Abstract
AIM
To determine a species-specific real-time polymerase chain reaction (PCR) assay to detect Pseudomonas aeruginosa (PA), a secondary DNA target for PA that may provide a universal target for other bacterial pathogens, and validate both assays for diagnostic testing.
METHODS
PCR detection was established against the ecfX PA gene and the 16S rRNA gene using known PA keratitis isolates. The outcome parameters for both assays were “limit of detection” (LOD), amplification efficiency (AE), and PAGE amplified product analysis. Both assays were validated against 20 true-positive clinical samples positive for PA DNA and 20 true-negative samples containing no PA DNA. Descriptive statistics and PAGE analysis were used as outcome parameters.
RESULTS
AE of the ecfX assay was 96.6%, and LOD was 33.6 copies of target DNA per microliter. AE of the 16S rRNA assay was 103.4%, and LOD was 8.12 copies per microliter. The sensitivity, specificity, positive predictive value, negative predictive value, and efficiency for the ecfX and 16S rRNA assays were [75%, 95%, 94%, 79%, and 85%], and [70%, 100%, 100%, 77%, and 85%], respectively. Both PCR assays were validated, followed by confirmation of DNA patterns from PAGE analysis.
CONCLUSION
The PCR methodology described here may be a useful adjunct to standard methods in the diagnosis of PA keratitis.
Keywords: Pseudomonas aeruginosa, cornea, real-time PCR, ecfX, 16S rRNA
Introduction
Pseudomonas aeruginosa (PA) keratitis is a serious infection that if not treated properly could result in visual impairment and eventual corneal transplantation[1]. Contact lens wear associated bacterial infections affect about 1 to 3 people per 10,000 people per year in the United States[2]. Each year, it is estimated that about 500,000 people fall victim to bacterial keratitis in the world, with about 30,000 of these cases occurring in the United States[3]. The optimal scenario for detecting PA is by culturing the cornea for isolation on standard culture media at the earliest stage after onset of infection, prior to empiric topical therapy. Once PA is isolated, in vitro anti-infective susceptibility testing can support the choice of empiric therapy or be used to change the selection of anti-infectives. Complications can occur when patients are treated without confirming the identification of the etiologic agent by laboratory studies. The clinical presentation might lead to treating the incorrect etiologic agent or the infecting agent may be resistant to therapy. Under these conditions, the keratitis may further digress to a worse situation with a referral to a tertiary care facility. At this point, standard laboratory techniques (i.e. culture isolation) will be less reliable, and other techniques will be required for a laboratory diagnosis (i.e. PCR).
A single PCR test to detect specifically PA from cornea specimens could be a useful laboratory adjunct. Even better, a single PCR test to detect PA and other bacterial pathogens would be more useful if the nature of the infectious agent was unknown. In this study, we chose two genetic sites for amplifying DNA to detect PA. The first genetic site was the species-specific ecfX gene and the second site was the 16S rRNA gene. Both targets detect PA, but the second site also has the potential to detect other bacterial pathogens. We hypothesize that PA can be detected from corneal samples using PCR testing directed to the species-specific ecfX gene and to the 16S rRNA gene that may be universal to other bacterial pathogens. The focus of the study will only be directed to corneal isolates of PA. The aims of the study are: 1) to establish a species-specific real-time PCR assay to detect PA; 2) to determine a secondary target for PA that may provide a universal target for other bacterial pathogens; and 3) to validate both assays for diagnostic testing with true positive and true negative clinical samples.
Materials and Methods
Test 1-Species-Specific PCR to Detect Pseudomonas aeruginosa
PCR was developed using the species-specific ecfX gene to detect PA. The ecfX gene is specific to PA and encodes an extracytoplasmic function sigma factor that might act as a virulence factor and assist in haem-uptake[4]. Anuj et al[5] utilized the 63 base pair (bp) ecfX gene as a target to detect PA isolates of clinical and environmental origin. Cattoir et al[6] used the ecfX target to detect PA DNA in blood samples. We used the ecfX gene in the development of a PCR assay to detect PA isolated from the corneas of keratitis patients. In brief, 1) PA and non-PA isolates were selected, 2) the DNA for PCR was extracted from the corneal pathogens, and 3) PCR was performed. The outcome parameters for assay development were: A) optimization of amplification efficiency, B) determination of a minimal limit of detection, and C) confirmation of amplified DNA product.
