The mineralization of cartilage and bone matrix requires adequate supplies of calcium and phosphate. Among the causes of defective mineralization of bone (osteomalacia) and defective mineralization of cartilage (rickets) are renal phosphate-wasting disorders that produce hypophosphatemia. Phosphate wasting is either inherited as X-linked hypophosphatemic rickets or autosomal dominant hypophosphatemic rickets, or acquired, as can occur in patients with a variety of benign mesenchymal tumors (hemangiopericytomas, fibromas, angiosarcomas, etc.) (1). Osteomalacia induced by tumors is invariably curable if the tumor can be found and resected, indicating that it has a humoral basis. A paper by Shimada et al. (2) in this issue of PNAS identifies a member of the fibroblast growth factor family, FGF23, as the humoral factor that is secreted by tumors to cause tumor-induced osteomalacia.
With the discovery that a protease mutation and a cleavage site mutation may cause the same disease, the puzzle pieces could fit together nicely.
Shimada et al. (2) cloned cDNAs from a hemangiopericytoma that caused hypophosphatemic osteomalacia (3) and found clones identical to FGF23, which has recently been identified by positional cloning as the gene responsible for autosomal dominant hypophosphatemic rickets (4). When injected into mice, recombinant FGF23 produced mild phosphaturia and hypophosphatemia, but CHO-FGF23 cells, when grown as tumors in nude mice, fully reproduced the human syndrome of severe hypophosphatemia, growth retardation, rickets in the growth plates, deformities of the skeleton, reduced mineralization of bone, and seams of unmineralized osteoid in bone (2). FGF23 was expressed at high levels in the tumor from which it was cloned, and as recently reported by another group, is also expressed at high levels in other tumors associated with acquired osteomalacia (5), but expression is barely detectable in normal tissues [liver, lymph node, thymus, heart, and the ventrolateral thalamic nucleus of the brain (2, 4, 6)], and notably absent in bone and bone cells.
At first glance a member of the matrix-binding FGF family is a surprising candidate as a humoral messenger, but FGF23 lacks several residues that are heparan-binding in FGF1 and conserved in other heparin-binding FGFs (7), and hence may be more soluble than other FGFs. With a 72-aa carboxyl-terminal domain not shared by other family members, FGF23 is the largest member of the FGF family. Insight into its function is gained by considering the mutations that cause autosomal dominant hypophosphatemic rickets. All four unrelated families who were studied had missense mutations in one of two closely spaced arginine residues (R176 and R179) that cosegregated with rickets, with two families sharing the same mutation (4). This clustering of missense mutations in a disorder with dominant inheritance strongly suggested they were gain-of-function mutations. It is therefore interesting that Shimada et al., when they expressed FGF23 in CHO cells, found in addition to the mature protein a fragment beginning with S180. The demonstration that R179/S180 is a processing site in FGF23 strongly suggests that mutations of the flanking arginines confer a gain of function on FGF23 by blocking its degradation. The cleavage site is at the boundary between the FGF-homologous region and the unique carboxyl terminus.
The other piece of the hypophosphatemia puzzle is X-linked hypophosphatemic rickets, the most common inherited disorder of renal phosphate transport. Positional cloning identified mutations that inactivate a gene called PHEX (8), predicted to encode a membrane-associated metalloprotease of the M13 family, which includes neutral endopeptidase 24.11, endothelin-converting enzymes 1 and 2, and the Kell blood group antigen (9). With the discovery that a protease mutation and a cleavage site mutation may cause the same disease, the puzzle pieces could fit together nicely. FGF23 may be the long-sought “phosphatonin,” the phosphaturic factor secreted by tumors that cause osteomalacia. It may be hypothesized that FGF23 is also secreted by one or more normal tissues as a phosphate-regulating hormone, and that the level of FGF23 in blood is determined in part by the rate of its cleavage by the PHEX protease at R179/S180. Overproduction of FGF23 by tumors, mutations that prevent cleavage of FGF23 (as in autosomal dominant hypophosphatemic rickets), or mutations that inactivate the responsible protease, PHEX (as in X-linked hypophosphatemic rickets), would all increase the level of FGF23, with resultant phosphaturia, hypophosphatemia, and rickets/osteomalacia (Fig. 1). It is a compelling picture, but do the pieces really fit?
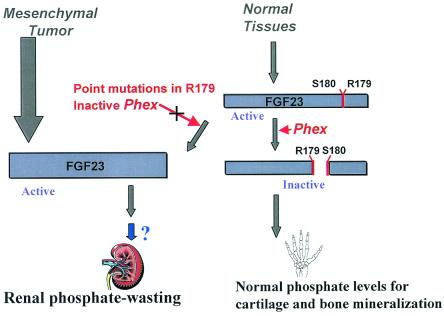
Figure 1.
