Abstract
Gene–lifestyle interactions have been suggested to contribute to the development of type 2 diabetes. Glucose levels 2 h after a standard 75-g glucose challenge are used to diagnose diabetes and are associated with both genetic and lifestyle factors. However, whether these factors interact to determine 2-h glucose levels is unknown. We meta-analyzed single nucleotide polymorphism (SNP) × BMI and SNP × physical activity (PA) interaction regression models for five SNPs previously associated with 2-h glucose levels from up to 22 studies comprising 54,884 individuals without diabetes. PA levels were dichotomized, with individuals below the first quintile classified as inactive (20%) and the remainder as active (80%). BMI was considered a continuous trait. Inactive individuals had higher 2-h glucose levels than active individuals (β = 0.22 mmol/L [95% CI 0.13–0.31], P = 1.63 × 10−6). All SNPs were associated with 2-h glucose (β = 0.06–0.12 mmol/allele, P ≤ 1.53 × 10−7), but no significant interactions were found with PA (P > 0.18) or BMI (P ≥ 0.04). In this large study of gene–lifestyle interaction, we observed no interactions between genetic and lifestyle factors, both of which were associated with 2-h glucose. It is perhaps unlikely that top loci from genome-wide association studies will exhibit strong subgroup-specific effects, and may not, therefore, make the best candidates for the study of interactions.
Glucose levels 2 h after a 75-g glucose challenge are used to diagnose diabetes and are associated with cardiovascular morbidity and mortality even below the diabetic threshold (1). A large number of type 2 diabetes–associated genetic variants have now been identified (2), and recent genome-wide meta-analyses identified five loci that were associated with postchallenge glucose at genome-wide levels of significance (3). Previously identified single nucleotide polymorphisms (SNPs) in TCF7L2 and GCKR were associated with 2-h glucose levels, as were newly identified loci in ADCY5, GIPR, and VPS13C. Risk alleles at each of these loci conferred elevated 2-h glucose levels with effect sizes ranging from 0.07 to 0.11 mmol/L per allele (3), although with some heterogeneity.
Age, BMI, and physical inactivity are all associated with glycemia and are key risk factors for type 2 diabetes (4–6). Glucose levels at 2 h appear more susceptible to age- and lifestyle-mediated increases than fasting glucose levels. For example, physical activity (PA) levels have been shown to be inversely associated with 2-h glucose but not with fasting glucose (7,8). Differences in 2-h glucose between individuals at either end of the PA spectrum are appreciable, with the most active individuals having a mean 2-h glucose level ∼1 mmol/L lower than those with low PA levels (7). Furthermore, lifestyle intervention trials including prescribed PA have been effective in decreasing the incidence of diabetes in individuals with impaired glucose tolerance at baseline (9,10). However, it is unclear whether these responses to PA are homogenous among those with genetically conferred elevations in 2-h glucose levels or whether genetic effects are similar across lifestyle strata. Identification of gene–lifestyle interactions will offer valuable insight into the etiologic processes leading to disease and the biologic pathways by which lifestyle modification can reduce the risk of diabetes.
Although gene–lifestyle interactions are suggested as being important in the etiology of type 2 diabetes, few consistently replicated examples have been identified (11) and methodologic difficulties limit the opportunity for literature-based meta-analyses (12). The association of 2-h glucose with lifestyle and genetic factors makes it a good trait for the study of gene–lifestyle interaction. Furthermore, the heterogeneity observed in the association between SNPs and 2-h glucose (3) is potentially attributable to factors such as gene–lifestyle interaction. Therefore, we investigated the presence of gene–lifestyle interactions at these five 2-h glucose-associated loci (in or near GCKR, ADCY5, TCF7L2, VPS13C, and GIPR) by meta-analyzing SNP×PA and SNP×BMI interactions on 2-h glucose in up to 54,884 individuals from 22 studies.
RESEARCH DESIGN AND METHODS
Participating cohort characteristics.
We meta-analyzed results from up to 22 Meta-Analyses of Glucose and Insulin Related Traits Consortium (MAGIC) studies (3) comprising up to 54,884 individuals. Study descriptives are detailed in Supplementary Table 1. Participants with known diabetes, those with fasting glucose ≥ 7 mmol/L, and individuals with a BMI <18.5 kg/m2 were excluded. All data were cross-sectional except for Atherosclerosis Risk in Communities Study (ARIC) where PA data were available at the visit ∼3 years before 2-h glucose measurement.