Samples
For optimal species-specific PCR detection of PA DNA from corneal samples, 20 PA isolates from 20 separate cases of clinical keratitis were selected for testing. The PA isolates were retrieved as de-identified isolates (patient information not available to researchers) from a clinical tissue bank used for the validation of in vitro antibiotic susceptibility testing and new diagnostic testing. An ATCC (American Type Culture Collection) isolate of PA (ATCC# 27853) was used as the positive control. An optimal species-specific PCR test for PA DNA should not detect DNA from other microbial species. To assure negative PCR for PA DNA, the following 35 de-identified non- PA non-ocular isolates (use for validation and susceptibility monitoring) were selected for the study: Achromobacter xylosoxidans, Acinetobacter baumanii, Bacillus species, Burkholderia cepacia, Chlamydia trachomatis, Chryseobacterium indologenes, Chryseobacterium meningosepticum, Deltfia acidovorans, Enterobacter aerogenes, Enterococcus faecalis, Escherichia coli, Haemophilus influenzae, Klebsiella oxytoca, Moraxella lacunata, Mycobacteria chelonae, Nocardia farcinica, nutritionally variant Streptococcus, Ochrobacterium anthropi, Propionibacterium acnes, Pseudomonas fluorescens, Pseudomonas putida, Serratia marcescens, Staphylococcus aureus, Streptococcus pneumoniae, Streptococcus viridans, and Stenotrophomonas maltophilia. The fungal isolates, Candida parapsilosis, Candida albicans, Aspegillus niger, Alternaria species, and Fusarium species were included, in addition to free-living amoebas (Acanthamoeba and Hartmanella species), and viruses (adenovirus and Herpes simplex virus type 1).
DNA extraction
The PA and non-PA isolates were subjected to DNA extraction techniques in order to isolate and remove the DNA from the cells. DNA from PA isolates was extracted using a DNA extraction heating method used by Anuj et al[5], as well as the QIAamp DNA Mini Kit (Valencia, CA). The non-PA isolates had been previously extracted for past work utilizing an Epicentre® DNA extraction method (Epicentre, Madison, WI) and had been stored at -20°C. In brief, bacteria were suspended to a turbidity of a 0.5-McFarland suspension for each isolate. A 300 µL aliquot of each sample was heated at 98°C for 10 minutes, and placed on ice for a few minutes, followed by the addition of 150 µL of Master Pure Complete (MPC) protein precipitation solution (Epicentre, Madison, WI). Each tube was vortexed for 10 seconds and centrifuged at 10000 r/min for 10 minutes at 4°C. The supernatant from each tube was decanted into a clean tube that contained 500 µL of isopropanol (DNase, RNase, Protease free) (Acros Organics, Fisher Scientific, Pittsburgh, PA). Each new tube was inverted 30 to 40 times and centrifuged at 10000r/min for 10 minutes at 4°C. Following centrifugation, the supernatants in each tube were discarded, with 500 µL of 75% ethanol (Spectrum Chemical Mfg. Corp., Gardena, CA) being added to each pellet. All tubes were spun and centrifuged at 10 000r/min for 5 minutes at 4°C, followed by the addition of 500 µL of 75% ethanol to the pellets and centrifugation at the same settings. Once centrifugation was complete, the pellets were dried using a vacuum system for 30 to 45 minutes. After drying, the pellets were suspended in 35 µL of TE buffer (10mmol/L Tris-HCL [pH 8.0], 1 mmol/L EDTA, Epicentre, Madison, WI).
PCR conditions
The Cepheid SmartCycler® II Real-Time PCR system (Sunnyvale, CA) was used for all real-time PCR testing. The primers and probe set for the ecfX amplification assay are listed in Table 1. The set targeted for the 63 bp ecfX gene was retrieved from Anuj et al[5] and the primers and probe were purchased from Integrated DNA Technologies (IDT) (Coralville, IA). The PCR reactions using the primers and probe targeted at the ecfX gene consisted of 18.1 µL of master mix and 6.9 µL of sample. The master mix used in this assay consisted of a forward primer, reverse primer, probe, and Takara Premix Ex Taq (Otsu, Shiga, Japan). The concentration of the primers used in all of the different real-time PCR reactions was set at 0.4 µmol/L, with probe concentration set at 0.2 µmol/L. The PCR reactions for the ecfX gene specific real-time PCR assay were set at the following parameters: Stage 1) 95.0°C for 900 seconds; Stage 2) 45 cycles of 95.0°C for 15 seconds and 60.0°C for 60 seconds.
