Abstract
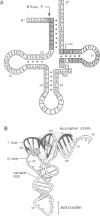
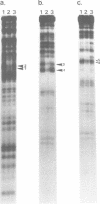
Precursor tRNAAsp molecules, containing a 26-base 5' leader, were treated with diethylpyrocarbonate, 50% hydrazine or anhydrous hydrazine/3M NaCl and then subjected to processing by RNase P RNAs from Escherichia coli or Bacillus subtilis. Fully processed tRNAs and material not successfully cleaved by the catalytic RNAs were analyzed for their content of chemically altered nucleotides. Several bases were identified as being required intact for optimal activity as substrate as judged by exclusion of chemically modified residues from processed molecules, and simultaneous enhancement in material that was not recognized as substrate. Such nucleotides cluster near the site of cleavage at the mature 5' end and in the T stem and loop. Purines at residues 1 and 2 adjacent to the site of cleavage, position 57 in the T loop, and site 64 in the T stem exhibited the most pronounced effects. These results suggest a model of recognition of substrate by RNase P RNAs in which the ribozyme interacts with the corner of the precursor tRNA's three dimensional structure, where the T- and D-loops are juxtaposed, and extends along the top of the molecule back towards the site of catalysis.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Cech T. R. The chemistry of self-splicing RNA and RNA enzymes. Science. 1987 Jun 19;236(4808):1532–1539. doi: 10.1126/science.2438771. [DOI] [PubMed] [Google Scholar]
- Chaconas G., van de Sande J. H. 5'-32P labeling of RNA and DNA restriction fragments. Methods Enzymol. 1980;65(1):75–85. doi: 10.1016/s0076-6879(80)65012-5. [DOI] [PubMed] [Google Scholar]
- England T. E., Bruce A. G., Uhlenbeck O. C. Specific labeling of 3' termini of RNA with T4 RNA ligase. Methods Enzymol. 1980;65(1):65–74. doi: 10.1016/s0076-6879(80)65011-3. [DOI] [PubMed] [Google Scholar]
- Gardiner K. J., Marsh T. L., Pace N. R. Ion dependence of the Bacillus subtilis RNase P reaction. J Biol Chem. 1985 May 10;260(9):5415–5419. [PubMed] [Google Scholar]
- Guerrier-Takada C., Gardiner K., Marsh T., Pace N., Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983 Dec;35(3 Pt 2):849–857. doi: 10.1016/0092-8674(83)90117-4. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Haydock K., Allen L., Altman S. Metal ion requirements and other aspects of the reaction catalyzed by M1 RNA, the RNA subunit of ribonuclease P from Escherichia coli. Biochemistry. 1986 Apr 8;25(7):1509–1515. doi: 10.1021/bi00355a006. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Lumelsky N., Altman S. Specific interactions in RNA enzyme-substrate complexes. Science. 1989 Dec 22;246(4937):1578–1584. doi: 10.1126/science.2480641. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., van Belkum A., Pleij C. W., Altman S. Novel reactions of RNAase P with a tRNA-like structure in turnip yellow mosaic virus RNA. Cell. 1988 Apr 22;53(2):267–272. doi: 10.1016/0092-8674(88)90388-1. [DOI] [PubMed] [Google Scholar]
- Hegg L. A., Thurlow D. L. Cytidines in tRNAs that are required intact by ATP/CTP:tRNA nucleotidyltransferases from Escherichia coli and Saccharomyces cerevisiae. Nucleic Acids Res. 1990 Oct 25;18(20):5975–5979. doi: 10.1093/nar/18.20.5975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hegg L. A., Thurlow D. L. Residual tRNA secondary structure in 'denaturing' 8M urea/TBE polyacrylamide gels: effects on electrophoretic mobility and dependency on prior chemical modification of the tRNA. Nucleic Acids Res. 1990 May 25;18(10):2993–3000. doi: 10.1093/nar/18.10.2993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- James B. D., Olsen G. J., Liu J. S., Pace N. R. The secondary structure of ribonuclease P RNA, the catalytic element of a ribonucleoprotein enzyme. Cell. 1988 Jan 15;52(1):19–26. doi: 10.1016/0092-8674(88)90527-2. [DOI] [PubMed] [Google Scholar]
