Abstract
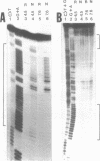
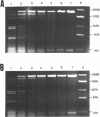
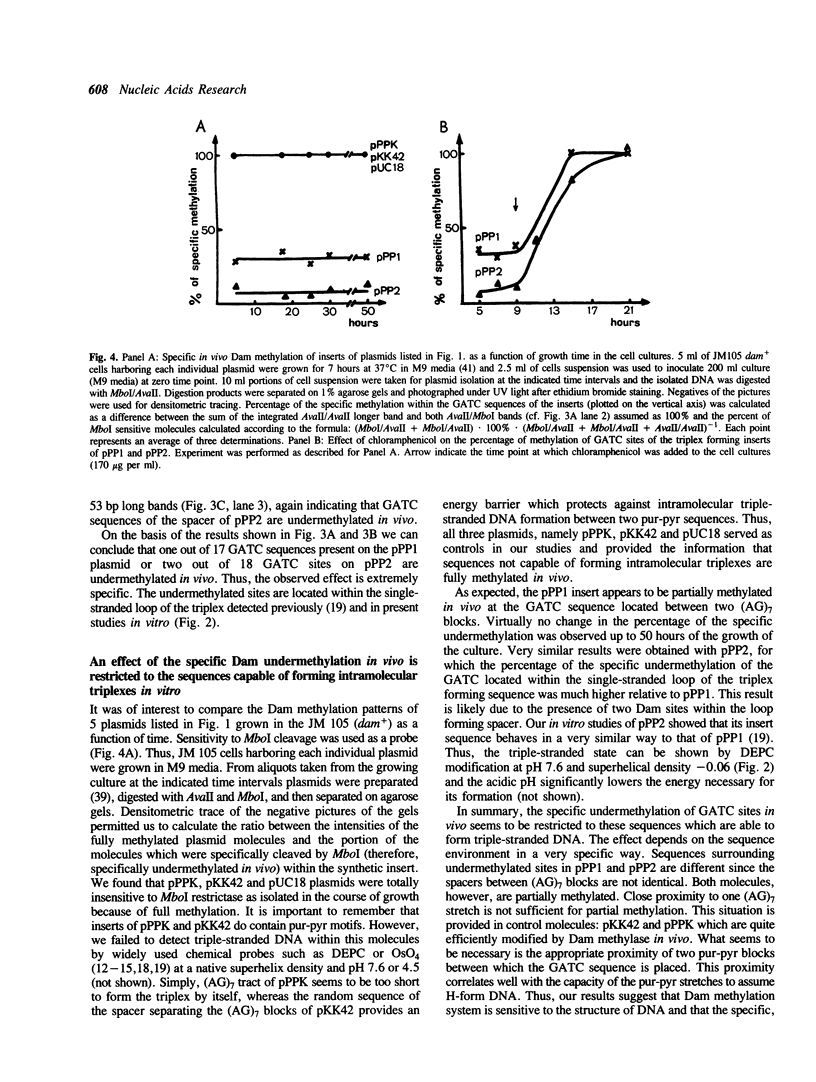
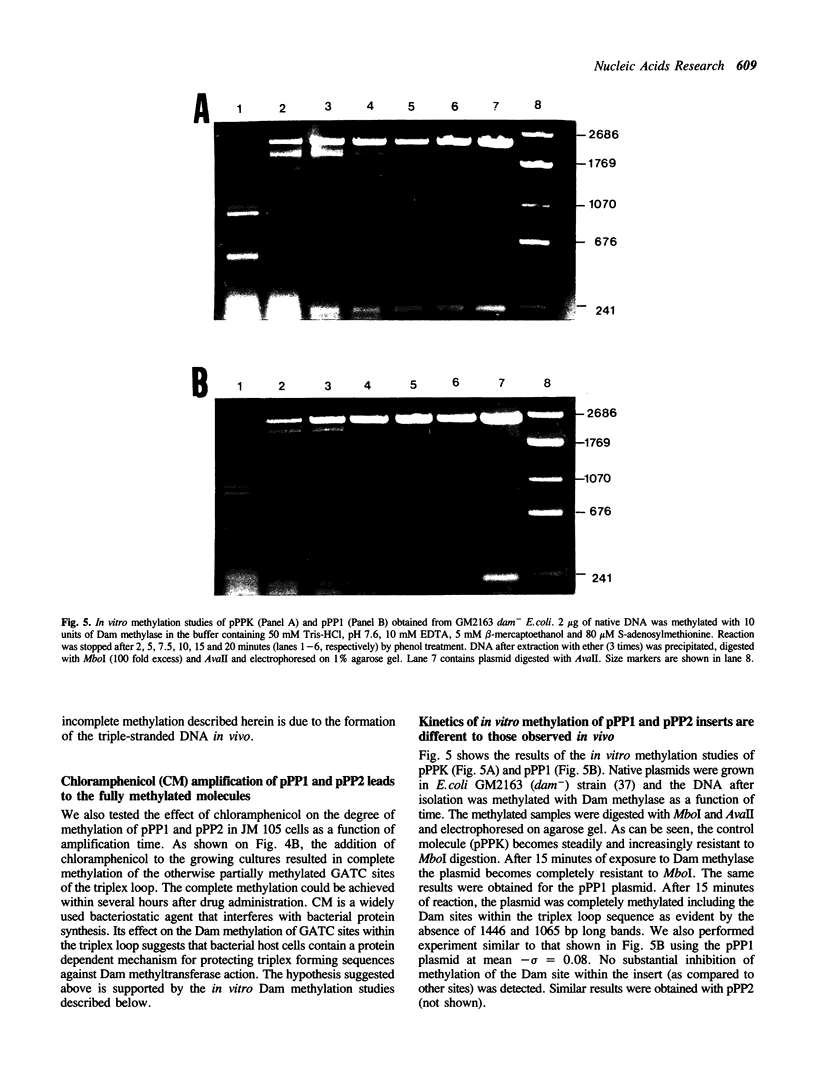
Several derivatives of pUC18 plasmid were constructed that contained oligopurine-oligopyrimidine (pur-pyr) motifs surrounded by Dam methylation sites. Inserts of two of the molecules (pPP1 and pPP2) were able to adopt the triple-stranded conformation in vitro and show in vivo a remarkable undermethylation of specific sites when grown in JM105 dam+ strain. Mapping experiments revealed that undermethylated GATC sequences were located exclusively within the single-stranded loop region of the sequence involved in H-DNA formation. Control molecules which either contained the pur-pyr tracts (pPPK and pKK42) or not (pUC18) and were not able to form the triple-stranded conformation were found to be normally methylated by the dam gene product in vivo. Location of GATC within the triplex forming sequence seems to be a prerequisite for achieving its in vivo undermethylation. E.coli host factors are involved in the observed phenomenon. This has been deduced from the fact that the undermethylated state of pPP1 and pPP2 does not depend on the phase of growth of host cells and is steadily maintained up to 50 hours, whereas the kinetics of Dam methylation in vitro of sites located within the triplex loop does not differ substantially from the kinetics of methylation of other sites on the vector. Full methylation can be readily achieved in vitro. Additional factor(s) that operate in vivo to control the undermethylated state are most likely proteins since the observed effect can be suppressed by chloramphenicol administration to the cell cultures.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christophe D., Cabrer B., Bacolla A., Targovnik H., Pohl V., Vassart G. An unusually long poly(purine)-poly(pyrimidine) sequence is located upstream from the human thyroglobulin gene. Nucleic Acids Res. 1985 Jul 25;13(14):5127–5144. doi: 10.1093/nar/13.14.5127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collier D. A., Griffin J. A., Wells R. D. Non-B right-handed DNA conformations of homopurine.homopyrimidine sequences in the murine immunoglobulin C alpha switch region. J Biol Chem. 1988 May 25;263(15):7397–7405. [PubMed] [Google Scholar]
- Davis M. M., Kim S. K., Hood L. E. DNA sequences mediating class switching in alpha-immunoglobulins. Science. 1980 Sep 19;209(4463):1360–1365. doi: 10.1126/science.6774415. [DOI] [PubMed] [Google Scholar]
