Abstract
Traditionally, image studies evaluating the effectiveness of computer-aided diagnosis (CAD) use a single label from a medical expert compared with a single label produced by CAD. The purpose of this research is to present a CAD system based on Belief Decision Tree classification algorithm, capable of learning from probabilistic input (based on intra-reader variability) and providing probabilistic output. We compared our approach against a traditional decision tree approach with respect to a traditional performance metric (accuracy) and a probabilistic one (area under the distance–threshold curve—AuCdt). The probabilistic classification technique showed notable performance improvement in comparison with the traditional one with respect to both evaluation metrics. Specifically, when applying cross-validation technique on the training subset of instances, boosts of 28.26% and 30.28% were noted for the probabilistic approach with respect to accuracy and AuCdt, respectively. Furthermore, on the validation subset of instances, boosts of 20.64% and 23.21% were noted again for the probabilistic approach with respect to the same two metrics. In addition, we compared our CAD system results with diagnostic data available for a small subset of the Lung Image Database Consortium database. We discovered that when our CAD system errs, it generally does so with low confidence. Predictions produced by the system also agree with diagnoses of truly benign nodules more often than radiologists, offering the possibility of reducing the false positives.
Keywords: Chest CT, Computer-aided diagnosis (CAD), Feature extraction, Image analysis, Machine learning, Radiographic image interpretation, Computer-assisted
Introduction
Consensus interpretation of imaging studies is defined as the agreement reached when two or more radiologists report the imaging findings [1]. In the clinical practice of radiology, consensus, when interpreting the medical image, is hardly ever reached, especially with diagnostic tests where interpreted components are subjective. Several studies showed substantial observer variability not only among non-specialized radiologists during standard clinical reporting [2, 3], but also at the expert level of image interpretation [4, 5].
In the computer-aided diagnosis (CAD) literature, consensus image interpretation is used as a standard of reference to which the CAD method is compared. Although there are several CAD studies [6–8] that looked at the performance of individual radiologists before and after using CAD, most CAD systems consider consensus as the reference standard for development and evaluation when interpretations of multiple radiologists are available. Within the consensus approach, either only the consensus opinion is known [9–11] or individual interpretations are known but a consensus opinion is formed [12–20].
In a comparison of standard reading and computer-aided detection on a national proficiency test of screening mammography [12], the CAD performance was compared with double reading emulated by combining evaluations of four experienced radiologists. The combination was performed in such a manner that a case was considered positive if at least one of the observers marked it as positive. In another study for comparing computer-aided detection versus independent double reading of masses in mammograms, Karssemeijer et al. [13] evaluated three reading conditions: a single radiologist’s interpretation, emulated double reading (emulation was performed by combining radiologist interpretations pair-wise and then averaging the results of produced pairs), and emulated CAD as a second reader by combining CAD results with single radiologist interpretations. Muratmatsu et al. [14] used the mean or mode to form a consensus when investigating a psychophysical measure for evaluation of similar images for mammographic masses. Tao et al. [16] described a system for joint segmentation and spiculation detection of mammographic masses. The system demonstrated overlap ratios between reference truth and produced segmentation of 0.766 and 0.642 for the whole mass and margin portion of mass correspondingly. In order to create a reference truth for the segmentation, the consensus region was formed out of five available radiologists boundaries in such way that the pixel was considered a part of the region if it was marked by at least three out of five radiologists. In the work of Sahiner et al. [17] dedicated to extraction of image features for mammographic mass characterization, artificial consensus was employed to build one of the reference standards: the ratings provided by the majority of radiologists (two out of three) were used to label the mass.
In the realm of CAD for pulmonary nodules, there are several studies that investigate the use of the consensus approach as a reference truth. The creation of the National Cancer Institute (NCI) Lung Image Database Consortium (LIDC) dataset [21] also allowed for investigating the variability among readers as it includes ratings for nine nodules characteristics and the boundary delineations by up to four radiologists. Two recent studies by [18, 19] looked at the impact on reader agreement on reported CAD systems when using the LIDC dataset. Furthermore, Armato et al. [20] assessed the radiologist performance in the detection of lung nodules and demonstrated the strong impact of the definition of “truth” on the lung nodule detection process. A total of 24 reference standards were created from the annotations of four radiologists in the following manner: 1—all possible pair-wise combinations of four radiologists (six combinations) using logical AND and OR operators (total of 6 × 2 reference standards); 2—all possible triplet combination of four radiologists (four combinations) using AND, OR, and majority operators (total of 4 × 3 reference standards). The number of ≥3 mm nodules of interest ranged from 15 to 89 in different reference standards. On the next step the performance of each of four radiologists was evaluated against all the available reference standards. The results showed mean sensitivities (across radiologists) ranging from 51% to 83.2% and mean false-positive rates ranging from 0.33 to 1.39 for different reference standards.
In summary, the use of a consensus standard does not show the extent to which a new technique is accurate in establishing a diagnosis, but only the extent to which a new technique agrees with the consensus reading. This becomes problematic since although a consensus might be reached, nothing guarantees that the consensual result is true, as mere consensus does not imply the correctness of a diagnostic decision [1]. The American College of Radiology Imaging Network (ACRIN) recently highlighted that if imaging research results are aimed to be translated into the clinical practice of radiology, then the potential variability among observers should require the same level of attention as the potential variability among study subjects and variability in the imaging devices [22]. As a result, ACRIN has recommended the incorporation of the expected variability between observers into sample size calculations and other fundamental parameters that determine a given experimental design [22]. Furthermore, in a Radiology journal editorial, Bankier et al. [1] concluded that the use of consensus readings should be regarded as a study limitation and recommended that researchers look into more robust and viable alternatives to reference standards based on consensus.
To the best of our knowledge, none of the previously published works proposed classification system for learning from and predicting the whole distribution of radiologists’ annotations. Such approach can be beneficial for CAD purposes for several reasons: learning from the distribution of annotations will help to avoid the loss of potentially important information when classification system has no knowledge on radiologists’ level of expertise; probabilistic prediction can carry important information besides the malignancy of the nodule (shape of the distribution can be used as an indicator of complexity of the particular classification task).