Table 1. Primer and probe sequences for the development of Pseudomonas aeruginosa real-time PCR.
| Pathogen | Target | Size (bp) | Primers and Probe |
| Pseudomonas aeruginosa | ecfX gene | 63 | Forward Primer: ecfX-F: 5′-CCC ATG CCT ATC ACC CCT T-3′ |
| Reverse Primer: ecfX-R: 5′-CAA CTG CCC AGG TGC TTG C-3′ | |||
| ecfX TM: 5′- /5G FAM/ATG GCG AGT TGC TGC GCT TCC T/3BHQ-1/-3′ | |||
| Pseudomonas aeruginosa | 16S rRNA | 144 | Forward Primer: F2: 5′-GCA CTT TAA GTT GGG AGG AA-3′ |
| Reverse Primer: 16S1RR-B: 5′-CTT TAC GCC CAR TRA WTC CG-3′ | |||
| 514-S: 5′- /56-FAM/TNT TAC CGC GGC TGC TGG CAC G/36-TAMSP/-3 |
Amplification efficiency determination
The ecfX open reading frame was amplified from PA strain PA14 using oligonucleotide primers and a high fidelity polymerase (Phusion, New England Biolabs). This plasmid was cloned into pMQ236 using yeast recombineering as previously described[7] . Diagnostic PCR and DNA sequencing were used to verify the cloning of ecfX. This plasmid was used for measuring amplification efficiency (AE) and limit of detection (LOD).
AE measures the doubling rate of a single copy of DNA. Optimally, one single DNA strand copies into another single strand per each PCR cycle. The AE was determined by PCR, testing serial dilutions (10−3 to 10−9) of the pMQ236 plasmid that contained ecfX, to determine the cycles of positive amplification (Ct). A regression curve was plotted (Minitab Inc., State College, PA) with the Ct value versus the dilution of the plasmid. The slope of the curve was used to determine the AE in the equation [AE = 10(−1/slope)-1] from the regression line equation using the simple algebraic equation y= ax + b (y=Ct, x = plasmid dilution, and a = slope). AE values between 90% and 105% represent optimal PCR testing[8].
LOD determination
The LOD (copies per µL) was determined by multiplying [the DNA concentration of the ecfX plasmid (mg/L)] by [the copy number of plasmid per ng] and [the dilution factor from the original sample]. Several factors were taken into consideration for the calculation of the LOD. The total length of the plasmid construct, measured in base pairs, was used to calculate the number of molecules (copies) in one nanogram (ng). The DNA concentration value of the plasmid construct was used to calculate the (ng) weight per one microliter of DNA. The real-time PCR results of the serial dilutions of the plasmid constructs were used to determine the dilution factor. The LOD determined the minimum number of DNA copies per liter required by the Cepheid SmartCycler® II Real-Time PCR system (Sunnyvale, CA) to detect PA DNA using the ecfX target.
PAGE amplified product analysis
The amplified product of the ecfX amplicon was confirmed using restriction enzyme and PAGE analysis. The DNA amplified from chromosomal DNA from the positive control strain of PA (ATCC# 27853) was first purified to remove primers and other impurities with the QIAquick PCR Purification kit (Valencia, CA) and stored in 30 µL of DNAse free water at -20°C. The NEBcutter V2.0 software (New England BioLabs, Inc.) was used to determine the available restriction enzyme site options for the 63 bp ecfX gene target. The amplified product was cut with the restriction enzyme, NcoI (New England BioLabs, Inc.), in order to obtain 44 bp and 19 bp fragments as predicted for the ecfX amplicon. The reaction consisted of the following reagents: 10 µL of DNA; 2 µL of Buffer 4; 7.5 µL of distilled water; and 0.5 µL of NcoI. Reactions were incubated overnight in a water bath at 37°C to ensure full digestion. Digested and non-digested control amplicons were subjected to PAGE analysis. The positive control PA sample (ATCC# 27853) was loaded to a 15% polyacrylamide gel to confirm the presence of the gene target, as well as the expected restriction digest patterns. The gel was run at 100V and stained with 10 µL of ethidium bromide prior to image collection using a Gel Logic Imaging System (Carestream Health Molecular Imaging, Woodbridge, CT).
