Abstract
Many studies have shown that in reconstituted chromatin model systems, containing only purified DNA and histone octamer, nucleosomes can adopt well defined locations with respect to DNA nucleotide sequence. Recently, nucleosome-nucleosome interactions were suggested as one of the factors underlying preferential nucleosomes positioning. In the present paper this aspect has been studied by topological analysis and electron microscopy visualization of minichromosomes reconstituted at different histone/DNA ratios. Both methods suggest that cooperativity plays a role in nucleosomes formation. A linear cooperative model in which nucleosomes are formed on discrete sites with cooperative interactions occurring only between nearest neighbours allows to calculate the cooperative constant. The reported results show that basic interactions, which are of relevance in the process of chromatin folding, are present also in very simple model system.
Full text
PDF

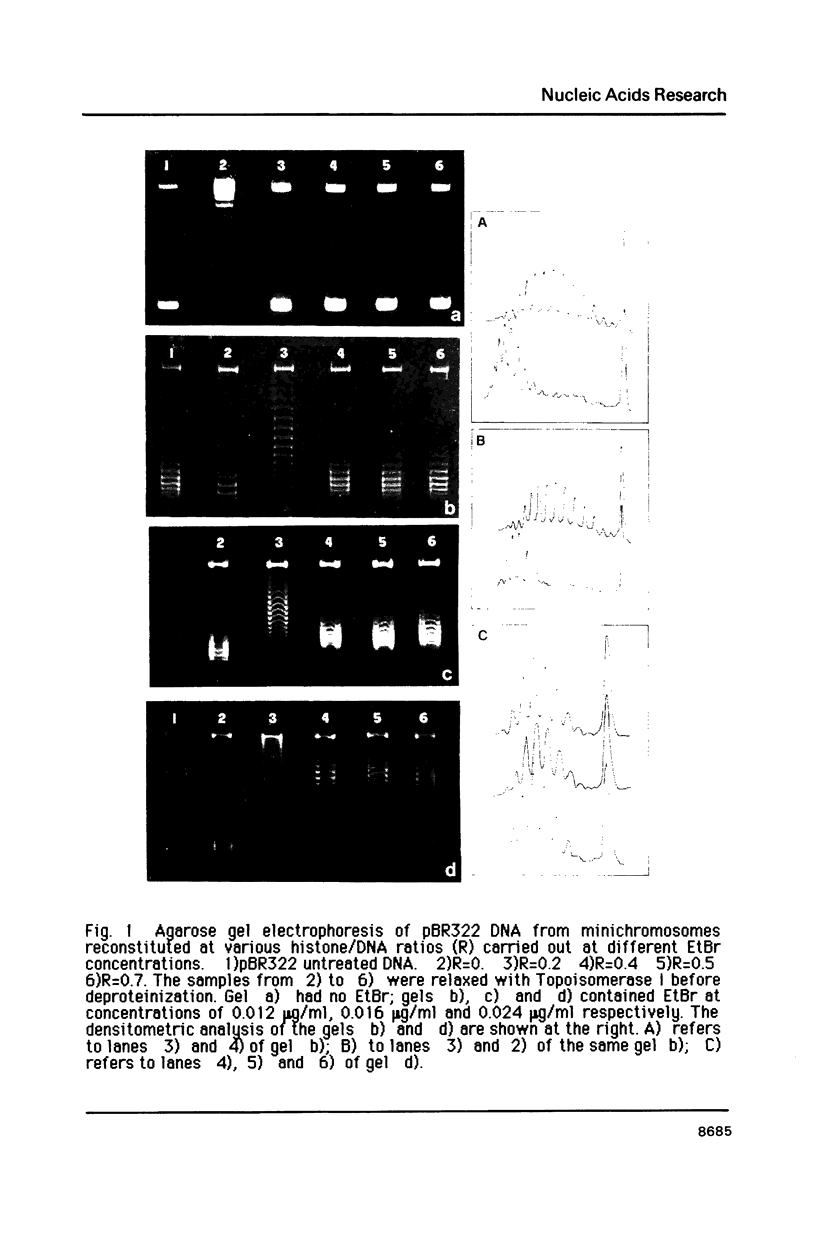
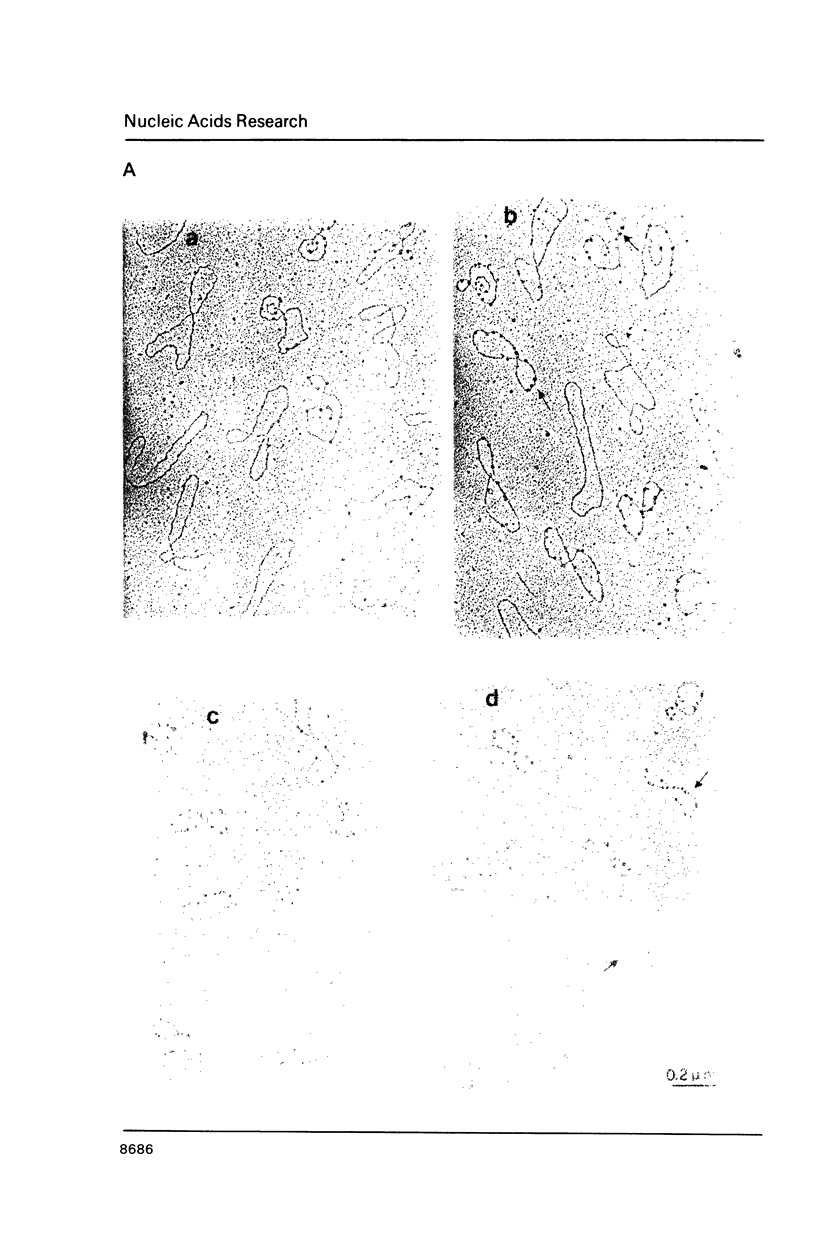




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bina-Stein M., Vogel T., Singer D. S., Singer M. F. H5 Histone and DNA-relaxing enzyme of chicken erythrocytes. Interaction with superhelical DNA. J Biol Chem. 1976 Dec 10;251(23):7363–7366. [PubMed] [Google Scholar]
- Caffarelli E., De Santis P., Leoni L., Palleschi A., Savino M. Preferential positioning of nucleosomes on pBR322 as evaluated via Fourier transform of data from electron microscopy. Eur J Biochem. 1988 Feb 1;171(3):497–501. doi: 10.1111/j.1432-1033.1988.tb13817.x. [DOI] [PubMed] [Google Scholar]
- Caffarelli E., Franzini C., Leoni L., Savino M. Nucleosome "phasing" and cruciform structures in circular supercoiled pBR322 DNA. Cell Biophys. 1984 Mar;6(1):23–31. doi: 10.1007/BF02788578. [DOI] [PubMed] [Google Scholar]
- Caffarelli E., Leoni L., Sampaolese B., Savino M. Persistence of cruciform structure and preferential location of nucleosomes on some regions of pBR322 and ColE 1 DNAs. Eur J Biochem. 1986 Apr 15;156(2):335–342. doi: 10.1111/j.1432-1033.1986.tb09587.x. [DOI] [PubMed] [Google Scholar]