In this work, we propose the investigation of CAD performance based on the distribution of radiologists’ interpretations rather than their consensus. Specifically, instead of considering the majority malignancy rating among R radiologist ratings  for a certain case, we consider the probability distribution of these ratings
for a certain case, we consider the probability distribution of these ratings  across each of the
across each of the  malignancy classes (k ≥ 2) where p(ci) is calculated as the number of radiologists who assigned the case to class ci over the total number of radiologists. To deal with the inter-observer variability quantified through a vector of class probabilities, we propose a probabilistic classification approach based on belief decision trees [23]. Although other approaches can be used such as those based on support vector machines with pair-wise coupling [24]; however, we chose to employ Decision Tree classification approach in this research because the decision rules can be easily visualized and understood; furthermore, the decision trees approach also has a feature selection algorithm embedded in the classification process which allows to understand which image features are more important.
malignancy classes (k ≥ 2) where p(ci) is calculated as the number of radiologists who assigned the case to class ci over the total number of radiologists. To deal with the inter-observer variability quantified through a vector of class probabilities, we propose a probabilistic classification approach based on belief decision trees [23]. Although other approaches can be used such as those based on support vector machines with pair-wise coupling [24]; however, we chose to employ Decision Tree classification approach in this research because the decision rules can be easily visualized and understood; furthermore, the decision trees approach also has a feature selection algorithm embedded in the classification process which allows to understand which image features are more important.
Furthermore, in addition to the inter-observer variance, the LIDC dataset presents another interesting challenge in dealing with degrees of uncertainty such as: 1 = ‘highly unlikely’, 2 = ‘moderately unlikely’, 3 = ‘indeterminate’, 4 = ‘moderately suspicious’, 5 = ‘highly suspicious’. This is a situation present in most biological systems where there is overlap or gradation between normal and abnormal [25]. Given that interpretation disagreement often occurs for these complex cases and that setting thresholds for transforming the problem to the two conventional states, malignant versus benign, is challenging as thresholds can vary among observers, we propose to also investigate this classification problem as a multi-class classification problem in which k is >2.
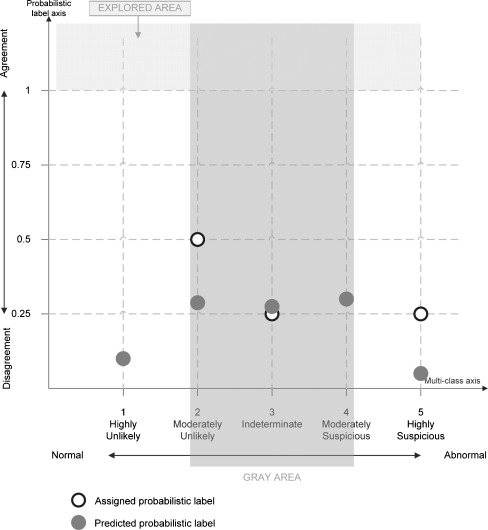
In conclusion, our proposed work aims to open new avenues of exploration for building and evaluating CAD systems in terms of including both the variability among readers—recently identified as a current limitation of the current CAD studies [1] and the degree of uncertainty (‘lack of diagnostic confidence’)—the perceived Achilles heel of the radiology report [26]. Figure 1 provides a visual representation of the probabilistic multi-class space we aim to explore.
Fig. 1.
The probabilistic multi-class space in which one nodule was interpreted by four radiologists: two assigned rating 2, one rating 3, and the fourth one label 5 (white points). The dark points represent the predicted probabilistic ratings for the same nodule. Explored area represents those cases that take into account agreement/consensus when predicting malignancy; the gray area represents those cases for which the nodules are not clearly benign or malignant
The rest of the paper is organized as follows: Section “Materials and Methods” presents the LIDC dataset, traditional decision trees, and the belief decision trees for our proposed approach; Section “Results” presents our results and findings; Section “Discussion” discusses the results in the context of a LIDC subset for which the gold truth (from biopsies and follow up studies) is available; and Section “Conclusions” summarizes our presented work and describes directions for future work.
Materials and Methods
This section describes the LIDC dataset employed in this research, provides details on image extraction process, and explains both traditional and multi-class probabilistic classification algorithms. The final sub-section discusses the performance evaluation technique for the multi-class probabilistic classification system.
LIDC Dataset
The publicly available LIDC database (downloadable through the National Cancer Institute’s Imaging Archive web site—http://ncia.nci.nih.gov/) provides the image data, the radiologists’ nodule outlines, and the radiologists’ subjective ratings of nodule characteristics for this study. The LIDC database currently contains complete thoracic CT scans for 400 patients acquired over different periods of time and with various scanners.
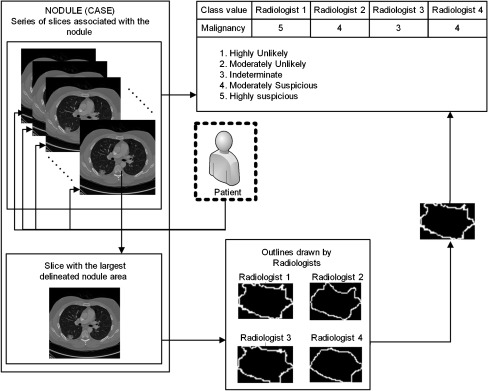
The Extensible Markup Language (XML) files accompanying the LIDC Digital Imaging and Communications in Medicine images contain the spatial locations of three types of lesions (nodules <3 mm in maximum diameter, but only if not clearly benign; nodules ≥3 mm but <30 mm regardless of presumed histology; and non-nodules ≥3 mm) as marked by a panel of four LIDC radiologists. For any lesion marked as a nodule ≥3 mm, the XML file contains the coordinates of nodule outlines constructed by any of the four LIDC radiologists who identified that structure as a nodule ≥3 mm. Since a nodule can appear on a different number of slices associated with that particular nodule (Fig. 2), each nodule is represented by a number of instances equal to the number of slices containing the nodule multiplied by the number of radiologists who marked that nodule on each slice. Moreover, any LIDC radiologist who identified a structure as a nodule ≥3 mm also provided subjective ratings for nine nodule characteristics: subtlety, internal structure, calcification, sphericity, margin, lobulation, spiculation, texture, and malignancy likelihood (1 = highly unlikely, 2 = moderately unlikely, 3 = indeterminate, 4 = moderately suspicious, and 5 = highly suspicious).
Fig. 2.
