Abstract
The passage from shore to marine life of juvenile penguins represents a major energetic challenge to fuel intense and prolonged demands for thermoregulation and locomotion. Some functional changes developed at this crucial step were investigated by comparing pre-fledging king penguins with sea-acclimatized (SA) juveniles (Aptenodytes patagonicus). Transcriptomic analysis of pectoralis muscle biopsies revealed that most genes encoding proteins involved in lipid transport or catabolism were upregulated, while genes involved in carbohydrate metabolism were mostly downregulated in SA birds. Determination of muscle enzymatic activities showed no changes in enzymes involved in the glycolytic pathway, but increased 3-hydroxyacyl-CoA dehydrogenase, an enzyme of the β-oxidation pathway. The respiratory rates of isolated muscle mitochondria were much higher with a substrate arising from lipid metabolism (palmitoyl-l-carnitine) in SA juveniles than in terrestrial controls, while no difference emerged with a substrate arising from carbohydrate metabolism (pyruvate). In vivo, perfusion of a lipid emulsion induced a fourfold larger thermogenic effect in SA than in control juveniles. The present integrative study shows that fuel selection towards lipid oxidation characterizes penguin acclimatization to marine life. Such acclimatization may involve thyroid hormones through their nuclear beta receptor and nuclear coactivators.
Keywords: in vivo, mitochondria, energy substrates, transcriptomic analysis
1. Introduction
Survival of endotherms in cold water is limited by the tremendous energetic challenge imposed by heat losses in water, a fluid carrying heat away from the body 25 times faster than air of the same temperature. While human survival is estimated within the range of minutes of immersion into cold water at temperature a few degrees above zero [1], some endothermic species, such as adult king penguins (Aptenodytes patagonicus), spend most of their living time foraging in cold circumpolar oceans [2,3]. Such capacity to withstand a prolonged energetic challenge depends on penguin ability to reduce heat losses and sustain high levels of heat production for long periods. However, the mechanisms enabling the development of such metabolic capacity when penguin juveniles depart to sea are unknown.
As marine birds, king penguins breed in huge colonies located on sub-Antarctic islands. At the end of their extended post-hatching development period lasting 12–14 months [4], terrestrial chicks moult to lose their thick down and acquire tight waterproof feathers. Moulting represents a key step before immature penguins face the intensive and prolonged energetic demand imposed by the passage from shore to marine life to reach nutritional emancipation [5]. At sea, the increased internal insulation allowed by a powerful peripheral vasoconstriction and the low oxygen consumption associated with a reduction in body temperature during diving bouts would reduce the daily metabolic cost of thermoregulation [6–8]. Yet daily field metabolic rates (MRs) of penguins are on average three to four times the basal MR [9,10], and immersions into moderately cold water (10°C) induce a two- to threefold increase in energy expenditure [6,11,12]. In line, adaptive thermal responses were described in several penguin species after repeated immersion into cold water with a progressive increase in thermogenic capacity after acclimatization to marine life [6,12]. However, no studies investigated the metabolic basis of such acclimatization to marine life that fuels the sustained energy expenditure imposed by prolonged immersion into cold sea water.
Cold is not the only energetic challenge encountered by penguins. King penguins are known to forage as far as the polar front or the ice edge area, 300–1600 km away from their colony, according to seasonal changes in local prey availability [2,13]. Such tremendous and sustained exercise may further enhance the cooling power of water by increasing convective heat losses. In birds, prolonged and energetically very demanding exercise occurring during flight migration is mainly fuelled by lipid oxidation [14–17] despite the high intensity of this type of exercise. Rapid lipid processing, therefore, appears as a crucial component of the avian migrant phenotype with high lipid fluxes made possible by accelerated lipid transport and upgraded metabolic machinery for lipolysis and lipid oxidation in active skeletal muscles [16]. In addition to endurance, swimming penguins must also dive to catch their prey and these birds are among the most accomplished of divers [7,18]. Anaerobic metabolism may thus be required when oxygen availability is reduced by prolonged apnoea. It is yet not known whether the exceptional capacities exhibited by foraging penguins to sustain intensive and enduring exercise in a harsh cold environment with frequent deep dives relies on coordinated increases in lipid oxidative catabolism and carbohydrate catabolism.
The aim of this study was, therefore, to determine whether the transition from shore to marine life of king penguin juveniles was associated with metabolic adjustments of lipid and carbohydrate metabolism in skeletal muscle directly involved in endurance exercise and cold-induced thermogenesis. Multi-disciplinary approaches were used from transcriptomic analysis, enzymatic activities, mitochondrial bioenergetics and whole body calorimetry.