- Dybvig K., Clark C. D., Aliperti G., Schlesinger M. J. A chicken repetitive DNA sequence that is highly sensitive to single-strand specific endonucleases. Nucleic Acids Res. 1983 Dec 10;11(23):8495–8508. doi: 10.1093/nar/11.23.8495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evans T., Efstratiadis A. Sequence-dependent S1 nuclease hypersensitivity of a heteronomous DNA duplex. J Biol Chem. 1986 Nov 5;261(31):14771–14780. [PubMed] [Google Scholar]
- Evans T., Schon E., Gora-Maslak G., Patterson J., Efstratiadis A. S1-hypersensitive sites in eukaryotic promoter regions. Nucleic Acids Res. 1984 Nov 12;12(21):8043–8058. doi: 10.1093/nar/12.21.8043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fowler R. F., Skinner D. M. Eukaryotic DNA diverges at a long and complex pyrimidine.purine tract that can adopt altered conformations. J Biol Chem. 1986 Jul 5;261(19):8994–9001. [PubMed] [Google Scholar]
- Glikin G. C., Gargiulo G., Rena-Descalzi L., Worcel A. Escherichia coli single-strand binding protein stabilizes specific denatured sites in superhelical DNA. Nature. 1983 Jun 30;303(5920):770–774. doi: 10.1038/303770a0. [DOI] [PubMed] [Google Scholar]
- Hanvey J. C., Klysik J., Wells R. D. Influence of DNA sequence on the formation of non-B right-handed helices in oligopurine.oligopyrimidine inserts in plasmids. J Biol Chem. 1988 May 25;263(15):7386–7396. [PubMed] [Google Scholar]
- Hanvey J. C., Shimizu M., Wells R. D. Intramolecular DNA triplexes in supercoiled plasmids. Proc Natl Acad Sci U S A. 1988 Sep;85(17):6292–6296. doi: 10.1073/pnas.85.17.6292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hentschel C. C. Homocopolymer sequences in the spacer of a sea urchin histone gene repeat are sensitive to S1 nuclease. Nature. 1982 Feb 25;295(5851):714–716. doi: 10.1038/295714a0. [DOI] [PubMed] [Google Scholar]
- Htun H., Dahlberg J. E. Single strands, triple strands, and kinks in H-DNA. Science. 1988 Sep 30;241(4874):1791–1796. doi: 10.1126/science.3175620. [DOI] [PubMed] [Google Scholar]
- Htun H., Dahlberg J. E. Topology and formation of triple-stranded H-DNA. Science. 1989 Mar 24;243(4898):1571–1576. doi: 10.1126/science.2648571. [DOI] [PubMed] [Google Scholar]
- Htun H., Lund E., Dahlberg J. E. Human U1 RNA genes contain an unusually sensitive nuclease S1 cleavage site within the conserved 3' flanking region. Proc Natl Acad Sci U S A. 1984 Dec;81(23):7288–7292. doi: 10.1073/pnas.81.23.7288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Htun H., Lund E., Westin G., Pettersson U., Dahlberg J. E. Nuclease S1-sensitive sites in multigene families: human U2 small nuclear RNA genes. EMBO J. 1985 Jul;4(7):1839–1845. doi: 10.1002/j.1460-2075.1985.tb03858.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaworski A., Zacharias W., Hsieh W. T., Blaho J. A., Larson J. E., Wells R. D. In vivo existence of left-handed DNA. Gene. 1988 Dec 25;74(1):215–220. doi: 10.1016/0378-1119(88)90290-9. [DOI] [PubMed] [Google Scholar]