Visual representation of the LIDC data structure; one nodule is exemplified through the differences in the nodule’s outlines and malignancy ratings
Until recently, diagnostic truth was not available for the LIDC dataset and therefore, the ratings supplied by radiologists had to be used for training the computer-aided diagnostic system and evaluating the results. However, LIDC radiologists are anonymous and represented by ID numbers in the XML files. ID numbers are not consistent across different nodules; therefore, it is not possible to extract a subset of outlines or ratings provided by a particular radiologist for all LIDC nodules. If radiologists’ IDs were known, then a Bayesian approach could have been employed to model the performance of each radiologist for nodule classification. Therefore, either a consensus approach (mode rating per nodule) had to be employed as proposed by Zinovev et al. [27] or a new probabilistic multi-class approach as proposed in this paper.
Image Feature Extraction
For each nodule greater than 5 × 5 pixels (around 3 × 3 mm)—nodules smaller than this would not have yielded meaningful texture data—we calculate a set of 63 two-dimensional (2D), low-level image features from four categories: shape, texture, intensity, and size. Although each nodule is present in a sequence of slices, in this study we are considering only the slice in which the nodule has the largest area with respect to the outlines provided by up to four radiologists who annotated the corresponding nodule. Therefore, only the largest outline is considered as the most representative for feature extraction. Future work will be looking into other ways to quantify the outline such as using probabilistic p-maps from manually generated outlines or computer-based generated ones.
Size Features
We use the following seven features to quantify the size of the nodules: area, ConvexArea, perimeter, ConvexPerimeter, EquivDiameter, MajorAxisLength, and MinorAxisLength. The area and perimeter image features measure the actual number of pixels in the region and on the boundary, respectively. The ConvexArea and ConvexPerimeter measure the number of pixels in the convex hull and on the boundary of the convex hull corresponding to the nodule region. EquivDiameter is the diameter of a circle with the same area as the region. Lastly, the MajorAxisLength and MinorAxisLength give the length (in pixels) of the major and minor axes of the ellipse that has the same normalized second central moments as the region.
Shape Features
We use seven common image shape features: circularity, roughness, elongation, compactness, eccentricity, extent, and the standard deviation of the radial distance. Circularity is measured by dividing the circumference of the equivalent area circle by the actual perimeter of the nodule. Roughness can be measured by dividing the perimeter of the region by the convex perimeter. A smooth convex object, such as a perfect circle, will have a roughness of 1.0. The eccentricity is obtained using the ellipse that has the same second-moments as the region. The eccentricity is the ratio of the distance between the foci of the ellipse and its major axis length. The value is between 0 (a perfect circle) and 1 (a line). Solidity is the proportion of the pixels in the region to the pixels in the convex hull of the region. Extent is the proportion of the pixels in the bounding box (the smallest rectangle containing the region) that are also in the region. Finally, the RadialDistanceSD is the standard deviation of the distances from every boundary pixel to the centroid of the region.
Intensity Features
We use a total of nine intensity features: the minimum, maximum, mean, and standard deviation of the gray-level intensity of every pixel in each segmented nodule and the same four values for every background pixel in the bounding box containing each segmented nodule. Another feature, IntensityDifference, is the absolute value of the difference between the mean of the gray-level intensity of the segmented nodule and the mean of the gray-level intensity of its background.
Texture Features
Normally, texture analysis can be grouped into four categories: model-based, statistical-based, structural-based, and transform-based methods. Structural approaches seek to understand the hierarchal structure of the image, while statistical methods describe the image using pure numerical analysis of pixel intensity values. Transform approaches generally perform some kind of modification to the image, obtaining a new “response” image that is then analyzed as a representative proxy for the original image. Model-based methods are based on the concept of predicting pixel values based on a mathematical model. In this research, we focus on three well-known texture analysis techniques: co-occurrence matrices (a statistical-based method) which produces 11 features in total—contrast, correlation, entropy, energy, homogeneity, third-order moment, inverse variance, sum average, variance, cluster tendency, maximum probability; Gabor filters (a transform-based method) which produces 24 features in total—mean and standard deviation of 12 Gabor response images produced by varying orientation = 0°, 45°, 90°, and 135° and frequency = 0.3, 0.4, and 0.5 of the corresponding filter; and Markov Random Fields (a model-based method) which captures the local contextual information of an image and produces five features in total—means of four different Markov response images produced by varying orientation = 0°, 45°, 90°, 135° of the corresponding filter, along with the variance response image. All extracted features were represented by continuous values and were normalized using min–max normalization to the range from 0 to 1.
After completion of the feature extraction process, each nodule is represented using a 63-dimensional low-level image and a probabilistic label  where
where  represent the malignancy classes and p(ci) is the probability of that class for that corresponding nodule based on the radiologists’ interpretation.
represent the malignancy classes and p(ci) is the probability of that class for that corresponding nodule based on the radiologists’ interpretation.
Traditional Decision Trees
Most classification approaches used to build computer-aided diagnosis systems take a set of features as input and a single value class (malignant versus benign) output. These approaches include neural networks, support vector machines, linear discriminant analysis, and decision trees. In this paper, we employ a classification approach based on decision trees, and therefore we will describe a traditional algorithm for the decision trees, C4.5, and later compare it with our proposed approach based on belief decision trees.