Lifestyle exposure classification.
Study-specific details of the measurement of PA are in Supplementary Table 1. Where a quantitative measure of PA was available, individuals below the first quintile were classified as inactive and the remainder as active (i.e., 20% inactive and 80% active). In studies where PA data were categoric, the proportion of inactive individuals was dependent on the questionnaire used and reported in Supplementary Table 1. Inactive individuals were coded as 0 and active individuals as 1 in the analyses. BMI was treated as a continuous variable in the primary analyses.
Genotyping and statistical analysis.
Genotyping methods are reported in Supplementary Table 1 and have been described in detail previously (3). Analysts from each study performed study-level analyses and submitted summary statistics to the meta-analysis group. We ran linear regression models testing the association of each SNP with 2-h glucose, adjusted for age, sex, fasting glucose, BMI, and PA (as a dichotomous variable), and any necessary study-specific variables. We also examined the association of each SNP with BMI, adjusted for age and sex. Given our exclusion of individuals on the basis of glycemia, we sought replication of BMI associations by lookup of those SNPs in previous Genetic Investigation of ANthropometric Traits (GIANT) meta-analyses (13). To investigate SNP×PA and SNP×BMI interactions, each study included interaction terms (e.g., SNP×PA) in the models and reported the estimated interaction effect and standard error. The interaction effect estimates were combined using inverse variance-weighted meta-analysis. Studies with genotypes extracted from genome-wide SNP arrays reported interaction terms with robust standard errors and are included in the meta-analysis as such. Additive genetic models were applied.
We performed meta-analyses using a fixed-effects, inverse-variance weighted approach via the metan command in Stata SE-11.1 software (StataCorp LP, College Station, TX) to study SNP main effects on 2-h glucose or BMI. To study interaction between SNPs and PA or BMI, as well as the association between PA and BMI with 2-h glucose, we used random effects meta-analyses to account for potential heterogeneity introduced by factors such as PA differences among studies (Supplementary Table 1).
RESULTS
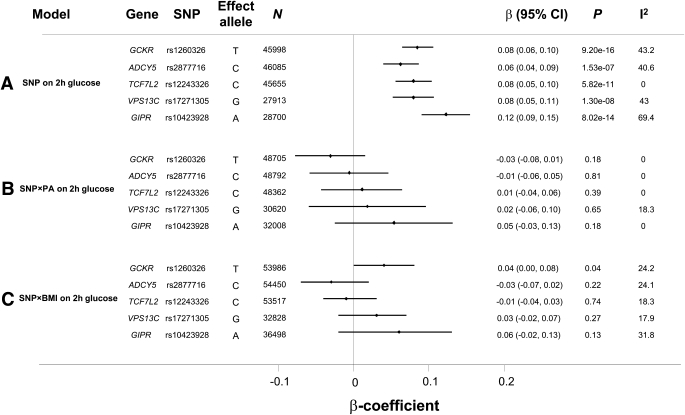
Study descriptives are reported in Supplementary Table 1. Inactive individuals had a higher 2-h glucose (β = 0.22 mmol/L [95% CI 0.13–0.31], P = 1.63 × 10−6) and BMI (β = 0.73 kg/m2 [0.51–0.95], P = 1.42 × 10−10) than active individuals. Higher BMI was also associated with higher 2-h glucose levels (β per 1 kg/m2 = 0.086 mmol/L [0.08–0.10], P = 1.04 × 10−47). SNP effects were consistent with those reported previously in overlapping studies (Fig. 1A) (3).
FIG. 1.
A: Effect of SNP is shown on 2-h glucose. The β-coefficient is the magnitude of the observed association. B: Shows the SNP×PA interaction effect in which the β-coefficient is the difference in SNP association effect between inactive and active individuals. Inactive individuals were coded as 0 and active individuals a 1; therefore, a value of 0 for the interaction coefficient reflects equivalent SNP effect in inactive and active strata, whereas a positive value reflects a larger SNP effect in active individuals. C: The SNP×BMI interaction is shown. Here, the β-coefficient is the difference in SNP effect per 10 kg/m2 difference in BMI. A positive value reflects a larger SNP effect in those with higher BMI. The 2-h glucose–raising allele in A is always the effect allele.