- Johnston B. H. The S1-sensitive form of d(C-T)n.d(A-G)n: chemical evidence for a three-stranded structure in plasmids. Science. 1988 Sep 30;241(4874):1800–1804. doi: 10.1126/science.2845572. [DOI] [PubMed] [Google Scholar]
- Kilpatrick M. W., Torri A., Kang D. S., Engler J. A., Wells R. D. Unusual DNA structures in the adenovirus genome. J Biol Chem. 1986 Aug 25;261(24):11350–11354. [PubMed] [Google Scholar]
- Lee J. S., Woodsworth M. L., Latimer L. J., Morgan A. R. Poly(pyrimidine) . poly(purine) synthetic DNAs containing 5-methylcytosine form stable triplexes at neutral pH. Nucleic Acids Res. 1984 Aug 24;12(16):6603–6614. doi: 10.1093/nar/12.16.6603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lyamichev V. I., Mirkin S. M., Frank-Kamenetskii M. D. A pH-dependent structural transition in the homopurine-homopyrimidine tract in superhelical DNA. J Biomol Struct Dyn. 1985 Oct;3(2):327–338. doi: 10.1080/07391102.1985.10508420. [DOI] [PubMed] [Google Scholar]
- Lyamichev V. I., Mirkin S. M., Frank-Kamenetskii M. D. Structures of homopurine-homopyrimidine tract in superhelical DNA. J Biomol Struct Dyn. 1986 Feb;3(4):667–669. doi: 10.1080/07391102.1986.10508454. [DOI] [PubMed] [Google Scholar]
- Mace H. A., Pelham H. R., Travers A. A. Association of an S1 nuclease-sensitive structure with short direct repeats 5' of Drosophila heat shock genes. Nature. 1983 Aug 11;304(5926):555–557. doi: 10.1038/304555a0. [DOI] [PubMed] [Google Scholar]
- Margot J. B., Hardison R. C. DNase I and nuclease S1 sensitivity of the rabbit beta 1 globin gene in nuclei and in supercoiled plasmids. J Mol Biol. 1985 Jul 20;184(2):195–210. doi: 10.1016/0022-2836(85)90373-0. [DOI] [PubMed] [Google Scholar]
- Marinus M. G., Carraway M., Frey A. Z., Brown L., Arraj J. A. Insertion mutations in the dam gene of Escherichia coli K-12. Mol Gen Genet. 1983;192(1-2):288–289. doi: 10.1007/BF00327681. [DOI] [PubMed] [Google Scholar]
- McKeon C., Schmidt A., de Crombrugghe B. A sequence conserved in both the chicken and mouse alpha 2(I) collagen promoter contains sites sensitive to S1 nuclease. J Biol Chem. 1984 May 25;259(10):6636–6640. [PubMed] [Google Scholar]
- Messer W., Noyer-Weidner M. Timing and targeting: the biological functions of Dam methylation in E. coli. Cell. 1988 Sep 9;54(6):735–737. doi: 10.1016/s0092-8674(88)90911-7. [DOI] [PubMed] [Google Scholar]