The C4.5 classification approach described by Quinlan [28] is a decision tree classification algorithm that is able to construct classifiers based on continuous attributes. The classifier constructed by the model is a series of rules that are used to make a decision about the class membership of a test case, based on its attributes. In order to create a classifier, the algorithm performs a series of consecutive splits, each resulting in two nodes. If no splits are created after a certain node, this node is considered a terminal node (leaf). Each split is based on a threshold value of a single attribute. The decision on which attribute/threshold value is optimal for a particular split is based on the current training subset of classification instances. After the split is determined, the current training subset of classification instances is also split into two parts, according to the attribute/threshold of the split. All the consecutive splits are based on the corresponding subset. Optimality of the split is determined by a selection measure which is, in case of C4.5 algorithm, the gain ratio—an information-based measure that takes into account different numbers and different probabilities of test outcomes. The decision on whether the particular split should or should not take place is based on some stopping criterion or on a combination of several stopping criteria such as the number of instances that reached the node, maximum achievable splitting measure, uniformity of all the instances that reached the node, etc. For every terminal node of the constructed classifier, the class probability of each class is calculated as a ratio of the instances of that class that reached the node to the total number of instances that reached the node. The class with the highest calculated probability is considered the predicted class of a terminal node:
 |
1 |
(where S is the subset of instances that reached the node and  is the count of instances from S for every class ci)
is the count of instances from S for every class ci)
Belief Decision Trees
In this work, we propose the adaptation of the decision-tree-based classification approach proposed by Elouedi et al. [23] that is able to handle data instances with uncertain labels. Classification is performed in a manner similar to the one of regular decision trees. At every node, the instance that is currently being classified is redirected to the right or the left child of the node depending on the value of the attribute corresponding to this node. The process is repeated until the instance reaches the leaf node, which has a class membership probability distribution or a basic belief assignment (BBA) associated with it. This BBA is considered to be the newly predicted class of a classified instance. The main difference lies in the way a tree is constructed. At every node of the tree, starting with the root, the algorithm attempts to perform a split based on every attribute/feature existing in the dataset. Out of all constructed splits it determines the best one and uses it for growing the tree further. In order to define a best split, the algorithm performs the following steps:
First the algorithm computes the pignistic probability (probability calculated from a belief) of instance Ij for each possible class Ci for every instance in the dataset. Due to the fact that all BBAs in the LIDC dataset are singletons (meaning that each radiologist has to pick one class and one class only when assigning the rating to a nodule), the pignistic probability of instance Ij for class Ci is the ratio of observers who assigned the instance to a given class to the total number of observers for that instance:
 |
2 |
(where λl = {0,1,2,3,4} is rater count for every class ci)
Second, the algorithm computes the average pignistic probability function  over the set of S instances present in the subset that reached the node to get the average probability on each class:
over the set of S instances present in the subset that reached the node to get the average probability on each class:
 |
3 |
where Θ is a set of all possible classes.
On the following steps the algorithm uses average pignistic probability to calculate the entropy of the node and finally calculate information gain and gain ratio values for every possible subsequent split from entropy of the parent node and entropies of resulting child nodes to determine the optimal split. Every newly created node is associated with a BBA that is constructed by the average of the BBAs of all training cases that reached that node. The initial BBA of a single training case is a set of pignistic probabilities of all classes that the case can belong to, where the pignistic probability for each class is calculated using Eq. 2. The process of creating decision rules and new nodes is repeated until one of the stopping criteria is reached: (1) there is only one instance that reached this node; (2) all BBAs of the instances which reached the node are equal; (3) all the available attributes/features are split; or (4) the gain ratio of all possible further splits is less than or equal to 0. Once one of the stopping criteria is reached, the newly created node is considered to be a leaf. The whole algorithm is described in detail in [23]. There were several modifications that we made to the original algorithm proposed in [23]. While the approach described by Elouedi et al. [23] assumes a categorical nature of the attributes, attributes present in the LIDC dataset are continuous. We modified the algorithm to work with continuous attributes by setting the threshold on attribute value that will divide a set of instances into the subset. In order to choose an appropriate threshold, we employed the approach proposed by Quinlan [28]. The approach extracts a separate threshold from every distinct pair of values in the sorted set of attribute values and uses the gain ratio maximization criterion to determine the most suitable one. Furthermore, we also noticed that the Gain Ratio splitting criterion in the case of the LIDC dataset tends to favor very unbalanced splits, assigning a very small ratio of training instances (as small as stopping rules allow) to one of the node’s children at every case. As a result the produced trees contained large numbers of terminal nodes and were over fitted. In order to avoid this we decided to use information gain instead of gain ratio as a splitting criterion.
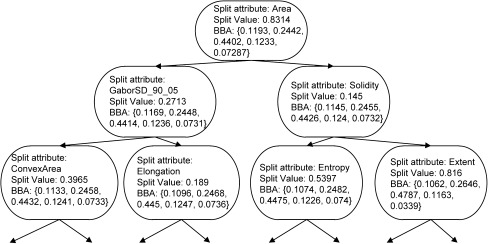
As the last change we modified one of the stopping rules setting the smallest number of instances that can reach any non-terminal node in a tree to 10 and setting the smallest number of instances that can reach the terminal node to 5. This change has also been implemented to avoid over fitting of the classification model. Figure 3 shows the first three levels of belief decision tree for malignancy semantic characteristic. The complete decision tree has not been shown due to its complexity. Each node on the figure shows the attribute and the value of this attribute on which the following split will be performed. The BBA associated with the node is also reported.
Fig. 3.
Upper 3 levels of belief decision tree constructed for malignancy semantic characteristic
Performance Evaluation
When evaluating a classification system that utilizes a probability distribution of ratings or classes as an input, and outputs a probability distribution of class membership, evaluation methods beyond accuracy should be used to better capture performance of the system. We propose the idea of a distance curve, in a similar vein to a receiver operating characteristic (ROC) curve [29], to assess the performance of our probabilistic multi-class classification approach. We were not able to construct ROC curves for the results that we obtained since the definitions of true-positive rate and false-positive rate are not directly applicable to the probabilistic multi-class classification task.
The distance curve is defined as follows: Let L be a sequence of instance labels, L = [L1,L2,…Lj…LN] where N is the number of instances and each Lj is a discrete probability density function over the label set λ.
Similarly, let P be a sequence of predicted labels, P = [P1,P2,…Pj…PN] where each Pj is discrete probability density function over the label set λ.
Let D be a normalized distance function defined on the instance/prediction pairs, D(Lj,Pj) ∈[0,1]. We define the distance–threshold curve as
 |
4 |
where x, threshold value for the distance, is defined from 0 to 1, and the [] are Iverson brackets, which equal 1 when the statement inside the brackets is true and 0 otherwise. It can be seen that the values of the curve itself are between 0 and 1 and that the curve is monotonically increasing.
We define the area under the distance–threshold curve simply as
 |
5 |
To generate the curve, we varied the thresholds of distance between the distributions for the classification to be considered “accurate.” For example, if we looked for nodules that have a normalized distance of 0, with 0 being a threshold value, between the input and output distributions, we would find little to none. As we increase the distance we find more and more nodules within that threshold. With a normalized distance–threshold of 1 between distributions, all the nodules would be considered correct or accurate. Once the curve is generated, the area under the distance–threshold curve (AuCdt) was used as the metric for comparison. For this study, we used the Jeffrey Divergence distance metric [30] to generate the D distance function for formula (6):
 |
6 |
Where r is uncertain label calculated from ratings assigned by radiologists, c is the label generated by the classification approach, i is the rating, and pi is the class probability for rating i.
