Abstract
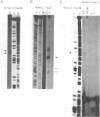
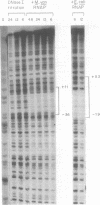
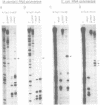
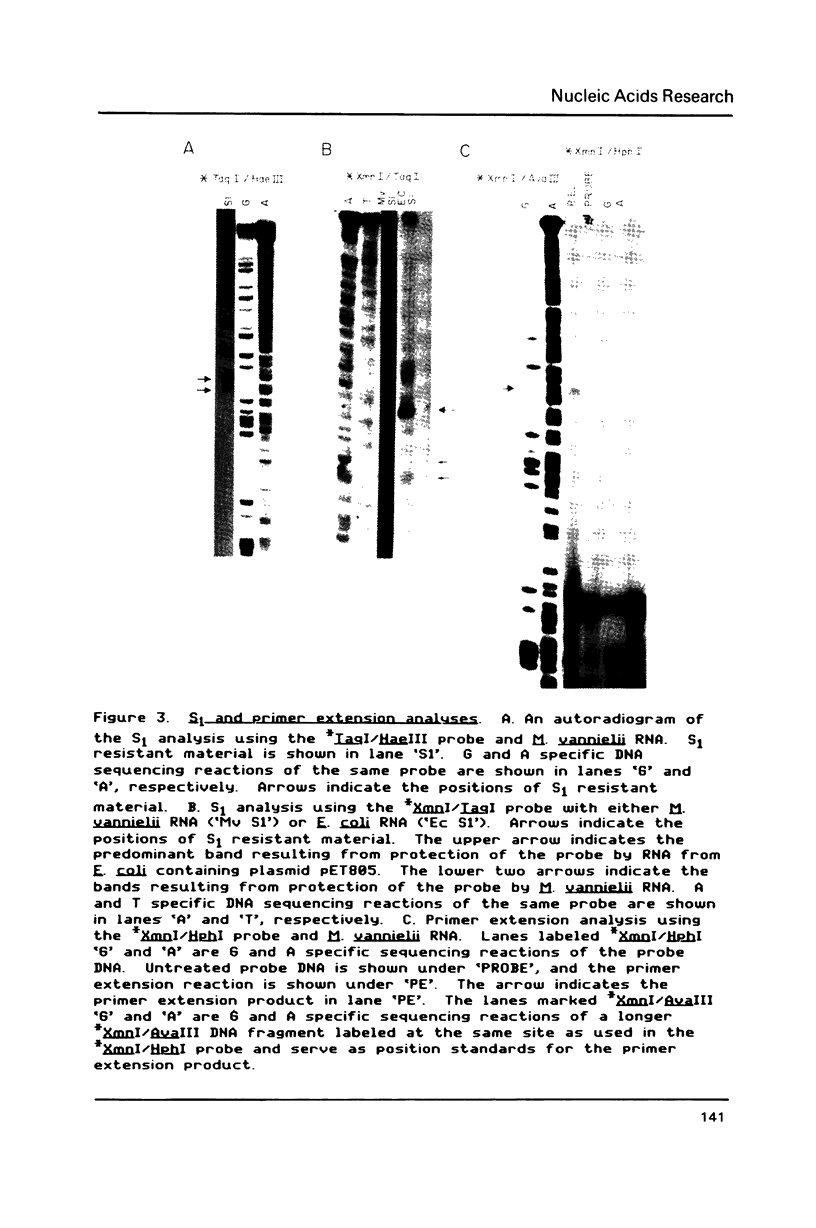
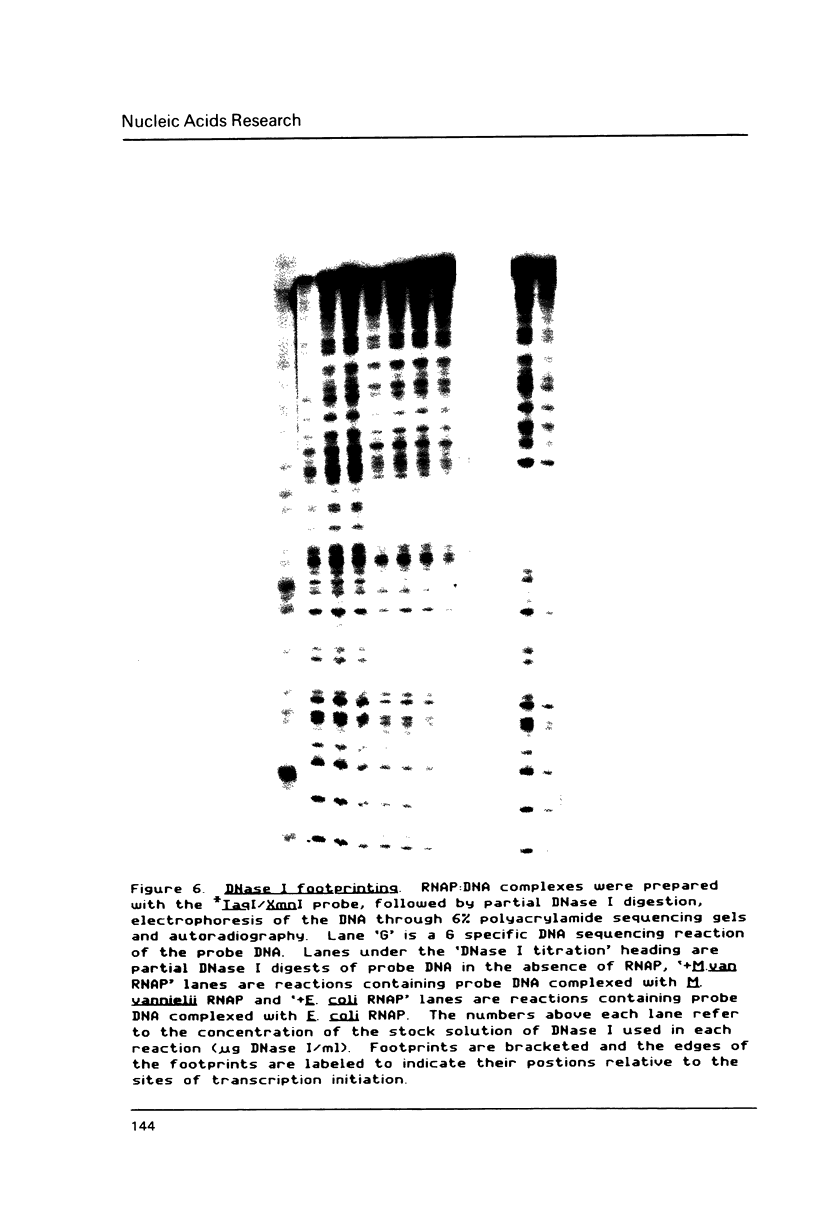
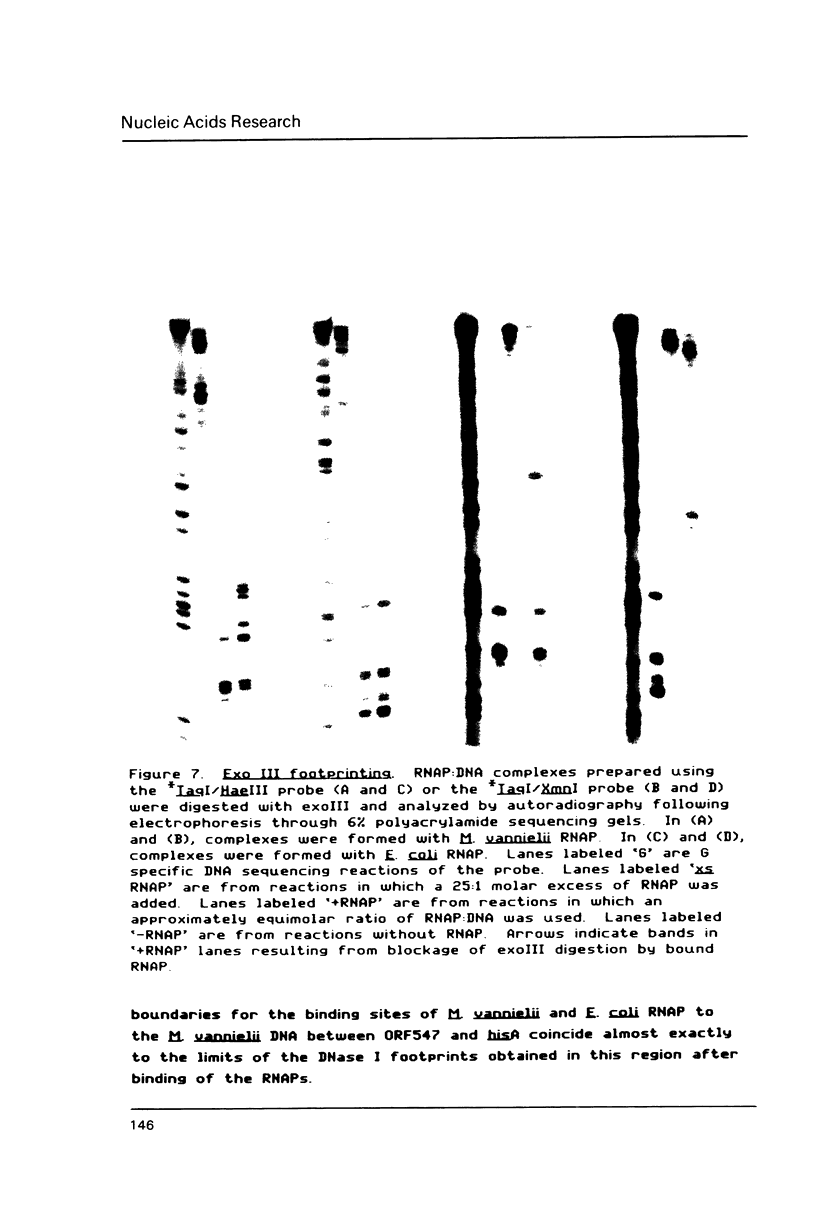
Transcription initiation of the hisA gene in vivo in the archaebacterium Methanococcus vannielii, as determined by nuclease S1 and primer extension analyses, occurs 73 base pairs (bp) upstream of the translation initiation site. Binding of M. vannielii RNA polymerase protects 43 bp of DNA, from 35 bp upstream (-35) to 8 bp downstream (+8) of the hisA mRNA initiation site, from digestion by DNase I and exonuclease III. An A + T rich region, with a sequence which conforms to the consensus sequence for promoters of stable RNA-encoding genes in methanogens, is found at the same location (-25) upstream of the polypeptide-encoding hisA gene. It appears therefore that a TATA-like sequence is also an element of promoters which direct transcription of polypeptide-encoding genes in this archaebacterium.