Proposed pathogenesis of renal phosphate wasting. Mesenchymal tumors produce renal phosphate wasting by overproduction of FGF23. FGF23 levels can also be increased by mutations in Phex, a protease that cleaves and inactivates the molecule, or by mutations at key arginine residues that render FGF23 resistant to cleavage by Phex. FGF23 excess causes phosphate wasting either directly or by inducing another phosphaturic factor.
The Na-phosphate cotransporter type IIa is responsible for the bulk of phosphate reabsorption in the proximal tubule (10). If FGF23 is a phosphatonin it should directly inhibit renal Na-phosphate cotransport. Yet Shimada et al. report that FGF23 had no effect on phosphate transport by OK cells, a cell line that expresses the renal Na-phosphate cotransporter IIa and is sensitive both to PTH and to changes in medium phosphate (2). It is conceivable that OK cells do not express the receptor for FGF23, and it will be important to determine whether FGF23 inhibits phosphate reabsorption in other systems in vitro (e.g., isolated renal proximal tubules). It is also possible that FGF23 requires further processing to a biologically active form. Finally, FGF23 may not be the final phosphatonin, but may stimulate secretion of a final phosphate-regulating factor. Candidate molecules include stanniocalcin-1 and -2. Stanniocalcin regulates calcium and phosphate homeostasis in fish. In mammals, stanniocalcin-1 is present in distal nephron segments of the kidney and stimulates phosphate reabsorption (11, 12); stanniocalcin-2 is found in bone cells and inhibits phosphate transport (13). A third candidate is MEPE, or matrix extracellular phosphoglycoprotein, which has also been isolated from tumors associated with osteomalacia (2, 14).
The pathogenesis of defective mineralization may require more than phosphate wasting. In the Hyp and Gy mouse models of hypophosphatemic rickets, both of which are caused by mutations of Phex (15), the renal conversion of vitamin D to calcitriol is not appropriately stimulated by hypophosphatemia, and serum calcitriol levels are inappropriately low. Knockout of the mouse Na-phosphate cotransporter gene Npt2 produces levels of phosphate wasting comparable to inactivation of the Phex gene, but in Npt2(−/−) mice calcitriol responds appropriately to the hypophosphatemic challenge, intestinal hyperabsorption of both phosphate and calcium ensues, and rickets and osteomalacia are absent (16). Comparison of these mouse models illustrates that renal phosphate wasting can be dissociated from defective synthesis of calcitriol, implying that phosphatonins have at least two independent renal effects, inhibition of phosphate reabsorption and impairment of the synthesis of calcitriol. The different phenotypes in Npt2(−/−) and Phex(o/−) mice also raise the possibility that FGF23 has direct effects on bone and cartilage that contribute, along with hypophosphatemia, to a defect in mineralization.
The discovery of inherited mutations in PHEX and FGF23, along with the work of Shimada et al. (2), uncovers what appear to be the pieces of a phosphate regulating system. But how do those pieces fit together? Phosphate can be virtually cleared from the urine in response to a low dietary intake, protecting against phosphate depletion (10). Decreased secretion of FGF23 could be the humoral arm of this response, coupling an as-yet unidentified phosphate sensor, possibly in the intestinal mucosa, to regulation of renal phosphate reabsorption. Consistent with this hypothesis, the Hyp mouse model of Phex inactivation responds to phosphate deprivation, albeit with continued phosphaturia relative to wild-type mice (17). It is also possible that the physiological role of FGF23 is local, perhaps even unrelated to phosphate homeostasis, and only when it is inappropriately secreted into blood does phosphate wasting occur. In this scenario, the phosphate wasting in tumor-induced osteomalacia would be analogous to the phosphate wasting that occurs when tumors overexpress the PTH-related protein, PTHrP. PTHrP is normally a local regulator of cell differentiation, but when overproduction gives it access to the circulation, it co-opts the PTH receptor in kidney to cause phosphaturia (18).
Whether its physiological role is local or humoral, FGF23 is an unusual hormone. Its degradation rate, rather than its rate of synthesis, can in some circumstances apparently determine its level in blood. It is the first FGF for which mutations are associated with a disease. And although the other 22 FGF's share only four known receptors, it is likely that FGF23 has a different receptor, because cleavage of its unique carboxyl terminus inactivates it. As our knowledge of it broadens, FGF23 will have an interesting future.