Test 2 - 16S rRNA PCR Target to Detect Pseudomonas aeruginosa
PCR was developed to a 16S rRNA gene target to detect PA, utilizing the forward primer, which was designed using MultAlin[9] software to work with the reverse primer and probe utilized by Schabereiter-Gurtner et al[10] . Table 1 describes the structure of the primers and probe. A 16S rRNA gene PCR assay has the potential to detect other bacterial corneal pathogens due to this gene being highly conserved in prokaryotes. Development was directed in detecting PA isolated from the corneas of keratitis patients and negative testing to non-bacterial pathogens isolated from corneal specimens. The focus of the aim was not directed to detect DNA of all bacterial pathogens. In brief, 1) PA and non-bacterial isolates were selected, 2) the DNA for PCR was extracted, and 3) PCR was performed. The outcome parameters for assay development were: A) optimization of amplification efficiency, B) determination of minimizing the limit of detection, and C) confirmation of amplified DNA product.
DNA extraction
As described previously for Test 1, DNA from PA isolates (same as for establishing PCR for the ecfX target under Test 1) was extracted using a DNA extraction boiling method used by Anuj et al[5], as well as the QIAamp DNA Mini Kit (Valencia, CA).
PCR conditions
The Cepheid SmartCycler® II Real-Time PCR system (Sunnyvale, CA) was used for all real-time PCR testing. The primer set, ordered from IDT (Coralville, IA), produced a target size of 144 bp. All forward and reverse primers were used at a concentration of 0.4 µmol/L, and the Taqman probe was used at a concentration of 0.2 µmol/L. The PCR reactions for the 16S rRNA primers and probe set consisted of 17.5 µL of master mix and 7.5 µL of sample. The master mix used in this assay consisted of a forward primer, reverse primer, probe, and Takara Premix Ex Taq™ (Otsu, Shiga, Japan). The PCR settings for the 16S rRNA gene assay were optimized at the following parameters: Stage 1) 95.0°C for 900 seconds; Stage 2) 45 cycles of 95.0°C for 15 seconds and 60.0°C for 30 seconds.
The plasmid containing the 16S rRNA gene, pGEM®-T Easy/PA UNI, was constructed with the pGEM®-T Easy Vector System (Promega Corporation, Madison, WI), following the pGEM®-T Easy Vector System Protocol (Promega, Corporation, Madison, WI). This plasmid was validated by PCR and sequencing. AE was calculated similarly as described for Test 1. The AE was determined by PCR, testing serial dilutions (10−3 to 10−9) of the pGEM®-T Easy plasmid that contained the 16S rRNA gene. The LOD was calculated similarly as described for Test 1.
PAGE amplified product analysis
The amplified product of the positive control PA (ATCC# 27853) was confirmed using restriction enzyme and PAGE analysis. The amplified positive control PA was first purified to remove primers and other impurities with the QIAGEN QIAquick PCR Purification kit (Valencia, CA) and stored in 30 µL of DNAse free water at -20°C. The NEBcutter V2.0 software (New England BioLabs, Inc.) was used to determine the available restriction enzyme site options for the 144 bp 16S rRNA gene target. The amplified product was cut with the restriction enzyme, SacII (New England BioLabs, Inc.), in order to obtain 97 bp and 47 bp fragments as predicted for the 16S rRNA amplicon. The reaction consisted of the following reagents: 10 µL of DNA; 2 µL of Buffer 4; 7.5 µL of distilled water; and 0.5 µL of SacII. Reactions were incubated overnight in a waterbath at 37°C to ensure full digestion. After the amplified PCR product had undergone restriction digestion overnight, the product was subjected to PAGE analysis. A 15% polyacrylamide gel was poured and the positive control PA sample (ATCC# 27853) was loaded to validate the presence of the 144 bp 16S rRNA gene target, as well as the expected restriction digest patterns. All gels were run at 100V and stained with 10 µL of ethidium bromide prior to image collection using a Gel Logic Imaging System (Carestream Health Molecular Imaging, Woodbridge, CT).