- Kahle D., Wehmeyer U., Char S., Krupp G. The methylation of one specific guanosine in a pre-tRNA prevents cleavage by RNase P and by the catalytic M1 RNA. Nucleic Acids Res. 1990 Feb 25;18(4):837–844. doi: 10.1093/nar/18.4.837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kahle D., Wehmeyer U., Krupp G. Substrate recognition by RNase P and by the catalytic M1 RNA: identification of possible contact points in pre-tRNAs. EMBO J. 1990 Jun;9(6):1929–1937. doi: 10.1002/j.1460-2075.1990.tb08320.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marsh T. L., Pace N. R. Ribonuclease P catalysis differs from ribosomal RNA self-splicing. Science. 1985 Jul 5;229(4708):79–81. doi: 10.1126/science.4012313. [DOI] [PubMed] [Google Scholar]
- McClain W. H., Guerrier-Takada C., Altman S. Model substrates for an RNA enzyme. Science. 1987 Oct 23;238(4826):527–530. doi: 10.1126/science.2443980. [DOI] [PubMed] [Google Scholar]
- Ogasawara N., Moriya S., von Meyenburg K., Hansen F. G., Yoshikawa H. Conservation of genes and their organization in the chromosomal replication origin region of Bacillus subtilis and Escherichia coli. EMBO J. 1985 Dec 1;4(12):3345–3350. doi: 10.1002/j.1460-2075.1985.tb04087.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pace N. R., Smith D. Ribonuclease P: function and variation. J Biol Chem. 1990 Mar 5;265(7):3587–3590. [PubMed] [Google Scholar]
- Peattie D. A. Direct chemical method for sequencing RNA. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1760–1764. doi: 10.1073/pnas.76.4.1760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reed R. E., Baer M. F., Guerrier-Takada C., Donis-Keller H., Altman S. Nucleotide sequence of the gene encoding the RNA subunit (M1 RNA) of ribonuclease P from Escherichia coli. Cell. 1982 Sep;30(2):627–636. doi: 10.1016/0092-8674(82)90259-8. [DOI] [PubMed] [Google Scholar]
- Reich C., Gardiner K. J., Olsen G. J., Pace B., Marsh T. L., Pace N. R. The RNA component of the Bacillus subtilis RNase P. Sequence, activity, and partial secondary structure. J Biol Chem. 1986 Jun 15;261(17):7888–7893. [PubMed] [Google Scholar]
- Reyes V. M., Abelson J. Substrate recognition and splice site determination in yeast tRNA splicing. Cell. 1988 Nov 18;55(4):719–730. doi: 10.1016/0092-8674(88)90230-9. [DOI] [PubMed] [Google Scholar]
- Rich A., RajBhandary U. L. Transfer RNA: molecular structure, sequence, and properties. Annu Rev Biochem. 1976;45:805–860. doi: 10.1146/annurev.bi.45.070176.004105. [DOI] [PubMed] [Google Scholar]
- Rould M. A., Perona J. J., Söll D., Steitz T. A. Structure of E. coli glutaminyl-tRNA synthetase complexed with tRNA(Gln) and ATP at 2.8 A resolution. Science. 1989 Dec 1;246(4934):1135–1142. doi: 10.1126/science.2479982. [DOI] [PubMed] [Google Scholar]
- Spacciapoli P., Doviken L., Mulero J. J., Thurlow D. L. Recognition of tRNA by the enzyme ATP/CTP:tRNA nucleotidyltransferase. Interference by nucleotides modified with diethyl pyrocarbonate or hydrazine. J Biol Chem. 1989 Mar 5;264(7):3799–3805. [PubMed] [Google Scholar]
- Sprinzl M., Moll J., Meissner F., Hartmann T. Compilation of tRNA sequences. Nucleic Acids Res. 1985;13 (Suppl):r1–49. doi: 10.1093/nar/13.suppl.r1. [DOI] [PMC free article] [PubMed] [Google Scholar]