Full text
PDF












Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Beckler G. S., Reeve J. N. Conservation of primary structure in the hisI gene of the archaebacterium, Methanococcus vannielii, the eubacterium Escherichia coli, and the eucaryote Saccharomyces cerevisiae. Mol Gen Genet. 1986 Jul;204(1):133–140. doi: 10.1007/BF00330200. [DOI] [PubMed] [Google Scholar]
- Betlach M., Friedman J., Boyer H. W., Pfeifer F. Characterization of a halobacterial gene affecting bacterio-opsin gene expression. Nucleic Acids Res. 1984 Oct 25;12(20):7949–7959. doi: 10.1093/nar/12.20.7949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chan P. T., Lebowitz J. The coupled use of 'footprinting' and exonuclease III methodology for RNA polymerase binding and initiation. Application for the analysis of three tandem promoters at the control region of colicin El. Nucleic Acids Res. 1983 Feb 25;11(4):1099–1116. doi: 10.1093/nar/11.4.1099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ciliberto G., Castagnoli L., Melton D. A., Cortese R. Promoter of a eukaryotic tRNAPro gene is composed of three noncontiguous regions. Proc Natl Acad Sci U S A. 1982 Feb;79(4):1195–1199. doi: 10.1073/pnas.79.4.1195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dynan W. S., Tjian R. The promoter-specific transcription factor Sp1 binds to upstream sequences in the SV40 early promoter. Cell. 1983 Nov;35(1):79–87. doi: 10.1016/0092-8674(83)90210-6. [DOI] [PubMed] [Google Scholar]
- Galas D. J., Schmitz A. DNAse footprinting: a simple method for the detection of protein-DNA binding specificity. Nucleic Acids Res. 1978 Sep;5(9):3157–3170. doi: 10.1093/nar/5.9.3157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinkle D. C., Chamberlin M. J. Studies of the binding of Escherichia coli RNA polymerase to DNA. I. The role of sigma subunit in site selection. J Mol Biol. 1972 Sep 28;70(2):157–185. doi: 10.1016/0022-2836(72)90531-1. [DOI] [PubMed] [Google Scholar]
- Huet J., Schnabel R., Sentenac A., Zillig W. Archaebacteria and eukaryotes possess DNA-dependent RNA polymerases of a common type. EMBO J. 1983;2(8):1291–1294. doi: 10.1002/j.1460-2075.1983.tb01583.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ptashne M., Backman K., Humayun M. Z., Jeffrey A., Maurer R., Meyer B., Sauer R. T. Autoregulation and function of a repressor in bacteriophage lambda. Science. 1976 Oct 8;194(4261):156–161. doi: 10.1126/science.959843. [DOI] [PubMed] [Google Scholar]
- Reiter W. D., Palm P., Voos W., Kaniecki J., Grampp B., Schulz W., Zillig W. Putative promoter elements for the ribosomal RNA genes of the thermoacidophilic archaebacterium Sulfolobus sp. strain B12. Nucleic Acids Res. 1987 Jul 24;15(14):5581–5595. doi: 10.1093/nar/15.14.5581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shalloway D., Kleinberger T., Livingston D. M. Mapping of SV40 DNA replication origin region binding sites for the SV40 T antigen by protection against exonuclease III digestion. Cell. 1980 Jun;20(2):411–422. doi: 10.1016/0092-8674(80)90627-3. [DOI] [PubMed] [Google Scholar]
- Thomm M., Madon J., Stetter K. O. DNA-dependent RNA polymerases of the three orders of methanogens. Biol Chem Hoppe Seyler. 1986 Jun;367(6):473–481. doi: 10.1515/bchm3.1986.367.1.473. [DOI] [PubMed] [Google Scholar]
- Thomm M., Stetter K. O. Transcription in methanogens. Evidence for specific in vitro transcription of the purified DNA-dependent RNA polymerase of Methanococcus thermolithotrophicus. Eur J Biochem. 1985 Jun 3;149(2):345–351. doi: 10.1111/j.1432-1033.1985.tb08932.x. [DOI] [PubMed] [Google Scholar]
- Wich G., Hummel H., Jarsch M., Bär U., Böck A. Transcription signals for stable RNA genes in Methanococcus. Nucleic Acids Res. 1986 Mar 25;14(6):2459–2479. doi: 10.1093/nar/14.6.2459. [DOI] [PMC free article] [PubMed] [Google Scholar]