Test 3 - Validation of Assays for Diagnostic Testing with True Positive and True Negative Clinical Samples
A total of 40 de-identified patient samples were used in this study, following the establishment of both real-time PCR assays with PA and non-PA isolates. All retrospective, de-identified clinical samples were part of a clinical bank, which is used for validation and susceptibility monitoring, and collected as excess specimens. No additional specimens were collected for this study. These samples, stored at -80°C, were collected for bacterial culture and/or viral testing. A sterile swab (Fisher Scientific, Houston, TX) or a kimura spatula were used to obtain corneal samples from patients, which were then added to 2.0 mL of Chlamydial Transport medium (Bartels, Bellevue, WA). In brief, 1) de-identified patient samples were selected, 2) the DNA for PCR was extracted from these samples, and 3) both PCR assays were performed, followed by A) calculations of descriptive statistics and B) confirmation of amplified DNA products.
Extraction of DNA
The 40 de-identified patient samples consisted of 20 samples positive for PA and 20 negative controls that were culture medium spiked with Aspergillus niger, Candida albicans, Fusarium species, HSV, and VZV. The negative controls, both viral and fungal samples, were chosen based on the fact that they did not contain the highly conserved 16S rRNA gene found in prokaryotes. Direct patient samples were also extracted using the Epicentre® DNA extraction technique (Epicentre, Madison, WI) described under Test 1. The only change in the protocol that was implemented was that the final pellets were suspended in 45 µL of TE buffer rather than 35 µL, in order to utilize and store more extracted DNA from the samples.
PCR conditions
The Cepheid SmartCycler® II Real-Time PCR system (Sunnyvale, CA) was used for all real-time PCR testing. All 40 de-identified patient samples were tested with both PCR assays, targeting the ecfX and 16S rRNA genes. The same procedures were carried out for both PCR assays, as described in Tests 1 and 2. Five different descriptive statistics (sensitivity, specificity, positive predictive value, negative predictive value, and efficiency) were calculated describing the real-time PCR assays. Sensitivity described the ability of PCR to detect DNA in true-positive samples and not in true-negative samples. In contrast, specificity described the ability of PCR not to detect DNA in negative samples and to detect it in true-positive samples. The positive predictive value determined the likelihood that a positive PCR test represented a positive test based on true-positive and false-positive testing. A negative predictive value determined the likelihood that a negative PCR test represented a negative test based on true-negative and false-negative testing. Efficiency determined the overall ability of PCR to correctly predict whether DNA is present or absent.
PAGE amplified product analysis
The QIAGEN QIAquick PCR Purification kit (Valencia, CA) was used to purify the amplified product of the true-positive patient samples from the ecfX gene assay and the 16S rRNA gene assay. The kit protocol was followed, with the exception of the final step where 30 µl of DNAse free water was added to each sample and allowed to sit in the hood for at least one minute, followed by centrifugation for one minute. In order to confirm that the amplified PCR products (from the de-identified patient samples) in the real-time PCR assays were the expected amplified PCR products, restriction enzyme digestion was carried out, as described by tests 1 and 2. After the amplified PCR products had undergone restriction digestion overnight, DNA fragments were subjected to PAGE analysis, as mentioned under Tests 1 and 2. 15% polyacrylamide gels were poured for the ecfX assay amplified products that were detected as positive for PA DNA, while 6% polyacrylamide gels were poured for the 16S rRNA assay amplified products. All gels were run at 100V and stained with 10 µL of ethidium bromide prior to image collection using a Gel Logic Imaging System (Carestream Health Molecular Imaging, Woodbridge, CT).