SNP×PA and SNP×BMI interactions on 2-h glucose.
Figure 1B shows the absence of any difference in SNP effect on 2-h glucose between inactive and active individuals (SNPxPA P ≥ 0.18 for interaction). Likewise, we did not observe any significant interaction effects when analyses were limited to those studies showing association between PA and 2-h glucose (SNPxPA P ≥ 0.1 for interaction). Figure 1C shows the difference in SNP effect on 2-h glucose per 10 kg/m2. Again, no statistically significant interaction effects were observed after correction for multiple testing (five tests for each hypothesis: α = 0.01), although rs1260326 in GCKR reached nominal levels of statistical significance (albeit with very small interaction effects). BMI-stratified results for rs1260326 showed that SNP effects were largest in the 30 to 34.9 kg/m2 group (Supplementary Fig. 1), although few individuals at >35 kg/m2 makes the smaller effect in this stratum difficult to interpret.
Association of SNPs with BMI.
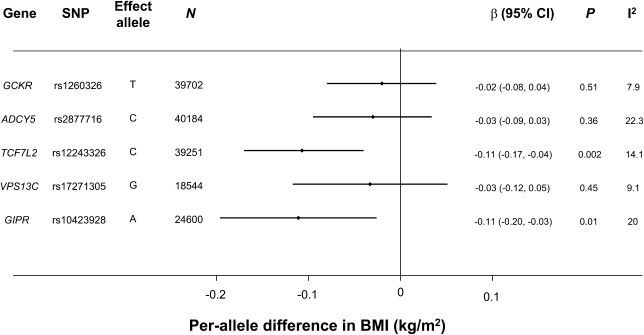
As can be seen from Fig. 2, TCF7L2 rs12243326 and GIPR rs10423928 were both associated with BMI: the alleles associated with increased 2-h glucose were associated with lower BMI. The TCF7L2 and GIPR SNPs were both associated with a 0.11 kg/m2 lower BMI per allele (95% CI −0.17 to −0.04 and −0.20 to −0.03, respectively; Fig. 2). These findings were directionally consistent with those from previous meta-analyses by the GIANT consortium (13) (rs12243326 P = 5.7 × 10−4; rs10423928 P = 1.9 × 10−6).
FIG. 2.
The SNP association with BMI is shown. The 2-h glucose–raising allele from Fig. 1A is shown as the effect allele.
DISCUSSION
Each of the gene variants investigated in the current study was robustly associated with 2-h glucose levels, as reported previously in overlapping studies (3). However, we observed no difference in effect of the gene variants studied among PA groups or with increasing BMI.
Previous studies have reported gene–lifestyle interactions, although often based on small sample sizes without independent replication (11). In light of the small effect sizes of most complex disease-associated SNPs, large sample sizes are important to investigate interactions (14). However, despite the large sample size in the current study, no significant interactions were observed between 2-h glucose-associated SNPs and established lifestyle correlates. Notably, however, in 6 of the 10 interaction meta-analyses we performed (i.e., 5 SNPs and interaction with PA or BMI), at least 1 individual study would have shown a significant interaction had it been studied in isolation. However, had we considered studies individually, for ∼200 tests of interaction we would have observed only nine interactions at P < 0.05, spread among a range of studies. We suggest that these associations reflect type I errors. Such a finding further supports our use of large sample sizes or independent replication to reduce the potential for type I error.
Although the variety of subjective measures and dichotomous classification of PA is an important limitation of the current study, inactive individuals had a higher 2-h glucose and a higher BMI than active individuals, suggesting the validity of our PA classification. Another factor may contribute to the absence of interactions: the SNPs we selected arose as top SNPs associated with 2-h glucose levels from a genome-wide meta-analysis (3). Although heterogeneity of associations was observed for ADCY5, TCF7L2, and GIPR (3), these SNPs had the strongest association P values in the genome by virtue of their effect size relative to the variation in effect size among samples. One may not, therefore, expect significant variation in genetic effect between subgroups of the population. A similar approach to ours was recently used to investigate interactions between breast cancer–associated genes and risk-altering lifestyle exposures and also failed to detect significant interactions between them (15), although there was a suggestion that a physically active lifestyle attenuated genetic predisposition to obesity (16).
