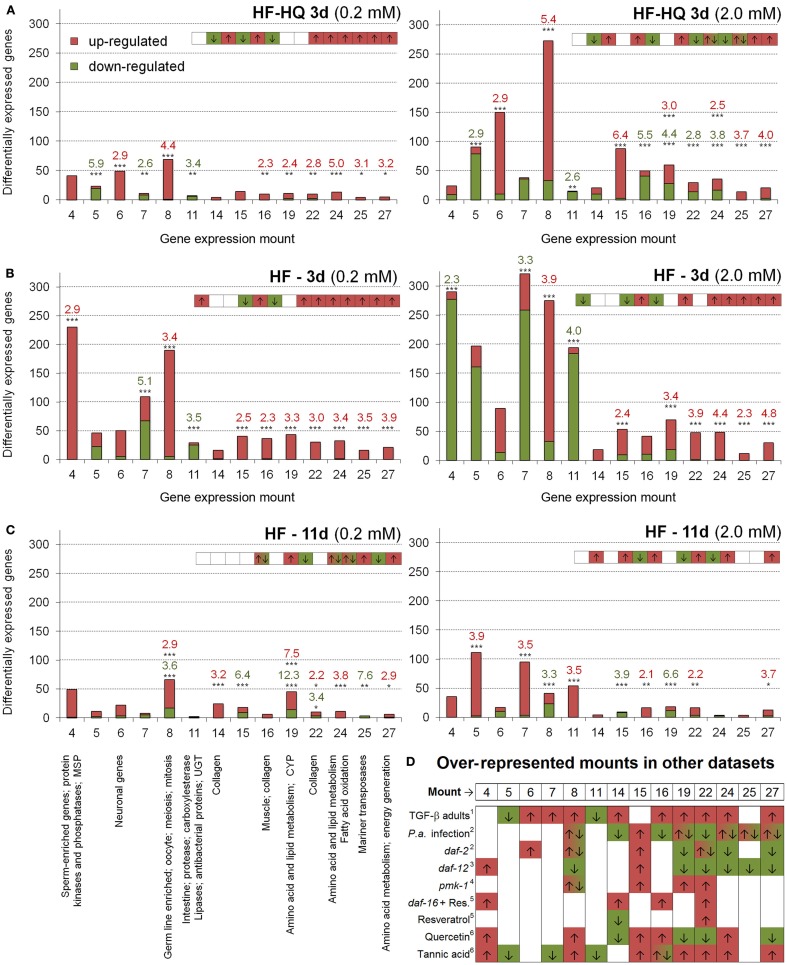
HF and HF-HQ derived transcriptional profiles were also analyzed by assigning DEGs to gene expression mountains originally assembled from 553 different C. elegans DNA microarray experiments (Kim et al. (2001). Table S7 in Supplementary Material displays the overlap of all gene expression mountains with the HF and HF-HQ derived datasets, respectively, as calculated by RFs. A summary is presented in Figure 2 and includes a graphical overview of the data obtained for 14 mounts, distinguishing between up- and down-regulated genes as well as a summary from selected published data sets. Young adult nematodes exposed to the lower concentration of HF-HQ (Figure 2A, left side), and both HF concentrations (Figure 2B) resembled the gene expression mount map characteristic for Tannic acid exposed nematodes or mutants of the TGF-β pathway. The higher concentration of HF-HQ on the other hand seems to overlap, in part, with the gene expression patterns following P. aeruginosa infection (Figure 2A, right side). No clear categorization was possible for old adults exposed to HF.
Figure 2.
Over-represented gene expression mountains. Identification of over-represented gene expression mountains of (A) HF-HQ (3 days), (B) HF (3 days), and (C) HF (11 days) treated nematodes; 0.2 mM DOC data are shown on the left, 2.0 mM data on the right. Only mounts which are significantly affected by at least one HS treatment are presented, red labels represent up-, green labels down-regulated genes. *P < 0.05, **P < 0.01, ***P < 0.001. (D) Presents the associated term names on the left and a comparison to selected published datasets on the right (only mounts relevant to HS are shown). Note the short graphical overview in the right upper corner of each diagram. The complete dataset (gene numbers and RFs for all 44 gene expression mountains of all six HS conditions as well as RFs for previously published datasets) can be found in Table S7 in Supplementary Material. 1Shaw et al. (2007), 2Evans et al. (2008), 3Fisher and Lithgow (2006), 4Troemel et al. (2006), 5Viswanathan et al. (2005), 6Pietsch et al. (2012).