RESULTS
Species-Specific PCR
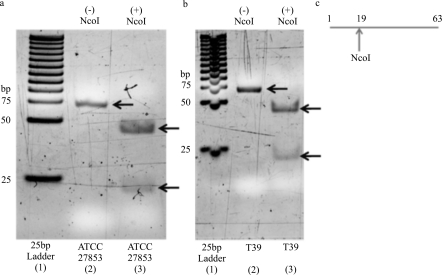
All PA strains (100%, 20 of 20) isolated from patient corneas were detected with the ecfX real-time PCR assay, while the non-PA bacterial, fungal, acanthamoebal, and viral isolates (0%, 0 of 35) were not detected. The AE of the ecfX assay was 96.6%. A sigmoidal, S-shaped curve appeared for each dilution value, which is a visual sign of an efficient real-time PCR reaction. The Ct values ranged from 18.51 to 38.67, with a cycle difference of about 3 between each measured dilution value. The slope of the regression plot (Ct values versus dilutions) was -3.40611. The LOD was 33.6 copies of target DNA per microliter. In Figure 1A, the positive control (ATCC# 27853) is depicted as the expected 63 bp band on a 15% polyacrylamide gel for the ecfX real-time assay. Restriction enzyme analysis depicted the expected digest patterns with a 44 bp and a 19 bp band.
Figure 1. Confirmation of the ecfX amplicon with polyacrylamide gel electrophoresis by restriction analysis.
Secondary PCR Target
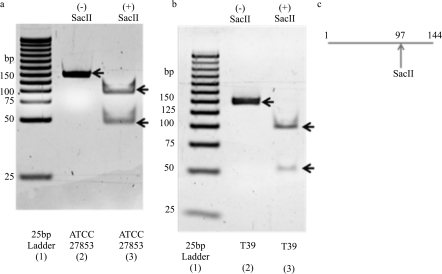
All PA strains (100%, 20 of 20) isolated from patient corneas were detected with the primer set in this real-time PCR assay. AE of the 16S rRNA assay was 103.4%. A sigmoidal, S-shaped type curve was present at each dilution value. The Ct values ranged from 18.89 to 35.69, with a cycle difference of about 3 between each measured dilution value. The slope of this regression plot was -3.24314. The LOD was 8.12 copies per microliter. In Figure 2A, the positive control (ATCC# 27853) is depicted as the expected 144 bp band on a 15% polyacrylamide gel. Restriction enzyme analysis depicted the expected digest patterns with a 97 bp and 47 bp band.
Figure 2. Confirmation of the 16S rRNA amplicon with polyacrylamide gel electrophoresis by restriction analysis.
Diagnostic validation
True positives were direct samples from the corneas of patients that were culture positive for PA. True negatives were samples that did not have PA DNA; instead, these samples were spiked with the following isolates: Varicella zoster virus, Herpes simplex virus, Fusarium species, Candida albicans, and Aspergillus niger. The sensitivity, specificity, positive predictive value, negative predictive value, and efficiency for the ecfX assay were 75% (15/20), 95% (19/20), 94% (15/16), 79% (19/24), and 85% (34/40), respectively (Table 2). Positive testing demonstrated sigmoidal, S-shaped curves. Ct values of the positive samples ranged from 24.89 cycles up to 44.86 cycles. The sensitivity, specificity, positive predictive value, negative predictive value, and efficiency for the 16S rRNA assay were 70% (14/20), 100% (20/20), 100% (14/14), 77% (20/26), and 85% (34/40), respectively (Table 2). Positive testing demonstrated sigmoidal, S-shaped curves. Ct values of the positive samples ranged from 25.04 cycles up to 39.41 cycles. The fifteen ecfX purified PCR-positive samples demonstrated the presence of the expected 63 bp band, as seen in the positive control gel and representative sample gel of the direct cornea sample from patient T39 (Figures 1A and 1B). After being cut with the NcoI restriction enzyme, fourteen of the fifteen samples demonstrated the expected restriction digest patterns of the 44 bp and 19 bp fragments. The expected cuts of the 63 bp amplicon by NcoI are depicted in the schematic diagram (Figure 1C.). The fourteen 16S rRNA purified PCR-positive products demonstrated the presence of the 144 bp band, as seen in the positive control gel and representative sample gel (Figure 2A,B). All fourteen samples had the expected digest patterns, after being cut with the SacII restriction enzyme, demonstrating the presence of a 97 and a 47 bp fragment. The expected cuts of the 144 bp amplicon by SacII are depicted in the schematic diagram (Figure 2C).