2. Results
(a). Microarray data: analysis of genes involved in energy metabolism
Relative changes in differentially expressed genes involved in energy metabolism and metabolic pathways are presented in table 1. By comparison with never-immersed (NI) immature penguins, genes encoding proteins involved in glucose transport, glycolysis or glycogenesis were significantly downregulated in sea-acclimatized (SA) birds (from −21 to −60%). The analysis of genes encoding proteins involved in lipid metabolism was performed considering the sequential steps of cellular uptake, transport and metabolism. Although the gene-encoding lipoprotein lipase was slightly downregulated in SA birds (−15% versus NI), both membrane (SLC27A1, +56%), cytosolic (FABP, +21%) and mitochondrial (CPT2, +25%) lipid transporter-encoding genes were upregulated by comparison with NI birds. In mitochondria, lipids are metabolized by enzymes of the β-oxidation pathway and the majority of genes encoding these enzymes were upregulated in SA by comparison with NI birds. Enzymes involved in fatty acid peroxysomal oxidation were also upregulated such as the acyl-CoA thioester hydroase (+38%) as was the gene encoding the carnitine O-octanolyltransferase (+66%) that catalyses the entry into peroxysomes or that encoding acyl-CoA thioesterase 2 that removes shorter free fatty acids and liberates CoA. Associated with oxidative metabolism, the gene encoding pyruvate dehydrogenase, which catalyses the transformation of pyruvate, the product of glycolysis, into acetyl-CoA, the main substrate of the citric acid cycle, was found to be upregulated (+25%). Interestingly, the gene encoding pyruvate dehydrogenase kinase, a negative regulator of pyruvate dehydrogenase, was downregulated (−31%). The few differentially expressed genes encoding enzymes of the citric acid cycle did not vary consistently as in SA penguins, isocitrate and malate dehydrogenase were slightly downregulated (−23 and −19%, respectively), while succinate dehydrogenase and succinate CoA ligase were upregulated (+23 and +30%, respectively) in comparison with NI birds. Altogether, present analysis of microarray data revealed that acclimatization to marine life of immature king penguin was associated with a global increase in the relative expression of genes involved in lipid metabolism. Known regulatory factors involved in metabolic adjustment such as peroxisome proliferator-activated receptors and PPAR-coactivator PGC-1 were not differentially expressed between penguin groups. However, genes encoding the β T3 receptor (+55%; not the α isoform) and a number of nuclear coactivators including cAMP-response-element-binding (CREB)-regulated transcription coactivator 1 (+47%), nuclear receptor coactivator MED1 (+28%) and steroid receptor coactivator (NcoA6, +102%) were upregulated in SA juveniles suggesting a regulatory role in the observed transcriptomic changes. The specific increase in the relative abundance of T3Rβ mRNA was confirmed by Q-PCR that showed a nearly 10-fold increase in the relative abundance of the transcript (electronic supplementary material, data 1).
Table 1.
Microarray data analysed by the MaxRS algorithm and focused on the genes encoding proteins involved in glycolytic and β oxidation pathways and energy metabolism. Transcriptomic analysis was performed from pectoral muscle biopsies collected on never-immersed (NI) or sea-acclimatized (SA) king penguin juveniles. All values of gene expression variations are expressed in percentage relative to NI birds. Genes in bold characters were also tested by Q-PCR. Results are shown in figure 1 and in electronic supplementary material, data.