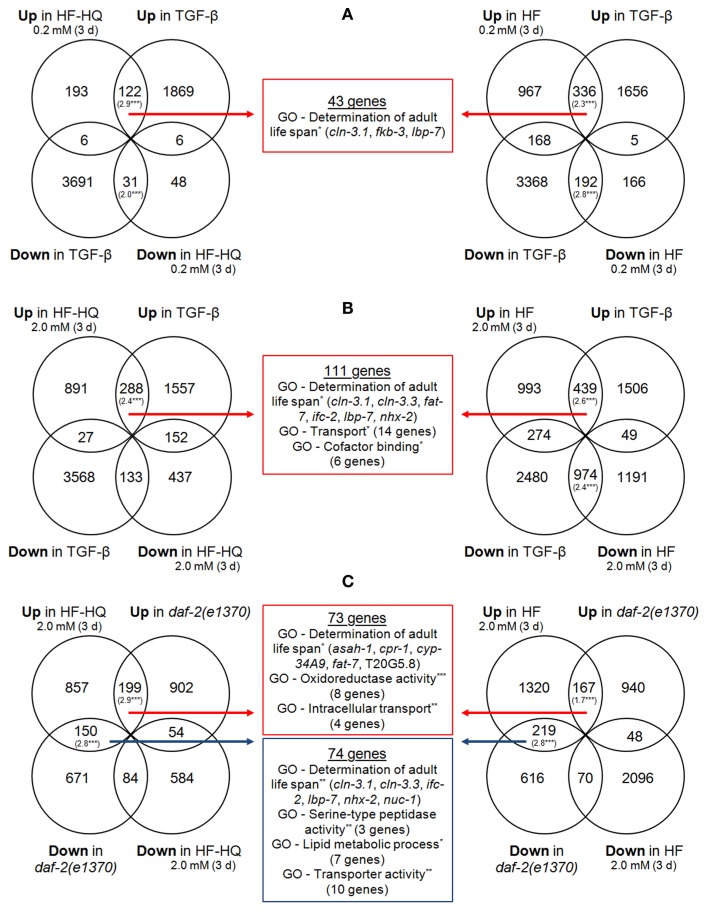
A stringent GO term analysis on genes belonging to each of the significantly overlapping sections made it possible to focus on similarities within and between HF, HF-HQ, and long-lived mutant strains. Only gene lists derived from TGF-β mutants as well as daf-2(e1370) mutants produced significant results. Figure 3 shows the Venn diagrams describing the overlap between TGF-β mutants and 0.2 mM (Figure 3A) as well as 2.0 mM (Figure 3B) HF or HF-HQ treated nematodes. The overlap between the profiles of daf-2(e1370), and 2.0 mM HF or HF-HQ treated nematodes is given in Figure 3C. All three analyses returned the term “Determination of adult life span,” moreover, common transport processes were identified. Some of these genes act downstream of DAF-16 (Murphy et al., 2003) and/or thought to be involved in lysosomal metabolism (e.g., asah-1, encoding a putative n-acylsphingosine amidohydrolase) or a potential drug metabolizer (e.g., cyp-34A9, a cytochrome P450 monooxygenase). Both mutant strains were selected for lifespan assays as described below.
Figure 3.
Overlap of similarly regulated genes. Shown are comparisons of DNA microarray data sets of HF (right) and HF-HQ (left) treatments with the data set derived from TGF-β adults (Shaw et al., 2007) – 0.2 mM DOC HS (A) and 2.0 mM DOC HS (B) – as well as from daf-2(e1370) mutants (Evans et al., 2008) (C). The RF values in brackets indicate a significant overlap between the data sets. The middle section represents the overlap of DEGs in both HF, HF-HQ, and the comparative condition, shown are significantly over-represented GO terms (Biological process), the associated number of genes, and the individual names of all genes which are part of the GO term “Determination of adult life span.” Red arrows/boxes are intersections of commonly up-regulated transcripts; blue arrows/boxes are intersections derived from genes with opposite transcriptional responses. *P < 0.05. **P < 0.005. ***P < 0.001
Identification of genes required for HF/HF-HQ mediated longevity
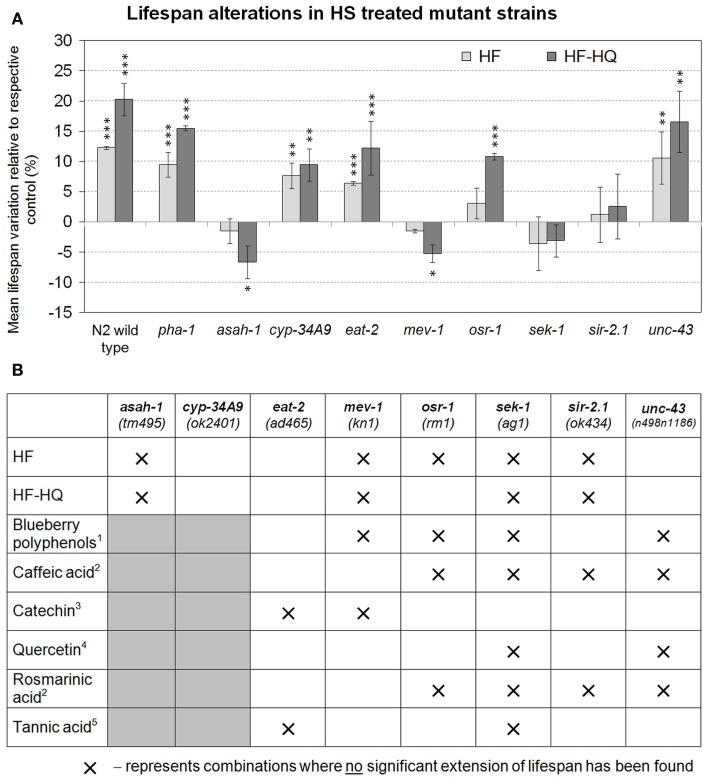
To substantiate the importance of stress response genes and genes relevant to aging, nine C. elegans mutants were tested for their ability to trigger longevity during humic substance exposure (Figure 4A). HF or HF-HQ exposure led to a significant increase in mean lifespan of N2 wild type and the pha-1(e2123) mutant strain, confirming that pha-1 was a suitable test strain for this analysis. In addition, cyp-34A9(ok2401), eat-2(ad465), and unc-43(n498n1186) responded to the exposure to HF or HF-HQ with an increase in lifespan. HF-HQ was seemingly a more effective trigger of longevity than the unmodified HF, a fact which may explain why osr-1(rm1) responded only to HF-HQ, but not to HF. No lifespan extension was observed in sek-1(ag1) and sir-2.1(ok434) and was even reduced in asah-1(tm495) and mev-1(kn1) (Figure 4A). An overview of comparative data from similar experiments testing the impact of individual polyphenols and one fruit extract suggest that the HF/HF-HQ mediated longevity align well with the key mechanisms identified for Caffeic acid and Rosmarinic acid as well as blueberry polyphenols. The overlap to Tannic acid treated nematodes was limited to sek-1, a frequently encountered effector in aging pathways (Figure 4B).