Table 2. Clinical descriptive statistics for Pseudomonas aeruginosa DNA detection by real-time PCR.
| Descriptive Statistics | ecfX gene | 16S rRNA gene |
| True-positives | 20 | 20 |
| True-negatives | 20 | 20 |
| Sensitivity | 75% (15/20) | 70% (14/20) |
| Specificity | 95% (19/20) | 100% (20/20) |
| Positive predictive value | 94% (15/16) | 100% (14/14) |
| Negative predictive value | 79% (19/24) | 77% (20/26) |
| Efficiency | 85% (34/40) | 85% (34/40) |
DISCUSSION
Validation of PCR testing for the detection of PA is necessary for laboratory accreditation and approval to conduct testing in each laboratory. Ophthalmic specialty laboratories may be able to produce the number of keratitis specimens to validate testing, but most general laboratories would not devote the time and effort to validate testing for an infrequent testing request. Each laboratory must decide the practicality of PCR with comparison to conventional testing. In the present study, culture on agar media (standard testing) was determined to be better than PCR. In addition, standard testing is more available to all clinicians and it provides the in vitro antibiotic susceptibility that directs the appropriate topical therapy for PA keratitis. It is true that PCR can be faster than culture if the specimen is processed immediately after collection, but the likelihood that a laboratory has the manpower and time to provide such an expedite service is slim. Although PCR testing for PA DNA appears less practical than standard microbiological techniques, it may be important to the tertiary care of keratitis in cases where an etiologic agent is unknown and eyesight is threatened. PA is a frequent bacterial pathogen especially in the contact lens wearer[11]. Pretreatment without laboratory studies can prevent a definitive diagnosis whether the differential is bacterial, viral, fungal, or amoebic. Delayed or incorrect antibiotic treatment can result in an extended chronic keratitis that may require eventual transplantation. In these cases, PCR detection of PA DNA may be very important. For example, using our validated PCR assay, we tested two patients who were culture negative for PA. One case was a patient that was culture positive in the validating study, but one month later after extensive treatment and a lack of clinical resolution, the culture was negative and tested PCR positive. Another patient that presented with no definitive diagnosis also tested PCR positive. These are the type of cases that PCR may provide as an important adjunct for detecting PA DNA from ocular infection.
The ecfX gene has been shown to be a target for identification of PA[4]-[6] but to our knowledge, not yet been tested against keratitis isolates and direct patient samples from the cornea. In the current study, all of the PA isolates collected from keratitis contained the ecfX PCR gene. The ecfX gene was not present in any of the non-PA isolates. The less than optimal sensitivity of 75% may be due to culture collection. Culturing the cornea may be precarious due to the fear of perforation and the lack of experience of the attending ophthalmologist. In general, multiple media are inoculated and the sample for PCR testing was allotted from the viral culture medium. Focusing on collecting a sample for PCR alone may increase the sensitivity. In the present study, we demonstrated (data not presented) that there were no factors inhibiting PCR by amplifying VZV in the true-positive samples that tested PCR negative for PA DNA. In addition to sampling error, the storage of the amplified PCR products and freeze-thaw issues could have affected the real-time PCR results. Also, DNA could have been lost during the DNA extraction process, leading to a concentration lower than that of the calculated limit of detection. We chose the universal primers and probes based on the size of the 144 bp target amplified product. The recommended target size for optimal real-time PCR results is between 75 and 200 bp[8]. As with the ecfX target, the sensitivity of detecting PA DNA was less than optimal at 70%. Once again, an increase in sensitivity could parallel a focus on sampling collection. In conclusion, real-time PCR provides clinical laboratories with a supplementary diagnostic tool to culture in the diagnosis of PA keratitis. Species-specific primers and primers targeted at the 16S rRNA gene have the ability to detect bacterial DNA extracted from isolates and direct patient samples. The ecfX real-time PCR assay[4] was adapted to the Cepheid SmartCycler® II real-time PCR system and was demonstrated to detect PA isolates and PA from direct de-identified patient samples. Universal primer real-time PCR assays targeting the highly conserved bacterial 16S rRNA gene, once validated, may be able to detect bacteria from the cornea that do not grow out in culture. Although both assays detected different genes of PA, real-time PCR testing needs to produce higher sensitivity values for clinical diagnostic purposes. Restriction digestion and PAGE analysis are tools that can be used to validate results from real-time PCR assays, as used in this study. By working to improve the sensitivity of PCR testing, clinical laboratories will have another diagnostic tool to use in diagnosing patients with keratitis.