- Drew H. R., Calladine C. R. Sequence-specific positioning of core histones on an 860 base-pair DNA. Experiment and theory. J Mol Biol. 1987 May 5;195(1):143–173. doi: 10.1016/0022-2836(87)90333-0. [DOI] [PubMed] [Google Scholar]
- Germond J. E., Hirt B., Oudet P., Gross-Bellark M., Chambon P. Folding of the DNA double helix in chromatin-like structures from simian virus 40. Proc Natl Acad Sci U S A. 1975 May;72(5):1843–1847. doi: 10.1073/pnas.72.5.1843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horowitz D. S., Wang J. C. Torsional rigidity of DNA and length dependence of the free energy of DNA supercoiling. J Mol Biol. 1984 Feb 15;173(1):75–91. doi: 10.1016/0022-2836(84)90404-2. [DOI] [PubMed] [Google Scholar]
- Keller W. Determination of the number of superhelical turns in simian virus 40 DNA by gel electrophoresis. Proc Natl Acad Sci U S A. 1975 Dec;72(12):4876–4880. doi: 10.1073/pnas.72.12.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leoni L., Morosetti S., Palermo C., Sampaolese B., Savino M. Specific interactions between DNA left-handed supercoils and actinomycin D. Biophys Chem. 1989 Mar;33(1):11–17. doi: 10.1016/0301-4622(89)80002-x. [DOI] [PubMed] [Google Scholar]
- Linxweller W., Hörz W. Reconstitution experiments show that sequence-specific histone-DNA interactions are the basis for nucleosome phasing on mouse satellite DNA. Cell. 1985 Aug;42(1):281–290. doi: 10.1016/s0092-8674(85)80123-9. [DOI] [PubMed] [Google Scholar]
- McGhee J. D., von Hippel P. H. Theoretical aspects of DNA-protein interactions: co-operative and non-co-operative binding of large ligands to a one-dimensional homogeneous lattice. J Mol Biol. 1974 Jun 25;86(2):469–489. doi: 10.1016/0022-2836(74)90031-x. [DOI] [PubMed] [Google Scholar]
- Ramsay N. Deletion analysis of a DNA sequence that positions itself precisely on the nucleosome core. J Mol Biol. 1986 May 5;189(1):179–188. doi: 10.1016/0022-2836(86)90389-x. [DOI] [PubMed] [Google Scholar]