Experimental Design
The experimental design is summarized in Table 1. The dataset used for training and testing of traditional decision tress contained 2,204 instances (up to four instances per nodule, depending on the number of radiologists who rated the nodule). The assigned class was drawn directly from each individual radiologist’s assessment; the predicted class was calculated as the majority class across all instances that reached the same leaf nodule as the nodule instance under consideration. In the consensus approach, the dataset of 2,204 nodules was reduced to 914 nodules by considering only one instance (the outline with the largest area) and one rating (the mode across all radiologists’ ratings for that particular nodule). The consensus approach produces comparable results in terms of having the same number of instances under consideration for building and validating the model.
Table 1.
Experimental design summary
| Approach | No. of instances | Assigned class | Predicted class |
|---|---|---|---|
| Traditional DTs | 2204 | Individual ratings assigned to corresponding individual outlines | Single rating carried by the majority of instances that reached the particular leaf node. |
| Traditional DTs with consensus | 914 | Single rating calculated as mode of all ratings assigned to that nodule | Single rating carried by the majority of instances that reached the particular leaf node. |
| BDTs | 914 | Distribution of ratings over panel of radiologists (probabilistic multi-class label) | Class probability distribution calculated by averaging assigned probability labels of the instances that reached the particular leaf node. (Resulting belief for each class of predicted BBA is calculated using Eq. 3.) |
The dataset for belief decision trees contained 914 instances (one instance per nodule - the outline with the largest area). The assigned probabilistic multi-class label of every instance (nodule) was constructed as a BBA, where the belief for each class was calculated as the ratio of radiologists who assigned the nodule to a given class (rating) to the total number of radiologists who rated that nodule as in Eq. 2. The predicted probabilistic multi-class label was calculated by averaging the assigned BBAs of instances in a leaf node. The belief for each class in a resulting BBA was calculated as an average of pignistic probabilities for this class over the set of instances that reached the leaf node as shown in Eq. 3. Table 2 shows an example of BBA calculations for a hypothetical single terminal node of a tree that was reached by three instances; it also shows how a probability distribution can be associated with a terminal node for the traditional decision tree to make the results comparable in terms of probabilities and therefore, AuCdt.
Table 2.
Example of calculating the probability distributions for a terminal node for DTs versus BDTs
| Malignancy | Highly unlikely | Moderately unlikely | Indeterminate | Moderately suspicious | Highly suspicious |
|---|---|---|---|---|---|
| Assigned class | |||||
| Nodule 1 | 0.25 | 0.75 | 0 | 0 | 0 |
| Nodule 2 | 0.75 | 0.25 | 0 | 0 | 0 |
| Nodule 3 | 0.75 | 0 | 0.25 | 0 | 0 |
| Predicted class | |||||
| Traditional DTs | 0. 67 | 0.33 | 0 | 0 | 0 |
| BDTs | 0.58 | 0.33 | 0.08 | 0 | 0 |
To build the three classification models, a 10-fold cross-validation technique was applied on 90% (training subset) of the data, and further validated on the remaining 10% (validation subset). The 90% and 10% subsets were formed in such a way that the nodule distributions of the validation subsets mimic the nodule distributions of the training subsets with respect to radiologist agreement and the number of radiologists who rated the nodule. For the 2,204 instances dataset the split was done in such manner that instances describing the same nodule could not appear in both 90% and 10% subsets simultaneously.
Furthermore, as diagnostic truth for an LIDC subset has recently become publically available, we explore further the impact of using probabilistic labels instead of deterministic/consensus ones. The diagnosis data from follow-ups or biopsy procedures is provided on a patient level for each nodule found in the CT series of that patient. Although the numbering of the nodules in the diagnosis data file is not consistent with numbering of nodules in LIDC xml files, we were able to reliably identify a correspondence between the LIDC dataset and the diagnosis file for a total of 18 nodules. This dataset of 18 nodules was obtained selecting those nodules that corresponded to patients for which there was only one nodule in the diagnosis dataset. This was the best way to get a reliable mapping with the LIDC data given the lack of nodule IDs in the diagnosis data.. We examined both radiologists’ ratings of malignancy and predictions provided by our system with respect to the diagnosis data. Since the diagnosis was provided on a 4-number scale (0—unknown, 1—benign or nonmalignant disease, 2—malignant, primary lung cancer, 3—malignant metastatic) and radiologists and predicted ratings were on 5-number scale, we created a 3-number scale translation metric described in Table 3. Rows of the table represent the correspondence between different ratings in three datasets. After the translation was performed the dataset contained eight truly malignant, nine truly benign and one truly indeterminate nodule.
Table 3.
Conversion table of the ratings for both LIDC and diagnosis data
| Rating value | New (3-number scale) | Diagnosis data file (4-number scale) | LIDC data Radiologist/computer (5-number scale) |
|---|---|---|---|
| Benign | 1 | 1 | 1.2 |
| Indeterminate/unknown | 2 | 0 | 3 |
| Malignant | 3 | 2.3 | 4.5 |
Results
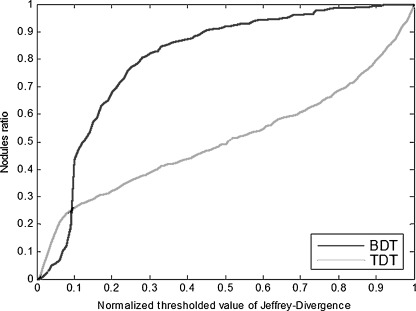
Table 4 shows the distance–threshold curve (AuCdt) and accuracy (ACC) for the two approaches. Although the definition of accuracy applies only for deterministic labels, we nevertheless used it to evaluate the proposed approach and compared it with the traditional approach by considering the consensus values (maximum probability) of the probabilistic labels as well. Figure 4 shows the example of distance–threshold curve for the first iteration of the cross-validation process on training dataset.
Table 4.
Comparison of the two approaches based on accuracy (ACC) and area under the distance–threshold curve (AuCdt) performance metrics
| Traditional decision tree | Belief decision tree | |||||
|---|---|---|---|---|---|---|
| Malignancy | % AuCdt (nodule based) | % ACC (nodule based) | % ACC (2204 instances) | % AuCdt (nodule based) | % ACC (nodule based) | % ACC (2,204 instances) |
| Training subset | 42.82 | 33.32 | 39.44 | 73.10 | 61.58 | −/− |
| Validation subset | 40.95 | 28.81 | 31.00 | 64.16 | 49.45 | −/− |
The nodule based denotes the consensus setup. The column for 2,204 instances is empty under the BDTs because BDTs assume probabilistic classes as part of their input
Fig. 4.