| name | affymetrix probe set ID | log2 (SA/NI) | residual variance | SA/NI (%) | p-value |
|---|---|---|---|---|---|
| glycolysis | |||||
| enolase 1, (alpha) | GgaAffx.12833.1.S1_s_at | −0.80 | 0.03 | −43 | 0.0002 |
| glucose phosphate isomerase | GgaAffx.20134.1.S1_s_at | −0.53 | 0.05 | −31 | 0.0097 |
| phosphofructokinase | GgaAffx.4338.1.S1_s_at | −1.34 | 0.42 | −60 | 0.0199 |
| phosphoglycerate kinase 1 | Gga.4503.1.S1_a_at | −1.07 | 0.09 | −52 | 0.0006 |
| phosphoglycerate mutase 1 | Gga.6033.1.S1_s_at | 0.36 | 0.03 | +28 | 0.0257 |
| glycogenesis | |||||
| glycosyltransferase 8 domain containing 2 | Gga.11247.1.S1_at | −0.54 | 0.05 | −31 | 0.0117 |
| glycosyltransferase 8 domain containing 3 | GgaAffx.12201.1.S1_at | −0.35 | 0.02 | −21 | 0.0130 |
| phosphoglucomutase 2-like 1 | GgaAffx.11386.1.S1_s_at | −0.59 | 0.05 | −33 | 0.0078 |
| phosphoglucomutase 5 | GgaAffx.9522.1.S1_at | −0.43 | 0.03 | −25 | 0.0082 |
| glucose transport | |||||
| solute carrier family 2 (glucose transporter) | Gga.871.1.S1_at | −0.34 | 0.03 | −21 | 0.0399 |
| β oxidation | |||||
| acetyl-CoA acyltransferase 2 (mitochondrial 3-oxoacyl-CoA thiolase) | GgaAffx.11719.1.S1_s_at | 0.36 | 0.04 | +29 | 0.0394 |
| acetyl-CoA acyltransferase 1 | Gga.17023.1.A1_at | 0.37 | 0.05 | +30 | 0.0230 |
| acyl-CoA dehydrogenase, short chain | Gga.16136.1.S1_at | 0.54 | 0.05 | +45 | 0.0092 |
| enoyl CoA hydratase domain containing 2 | Gga.9702.2.S1_a_at | 0.60 | 0.03 | +52 | 0.0011 |
| hydroxyacyl-CoA dehydrogenase | Gga.3036.1.S1_at | −0.47 | 0.02 | −27 | 0.0028 |
| other enzymes involved in lipid metabolism | |||||
| acyl-CoA thioesterase 2 | Gga.11254.1.S1_at | 0.31 | 0.02 | +24 | 0.0264 |
| diacylglycerol lipase, beta | Gga.5540.1.S1_at | 0.52 | 0.07 | +43 | 0.0286 |
| lipase A, lysosomal acid, cholesterol esterase | GgaAffx.11785.1.S1_s_at | 0.42 | 0.03 | +33 | 0.0117 |
| peroxisomal acyl-CoA thioester hydrolase 2a | Gga.12913.2.S1_a_at | 0.47 | 0.06 | +38 | 0.0378 |
| thioesterase B | Gga.8312.1.S1_at | 0.87 | 0.14 | +83 | 0.0114 |
| lipid transport | |||||
| carnitine acetyltransferase | GgaAffx.2762.2.S1_at | 0.64 | 0.09 | +56 | 0.0178 |
| carnitine O-octanoyltransferase | Gga.9995.1.S1_at | 0.73 | 0.12 | +66 | 0.0192 |
| carnitine palmitoyltransferase II | Gga.16218.1.S1_s_at | 0.32 | 0.03 | +25 | 0.0351 |
| fatty acid binding protein 3 | Gga.12266.1.S1_at | −1.12 | 0.28 | −54 | 0.0184 |
| fatty acid binding protein 4 | Gga.4939.1.S1_s_at | 0.27 | 0.03 | +21 | 0.0265 |
| lipoprotein lipase | GgaAffx.21769.1.S1_s_at | −0.25 | 0.02 | −16 | 0.0433 |
| solute carrier family 27 (fatty acid transporter) | GgaAffx.8475.2.S1_s_at | 0.64 | 0.06 | +56 | 0.0101 |
| citric Acid Cycle | |||||
| isocitrate dehydrogenase 3 | Gga.5623.2.S1_s_at | −0.38 | 0.05 | −23 | 0.0437 |
| malate dehydrogenase 1 | Gga.1141.1.S1_at | −0.76 | 0.05 | −40 | 0.0012 |
| malate dehydrogenase 2 | Gga.4839.1.S1_a_at | −0.31 | 0.02 | −19 | 0.0302 |
| malic enzyme 3 | Gga.6003.1.S1_a_at | 0.91 | 0.04 | +88 | 0.0001 |
| pyruvate dehydrogenase complex | Gga.15677.1.S1_at | 0.32 | 0.02 | +25 | 0.0196 |
| pyruvate dehydrogenase kinase 1 | GgaAffx.8426.1.S1_s_at | −0.53 | 0.13 | −31 | 0.0233 |
| succinate dehydrogenase complex | Gga.1161.1.S1_at | 0.30 | 0.02 | +23 | 0.0307 |
| succinate-CoA ligase, GDP-forming | Gga.2557.1.S1_at | 0.38 | 0.03 | +30 | 0.0170 |
| regulatory factors | |||||
| CREB-regulated transcription coactivator 1 | GgaAffx.25810.2.S1_s_at | 0.55 | 0.04 | +46 | 0.0060 |
| nuclear receptor coactivator (MED1) | GgaAffx.24919.1.S1_at | 0.35 | 0.03 | +28 | 0.0364 |
| steroid receptor coactivator (NCOA6) | GgaAffx.783.1.S1_at | 1.01 | 0.08 | +102 | 0.0007 |
| thyroid hormone receptor beta | Gga.699.1.S1_at | 0.64 | 0.08 | +56 | 0.0081 |
(b). Enzyme activities
We measured the activities of four enzymes involved in either carbohydrate or lipid metabolism. The activities of hexokinase (HK: 5.8 ± 0.5 versus 6.1 ± 0.9) and lactate dehydrogenase (LDH: 1598 ± 99 versus 1718 ± 222), both involved in the glycolytic pathway, were similar in NI and SA birds. The activities of 3-hydroxyacyl-CoA dehydrogenase (HAD: 20.7 ± 2.1 versus 15.4 ± 0.8, t = 2.25, p < 0.05), an enzyme of the β-oxidation pathway and citrate synthase (CS: 235 ± 28 versus 175 ± 12, t = 1.94, p = 0.08), an enzyme of the citric acid cycle, were overall 34 per cent higher in SA than in NI birds, although CS activities did not reach statistical significance.