Although it has been suggested that the search for interactions should be informed by biologic plausibility (11), experience from the study of genetic main effects, where hypothesis-generating discovery approaches revolutionized the field (2), suggests that such an approach, not limited to those SNPs with extremely significant main effects, may be valuable in detecting gene–lifestyle interactions. Such approaches have been proposed and efforts are underway, but whether current analytic methods will yield success in the genome-wide search for gene–lifestyle interactions remains unclear.
Data from large-scale trials, such as the Finnish Diabetes Prevention Study (DPS) and Diabetes Prevention Program (DPP), have shown suggestions of differential response to intervention by genotype (17–22), although not always reaching statistical significance for interaction (17–19). However, lifestyle interventions in such studies often contain numerous lifestyle modifications, making interpretation of any interaction difficult, whereas large-scale genotype-stratified lifestyle intervention trials are not feasible. Therefore, prospective nested approaches will likely offer the most efficient approach for the study of gene–lifestyle interaction (23), allowing standardized measures of lifestyle at baseline and also the opportunity to study large numbers of individuals. Refined and standardized lifestyle exposure measurement will also represent a valuable alternative to straightforward increases in sample size (24).
Variants in TCF7L2 have previously been associated with diabetes (2,17) and a number of related traits (3,25). The diabetogenic, glucose-raising allele was previously associated with lower BMI, although not conclusively (26), and principally in individuals with diabetes (27,28). Here, we replicate this association in a larger sample size of participants without diabetes, where the glucose-raising allele at rs12243326 was associated with a 0.11 kg/m2–lower BMI (Fig. 2). Similarly, we report that the glucose-raising allele at GIPR rs10423928 is associated with lower BMI (−0.11 kg/m2 per allele). Lookup results from the GIANT consortium suggest that these associations are unlikely to arise from ascertainment bias.
Although the association with BMI highlights a genetic predisposition on BMI and the risk of confusing gene–gene and gene–environment interactions, the small proportion of variance in BMI explained by such SNPs is likely to limit the effect of this concern in our study. Because BMI is a major risk factor for diabetes and has a strong positive association with 2-h glucose, it seems counterintuitive that 2-h glucose-raising alleles at TCF7L2 and GIPR are associated with lower BMI and highlights the etiologic complexity of type 2 diabetes.
In conclusion, in our study of up to 54,884 individuals from 22 studies, we found no evidence of gene–lifestyle interaction among the variants studied. This was despite the clear association of 2-h glucose with PA, BMI, and genetic exposures. Although the descriptive epidemiology of diabetes suggests an influence of gene–lifestyle interaction in its etiology, our study finds no evidence to that effect for SNPs known to be associated with 2-h glucose. Further, our study supports the use of large-scale analyses to robustly investigate gene–lifestyle interaction. In future, hypothesis-generating approaches may offer a valuable opportunity to detect gene–lifestyle interactions in type 2 diabetes and related traits.
Supplementary Material
ACKNOWLEDGMENTS
A complete list of disclosures and acknowledgments is included in the Supplementary Data online.
I.B. owns stock in GlaxoSmithKline and Incyte. No other potential conflicts of interest relevant to this article were reported.
R.A.S., A.Y.C., N.G., A.K.M., M.-F.H., A.T., O.P., J.B.M., E.I., I.B., J.C.F., P.W.F., J.D., N.J.W., and C.La. wrote the manuscript. R.A.S. and A.Y.C. were involved in management of the project and/or involved studies and in the project design and performed statistical analyses. N.G., A.K.M., M.F.-H., D.S., A.Y., D.B., Z.K., D.M., L.J.R.-T., H.M.S., V.La., S.G., and T.M.T. performed statistical analyses. N.B.-N., C.L.