Abstract
We compared cleavage efficiencies of mono-molecular and bipartite model RNAs as substrates for RNase P RNAs (M1 RNAs) and holoenzymes from E. coli and Thermus thermophilus, an extreme thermophilic eubacterium. Acceptor stem and T arm of pre-tRNA substrates are essential recognition elements for both enzymes. Impairing coaxial stacking of acceptor and T stems and omitting the T loop led to reduced cleavage efficiencies. Small model substrates were less efficiently cleaved by M1 RNA and RNase P from T. thermophilus than by the corresponding E. coli activities. Competition kinetics and gel retardation studies showed that truncated tRNA substrates are less tightly bound by RNase P and M1 RNA from both bacteria. Our data further indicate that (pre-)tRNA interacts stronger with E. coli than T. thermophilus M1 RNA. Thus, low cleavage efficiencies of truncated model substrates by T. thermophilus RNase P or M1 RNA could be explained by a critical loss of important contact points between enzyme and substrate. In addition, acceptor stem--T arm substrates, composed of two synthetic RNA fragments, have been designed to mimic internal cleavage of any target RNA molecule available for base pairing.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bhattacharyya A., Murchie A. I., Lilley D. M. RNA bulges and the helical periodicity of double-stranded RNA. Nature. 1990 Feb 1;343(6257):484–487. doi: 10.1038/343484a0. [DOI] [PubMed] [Google Scholar]
- Burkard U., Willis I., Söll D. Processing of histidine transfer RNA precursors. Abnormal cleavage site for RNase P. J Biol Chem. 1988 Feb 15;263(5):2447–2451. [PubMed] [Google Scholar]
- Castenholz R. W. Thermophilic blue-green algae and the thermal environment. Bacteriol Rev. 1969 Dec;33(4):476–504. doi: 10.1128/br.33.4.476-504.1969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciesiołka J., Lorenz S., Erdmann V. A. Structural analysis of three prokaryotic 5S rRNA species and selected 5S rRNA--ribosomal-protein complexes by means of Pb(II)-induced hydrolysis. Eur J Biochem. 1992 Mar 1;204(2):575–581. doi: 10.1111/j.1432-1033.1992.tb16670.x. [DOI] [PubMed] [Google Scholar]
- Ciesiołlka J., Lorenz S., Erdmann V. A. Different conformational forms of Escherichia coli and rat liver 5S rRNA revealed by Pb(II)-induced hydrolysis. Eur J Biochem. 1992 Mar 1;204(2):583–589. doi: 10.1111/j.1432-1033.1992.tb16671.x. [DOI] [PubMed] [Google Scholar]
- Cronenberger J. H., Erdmann V. A. Stimulation of polypeptide polymerization by blocking of free sulphydryl groups in Escherichia coli ribosomal proteins. J Mol Biol. 1975 Jun 15;95(1):125–137. doi: 10.1016/0022-2836(75)90340-x. [DOI] [PubMed] [Google Scholar]
- Ehresmann C., Baudin F., Mougel M., Romby P., Ebel J. P., Ehresmann B. Probing the structure of RNAs in solution. Nucleic Acids Res. 1987 Nov 25;15(22):9109–9128. doi: 10.1093/nar/15.22.9109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forster A. C., Altman S. External guide sequences for an RNA enzyme. Science. 1990 Aug 17;249(4970):783–786. doi: 10.1126/science.1697102. [DOI] [PubMed] [Google Scholar]
- Green C. J., Vold B. S. Structural requirements for processing of synthetic tRNAHis precursors by the catalytic RNA component of RNase P. J Biol Chem. 1988 Jan 15;263(2):652–657. [PubMed] [Google Scholar]
- Grosshans C. A., Cech T. R. A hammerhead ribozyme allows synthesis of a new form of the Tetrahymena ribozyme homogeneous in length with a 3' end blocked for transesterification. Nucleic Acids Res. 1991 Jul 25;19(14):3875–3880. doi: 10.1093/nar/19.14.3875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guerrier-Takada C., Gardiner K., Marsh T., Pace N., Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983 Dec;35(3 Pt 2):849–857. doi: 10.1016/0092-8674(83)90117-4. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Lumelsky N., Altman S. Specific interactions in RNA enzyme-substrate complexes. Science. 1989 Dec 22;246(4937):1578–1584. doi: 10.1126/science.2480641. [DOI] [PubMed] [Google Scholar]
- Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983 Jun 5;166(4):557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- Hartmann R. K., Erdmann V. A. Analysis of the gene encoding the RNA subunit of ribonuclease P from T. thermophilus HB8. Nucleic Acids Res. 1991 Nov 11;19(21):5957–5964. doi: 10.1093/nar/19.21.5957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartmann R. K., Toschka H. Y., Erdmann V. A. Processing and termination of 23S rRNA-5S rRNA-tRNA(Gly) primary transcripts in Thermus thermophilus HB8. J Bacteriol. 1991 Apr;173(8):2681–2690. doi: 10.1128/jb.173.8.2681-2690.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartmann R. K., Ulbrich N., Erdmann V. A. An unusual rRNA operon constellation: in Thermus thermophilus HB8 the 23S/5S rRNA operon is a separate entity from the 16S rRNA operon. Biochimie. 1987 Oct;69(10):1097–1104. doi: 10.1016/0300-9084(87)90009-5. [DOI] [PubMed] [Google Scholar]
- Holm P. S., Krupp G. The acceptor stem in pre-tRNAs determines the cleavage specificity of RNase P. Nucleic Acids Res. 1992 Feb 11;20(3):421–423. doi: 10.1093/nar/20.3.421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kahle D., Wehmeyer U., Krupp G. Substrate recognition by RNase P and by the catalytic M1 RNA: identification of possible contact points in pre-tRNAs. EMBO J. 1990 Jun;9(6):1929–1937. doi: 10.1002/j.1460-2075.1990.tb08320.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirsebom L. A., Svärd S. G. The kinetics and specificity of cleavage by RNase P is mainly dependent on the structure of the amino acid acceptor stem. Nucleic Acids Res. 1992 Feb 11;20(3):425–432. doi: 10.1093/nar/20.3.425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koski R. A., Clarkson S. G. Synthesis and maturation of Xenopus laevis methionine tRNA gene transcripts in homologous cell-free extracts. J Biol Chem. 1982 Apr 25;257(8):4514–4521. [PubMed] [Google Scholar]
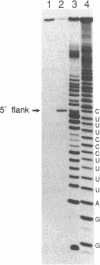
- Krupp G., Kahle D., Vogt T., Char S. Sequence changes in both flanking sequences of a pre-tRNA influence the cleavage specificity of RNase P. J Mol Biol. 1991 Feb 20;217(4):637–648. doi: 10.1016/0022-2836(91)90522-8. [DOI] [PubMed] [Google Scholar]
- Leontis N., DaLio A., Strobel M., Engelke D. Effects of tRNA-intron structure on cleavage of precursor tRNAs by RNase P from Saccharomyces cerevisiae. Nucleic Acids Res. 1988 Mar 25;16(6):2537–2552. doi: 10.1093/nar/16.6.2537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y., Guerrier-Takada C., Altman S. Targeted cleavage of mRNA in vitro by RNase P from Escherichia coli. Proc Natl Acad Sci U S A. 1992 Apr 15;89(8):3185–3189. doi: 10.1073/pnas.89.8.3185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mans R. M., Guerrier-Takada C., Altman S., Pleij C. W. Interaction of RNase P from Escherichia coli with pseudoknotted structures in viral RNAs. Nucleic Acids Res. 1990 Jun 25;18(12):3479–3487. doi: 10.1093/nar/18.12.3479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClain W. H., Guerrier-Takada C., Altman S. Model substrates for an RNA enzyme. Science. 1987 Oct 23;238(4826):527–530. doi: 10.1126/science.2443980. [DOI] [PubMed] [Google Scholar]
- Milligan J. F., Uhlenbeck O. C. Synthesis of small RNAs using T7 RNA polymerase. Methods Enzymol. 1989;180:51–62. doi: 10.1016/0076-6879(89)80091-6. [DOI] [PubMed] [Google Scholar]
- Pyle A. M., McSwiggen J. A., Cech T. R. Direct measurement of oligonucleotide substrate binding to wild-type and mutant ribozymes from Tetrahymena. Proc Natl Acad Sci U S A. 1990 Nov;87(21):8187–8191. doi: 10.1073/pnas.87.21.8187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quigley G. J., Rich A. Structural domains of transfer RNA molecules. Science. 1976 Nov 19;194(4267):796–806. doi: 10.1126/science.790568. [DOI] [PubMed] [Google Scholar]
- Reich C., Olsen G. J., Pace B., Pace N. R. Role of the protein moiety of ribonuclease P, a ribonucleoprotein enzyme. Science. 1988 Jan 8;239(4836):178–181. doi: 10.1126/science.3122322. [DOI] [PubMed] [Google Scholar]
- Smith D., Burgin A. B., Haas E. S., Pace N. R. Influence of metal ions on the ribonuclease P reaction. Distinguishing substrate binding from catalysis. J Biol Chem. 1992 Feb 5;267(4):2429–2436. [PubMed] [Google Scholar]
- Thurlow D. L., Shilowski D., Marsh T. L. Nucleotides in precursor tRNAs that are required intact for catalysis by RNase P RNAs. Nucleic Acids Res. 1991 Feb 25;19(4):885–891. doi: 10.1093/nar/19.4.885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vioque A., Arnez J., Altman S. Protein-RNA interactions in the RNase P holoenzyme from Escherichia coli. J Mol Biol. 1988 Aug 20;202(4):835–848. doi: 10.1016/0022-2836(88)90562-1. [DOI] [PubMed] [Google Scholar]