- Ramsay N., Felsenfeld G., Rushton B. M., McGhee J. D. A 145-base pair DNA sequence that positions itself precisely and asymmetrically on the nucleosome core. EMBO J. 1984 Nov;3(11):2605–2611. doi: 10.1002/j.1460-2075.1984.tb02181.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rhodes D. Structural analysis of a triple complex between the histone octamer, a Xenopus gene for 5S RNA and transcription factor IIIA. EMBO J. 1985 Dec 16;4(13A):3473–3482. doi: 10.1002/j.1460-2075.1985.tb04106.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Satchwell S. C., Drew H. R., Travers A. A. Sequence periodicities in chicken nucleosome core DNA. J Mol Biol. 1986 Oct 20;191(4):659–675. doi: 10.1016/0022-2836(86)90452-3. [DOI] [PubMed] [Google Scholar]
- Shore D., Langowski J., Baldwin R. L. DNA flexibility studied by covalent closure of short fragments into circles. Proc Natl Acad Sci U S A. 1981 Aug;78(8):4833–4837. doi: 10.1073/pnas.78.8.4833. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simpson R. T., Stafford D. W. Structural features of a phased nucleosome core particle. Proc Natl Acad Sci U S A. 1983 Jan;80(1):51–55. doi: 10.1073/pnas.80.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thoma F., Koller T. Influence of histone H1 on chromatin structure. Cell. 1977 Sep;12(1):101–107. doi: 10.1016/0092-8674(77)90188-x. [DOI] [PubMed] [Google Scholar]
- Thoma F. Protein-DNA interactions and nuclease-sensitive regions determine nucleosome positions on yeast plasmid chromatin. J Mol Biol. 1986 Jul 20;190(2):177–190. doi: 10.1016/0022-2836(86)90291-3. [DOI] [PubMed] [Google Scholar]
- Thoma F., Simpson R. T. Local protein-DNA interactions may determine nucleosome positions on yeast plasmids. Nature. 1985 May 16;315(6016):250–252. doi: 10.1038/315250a0. [DOI] [PubMed] [Google Scholar]
- Travers A. A., Klug A. The bending of DNA in nucleosomes and its wider implications. Philos Trans R Soc Lond B Biol Sci. 1987 Dec 15;317(1187):537–561. doi: 10.1098/rstb.1987.0080. [DOI] [PubMed] [Google Scholar]
- Voordouw G. On the analysis of DNA-protein interactions: nucleosome formation by the interaction of SV-40 DNA with histones. Biopolymers. 1977 Jun;16(6):1363–1366. doi: 10.1002/bip.1977.360160616. [DOI] [PubMed] [Google Scholar]
- Widom J. Physicochemical studies of the folding of the 100 A nucleosome filament into the 300 A filament. Cation dependence. J Mol Biol. 1986 Aug 5;190(3):411–424. doi: 10.1016/0022-2836(86)90012-4. [DOI] [PubMed] [Google Scholar]