- Moos M., Gallwitz D. Structure of two human beta-actin-related processed genes one of which is located next to a simple repetitive sequence. EMBO J. 1983;2(5):757–761. doi: 10.1002/j.1460-2075.1983.tb01496.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Panayotatos N., Fontaine A. A native cruciform DNA structure probed in bacteria by recombinant T7 endonuclease. J Biol Chem. 1987 Aug 15;262(23):11364–11368. [PubMed] [Google Scholar]
- Parniewski P., Galazka G., Wilk A., Klysik J. Complex structural behavior of oligopurine-oligopyrimidine sequence cloned within the supercoiled plasmid. Nucleic Acids Res. 1989 Jan 25;17(2):617–629. doi: 10.1093/nar/17.2.617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pulleyblank D. E., Haniford D. B., Morgan A. R. A structural basis for S1 nuclease sensitivity of double-stranded DNA. Cell. 1985 Aug;42(1):271–280. doi: 10.1016/s0092-8674(85)80122-7. [DOI] [PubMed] [Google Scholar]
- Schon E., Evans T., Welsh J., Efstratiadis A. Conformation of promoter DNA: fine mapping of S1-hypersensitive sites. Cell. 1983 Dec;35(3 Pt 2):837–848. doi: 10.1016/0092-8674(83)90116-2. [DOI] [PubMed] [Google Scholar]
- Singleton C. K., Wells R. D. The facile generation of covalently closed, circular DNAs with defined negative superhelical densities. Anal Biochem. 1982 May 15;122(2):253–257. doi: 10.1016/0003-2697(82)90277-9. [DOI] [PubMed] [Google Scholar]
- Vojtísková M., Mirkin S., Lyamichev V., Voloshin O., Frank-Kamenetskii M., Palecek E. Chemical probing of the homopurine.homopyrimidine tract in supercoiled DNA at single-nucleotide resolution. FEBS Lett. 1988 Jul 18;234(2):295–299. doi: 10.1016/0014-5793(88)80102-9. [DOI] [PubMed] [Google Scholar]
- Voloshin O. N., Mirkin S. M., Lyamichev V. I., Belotserkovskii B. P., Frank-Kamenetskii M. D. Chemical probing of homopurine-homopyrimidine mirror repeats in supercoiled DNA. Nature. 1988 Jun 2;333(6172):475–476. doi: 10.1038/333475a0. [DOI] [PubMed] [Google Scholar]
- Wells R. D., Collier D. A., Hanvey J. C., Shimizu M., Wohlrab F. The chemistry and biology of unusual DNA structures adopted by oligopurine.oligopyrimidine sequences. FASEB J. 1988 Nov;2(14):2939–2949. [PubMed] [Google Scholar]
- Wohlrab F., McLean M. J., Wells R. D. The segment inversion site of herpes simplex virus type 1 adopts a novel DNA structure. J Biol Chem. 1987 May 5;262(13):6407–6416. [PubMed] [Google Scholar]
- Yagil G., Sussman J. L. Structural models for non-helical DNA. EMBO J. 1986 Jul;5(7):1719–1725. doi: 10.1002/j.1460-2075.1986.tb04416.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]