Abstract
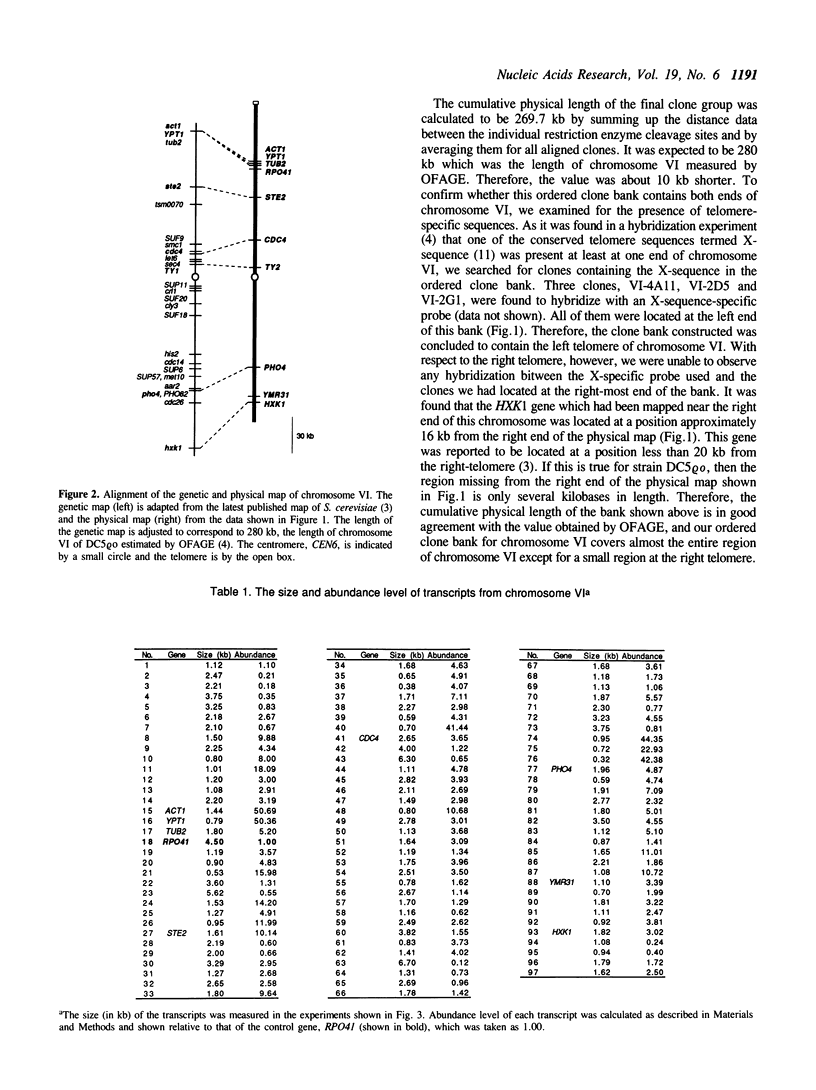
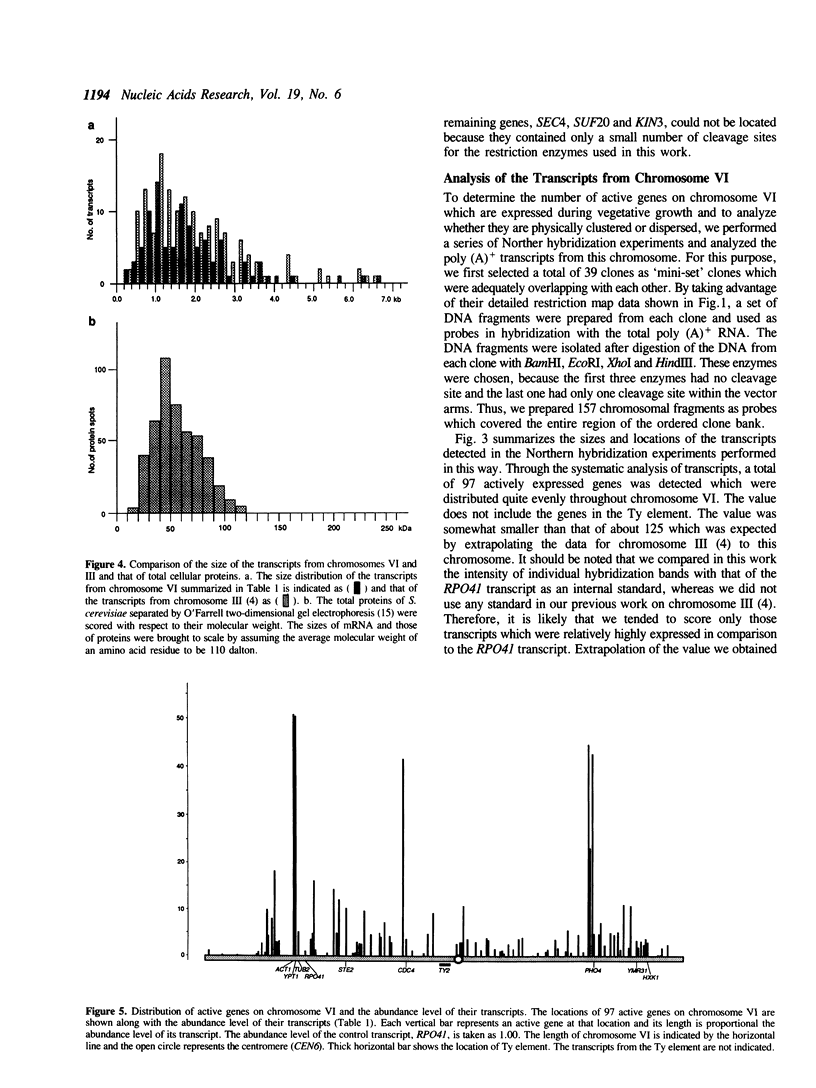
By comparing sequences of restriction enzyme cleavage sites and their distance data, we sorted 384 lambda phage clones containing segments of chromosome VI of S. cerevisiae and constructed an ordered clone bank for this chromosome. The physical length of this bank is 269.7 kb. The bank contains the entire chromosome including the left telomere, but it is not certain whether it contains the right telomere as well. To estimate the number of genes present on this chromosome, we performed a series of Northern hybridization experiments using 157 restriction enzyme fragments prepared from the bank as hybridization probes and total poly(A)+ RNA from vegetatively growing cells. Thus, 97 distinct transcripts were identified. The relative abundance levels of individual transcripts were measured by comparing their band intensity with that of the RPO41 transcript. It was found that the transcripts from the genes located in the telomeric and centromeric regions are less abundant as compared to those from the genes in the central regions of both arms.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Birren B. W., Lai E., Clark S. M., Hood L., Simon M. I. Optimized conditions for pulsed field gel electrophoretic separations of DNA. Nucleic Acids Res. 1988 Aug 11;16(15):7563–7582. doi: 10.1093/nar/16.15.7563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carle G. F., Frank M., Olson M. V. Electrophoretic separations of large DNA molecules by periodic inversion of the electric field. Science. 1986 Apr 4;232(4746):65–68. doi: 10.1126/science.3952500. [DOI] [PubMed] [Google Scholar]
- Carle G. F., Olson M. V. An electrophoretic karyotype for yeast. Proc Natl Acad Sci U S A. 1985 Jun;82(11):3756–3760. doi: 10.1073/pnas.82.11.3756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Gallwitz D., Donath C., Sander C. A yeast gene encoding a protein homologous to the human c-has/bas proto-oncogene product. Nature. 1983 Dec 15;306(5944):704–707. doi: 10.1038/306704a0. [DOI] [PubMed] [Google Scholar]