Sample distance–threshold curve for training dataset (first iteration of cross-validation). BDT stands for belief decision trees, TDT stands for traditional decision trees
Tables 5, 6, and 7 presents the nodules (10 out of 18) on which the CAD is in agreement with radiologists and diagnosis (Table 5), nodules (11 out of 18) on which the diagnosis coincides with the agreement between the radiologists and the computer (Table 6), and the nodules (10 out of 18) on which the radiologists’ interpretation coincides with the agreement between diagnosis and computer (Table 7). Since maximum probabilities for each probability distribution were taken into account to produce these tables, we also show the assigned and predicted distributions for these nodules in Figs. 6 and 7 organized by the nodule category. The misclassified nodules from this subset are shown in Fig. 5.
Table 5.
Nodules where the CAD is in agreement with radiologists and diagnosis
| Predicted | Summary | |||||
|---|---|---|---|---|---|---|
| 1 | 2 | 3 | Sum | |||
| Radiologists and diagnosis agreement | 1 | 1 | 1 | 1 | 3 | 7/10 = 70% |
| 2 | 1 | 1 | FN:1/10 | |||
| 3 | 1 | 5 | 6 | FP: 1/10 | ||
| Sum | 2 | 2 | 6 | 10 | ||
Table 6.
Nodules where the diagnosis coincides with the agreement between the radiologists and the computer
| Diagnosis | Summary | |||||
|---|---|---|---|---|---|---|
| 1 | 2 | 3 | Sum | |||
| Radiologists and predicted agreement | 1 | 1 | 1 | 2 | 7/11 = 63.63% | |
| 2 | 2 | 1 | 1 | 4 | FN:1/11 | |
| 3 | 5 | 5 | FP:0/10 | |||
| Sum | 3 | 1 | 7 | 11 | ||
Table 7.
Nodules where the radiologists’ interpretation coincides with the agreement between diagnosis and computer
| Radiologists | Summary | |||||
|---|---|---|---|---|---|---|
| 1 | 2 | 3 | Sum | |||
| Diagnosis and predicted agreement | 1 | 1 | 1 | 2 | 4 | 7/10 = 70% |
| 2 | 1 | 1 | FN:0/10 | |||
| 3 | 5 | 5 | FP:2/10 | |||
| Sum | 1 | 2 | 7 | 10 | ||
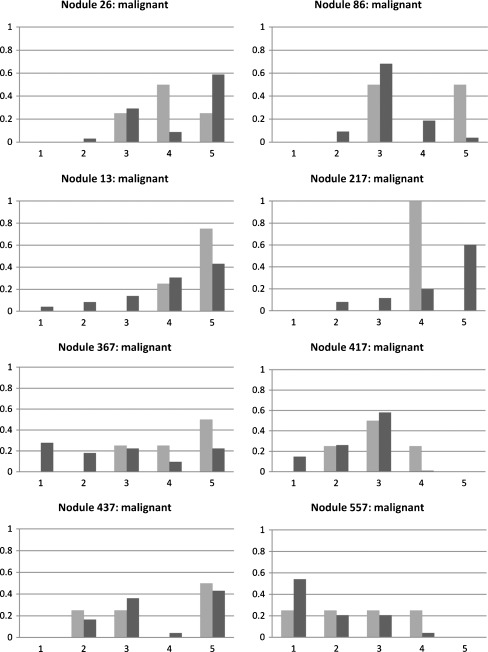
Fig. 6.
Probabilistic interpretation of eight malignant nodules by the computer and radiologists; dark bars denote the computer-based ratings and the light ones represent the radiologists’ ratings; the y-axis represents the probability and the x-axis represents the malignancy class
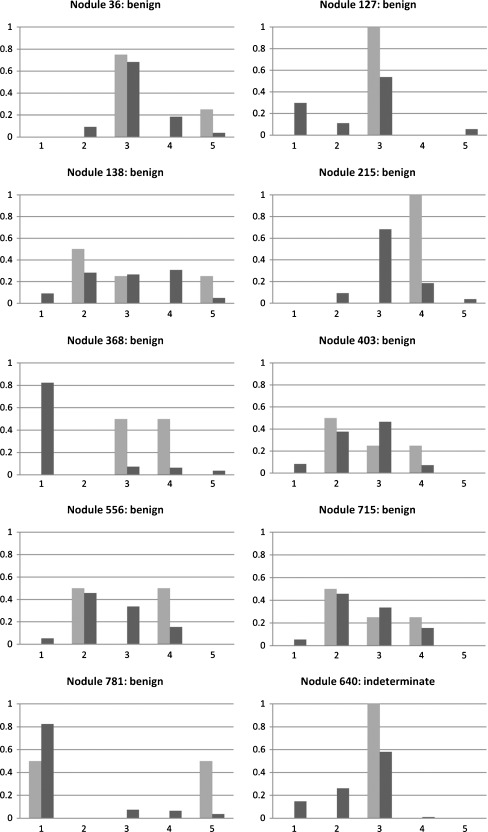
Fig. 7.
Probabilistic interpretation of nine benign and one indeterminate nodules by the computer and radiologists; dark bars denote the computer-based ratings and the light ones represent the radiologists’ ratings; the y-axis represents the probability and the x-axis represents the malignancy class
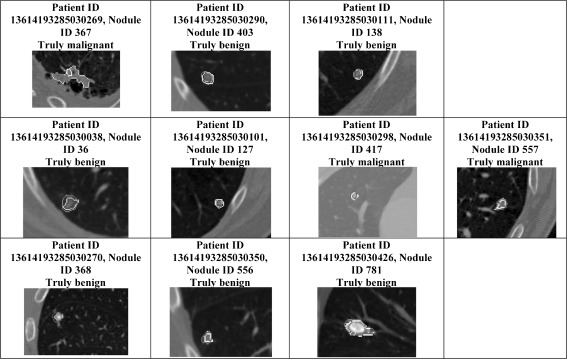
Fig. 5.