(c). Mitochondrial respiration rates
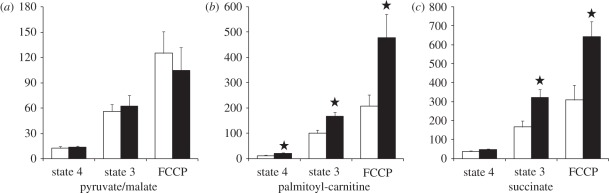
Figure 1 reports mitochondrial respiration using either carbohydrate- or lipid-derived substrates. When energized with pyruvate, a standard carbohydrate substrate that provides electrons to complex I of the respiratory chain, mitochondria of both penguin groups showed similar respiratory activities. When providing electrons to the same complex, but with a donor resulting from lipid metabolism (palmitoyl-l-carnitine), mitochondria from SA penguins exhibited much higher respiration rates in non-phosphorylating (state 4: +60%, t = 2.87, p < 0.05), phosphorylating (state 3: +143%, t = 3.04, p < 0.05) and maximal uncoupled states (FCCP: +197%, t = 4.34, p < 0.05) when compared with mitochondria from NI birds. Results were similar with succinate, providing electrons directly to complex II of the respiratory chain, with phosphorylating (state 3: +75%, t = 2.85, p < 0.05) and maximal uncoupled respiration rates (FCCP state: +68%, t = 2.82, p < 0.05) being higher in mitochondria from SA than from NI birds. On account of the large increase in mitochondrial cytochrome oxidase activity in SA penguins (54.9 ± 6.0 versus 34.3 ± 2.5 natom oxygen per minute per micro gram muscle in SA and NI juveniles, respectively, p < 0.05), the amount of mitochondrial protein per unit weight of skeletal muscle and thus the whole muscle ability to oxidize substrates were likely to be increased after acclimatization to marine life.
Figure 1.
Respiration rates of mitochondria isolated from pectoralis muscle of never-immersed (NI, open bars) or sea-acclimatized (SA, filled bars) king penguin juveniles. Oxygen uptake was determined with a Clark oxygen electrode (Rank Brother LTD) placed into a glass chamber thermostated at 38°C. State 4 was initiated with either: (a) 5 mM pyruvate and 2.5 mM malate, (b) 50 µM palmitoyl-l-carnitine and 2.5 mM malate, (c) 5 mM succinate in the presence of 5 µM rotenone. Phosphorylating respiration (state 3) was initiated with 500 µM ADP. Non-phosphorylating respiration (state 4) was obtained by addition of 1 µg ml−1 oligomycin and fully uncoupled respiration was obtained with 2 µM FCCP. Means ± s.e.m. *p < 0.05 versus NI birds.