- Gallwitz D., Sures I. Structure of a split yeast gene: complete nucleotide sequence of the actin gene in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1980 May;77(5):2546–2550. doi: 10.1073/pnas.77.5.2546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greenleaf A. L., Kelly J. L., Lehman I. R. Yeast RPO41 gene product is required for transcription and maintenance of the mitochondrial genome. Proc Natl Acad Sci U S A. 1986 May;83(10):3391–3394. doi: 10.1073/pnas.83.10.3391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones D. G., Rosamond J. Isolation of a novel protein kinase-encoding gene from yeast by oligodeoxyribonucleotide probing. Gene. 1990 May 31;90(1):87–92. doi: 10.1016/0378-1119(90)90442-t. [DOI] [PubMed] [Google Scholar]
- Kaback D. B., Angerer L. M., Davidson N. Improved methods for the formation and stabilization of R-loops. Nucleic Acids Res. 1979 Jun 11;6(7):2499–2317. doi: 10.1093/nar/6.7.2499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein H. L., Petes T. D. Genetic mapping of Ty elements in Saccharomyces cerevisiae. Mol Cell Biol. 1984 Feb;4(2):329–339. doi: 10.1128/mcb.4.2.329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohara Y., Akiyama K., Isono K. The physical map of the whole E. coli chromosome: application of a new strategy for rapid analysis and sorting of a large genomic library. Cell. 1987 Jul 31;50(3):495–508. doi: 10.1016/0092-8674(87)90503-4. [DOI] [PubMed] [Google Scholar]
- Kopetzki E., Entian K. D., Mecke D. Complete nucleotide sequence of the hexokinase PI gene (HXK1) of Saccharomyces cerevisiae. Gene. 1985;39(1):95–101. doi: 10.1016/0378-1119(85)90113-1. [DOI] [PubMed] [Google Scholar]
- Lauer G. D., Roberts T. M., Klotz L. C. Determination of the nuclear DNA content of Saccharomyces cerevisiae and implications for the organization of DNA in yeast chromosomes. J Mol Biol. 1977 Aug 25;114(4):507–526. doi: 10.1016/0022-2836(77)90175-9. [DOI] [PubMed] [Google Scholar]
- Legrain M., De Wilde M., Hilger F. Isolation, physical characterization and expression analysis of the Saccharomyces cerevisiae positive regulatory gene PHO4. Nucleic Acids Res. 1986 Apr 11;14(7):3059–3073. doi: 10.1093/nar/14.7.3059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ludwig J. R., 2nd, Foy J. J., Elliott S. G., McLaughlin C. S. Synthesis of specific identified, phosphorylated, heat shock, and heat stroke proteins through the cell cycle of Saccharomyces cerevisiae. Mol Cell Biol. 1982 Feb;2(2):117–126. doi: 10.1128/mcb.2.2.117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Masters B. S., Stohl L. L., Clayton D. A. Yeast mitochondrial RNA polymerase is homologous to those encoded by bacteriophages T3 and T7. Cell. 1987 Oct 9;51(1):89–99. doi: 10.1016/0092-8674(87)90013-4. [DOI] [PubMed] [Google Scholar]