Nodules found in both the LIDC dataset and the diagnosis dataset which correspond to the analysis of Tables 5, 6, and 7. Each row shows the nodules on which there was disagreement between radiologists, computer, and diagnosis as identified in Tables 5, 6, and 7
The main diagonal of Table 5 contains seven nodules out of 10 in total on which ratings provided by the classification system agreed with joint diagnosis/radiologists’ ratings. There were three misclassified nodules by the computer: one of the truly benign nodules was indeterminate with a confidence of ~45% by the computer (nodule no. 403), one of the truly malignant nodules was predicted as benign (nodule no. 367), and one of the truly benign nodules was predicted as malignant (nodule no. 138) but the confidence of both predictions was very low (~28%).
The main diagonal of Table 6 contains seven nodules out of 11 in total on which diagnosis information coincided with the agreement between the radiologists and computer (Figs. 6–7).
The main diagonal Table 7 contains seven nodules out of 10 in total on which ratings provided by the radiologists coincided with the agreement between diagnosis and computer. There were three nodules on which the radiologists’ interpretation was different; in particular, two of the truly benign nodules were classified by radiologists as malignant. The computer predictions for these two cases were made with confidences of ~50% (nodule no. 556) and ~85% (nodule no. 781). The computer prediction for the case where a truly benign nodule was predicted by radiologists as indeterminate was also made with confidences of ~85% (nodule no. 368).
Discussion
The results demonstrate that the belief decision tree approach outperforms the traditional decision tree algorithm with respect to both accuracy and area under the distance–threshold curve. On the 90% training subset using cross-validation evaluation technique, the relative performance boost for AuCdt was 30.28% in comparison with traditional decision trees based on consensus; for accuracy (ACC), the increase was 28.26%. With respect to the 10% validation subset, the belief decision trees also outperformed traditional decision tree with respect to both AuCdt (23.21% boost) and accuracy (20.64% boost) which indicates a higher generalization power for the belief decision trees.
When studying the behavior of distance–threshold curves on Fig. 4 we noticed that belief decision tree technique demonstrates higher performance than traditional decision tree algorithm starting from normalized threshold of 0.1. After this intersection point, the BDT curve shows steep growth; for example, for 80% of the nodules the distance between predicted and original probabilistic labels was less than 0.3 distance–threshold, while for the TDT curve that shows only gentle growth, only 35% of the nodules had the distances smaller than the 0.3 threshold.
When examining CT series in a hospital environment, radiologists cannot afford to produce false-negatives, therefore ratings that they provide are biases towards a high possibility of malignancy. Our system demonstrates the ability to correct diagnosis errors caused by this bias with a high confidence, therefore having the potential to reduce the amount of false-positive diagnoses when used as an aid by the radiologist expert. On the other hand, the false-negatives produced by the classification system are associated with a very low confidence and should not affect the opinion of a human expert toward the incorrect diagnosis, since the radiologist will be provided with the probabilistic label as opposed to a “crisp” (deterministic) decision. We suspect that multimodal probabilistic labels for a particular nodule will act as an alarm for the radiologist, signaling that the nodule requires additional attention and therefore helping to avoid incorrect diagnosis.
Conclusions
In this paper we adapted and evaluated a probabilistic multi-class belief decision tree and further compared its performance with another classification approach that is not probabilistic by its nature. We determined that the belief decision trees significantly outperformed the traditional decision trees in terms of both the accuracy and the area under the distance–threshold curve. We examined the performance of our system as well as the performance of human readers with respect to available ground truth and revealed several interesting trends such as high confidence for correctly classified cases and low confidence for incorrectly classified cases. We also determined that the classification system was more in agreement with the diagnosis of the truly benign cases than the radiologists, and therefore, the proposed system has the potential to reduce the number of false positives.
In terms of future work, we plan to expand this work as follows: first, we will include 3D image features in addition to the current 2D features; second, we will look at combining radiologist outlines using p-map approaches instead of considering just the largest outline; and lastly, we will look at incorporating belief classifiers such as belief decision trees described in this work, support vector machines with pair-wise coupling or logistic regression with thresholding into active ensemble learning workflow to take advantage of their classification capabilities in a combined rather than individual way. Additionally, when more diagnosis data becomes available, we will be able to conduct a statistical analysis to support the conclusions that we made from observing Tables 5, 6, and 7. We will also investigate the applicability of the developed CAD system to other radiological tasks such as interpretation of medical images produced by magnetic resonance imaging.
References
- 1.Bankier AA, Levin D, Halpern EF, Kressel HY. Consensus interpretation in imaging research: is there a better way? Radiology. 2010;257:14–17. doi: 10.1148/radiol.10100252. [DOI] [PubMed] [Google Scholar]
- 2.Mower WR. Evaluating bias and variability in diagnostic test reports. Ann Emerg Med. 1999;33(1):85–91. doi: 10.1016/S0196-0644(99)70422-1. [DOI] [PubMed] [Google Scholar]
- 3.Turner DA. Observer variability: what to do until perfect diagnostic tests are invented. J Nucl Med. 1978;19(4):435–437. [PubMed] [Google Scholar]