Figure 4.
Genetic modulators. (A) Lifespan alterations in nine mutant strains following the treatment with either 0.4 mM DOC HF (light gray) or 0.4 mM DOC HF-HQ (dark gray). Shown are the mean lifespan variations compared to the respective untreated control. Error bars represent SEM, **P < 0.01, ***P < 0.001. (B) provides a graphical comparison of previously published datasets on polyphenols. Crosses mark combinations where no significant extension of lifespan was observed; gray shaded boxes are untested combinations. 1Wilson et al. (2006), 2Pietsch et al. (2011), 3Saul et al. (2009), 4Pietsch et al. (2009), 5Saul et al. (2010).
Discussion
Caenorhabditis elegans exposed to the humic substance HF were shown to live longer and to be more stress resistant, but were impaired in their reproductive performance and growth (Menzel et al., 2011). Therefore, HF’s impact resembles other polyphenol monomers, such as Quercetin (Kampkötter et al., 2008; Pietsch et al., 2009) and Tannic acid (Saul et al., 2010). Indeed, HF was found to be rich in functional group content, with possibly hydroxybenzenes as effective building blocks (Meinelt et al., 2007). First experimental evidence was offered by Menzel et al. (2011) who were able to demonstrate that a chemical modification of HF (an enrichment in phenolic and quinonoid functional groups) resulted in the amplification of the biological effects. The current study aimed to extend this knowledgebase by applying global transcriptomics to compare the impact of HF and a modified HF (HF-HQ).
Quantitative analysis of transcript profiles and validation by qRT-PCR
Exposure to HF or HF-HQ had a profound impact on global gene expression patterns. Clearly, the return of over 1000 DEGs can be attributed, in part, to the non-stringent cut-off value of 1.25-fold, which was purposefully chosen to facilitate a comprehensive secondary analysis. The overlap between the HF and the HF-HQ DEGs-lists is considerable, even though HF returned more DEGs than HF-HQ. A 10-fold increase in concentration (from 0.2 to 2.0 mM DOC) amplified the number of DEGs approximately twofold for HF and threefold for HF-HQ. Moreover, at least for HF exposure, it was apparent that the transcriptome was more responsive in young adults than old adults. This finding together with the observed delay in reproduction and the retarded growth (Menzel et al., 2011) suggests that the reproductive development is a target following HF exposure. The high level of overlap between the two concentrations in young adults was not observed in old adults, indicating a shift in gene expression dynamics, which in accordance with Van Straalen and Feder (2012) is separated by effects (concentration/dose) rather than different exposures (compounds). Given that the primary objective of this work was to distinguish between the two preparations of HF and compare the results to the genomics literature, the scenario “exposure” is more relevant. Therefore we focused on the 3-days derived data.
The Affymetrix DNA microarray system is a robust, reliable, and well established system (Dalma-Weiszhausz et al., 2006). Nonetheless, we analyzed the expression level of 10 genes by qRT-PCR, however in the N2 wild type rather than pha-1(e2123) which was used for the microarray experiments. By doing so, we were able to confirm the microarray data but also the validity of the mutant strain, which was required for the production of a large population of age-synchronized old adults. The addition of two additional time points, namely shorter exposure times of 24 and 48 h, revealed complex time resolved differences in transcription. Many genes were initially repressed or transcriptionally inactive but induced at 72 h. In contrast, skn-1 was induced at 24 h but returned to base line levels thereafter. Interestingly, skn-1 encodes a longevity-promoting transcription factor and is positioned in the p38 MAPK pathway. Its expression has been shown to be enhanced under conditions of stress (An and Blackwell, 2003) or reduced DAF-2 signaling.
Qualitative analysis of transcript profiles by abundance screens
The generation of a dataset that describes quantitative changes in gene expression upon a certain condition, e.g., exposure to a chemical compound, is less challenging than the interpretation of its relevance, significance, and contribution to the physiology of the organism. Statistical methods aid in the identification of over- or under-represented transcripts which can be aligned to biological processes or functions via KEGG- and GO-term screens.
The over-representation of DEGs related to lipid metabolism and biotransformation in HF and HF-HQ derived transcript profiles suggests the presence of enhanced catabolism, possibly of toxic intermediates. This may contribute to the lifespan extension which, according to the green theory of aging, is due to the investment in cellular waste disposal and protein conservation (Gems and McElwee, 2005). The induction of several signaling pathways (MAPK, Wnt, TGF-β, neuropeptide) might reflect transcriptional changes of downstream targets. However, changes in heat shock protein (HSP) gene expression were not observed. Clearly, other transcripts responded to the HF/HF-HQ challenge, some are possibly involved in the observed longevity phenotype. For example the pronounced over-representation of lysosome specific genes may be linked to the process of autophagy, which mediates the degradation of cellular components, including whole organelles and protein aggregates. The importance of an efficient lysosomal activity is indicated by the finding that long-lived C. elegans mutants frequently display increased autophagy (Melendez et al., 2003; Hars et al., 2007).