-M., and K.R. were involved in management of the project and/or involved studies and in genotyping and performed statistical analyses. N.L.G., A.U.J., and I.P. were involved in genotyping and performed statistical analyses. T.T., S.Be., S.R.B., S.Br., N.F.D., U.E., R.J.F.L., J.S.P., D.S.S., E.I., R.M.W., O.P., J.C.F., and C.La. were involved in management of the project and/or involved studies and in the project design. E.Bo., G.M., M.O.G., and C.H.S. contributed to data collection and phenotyping, were involved in genotyping, and performed statistical analyses. L.L.B., S.J.B., H.C., M.R.E., A.J., P.K., G.L., M.A.M., A.R.W., and W.X. were involved in genotyping. A.T., M.A., O.D.C., J.S.E., J.G., F.B.H., P.M.-V., F.P., and A.S. contributed to data collection and phenotyping. P.S.C. and M.S. contributed to data collection and phenotyping and were involved in genotyping. D.E.A., E.Br., Y.-D.I.C., F.S.C., D.J.C., N.G.F., T.H., A.H., T.J., C.Le., V.Ly., M.M., K.E.N., F.R., D.R., P.S., G.W., D.R.W., M.B., and M.W. were involved in management of the project and/or involved studies. E.M.D., P.F., L.G., W.H.L.K., I.B., and P.W.F. were involved in management of the project and/or involved studies and in the project design, contributed to data collection and phenotyping, and were involved in genotyping. C.S.F., G.H., I.J., P.E.H.S., G.H.W., J.W.H., K.A.J., A.A.S., C.C., B.M.P., J.I.R., L.F., and P.V. were involved in management of the project and/or involved studies and in data collection and phenotyping. B.I., M.Ki., J.K., M.L., and J.B.M. were involved in management of the project and/or involved studies and in the project design, data collection, and phenotyping. M.Ku. was involved in management of the project and/or involved studies and in the project design, data collection, and phenotyping, and genotyping. J.D. was involved in management of the project and/or involved studies, in project design, and in genotyping, and performed statistical analyses. N.J.W. was involved in management of the project and/or involved studies, in project design, and in data collection and phenotyping. R.A.S. is the guarantor of this work and, as such, had full access to all of the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Footnotes
This article contains Supplementary Data online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db11-0973/-/DC1.
REFERENCES
- 1.Qiao Q, Tuomilehto J, Borch-Johnsen K. Post-challenge hyperglycaemia is associated with premature death and macrovascular complications. Diabetologia 2003;46(Suppl 1):M17–M21 [DOI] [PubMed] [Google Scholar]
- 2.Voight BF, Scott LJ, Steinthorsdottir V, et al. MAGIC investigators. GIANT Consortium Twelve type 2 diabetes susceptibility loci identified through large-scale association analysis. Nat Genet 2010;42:579–589 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Saxena R, Hivert MF, Langenberg C, et al. GIANT consortium. MAGIC investigators Genetic variation in GIPR influences the glucose and insulin responses to an oral glucose challenge. Nat Genet 2010;42:142–148 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Narayan KM, Boyle JP, Thompson TJ, Gregg EW, Williamson DF. Effect of BMI on lifetime risk for diabetes in the U.S. Diabetes Care 2007;30:1562–1566 [DOI] [PubMed] [Google Scholar]
- 5.Manson JE, Rimm EB, Stampfer MJ, et al. Physical activity and incidence of non-insulin-dependent diabetes mellitus in women. Lancet 1991;338:774–778 [DOI] [PubMed] [Google Scholar]
- 6.DECODE Study Group Age- and sex-specific prevalences of diabetes and impaired glucose regulation in 13 European cohorts. Diabetes Care 2003;26:61–69 [DOI] [PubMed] [Google Scholar]
- 7.Wareham NJ, Wong MY, Day NE. Glucose intolerance and physical inactivity: the relative importance of low habitual energy expenditure and cardiorespiratory fitness. Am J Epidemiol 2000;152:132–139 [DOI] [PubMed] [Google Scholar]
- 8.Healy GN, Dunstan DW, Shaw JE, Zimmet PZ, Owen N. Beneficial associations of physical activity with 2-h but not fasting blood glucose in Australian adults: the AusDiab study. Diabetes Care 2006;29:2598–2604 [DOI] [PubMed] [Google Scholar]
- 9.Knowler WC, Barrett-Connor E, Fowler SE, et al. Diabetes Prevention Program Research Group Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N Engl J Med 2002;346:393–403 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tuomilehto J, Lindström J, Eriksson JG, et al. Finnish Diabetes Prevention Study Group Prevention of type 2 diabetes mellitus by changes in lifestyle among subjects with impaired glucose tolerance. N Engl J Med 2001;344:1343–1350 [DOI] [PubMed] [Google Scholar]
- 11.Franks PW, Mesa JL, Harding AH, Wareham NJ. Gene-lifestyle interaction on risk of type 2 diabetes. Nutr Metab Cardiovasc Dis 2007;17:104–124 [DOI] [PubMed] [Google Scholar]