- 4.Jarvik JG, Deyo RA. Moderate versus mediocre: the reliability of spine MR data interpretations. Radiology. 2009;250(1):15–17. doi: 10.1148/radiol.2493081458. [DOI] [PubMed] [Google Scholar]
- 5.Carrino JA, Lurie JD, Tosteson AN, et al. Lumbar spine: reliability of MR imaging findings. Radiology. 2009;250(1):161–170. doi: 10.1148/radiol.2493071999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.MacMahon H, Engelmann R, Behlen F, Hoffmann K, Ishida T, Roe C, Metz C, Doi K. Computer-aided diagnosis of pulmonary nodules: Results of a large-scale observer test. Radiology. 1999;13:723–726. doi: 10.1148/radiology.213.3.r99dc27723. [DOI] [PubMed] [Google Scholar]
- 7.Matsuki Y, Nakamura K, Watanabe H, Aoki T, Nakata H, Katsuragawa S, Doi K. Usefulness of an artificial neural network for differentiating benign from malignant pulmonary nodules on high-resolution CT: evaluation with receiver operating characteristic analysis. Am J Roentgenol. 2002;178(3):657–663. doi: 10.2214/ajr.178.3.1780657. [DOI] [PubMed] [Google Scholar]
- 8.Li F, Aoyama M, Shiraishi J, et al. Radiologists’ performance for differentiating benign from malignant lung nodules on high-resolution CT using computer estimated likelihood of malignancy. Am J Roentgenol. 2004;183:1209–1215. doi: 10.2214/ajr.183.5.1831209. [DOI] [PubMed] [Google Scholar]
- 9.Marten K, Grillhösl A, Seyfarth T, Obenauer S, Rummeny EJ, Engelke C. Computer-assisted detection of pulmonary nodules: evaluation of diagnostic performance using an expert knowledge-based detection system with variable reconstruction slice thickness settings. Eur Radiol. 2005;15:203–212. doi: 10.1007/s00330-004-2544-5. [DOI] [PubMed] [Google Scholar]
- 10.Peldschus K, Herzog P, Wood SA, Cheema JI, Costello P, Schoepf UJ. Computer-aided diagnosis as a second reader—spectrum of findings in CT studies of the chest interpreted as normal. Chest Journal. 2005;128:1517–1523. doi: 10.1378/chest.128.3.1517. [DOI] [PubMed] [Google Scholar]
- 11.Baker JA, Rosen EL, Lo JY, Gimenez EI, Walsh R, Soo MS. Computer-aided detection (CAD) in screening mammography: sensitivity of commercial CAD systems for detecting architectural distortion. Am J Roentgenol. 2003;181:1083–1088. doi: 10.2214/ajr.181.4.1811083. [DOI] [PubMed] [Google Scholar]
- 12.Ciatto S, Turco MR, Risso G, et al. Comparison of standard reading and computer-aided detection (CAD) on a national proficiency test of screening mammography. Eur J Radiol. 2003;45:135–138. doi: 10.1016/S0720-048X(02)00011-6. [DOI] [PubMed] [Google Scholar]
- 13.Karssemeijer N, Risso G, Catarzi S, et al. Computer-aided detection versus independent double reading of masses on mammograms. Radiology. 2003;227:192–200. doi: 10.1148/radiol.2271011962. [DOI] [PubMed] [Google Scholar]
- 14.Muramatsu C, Li Q, Suzuki K, et al. Investigation of psychophysical measure for evaluation of similar images for mammographic masses: Preliminary results. Medical Physics. 2005;32:2295–2304. doi: 10.1118/1.1944913. [DOI] [PubMed] [Google Scholar]
- 15.Fletcher JW, Kymes SM, Gould M, Alazraki N, Coleman RE, Lowe VJ, et al. A comparison of the diagnostic accuracy of 18FFDG PET and CT in the characterization of solitary pulmonary nodules. J Nucl Med. 2008;49:179–185. doi: 10.2967/jnumed.107.044990. [DOI] [PubMed] [Google Scholar]
- 16.Tao Y, Lo S-C B, Freedman M T, Xuan J: Joint segmentation and spiculation detection for ill-defined and spiculated mammographic masses. Proc. SPIE, doi:10.1117/12.844045, February 16, 2010
- 17.Sahiner B, Hadjiiski L M, Chan H P, Paramagul C, Nees A, Helvie M, Shi J: Concordance of Computer-Extracted Image Features with BI-RADS Descriptors for Mammographic Mass Margin. Proc. SPIE, doi: 10.1117/12.770752, March 17, 2008
- 18.Ochs R, Kimb HJ, Angel E, Panknin C, McNitt-Gray M, Brown M: Forming a reference standard from LIDC data: impact of reader agreement on reported CAD performance. Proc. SPIE, DOI: 10.1117/12.707916, March 30, 2007
- 19.Opfer R, Wiemker RD: Performance Analysis For Computer-Aided Lung Nodule Detection On LIDC Data. Proc. SPIE, DOI: 10.1117/12.708210, February 21, 2007
- 20.Armato SG, III, Roberts RY, Kocherginsky M, Aberle DR, Kazerooni EA, MacMahon H, Beek EJR, Yankelevitz DF, McLennan G, McNitt-Gray MF, Meyer CR, Reeves AP, Caligiuri P, Quint LE, Sundaram B, Croft BY, Clarke LP. Assessment of radiologist performance in the detection of lung nodules: dependence on the definition of “truth”. Acad Radiol. 2009;16:28–38. doi: 10.1016/j.acra.2008.05.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Armato SG, III, et al. Lung Image Database Consortium: developing a resource for the medical imaging research community. Radiology. 2004;232:739–748. doi: 10.1148/radiol.2323032035. [DOI] [PubMed] [Google Scholar]
- 22.Hillman BJ. ACRIN—lessons learned in conducting multi-center trials of imaging and cancer. Cancer Imaging. 2005;5(Spec No A):S97–S101. doi: 10.1102/1470-7330.2005.0026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Elouedi Z, Mellouli K, Smets P. Belief decision trees: theoretical foundations. International Journal of Approximate Reasoning. 2001;28:91–124. doi: 10.1016/S0888-613X(01)00045-7. [DOI] [Google Scholar]
- 24.Wu TF, Lin CJ, Weng RC. Probability estimates for multi-class classification by pairwise coupling. J Mach Learn Res. 2004;5(August):975–1005. [Google Scholar]
- 25.Robinson PJA. Radiology’s Achilles’ heel: error and variation in the interpretation of the Rontghen image. Br J Radiol. 1997;70:1085–1098. doi: 10.1259/bjr.70.839.9536897. [DOI] [PubMed] [Google Scholar]
- 26.Reiner B. Uncovering and improving upon the inherent deficiencies of radiology reporting through data mining. J Digit Imaging. 2010;23:109–118. doi: 10.1007/s10278-010-9279-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zinovev D, Raicu D, Furst J, Armato SG., III Predicting radiological panel opinions using a panel of machine learning classifiers. Algorithms Journal. 2009;2:1473–1502. doi: 10.3390/a2041473. [DOI] [Google Scholar]
- 28.Quinlan JR. Improved use of continuous attributes in C4.5. Journal of Artificial Intelligence Research. 1996;4:77–90. [Google Scholar]
- 29.Spackman KA: Signal detection theory: Valuable tools for evaluating inductive learning. Proc. 6th Int. Workshop on Machine Learning 160–163, 1989
- 30.Liu H, Song D, Rüger S, Hu R, Uren V: Comparing dissimilarity measures for content-based image retrieval. Proc. 4th Asia Inf. Ret. Conf. on Information Retrieval Technology 44–50, 2008