The down-regulation of the GO terms “reproductive developmental process,” “gamete generation,” and “cell cycle” corroborate the notion that the reproductive development slows down in HF/HF-HQ exposed nematodes. Interestingly, this process seems to be dynamic in nature as “gamete generation” and “cell cycle” appear to be up-regulated (at least in 2.0 mM DOC HF) in old adults. In contrast, transcripts involved in the constitution of the cuticle (also the gene class “cell structure”) or active in the extracellular region and the pseudopodium (here in particular MSPs) are consistently up-regulated. The cuticle of C. elegans can differ in layer numbers, relative thickness, and composition during development (particularly in larvae) and changing environmental conditions. Indeed, genes encoding for cuticle collagens were found to be induced in response to several bacterial species (Coolon et al., 2009) and under oxidative stress (Shin et al., 2011). In aging research, studies identified a large number of collagens as age regulated genes (Halaschek-Wiener et al., 2005; Budovskaya et al., 2008). These data suggest that cuticle collagens may be differentially regulated indirectly in defense against environmental perturbations and potentially in longevity. A comparable up-regulation of msp-genes were observed in the long-lived daf-12(rh273) mutant (Fisher and Lithgow, 2006), but also in Quercetin or Tannic acid exposed wild type worms (Pietsch et al., 2012).
The HF and HF-HQ derived data were remarkably similar. However, only HF-HQ returned an up-regulation of the Biological processes “oxidation/reduction activity” possibly due to the previously observed increase in oxidoreductive activity (Menzel et al., 2011).
Qualitative analysis of transcript profiles by literature comparison
The main problem of abundance screens (e.g., GO-term profiling) is the incomplete gene annotation of genomes and the risk of over-interpretation, as enrichment values can occur by chance (Rhee et al., 2008). The application of appropriate statistical tools minimizes, but cannot exclude, the frequency of false-positives. To offer a more independent verification, we conducted a meta-analysis to include published data and searched for overlapping gene clusters via the gene expression mount map created by Kim et al. (2001). This revealed that Tannic acid exposed wild type worms (Pietsch et al., 2012), TGF-β mutants (Shaw et al., 2007), and worms subjected to humic substances produce similar expression pattern mountains. The comparison of DEGs-lists of either up- or down-regulated genes confirmed this result. The overlap to worms with a challenged immunity in response to an infection with P. aeruginosa, Quercetin treated nematodes as well as long-lived and more stress-resistant mutants [daf-2(e1370) and daf-12(rh273)] was, at large, restricted to the section of up-regulated DEGs. Both HF preparations and TGF-β mutants (and daf-2 for 2.0 mM DOC) shared GO terms, suggesting that the negative regulation of the TGF-β pathway and, less pronounced, the insulin-like signaling (ILS) pathway, play prominent roles in the lifespan extension due to HF/HF-HQ. Both signaling cascades control, by responding to environmental conditions, whether C. elegans larvae grow to adults or to long-lived and stress-resistant dauer larvae. Based on our results, it seems that HF preparations are able to modulate these pathways, thereby facilitating the observed increase in stress resistance and longevity.
Genetic players that promote HF-dependent longevity
HF or HF-HQ treatment was not able to prolong the lifespan of the C. elegans mutants sek-1, sir-2.1, mev-1, asah-1 and (in the case of HF only) osr-1. SEK-1, a MAP2K, is part of the p38 MAP kinase pathway and acts downstream of TIR-1 (toll and interleukin receptor) and NSY-1 (MAP3K). It phosphorylates the MAP kinases JNK-1 and PMK-1 (Tanaka-Hino et al., 2002), the latter results in an elevated immune response to pathogen infection (Kim et al., 2002) and also functions via SKN-1 to control resistance against metals, such as arsenic (An and Blackwell, 2003). OSR-1 is coupled to SEK-1 (through UNC-43) and regulates the osmotic stress response and survival in hyper-osmotic environments, where viability depends on activity of the CaMKII pathway (Solomon et al., 2004). Both sek-1 and osr-1 were shown to be essential for the blueberry polyphenol induced longevity (Wilson et al., 2006) as well as the Caffeic acid and Rosmarinic acid (Pietsch et al., 2011) mediated longevity. Surprisingly, unc-43 was not found to be essential for the longevity effect by HF or HF-HQ.
A genetic analysis suggested that sir-2.1 (which encodes the NAD+-dependent deacetylase) extends lifespan via the ILS pathway and requires daf-16 (Tissenbaum and Guarente, 2001). Berdichevsky et al. (2006) proposed the existence of a stress-dependent pathway in which SIR-2.1 acts in parallel to the ILS pathway, but still via an activation of DAF-16. Likewise, Caffeic acid, Rosmarinic acid, and Resveratrol mediated lifespan extension were all shown to be dependent on SIR-2.1. Viswanathan et al. (2005) described the involvement of sir-2.1 in the up-regulation of stress response genes (especially abu-11), a mechanism which is thought to aid protein folding in the endoplasmic reticulum. SIR-2.1’s involvement in stress response pathways during aging and HF/HF-HQ challenge may thus promote longevity and is possibly linked to the ILS-signaling cascades.
To further examine whether HF preparations could protect against acute oxidative stress, we examined mev-1(kn1), a nematode which harbors a mutation in the cytochrome b large subunit of mitochondrial complex II (Ishii et al., 1998). The mutation causes an overproduction of superoxide and increased oxidative stress, resulting in accelerated aging and a reduced lifespan (Hosokawa et al., 1994; Senoo-Matsuda et al., 2001). Neither HF/HF-HQ, blueberry polyphenol (Wilson et al., 2006) nor Catechin (Saul et al., 2009) treatment were able to revert or protect against the oxidative stress encountered by the mev-1(kn1) mutants. The pro-oxidant properties, as described for different polyphenols by Akagawa et al. (2003) and Wiegant et al. (2009), may explain this result. Indeed, HF-HQ induced antioxidant enzymes, such as catalases (ctl-2, ctl-3) and a superoxide dismutase (sod-3).