(d). In vivo lipid oxidation
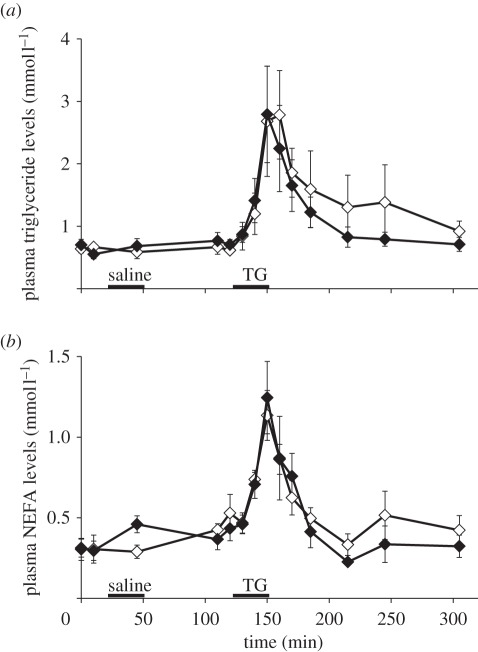
Molecular and biochemical data prompted us to investigate the capacity to oxidize lipid substrates at the whole organism level. To test the hypothesis of an increased capacity to oxidize lipids in vivo, we measured the thermogenic effect of lipid infusion in NI and SA king penguin juveniles. This was achieved by indirect calorimetry (figure 2a), a non-invasive technique that enables the determination of changes in MR in individual birds over time. We first verified that saline infusion was without effect on MR (tNI = 1.81 and tSA = −1.38, p > 0.05). After the end of the saline infusion, resting MR remained constant for more than 1 h and was similar between NI and SA juveniles (25.4 ± 0.2 versus 26.2 ± 0.3 W, respectively) as was the respiratory quotient (0.70 ± 0.06 and 0.70 ± 0.02, respectively). Infusion of lipid emulsion (8.4 kJ min−1) led to a rise in MR in both NI and SA juveniles. In NI juveniles, lipid infusion transiently increased MR that peaked at 28.9 ± 1.3 W (+13% above resting MR, t = 4.1, p < 0.05) 10 min after the start of the infusion and then returned to basal level 45 min after infusion. In SA juveniles, MR peaked at 31.7 ± 3.1 W (+21% above resting MR, t = 5.8, p < 0.05) and remained significantly elevated 2.30 h after infusion (28.7 ± 2.8 W, +9% above resting MR, t = 2.5, p < 0.05). The total lipid-induced thermogenic effect was more than four times higher in SA than in NI penguins (z = −2.7, p < 0.05, figure 2b). Figure 3 shows that lipid infusion led to an expected surge in plasma concentration of triglycerides (TGs; figure 3a) with a concomitant rise in plasma non-esterified fatty acids (NEFAs; figure 3b), indicating that infused TGs were hydrolysed into their molecular constituents, i.e. fatty acids and glycerol. After an initial rise, the plasma concentration of metabolites decreased and returned to basal level on completion of protocol. Despite a trend for a more rapid return to basal levels in SA birds, the kinetic patterns were not significantly different between groups indicating that lipids were similarly cleared by NI or SA juveniles, but the fate of these substrates was different between groups. Indeed, taking into account that respiratory quotient remained near 0.7 throughout the infusion, indicating a constant reliance on lipid catabolism in both SA and NI birds, the differential effects of lipid infusion upon energy expenditure (figure 2) show that SA oxidized a larger amount of the infused lipids than NI birds.
Figure 2.
Metabolic response to triglyceride (TG) infusion in NI (open symbols) or SA (filled symbols) king penguin immatures. (a) Changes in metabolic rate (MR) at 10°C (thermoneutrality) in NI (n = 5) or SA (n = 6) birds. Horizontal black lines represent infusions of either saline or TG emulsion for 30 min. Mean body mass was 9.5 ± 0.2 kg in NI and 8.9 ± 0.3 kg in SA birds. Means ± s.e.m. *p < 0.05 indicates that all of the MR values after the end of TG infusion were significantly different from basal level in SA birds. Shaded areas represent the points incorporated into the AUC and dashed horizontal lines mark the level of resting MR determined after saline infusion. (b) Bars represent the integrated total effect of TG infusion on MR (area under the curve) in NI (open bar) or SA (filled bar) juveniles. Means ± s.e.m. *p < 0.05 significantly different from NI birds. Open diamonds with solid line, NI; filled diamonds with solid line, SA.
Figure 3.
Changes in plasma lipid levels in NI (open symbols, n = 5) or SA (filled symbols, n = 6) king penguin juveniles. (a) Plasma TG levels. (b) Plasma non-esterified fatty acid (NEFA) levels. Black lines represent infusions of either saline solution or TG emulsion for 30 min. Blood aliquots were taken at different time point (0, 10, 45, 110, 120, 130, 140, 150, 160, 170, 185, 215, 245, 305 min). Values are means ± s.e.m.
3. Discussion
To our knowledge, this is the first study based on an integrative approach using transcriptomic, biochemical and physiological analysis to investigate changes in energy metabolism at a key step of penguin life. Results showed that the transition from shore to marine life leads to a selective increase in the capacity to oxidize lipids as a fuel for energy metabolism. Such global shift towards lipid oxidation may involve thyroid hormones interacting with their nuclear beta receptors and a number of nuclear coactivators.