- 12.Palla L, Higgins JP, Wareham NJ, Sharp SJ. Challenges in the use of literature-based meta-analysis to examine gene-environment interactions. Am J Epidemiol 2010;171:1225–1232 [DOI] [PubMed] [Google Scholar]
- 13.Speliotes EK, Willer CJ, Berndt SI, et al. MAGIC. Procardis Consortium Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat Genet 2010;42:937–948 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Smith PG, Day NE. The design of case-control studies: the influence of confounding and interaction effects. Int J Epidemiol 1984;13:356–365 [DOI] [PubMed] [Google Scholar]
- 15.Travis RC, Reeves GK, Green J, et al. Million Women Study Collaborators Gene-environment interactions in 7610 women with breast cancer: prospective evidence from the Million Women Study. Lancet 2010;375:2143–2151 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Li S, Zhao JH, Luan J, et al. Physical activity attenuates the genetic predisposition to obesity in 20,000 men and women from EPIC-Norfolk prospective population study. PLoS Med 2010;7: e1000332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Florez JC, Jablonski KA, Bayley N, et al. Diabetes Prevention Program Research Group TCF7L2 polymorphisms and progression to diabetes in the Diabetes Prevention Program. N Engl J Med 2006;355:241–250 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jablonski KA, McAteer JB, de Bakker PI, et al. Diabetes Prevention Program Research Group Common variants in 40 genes assessed for diabetes incidence and response to metformin and lifestyle intervention in the diabetes prevention program. Diabetes 2010;59:2672–2681 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Florez JC, Jablonski KA, Sun MW, et al. Diabetes Prevention Program Research Group Effects of the type 2 diabetes-associated PPARG P12A polymorphism on progression to diabetes and response to troglitazone. J Clin Endocrinol Metab 2007;92:1502–1509 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Franks PW, Jablonski KA, Delahanty L, et al. Diabetes Prevention Program Research Group The Pro12Ala variant at the peroxisome proliferator-activated receptor gamma gene and change in obesity-related traits in the Diabetes Prevention Program. Diabetologia 2007;50:2451–2460 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kilpeläinen TO, Lakka TA, Laaksonen DE, et al. Finnish Diabetes Prevention Study Group Interaction of single nucleotide polymorphisms in ADRB2, ADRB3, TNF, IL6, IGF1R, LIPC, LEPR, and GHRL with physical activity on the risk of type 2 diabetes mellitus and changes in characteristics of the metabolic syndrome: The Finnish Diabetes Prevention Study. Metabolism 2008;57:428–436 [DOI] [PubMed] [Google Scholar]
- 22.Todorova B, Kubaszek A, Pihlajamäki J, et al. Finnish Diabetes Prevention Study The G-250A promoter polymorphism of the hepatic lipase gene predicts the conversion from impaired glucose tolerance to type 2 diabetes mellitus: the Finnish Diabetes Prevention Study. J Clin Endocrinol Metab 2004;89:2019–2023 [DOI] [PubMed] [Google Scholar]
- 23.Manolio TA, Bailey-Wilson JE, Collins FS. Genes, environment and the value of prospective cohort studies. Nat Rev Genet 2006;7:812–820 [DOI] [PubMed] [Google Scholar]
- 24.Wong MY, Day NE, Luan JA, Chan KP, Wareham NJ. The detection of gene-environment interaction for continuous traits: should we deal with measurement error by bigger studies or better measurement? Int J Epidemiol 2003;32:51–57 [DOI] [PubMed] [Google Scholar]
- 25.Dupuis J, Langenberg C, Prokopenko I, et al. DIAGRAM Consortium. GIANT Consortium. Global BPgen Consortium. Anders Hamsten on behalf of Procardis Consortium. MAGIC investigators New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet 2010;42:105–116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Stolerman ES, Manning AK, McAteer JB, et al. TCF7L2 variants are associated with increased proinsulin/insulin ratios but not obesity traits in the Framingham Heart Study. Diabetologia 2009;52:614–620 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lyssenko V, Lupi R, Marchetti P, et al. Mechanisms by which common variants in the TCF7L2 gene increase risk of type 2 diabetes. J Clin Invest 2007;117:2155–2163 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Helgason A, Pálsson S, Thorleifsson G, et al. Refining the impact of TCF7L2 gene variants on type 2 diabetes and adaptive evolution. Nat Genet 2007;39:218–225 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.