The asah-1 gene encodes a putative acid ceramidase, a lysosomal enzyme which catalyses the hydrolysis of ceramide to sphingosine and free fatty acid. Although the annotation data for asah-1 is patchy, it is thought to act downstream from DAF-16 (Murphy et al., 2003), has been assigned to the GO-term “Determination of adult life span,” and leads to Farber lipogranulomatosis, when mutated in humans. In affected individuals, harmful amounts of lipids accumulate in cells and tissues throughout the body (Mao and Obeid, 2008). Because several gene clusters coding for lysosome components, ceramidases, and sphingolipid metabolizing enzymes were found to be up-regulated by HF/HF-HQ, these processes may be essential for the effectiveness of humic substances, a hypothesis that requires further investigation.
Conclusion
The humic substance preparation HF significantly extends the lifespan of the nematode C. elegans. Here we were able to show that HF’s mode of action resembles other polyphenol monomers: (i) the transcript profiles of HF are very similar to Tannic acid and, but less pronounced, to Quercetin; (ii) the enrichment of HF with hydroquinones (HF-HQ) enhances its phenotypic effects (Menzel et al., 2011) and returns a more streamlined transcript profile (i.e., HF-HQ and HF affected common gene clusters, even though the total number of DEGs was lower in the HF-HQ sample); and (iii) HF/HF-HQ and polyphenols induce similar effects on key mutant nematodes.
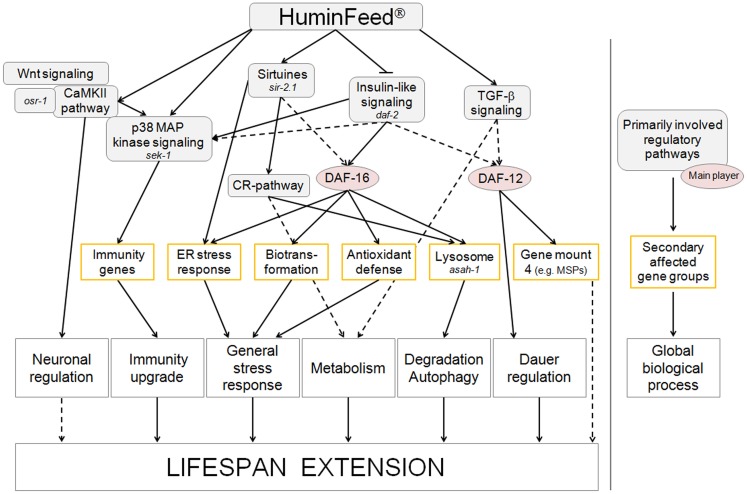
Despite these similarities, the mode of action of HF is complex (Figure 5) and is heavily influenced by the negative regulation of TGF-β- and ILS signaling as well as increased lysosomal activity. Longevity, according to Kirkwood and Austad (2000), is driven by an organism’s ability to cope with extrinsic or intrinsic stressors. Clearly stress response pathways do not function in isolation but act, in concert, within a stress network where multiple hubs serve as coordinators of various modules (Kourtis and Tavernarakis, 2011). The process of aging both influences and is influenced by this stress network. Mild environmental stress, as triggered by low concentrations of polyphenols or polyphenol-containing humic substances, primes response pathways which in turn increase stress resistance and longevity. These mechanisms are multidimensional but one of the prime candidates involved in the impact of polyphenols and humic substances is, at least in the nematode C. elegans, sir-2.1.
Figure 5.
Flow chart summarizing the multidimensional modes of action of HF/HF-HQ. Solid arrows denote direct, well-known connections; dotted lines highlight suggested relations. Genes identified in lifespan assays or selected by the literature meta-analysis are annotated in italics.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We would like to thank members of the King’s College Genomics Center (Dr. Matthew Arno and Dr. Estibaliz Aldecoa-otalora Astarloa) for assistance in our DNA microarray experiments. Furthermore, we thank the Caenorhabditis Genetics Center, which is funded by the National Institutes of Health National Centre for Research Resources (USA), for the supply of all Caenorhabditis elegans strains, except for asah-1(tm495), which was kindly provided by the Mitani Lab (Department of Physiology, Tokyo Women’s Medical University, School of Medicine, Japan) an its National BioResource Project for the nematode. Suresh C. Swain and Stephen R. Stürzenbaum were supported by the BBSRC (BB/E025099), Ralph Menzel, Sophie Tiedt, and Christian E. W. Steinberg by a DFG grant (STE673/16-1).