Analysis of transcriptomic data in penguins was made possible by the use of MaxRS, a new method for the analysis of heterologous hybridization on Affymetrix GeneChip chicken genome arrays despite the large phylogenetic distance between these two birds [19]. The differential expression of a large panel of genes detected by this approach was confirmed by quantitative PCR using penguin-specific primers ([19] and electronic supplementary material, data). With the present large-scale analysis, we highlighted a global increase in the relative expression of genes involved in lipid metabolism, while there was a global slight decrease in most genes encoding proteins involved in carbohydrate metabolism. This coordinated change in gene expression suggests a selective shift in metabolic pathways favouring the use of lipids as fuel to sustain energy expenditure. It may be ascribed to the effects of acclimatization to marine life as both groups of juveniles were exposed to similar thermal conditions and had the same nutritional status at the time of biopsy. Variations in transcripts were consistent with changes in enzymatic activity, leading to a relative increased capacity for mitochondrial β oxidation versus glycolysis. This supports the hypothesis that most changes in transcripts may lead to changes in functional activity here leading to an increased activity of lipid oxidation in skeletal muscles of SA juveniles. This hypothesis was also supported by the higher capacity of isolated muscle mitochondria to oxidize substrates derived from fatty acids in SA juveniles than in controls. Such change was not observed when a substrate derived from glycolysis (pyruvate) was used. These observations are consistent with a better utilization of lipid substrates after experiencing marine life in relation with an increased activity of the β-oxidation pathway and an enhanced substrate import into mitochondria, as detected with the microarray and enzymatic methods. The activation of mitochondrial respiration by lipid substrates may represent a major step towards natural sea-acclimatization in king penguin juveniles and could be related with an increased need in ATP production essential for counterbalancing the energetic depletion generated by intense physical activity and thermogenesis during foraging.
The fact that king penguin juveniles complete their development at sea raises the question whether the shift in metabolism could be due to developmental rather than environmentally induced changes. Although we cannot reach a definitive conclusion with the present data, it remains that NI king penguin juveniles had a similar body weight to SA birds and neither group has reached sexual maturity, ruling out major developmental differences between groups. Further, NI birds had overcome a long-term moulting fast before to be included in the present study [5], implying that they had already developed good capacities to oxidize lipids as lipids are the main energetic substrates in fasting penguins [20,21]. Since lipids are also the main substrates to fuel increased regulatory thermogenesis [22] and endurance metabolism in birds [15–17], the two experimental groups would then differ by differences in energetic demands imposed by their respective physiological situation, i.e. low energy expenditure in NI while resting ashore versus increased energy expenditure in SA while swimming in cold sea water. Therefore, it is reasonable to assume that most of the metabolic shift reported herein is likely to be induced by marine life components, such as cold and/or endurance exercise. In this respect, several studies have suggested similar enzyme alterations in the pectoral muscles of migrating birds to allow enhanced use of fatty acids in the absence of cold exposure [15–17]. Acclimatization to marine life, therefore, parallels the metabolic adjustment to endurance exercise encountered during flight migration and that are mainly fuelled by lipid oxidation [15–17], in association with cold stress which would further reinforce this metabolic adjustment [14].
The increased capacity to oxidize lipid substrates after experiencing marine life was confirmed at the whole animal level by an increased thermogenic effect of lipid infusion into birds kept at thermoneutrality. Present data suggest that SA juveniles oxidize a larger part of infused substrates in keeping with the higher oxidative potential exhibited at the molecular, enzymatic and mitochondrial levels. This lipid-induced extra MR in resting juveniles cannot be solely ascribed to skeletal muscle, and other tissues such as liver probably mediate part of this lipid-dependent rise in bird energy expenditure. However, by virtue of its large contribution to body mass and its high oxidative capacity, skeletal muscle can take up large amount of fatty acids and supply considerable metabolic heat from their oxidation. If such heat dissipation is conceivable when birds are immersed into cold water, present infusion experiments were performed in resting birds at thermoneutrality and therefore the extra heat production stimulated by lipid infusion is not related to regulatory thermogenic or muscle activity needs. It follows that the increased lipid availability artificially induced by infusion may have stimulated thermogenic processes that were more developed in SA than in NI juveniles. Increased thermogenic effects mediated by fatty acids through an uncoupling of oxidative phosphorylation processes have already been described in vitro in skeletal muscle mitochondria from SA king penguin juveniles [22] or from cold-acclimated ducklings [23]. Interestingly, an upregulation of a skeletal muscle uncoupling protein (UCP) was reported in SA juvenile penguins [22]. Although their physiological functions are still unclear, mammalian muscle UCPs have been associated with increased fatty acid oxidation through still undefined molecular mechanism [24]. Present results suggest a link between muscle UCP and thermogenic oxidation of fatty acids in SA juveniles [22,25].