Footnotes
See companion article on page 6500.
References
- 1.DiMeglio L A, White K E, Econs M J. Endocrinol Metab Clin North Am. 2000;29:591–609. doi: 10.1016/s0889-8529(05)70152-3. [DOI] [PubMed] [Google Scholar]
- 2.Shimada T, Mizutani S, Muto T, Yoneya T, Hino R, Takeda S, Takeuchi Y, Fujita T, Fukumoto S, Yamashita T. Proc Natl Acad Sci USA. 2001;98:6500–6505. doi: 10.1073/pnas.101545198. . (First Published May 8, 2001; 10.1073/pnas.101545198) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Fukumoto S, Takeuchi Y, Nagano A, Fujita T. Bone. 1999;25:375–377. doi: 10.1016/s8756-3282(99)00170-2. [DOI] [PubMed] [Google Scholar]
- 4.White K E, Evans W E, O'Riordan J L, Speer M C, Econs M J, Lorenz-Depiereux B, Grabowski M, Meitinger T, Strom T M. Nat Genet. 2000;26:345–348. [Google Scholar]
- 5.White K E, Jonsson K B, Carn G, Hampson G, Spector T D, Mannstadt M, Lorenz-Depiereux B, Miyauchi A, Yang I M, Ljung-gren O, et al. J Clin Endocrinol Metab. 2001;86:497–500. doi: 10.1210/jcem.86.2.7408. [DOI] [PubMed] [Google Scholar]
- 6.Yamashita T, Yoshioka M, Itoh N. Biochem Biophys Res Commun. 2000;277:494–498. doi: 10.1006/bbrc.2000.3696. [DOI] [PubMed] [Google Scholar]
- 7.Pellegrini L, Burke D F, von Delft F, Mulloy B, Blundell T L. Nature (London) 2000;407:1029–1034. doi: 10.1038/35039551. [DOI] [PubMed] [Google Scholar]
- 8.The HYP Consortium. Nat Genet. 1995;11:130–136. doi: 10.1038/ng1095-130. [DOI] [PubMed] [Google Scholar]
- 9.Turner A J, Tanzawa K. FASEB J. 1997;11:355–364. doi: 10.1096/fasebj.11.5.9141502. [DOI] [PubMed] [Google Scholar]
- 10.Murer H, Hernando N, Forster I, Biber J. Physiol Rev. 2000;80:1373–1409. doi: 10.1152/physrev.2000.80.4.1373. [DOI] [PubMed] [Google Scholar]
- 11.Wong C K, Ho M A, Wagner G F. J Endocrinol. 1998;158:183–189. doi: 10.1677/joe.0.1580183. [DOI] [PubMed] [Google Scholar]
- 12.Wagner G F, Vozzolo B L, Jaworski E, Haddad M, Kline R L, Olsen H S, Rosen C A, Davidson M B, Renfro J L. J Bone Miner Res. 1997;12:165–171. doi: 10.1359/jbmr.1997.12.2.165. [DOI] [PubMed] [Google Scholar]
- 13.Ishibashi K, Miyamoto K, Taketani Y, Morita K, Takeda E, Sasaki S, Imai M. Biochem Biophys Res Commun. 1998;250:252–258. doi: 10.1006/bbrc.1998.9300. [DOI] [PubMed] [Google Scholar]
- 14.Rowe P S, de Zoysa P A, Dong R, Wang H R, White K E, Econs M J, Oudet C L. Genomics. 2000;67:54–68. doi: 10.1006/geno.2000.6235. [DOI] [PubMed] [Google Scholar]
- 15.Strom T M, Francis F, Lorenz B, Boddrich A, Econs M J, Lehrach H, Meitinger T. Hum Mol Genet. 1997;6:165–171. doi: 10.1093/hmg/6.2.165. [DOI] [PubMed] [Google Scholar]
- 16.Beck L, Karaplis A C, Amizuka N, Hewson A S, Ozawa H, Tenenhouse H S. Proc Natl Acad Sci USA. 1998;95:5372–5377. doi: 10.1073/pnas.95.9.5372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Tenenhouse H S, Martel J, Biber J, Murer H. Am J Physiol. 1995;268:F1062–F1069. doi: 10.1152/ajprenal.1995.268.6.F1062. [DOI] [PubMed] [Google Scholar]
- 18.Strewler G J. N Engl J Med. 2000;342:177–185. doi: 10.1056/NEJM200001203420306. [DOI] [PubMed] [Google Scholar]