Footnotes
Supplementary Material
The Supplementary Material for this article can be found online at http://www.frontiersin.org/Toxicogenomics_/10.3389/fgene.2012.00050/abstract
References
- Aitlhadj L., Stürzenbaum S. R. (2010). The use of FUdR can cause prolonged longevity in mutant nematodes. Mech. Ageing Dev. 131, 364–365 10.1016/j.mad.2010.03.002 [DOI] [PubMed] [Google Scholar]
- Akagawa M., Shigemitsu T., Suyama K. (2003). Production of hydrogen peroxide by polyphenols and polyphenol-rich beverages under quasi-physiological conditions. Biosci. Biotechnol. Biochem. 67, 2632–2640 10.1271/bbb.67.2632 [DOI] [PubMed] [Google Scholar]
- An J. H., Blackwell T. K. (2003). SKN-1 links C. elegans mesendodermal specification to a conserved oxidative stress response. Genes Dev. 17, 1882–1893 10.1101/gad.1107803 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ashburner M., Ball C. A., Blake J. A., Botstein D., Butler H., Cherry J. M., Davis A. P., Dolinski K., Dwight S. S., Eppig J. T., Harris M. A., Hill D. P., Issel-Tarver L., Kasarskis A., Lewis S., Matese J. C., Richardson J. E., Ringwald M., Rubin G. M., Sherlock G. (2000). Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Azen S. P., Roy S., Pike M. C., Casagrande J., Mishell D. R., Jr. (1977). A new procedure for the statistical evaluation of intrauterine contraception. Am. J. Obstet. Gynecol. 128, 329–335 [DOI] [PubMed] [Google Scholar]
- Benjamini Y., Hochberg Y. (1995). Controlling the false discovery rate – a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Stat. Methodol. 57, 289–300 [Google Scholar]
- Berdichevsky A., Viswanathan M., Horvitz H. R., Guarente L. (2006). C. elegans SIR-2.1 interacts with 14-3-3 proteins to activate DAF-16 and extend life span. Cell 125, 1165–1177 [DOI] [PubMed] [Google Scholar]
- Brenner S. (1974). The genetics of Caenorhabditis elegans. Genetics 77, 71–94 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Budovskaya Y. V., Wu K., Southworth L. K., Jiang M., Tedesco P., Johnson T. E., Kim S. K. (2008). An elt-3/elt-5/elt-6 GATA transcription circuit guides aging in C. elegans. Cell 134, 291–303 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coolon J. D., Jones K. L., Todd T. C., Carr B. C., Herman M. A. (2009). Caenorhabditis elegans genomic response to soil bacteria predicts environment-specific genetic effects on life history traits. PLoS Genet. 5, e1000503. 10.1371/journal.pgen.1000503 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dalma-Weiszhausz D. D., Warrington J., Tanimoto E. Y., Miyada C. G. (2006). The Affymetrix GeneChip platform: an overview. Meth. Enzymol. 410, 3–28 10.1016/S0076-6879(06)10001-4 [DOI] [PubMed] [Google Scholar]
- Davies S. K., Leroi A. M., Bundy J. G. (2012). Fluorodeoxyuridine affects the identification of metabolic responses to daf-2 status in Caenorhabditis elegans. Mech. Ageing Dev. 133, 46–49 10.1016/j.mad.2011.11.002 [DOI] [PubMed] [Google Scholar]
- Evans E. A., Kawli T., Tan M. W. (2008). Pseudomonas aeruginosa suppresses host immunity by activating the DAF-2 insulin-like signaling pathway in Caenorhabditis elegans. PLoS Pathog. 4, e1000175. 10.1371/journal.ppat.1000175 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher A. L., Lithgow G. J. (2006). The nuclear hormone receptor DAF-12 has opposing effects on Caenorhabditis elegans lifespan and regulates genes repressed in multiple long-lived worms. Aging Cell 5, 127–138 10.1111/j.1474-9726.2006.00203.x [DOI] [PubMed] [Google Scholar]
- Gems D., McElwee J. J. (2005). Broad spectrum detoxification: the major longevity assurance process regulated by insulin/IGF-1 signaling? Mech. Ageing Dev. 126, 381–387 10.1016/j.mad.2004.09.001 [DOI] [PubMed] [Google Scholar]
- Granato M., Schnabel H., Schnabel R. (1994a). Genesis of an organ: molecular analysis of the pha-1 gene. Development 120, 3005–3017 [DOI] [PubMed] [Google Scholar]
- Granato M., Schnabel H., Schnabel R. (1994b). pha-1, a selectable marker for gene transfer in C. elegans. Nucleic Acids Res. 22, 1762–1763 10.1093/nar/22.9.1762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grigoryev D. N., Ma S. F., Irizarry R. A., Ye S. Q., Quackenbush J., Garcia J. G. (2004). Orthologous gene-expression profiling in multi-species models: search for candidate genes. Genome Biol. 5, R34. 10.1186/gb-2004-5-5-r34 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halaschek-Wiener J., Khattra J. S., Mckay S., Pouzyrev A., Stott J. M., Yang G. S., Holt R. A., Jones S. J. M., Marra M. A., Brooks-Wilson A. R., Riddle D. L. (2005). Analysis of long-lived C. elegans daf-2 mutants using serial analysis of gene expression. Genome Res. 15, 603–615 10.1101/gr.3274805 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hars E. S., Qi H., Ryazanov A. G., Jin S., Cai L., Hu C., Liu L. F. (2007). Autophagy regulates ageing in C. elegans. Autophagy 3, 93–95 [DOI] [PubMed] [Google Scholar]