Present data suggest potential mechanisms triggering the adaptive increase in lipid oxidation exhibited by penguin juveniles on the basis of the specific upregulation of the gene-encoding nuclear β T3 receptors. Most effects of T3 are mediated by interactions with nuclear T3R, which are ligand-dependent transcriptional factors regulated by a number of environmental, nutritional and endocrine factors in a tissue- and isoform-specific manner suggesting isoform-specific effects in gene regulation [26]. Interestingly, high expression of the T3Rβ1 isoform was also observed in liver of a rat strain exhibiting resistance to obesity, elevated energy expenditure, high spontaneous locomotor activity and a preference for fat oxidation [27]. This suggests specific roles of the T3Rβ isoform in activating genes related to energy metabolism. A stimulation of these nuclear T3Rβ in juvenile penguins at sea is conceivable as increased levels of plasma T3 and T3/T4 ratio have been reported in SA king penguin juveniles when compared with terrestrial juveniles [4]. This is also consistent with the high expression of malic enzyme (table 1) and avian UCP [22] in pectoralis muscle of SA juveniles as these genes are known to be controlled by T3 [28–30]. Present data also suggest the implication of nuclear coactivators such as CREB-regulated transcription coactivator 1, nuclear receptor coactivator MED1 and steroid receptor coactivator (NcoA6) that were simultaneously upregulated in SA juveniles. Experiments in transgenic mice have indeed indicated that these nuclear coreceptors might be involved in muscle energy metabolism [31,32]. Interestingly, MED1 was recently shown to dynamically interact with nuclear T3R in the activation of brown-fat-specific thermogenic UCP1 [33] further underlying its role in metabolic adjustments.
Alternatively, activation of T3Rβ expression in the pectoralis muscle of SA immature penguins might also be related to the physiological constraint of diving. Indeed, adult penguins are breath-hold divers that undergo critical depletion of muscle oxygen during dives [8], a condition we would expect to activate hypoxia-inducible factor 1 (HIF1). Recently, it has been suggested that T3Rβ/retinoid X receptor alpha heterodimer could activate HIF-1α [34], linking T3 with HIF1-induced expression of genes involved in angiogenesis, erythropoiesis and glucose homeostasis [35,36]. The upregulation of T3 response in SA king penguin juveniles could, therefore, confer skeletal muscle with hypoxia tolerance through augmented accumulation of HIF by allowing the maintenance of most of the activity of glycolytic enzymes while lipid catabolism was enhanced. However, it remains that direct HIF target genes [35] associated with glycolysis, such as phosphoglycerate kinase 1, phosphofructokinase and pyruvate dehydrogenase kinase, were found downregulated in SA penguins. It would be of utmost interest to investigate whether HIF abundance is altered after sea experience and whether the dives of juvenile king penguins are kept within aerobic limits while the adults may be pushing the physiological limits of aerobic diving.
In conclusion, the present integrative study has shown for the first time that fuel selection towards lipid oxidation is a key characteristic of penguin acclimatization to the constraints of sub-Antarctic marine life. It is suggested that such acclimatization may involve thyroid hormones through their interactions with nuclear beta receptor and nuclear coactivators. Present data not only improve our fundamental knowledge on comparative biology, but also have important implications in foraging and migration ecology.
4. Material and methods
(a). Animals
Field experiments were conducted on the Crozet archipelago (Possession Island, 46°25′ S, 51°45′ E) at the French Alfred Faure Station during two austral summer campaigns (2007–2008 and 2008–2009). According to the Agreed Measures for the Preservation of Antarctic and Sub-Antarctic Fauna, the project received the agreements of the French Committee for Antarctic Research (Programme 131). Pre-fledging juvenile king penguins (Aptenodytes patagonicus) of both sexes (13–16 months old) were captured on the nearby breeding colony of Baie du Marin before they had completed moulting, which is a pre-requisite for departing to sea [5,12]. Captured birds finished their moult in an outside enclosure near the laboratory and constituted the NI group. A second group of juvenile birds (25–28 months old) of both sexes was caught soon after their arrival at the beach after a foraging trip ensuring that they had fully accomplished their acclimatization to marine life. They constituted the SA group. All birds were kept in an outside enclosure, weighed every day, fed twice daily on Atlantic mackerel (Scomber vernalis up to 1 kg d−1) for 3 days and then fasted for 48 h before experiments. This protocol tended to minimize confounding effects of potential differences in nutritional status. On completion of the study, all penguins were fed for one week until full recovery and then released at the site of their capture.