Acknowledgments
We are grateful to The Pennsylvania Lions Club and The Charles T. Campbell Foundation for financing the purchase of equipment and essential materials. A core grant for Vision Research NIH EY008098 provided expertise within the molecular module, and Research to Prevent Blindness has provided continued support of the ophthalmology department. We would like to thank Jean-Paul Vergnes, Eric Kalivoda, and Paul R. Kinchington for their expertise and aid in this project, as well as the Graduate School of Public Health at the University of Pittsburgh.
Footnotes
Foundation item: Supported by The Pennsylvania Lions Club and The Charles T. Campbell Foundation. A core grant for Vision Research NIH EY008098 provided expertise within the molecular module, and Research to Prevent Blindness has provided continued support of the ophthalmology department.
REFERENCES
- 1.Choy M H, Stapleton F, Willcox M D, Zhu H. Comparison of virulence factors in Pseudomonas aeruginosa strains isolated from contact lens- and non-contact lens-related keratitis. J Med Microbiol. 2008;57:1539–1546. doi: 10.1099/jmm.0.2008/003723-0. [DOI] [PubMed] [Google Scholar]
- 2.Huang AJW, Wichiensin P, Yang MC. Bacterial Keraitis. In: Krachmer J H, Mannis M J, Holland E J, editors. Cornea. 2nd Ed. Vol 1. Elsevier Mosby; 2004. p. 1005. [Google Scholar]
- 3.Wilhelmus K R. Indecision about corticosteroids for bacterial keratitis: an evidence-based update. Ophthalmology. 2002;109:835–842. doi: 10.1016/s0161-6420(02)00963-6. [DOI] [PubMed] [Google Scholar]
- 4.Lavenir R, Jocktane D, Laurent F, Nazaret S, Cournoyer B. Improved reliability of Pseudomonas aeruginosa PCR detection by the use of the species-specific ecfX gene target. J Microbiol Methods. 2007;70:20–29. doi: 10.1016/j.mimet.2007.03.008. [DOI] [PubMed] [Google Scholar]
- 5.Anuj S N, Whiley D M, Kidd T J, Bell S C, Wainwright C E, Nissen M D, Sloots T P. Identification of Pseudomonas aeruginosa by a duplex real-time polymerase chain reaction assay targeting the ecfX and the gyrB genes. Diagn Microbiol Infect Dis. 2009;63:127–131. doi: 10.1016/j.diagmicrobio.2008.09.018. [DOI] [PubMed] [Google Scholar]
- 6.Cattoir V, Gilibert A, Le Glauec J-M, Launay N, Bait-Merabet L, Legrand P. Rapid detection of Pseudomonas aeruginosa from positive blood cultures by quantitative PCR. Ann Clin Microbiol & Antimicrob. 2010;9:21–25. doi: 10.1186/1476-0711-9-21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Shanks R M, Kadouri D E, MacEachran D P, O'Toole G A. New yeast recombineering tools for bacteria. Plasmid. 2009;62:88–97. doi: 10.1016/j.plasmid.2009.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bio-Rad Laboratories, Inc. Real-time PCR applications guide. 2006. pp. 2–24.
- 9.Corpet F. Multiple sequence alignment with hierarchical clustering. Nucleic Acids Res. 1988;16:10881–90. doi: 10.1093/nar/16.22.10881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Schabereiter-Gurtner C, Nehr M, Apfalter P, Makristathis A, Rotter M L, Hirschl A M. Evaluation of a protocol for molecular broad-range diagnosis of culture-negative bacterial infections in clinical routine diagnosis. J Appl Microbiol. 2008;104:1228–1237. doi: 10.1111/j.1365-2672.2007.03648.x. [DOI] [PubMed] [Google Scholar]
- 11.Mueller JB, McStay CM. Ocular infection and inflammation. Emerg Med Clin North Am. 2008;26:57–72. doi: 10.1016/j.emc.2007.10.004. [DOI] [PubMed] [Google Scholar]