- Herrero J., Al-Shahrour F., Diaz-Uriarte R., Mateos A., Vaquerizas J. M., Santoyo J., Dopazo J. (2003). GEPAS: a web-based resource for microarray gene expression data analysis. Nucleic Acids Res. 31, 3461–3467 10.1093/nar/gkg591 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hosokawa H., Ishii N., Ishida H., Ichimori K., Nakazawa H., Suzuki K. (1994). Rapid accumulation of fluorescent material with aging in an oxygen-sensitive mutant mev-1 of Caenorhabditis elegans. Mech. Ageing Dev. 74, 161–170 10.1016/0047-6374(94)90099-X [DOI] [PubMed] [Google Scholar]
- Huang D. W., Sherman B. T., Tan Q., Kir J., Liu D., Bryant D., Guo Y., Stephens R., Baseler M. W., Lane H. C., Lempicki R. A. (2007). DAVID Bioinformatics Resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35, W169–W175 10.1093/nar/gkm415 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishii N., Fujii M., Hartman P. S., Tsuda M., Yasuda K., Senoo-Matsuda N., Yanase S., Ayusawa D., Suzuki K. (1998). A mutation in succinate dehydrogenase cytochrome b causes oxidative stress and ageing in nematodes. Nature 394, 694–697 10.1038/29331 [DOI] [PubMed] [Google Scholar]
- Kampkötter A., Timpel C., Zurawski R. F., Ruhl S., Chovolou Y., Proksch P., Watjen W. (2008). Increase of stress resistance and lifespan of Caenorhabditis elegans by quercetin. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 149, 314–323 10.1016/j.cbpb.2007.10.004 [DOI] [PubMed] [Google Scholar]
- Kanehisa M. (2002). The KEGG database. Novartis Found. Symp. 247, 91–101 [Discussion 101–103, 119–128, 244–152]. 10.1002/0470857897.ch8 [DOI] [PubMed] [Google Scholar]
- Kim D. H., Feinbaum R., Alloing G., Emerson F. E., Garsin D. A., Inoue H., Tanaka-Hino M., Hisamoto N., Matsumoto K., Tan M. W., Ausubel F. M. (2002). A conserved p38 MAP kinase pathway in Caenorhabditis elegans innate immunity. Science 297, 623–626 10.1126/science.1073759 [DOI] [PubMed] [Google Scholar]
- Kim S. K., Lund J., Kiraly M., Duke K., Jiang M., Stuart J. M., Eizinger A., Wylie B. N., Davidson G. S. (2001). A gene expression map for Caenorhabditis elegans. Science 293, 2087–2092 10.1126/science.1059805 [DOI] [PubMed] [Google Scholar]
- Kirkwood T. B. L., Austad S. N. (2000). Why do we age? Nature 408, 233–238 10.1038/35041682 [DOI] [PubMed] [Google Scholar]
- Kourtis N., Tavernarakis N. (2011). Cellular stress response pathways and ageing: intricate molecular relationships. EMBO J. 30, 2520–2531 10.1038/emboj.2011.162 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levine M. T., Eckert M. L., Begun D. J. (2011). Whole-genome expression plasticity across tropical and temperate Drosophila melanogaster populations from Eastern Australia. Mol. Biol. Evol. 28, 249–256 10.1093/molbev/msq197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Livak K. J., Schmittgen T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) Method. Methods 25, 402–408 10.1006/meth.2001.1262 [DOI] [PubMed] [Google Scholar]
- Mao C., Obeid L. M. (2008). Ceramidases: regulators of cellular responses mediated by ceramide, sphingosine, and sphingosine-1-phosphate. Biochim. Biophys. Acta 1781, 424–434 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCarthy D. J., Smyth G. K. (2009). Testing significance relative to a fold-change threshold is a TREAT. Bioinformatics 25, 765–771 10.1093/bioinformatics/btp053 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meinelt T., Paul A., Phan T. M., Zwirnmann E., Krüger A., Wienke A., Steinberg C. E. W. (2007). Reduction in vegetative growth of the water mold Saprolegnia parasitica (Coker) by humic substance of different qualities. Aquat. Toxicol. 83, 93–103 10.1016/j.aquatox.2007.03.013 [DOI] [PubMed] [Google Scholar]
- Melendez A., Talloczy Z., Seaman M., Eskelinen E. L., Hall D. H., Levine B. (2003). Autophagy genes are essential for dauer development and life-span extension in C. elegans. Science 301, 1387–1391 [DOI] [PubMed] [Google Scholar]
- Menzel R., Menzel S., Tiedt S., Kubsch G., Stößer R., Bährs H., Putschew A., Saul N., Steinberg C. E. W. (2011). Enrichment of humic material with hydroxybenzene moieties intensifies its physiological effects on the nematode Caenorhabditis elegans. Environ. Sci. Technol. 45, 8707–8715 10.1021/es2023237 [DOI] [PubMed] [Google Scholar]
- Menzel R., Rödel M., Kulas J., Steinberg C. E. W. (2005). CYP35: xenobiotically induced gene expression in the nematode Caenorhabditis elegans. Arch. Biochem. Biophys. 438, 93–102 10.1016/j.abb.2005.03.020 [DOI] [PubMed] [Google Scholar]
- Murphy C. T., Mccarroll S. A., Bargmann C. I., Fraser A., Kamath R. S., Ahringer J., Li H., Kenyon C. (2003). Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans. Nature 424, 277–283 10.1038/nature01789 [DOI] [PubMed] [Google Scholar]
- Pietsch K., Saul N., Chakrabarti S., Stürzenbaum S. R., Menzel R., Steinberg C. E. W. (2011). Hormetins, antioxidants and prooxidants: defining quercetin-, caffeic acid- and rosmarinic acid-mediated life extension in C. elegans. Biogerontology 12, 329–347 10.1007/s10522-011-9334-7 [DOI] [PubMed] [Google Scholar]
- Pietsch K., Saul N., Menzel R., Stürzenbaum S. R., Steinberg C. E. W. (2009). Quercetin mediated lifespan extension in Caenorhabditis elegans is modulated by age-1, daf-2, sek-1 and unc-43. Biogerontology 10, 565–578 10.1007/s10522-008-9199-6 [DOI] [PubMed] [Google Scholar]