- Matsushita Y., Kitakawa M., Isono K. Cloning and analysis of the nuclear genes for two mitochondrial ribosomal proteins in yeast. Mol Gen Genet. 1989 Oct;219(1-2):119–124. doi: 10.1007/BF00261166. [DOI] [PubMed] [Google Scholar]
- Mortimer R. K., Schild D., Contopoulou C. R., Kans J. A. Genetic map of Saccharomyces cerevisiae, edition 10. Yeast. 1989 Sep-Oct;5(5):321–403. doi: 10.1002/yea.320050503. [DOI] [PubMed] [Google Scholar]
- Nakayama N., Miyajima A., Arai K. Nucleotide sequences of STE2 and STE3, cell type-specific sterile genes from Saccharomyces cerevisiae. EMBO J. 1985 Oct;4(10):2643–2648. doi: 10.1002/j.1460-2075.1985.tb03982.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neff N. F., Thomas J. H., Grisafi P., Botstein D. Isolation of the beta-tubulin gene from yeast and demonstration of its essential function in vivo. Cell. 1983 May;33(1):211–219. doi: 10.1016/0092-8674(83)90350-1. [DOI] [PubMed] [Google Scholar]
- Panzeri L., Philippsen P. Centromeric DNA from chromosome VI in Saccharomyces cerevisiae strains. EMBO J. 1982;1(12):1605–1611. doi: 10.1002/j.1460-2075.1982.tb01362.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Warmington J. R., Anwar R., Newlon C. S., Waring R. B., Davies R. W., Indge K. J., Oliver S. G. A 'hot-spot' for Ty transposition on the left arm of yeast chromosome III. Nucleic Acids Res. 1986 Apr 25;14(8):3475–3485. doi: 10.1093/nar/14.8.3475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yochem J., Byers B. Structural comparison of the yeast cell division cycle gene CDC4 and a related pseudogene. J Mol Biol. 1987 May 20;195(2):233–245. doi: 10.1016/0022-2836(87)90646-2. [DOI] [PubMed] [Google Scholar]
- Yoshikawa A., Isono K. Chromosome III of Saccharomyces cerevisiae: an ordered clone bank, a detailed restriction map and analysis of transcripts suggest the presence of 160 genes. Yeast. 1990 Sep-Oct;6(5):383–401. doi: 10.1002/yea.320060504. [DOI] [PubMed] [Google Scholar]
- Zakian V. A., Blanton H. M. Distribution of telomere-associated sequences on natural chromosomes in Saccharomyces cerevisiae. Mol Cell Biol. 1988 May;8(5):2257–2260. doi: 10.1128/mcb.8.5.2257. [DOI] [PMC free article] [PubMed] [Google Scholar]