- Pietsch K., Saul N., Swain S., Menzel R., Steinberg C. E. W., Stürzenbaum S. R. (2012). Ruling aging with polyphenols: meta-analysis of global transcriptional responses reveals conserved genetic pathways as mediators for quercetin and tannic acid effected longevity in C. elegans. Front. Genet. 3, 48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rainer J., Sanchez-Cabo F., Stocker G., Sturn A., Trajanoski Z. (2006). CARMAweb: comprehensive R- and bioconductor-based web service for microarray data analysis. Nucleic Acids Res. 34, W498–W503 10.1093/nar/gkl038 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rhee S. Y., Wood V., Dolinski K., Draghici S. (2008). Use and misuse of the gene ontology annotations. Nat. Rev. Genet. 9, 509–515 10.1038/nrg2363 [DOI] [PubMed] [Google Scholar]
- Saul N., Pietsch K., Menzel R., Stürzenbaum S. R., Steinberg C. E. W. (2009). Catechin induced longevity in C. elegans: from key regulator genes to disposable soma. Mech. Ageing Dev. 130, 477–486 10.1016/j.mad.2009.05.005 [DOI] [PubMed] [Google Scholar]
- Saul N., Pietsch K., Menzel R., Stürzenbaum S. R., Steinberg C. E. W. (2010). The longevity effect of tannic acid in Caenorhabditis elegans: disposable soma meets hormesis. J. Gerontol. A Biol. Sci. Med. Sci. 65, 626–635 10.1093/gerona/glq051 [DOI] [PubMed] [Google Scholar]
- Senoo-Matsuda N., Yasuda K., Tsuda M., Ohkubo T., Yoshimura S., Nakazawa H., Hartman P. S., Ishii N. (2001). A defect in the cytochrome b large subunit in complex II causes both superoxide anion overproduction and abnormal energy metabolism in Caenorhabditis elegans. J. Biol. Chem. 276, 41553–41558 10.1074/jbc.M104718200 [DOI] [PubMed] [Google Scholar]
- Shaw W. M., Luo S., Landis J., Ashraf J., Murphy C. T. (2007). The C. elegans TGF-beta dauer pathway regulates longevity via insulin signaling. Curr. Biol. 17, 1635–1645 10.1016/j.cub.2007.08.058 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shin H., Lee H., Fejes A. P., Baillie D. L., Koo H. S., Jones S. J. (2011). Gene expression profiling of oxidative stress response of C. elegans aging defective AMPK mutants using massively parallel transcriptome sequencing. BMC Res. Notes 4, 34. 10.1186/1756-0500-4-34 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Solomon A., Bandhakavi S., Jabbar S., Shah R., Beitel G. J., Morimoto R. I. (2004). Caenorhabditis elegans OSR-1 regulates behavioral and physiological responses to hyperosmotic environments. Genetics 167, 161–170 10.1534/genetics.167.1.161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinberg C. E. W., Saul N., Pietsch K., Meinelt T., Rienau S., Menzel R. (2007). Dissolved humic substances facilitate fish life in extreme aquatic environments and have the potential to extend lifespan of Caenorhabditis elegans. Ann. Environ. Sci. 1, 81–90 [Google Scholar]
- Sulston J. E., Hodgkin J. (1988). “Methods,” in The nematode Caenorhabditis elegans, ed. Wood W. B. (Cold Spring Harbor: CSHL Press; ), 587–606 [Google Scholar]
- Swain S., Wren J. F., Stürzenbaum S. R., Kille P., Morgan A. J., Jager T., Jonker M. J., Hankard P. K., Svendsen C., Owen J., Hedley B. A., Blaxter M., Spurgeon D. J. (2010). Linking toxicant physiological mode of action with induced gene expression changes in Caenorhabditis elegans. BMC Syst. Biol. 4, 32. 10.1186/1752-0509-4-32 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka-Hino M., Sagasti A., Hisamoto N., Kawasaki M., Nakano S., Ninomiya-Tsuji J., Bargmann C. I., Matsumoto K. (2002). SEK-1 MAPKK mediates Ca2+ signaling to determine neuronal asymmetric development in Caenorhabditis elegans. EMBO Rep. 3, 56–62 10.1093/embo-reports/kvf001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tissenbaum H. A., Guarente L. (2001). Increased dosage of a sir-2 gene extends lifespan in Caenorhabditis elegans. Nature 410, 227–230 10.1038/35065638 [DOI] [PubMed] [Google Scholar]
- Troemel E. R., Chu S. W., Reinke V., Lee S. S., Ausubel F. M., Kim D. H. (2006). p38 MAPK regulates expression of immune response genes and contributes to longevity in C. elegans. PLoS Genet. 2, e183. 10.1371/journal.pgen.0020183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Straalen N. M., Feder M. E. (2012). Ecological and evolutionary functional genomics-how can it contribute to the risk assessment of chemicals? Environ. Sci. Technol. 46, 3–9 10.1021/es204242a [DOI] [PubMed] [Google Scholar]
- Viswanathan M., Kim S. K., Berdichevsky A., Guarente L. (2005). A role for SIR-2.1 regulation of ER stress response genes in determining C. elegans life span. Dev. Cell 9, 605–615 10.1016/j.devcel.2005.09.017 [DOI] [PubMed] [Google Scholar]
- Wiegant F. A. C., Surinova S., Ytsma E., Langelaar-Makkinje M., Wikman G., Post J. A. (2009). Plant adaptogens increase lifespan and stress resistance in C. elegans. Biogerontology 10, 27–42 10.1007/s10522-008-9151-9 [DOI] [PubMed] [Google Scholar]
- Wilson M. A., Shukitt-Hale B., Kalt W., Ingram D. K., Joseph J. A., Wolkow C. A. (2006). Blueberry polyphenols increase lifespan and thermotolerance in Caenorhabditis elegans. Aging Cell 5, 59–68 10.1111/j.1474-9726.2006.00192.x [DOI] [PMC free article] [PubMed] [Google Scholar]