(b). Experimental design
The first austral summer was devoted to biochemical and molecular analysis in vitro. A total of 12 birds were captured, six pre-fledging (NI) and six SA juveniles. Superficial pectoralis muscle was surgically biopsied under general isoflurane anaesthesia as described previously [25]. A small sample (approx. 200 mg) was immediately frozen in liquid nitrogen and kept at −80°C until assayed for enzymatic and transcriptomic analysis; the remainder of the biopsy (3–4 g) was freshly used for mitochondrial extraction and bioenergetics analysis. The second austral summer was devoted to study the effect of lipid infusion upon energy expenditure in vivo. A total of 20 birds were captured, 10 pre-fledging (NI) and 10 SA juveniles. No tissue sampling was performed during this second campaign.
(c). Transcriptomic analysis
Total RNA was purified from pectoralis muscle biopsies using TriReagent procedure (Invitrogen, France) and the quality of extracted RNA was assessed using a bioanalyzer 2100 (Agilent technologies, Inc, Palto Alto, CA, USA). Labelling and hybridization were performed with the Affymetrix protocol (http://www.affymetrix.com) by the ProfileXpert platform (Lyon, France) on Affymetrix GeneChip chicken genome array. Microarray data are available in Gene Expression Omnibus (GSE17725). Analysis of the dataset was performed using MaxRS method [19].
(d). Enzymatic activities
HK (EC 2.7.1.1), LDH (EC 1.1.1.27) and HAD (EC 1.1.1.35) were fluorimetrically determined (n = 6 per group) on muscle homogenates [37]. CS (EC 4.1.3.7) was measured using 5–5′-dithiobis-2-nitrobenzoic acid. Enzyme activities were expressed in U per gram of protein. All assays were run in duplicates.
(e). Mitochondrial respiration
The bioenergetics of mitochondria was investigated in vitro in six SA and six NI birds as already described for penguins [25,30]. Briefly, mitochondria were isolated by a standard extraction protocol, involving potter homogenization, partial protease digestion and differential centrifugations. Mitochondria were pelleted at 8700 g and mitochondrial proteins were determined by a Biuret method. Cytochrome c oxidase was determined polarographically at 38°C on tissue homogenates as described previously [25].
(f). In vivo lipid oxidation
The metabolic effect of a TG infusion was determined in vivo at 10°C within the thermoneutral zone for these birds [38]. Eleven birds were included in this protocol (five NI and six SA). The day before experimentation, birds were equipped with intravenous catheters as described previously [39]. After stabilization within the thermostatic chamber for a night, MR was continuously recorded by indirect calorimetry as already described in penguins [6,12,25,38] following infusions at 1 ml min−1 of saline (0.9% NaCl) then Intralipid 20 per cent emulsion (Fresenius Kabi AB, Sweden). Each infusion time was 30 min long. Blood aliquots (700 µl) taken at different time points were centrifuged at 3500 g for 5 min at 4°C and the resulting plasma samples stored at −80°C until analysis of plasma TG and NEFA concentrations using commercially available kits from Biomérieux or Wako chemicals, respectively. Because the Intralipid emulsion also contains high levels of glycerol (2.3 g per 100 ml), control infusion experiments were also conducted with a glycerol solution in nine birds (five NI and four SA) that showed no significant thermogenic effect of the glycerol perfusion. In order to estimate the total effect of TGs upon MR, area under the curve (AUC) was calculated for each bird using trapezoidal integration minus the individual resting MR.
(g). Statistical analysis
Data were expressed as means ± s.e.m. In transcriptomic analysis, after normalization of microarray data with the quantile method [40], differentially expressed genes between NI and SA penguin juveniles were highlighted using the empirical Bayes-moderated t-statistics implemented in the Bioconductor package limma [41]. Mitochondrial enzymatic activities and respiration rates were tested with one-way analysis of variance followed by Student's t-tests (Statview v. 4.5 software). Effects of groups, saline infusion and TG infusion on MR and kinetics of plasma TG and NEFA levels were investigated using general linear mixed models. Individual were included as random factor. Time was considered as factor in kinetic statistical analysis. AUC values were tested with a non-parametric Wilcoxon rank sum test. These statistical analyses were performed using R with linear mixed effects function in the ‘nlme’ library for linear mixed models. Factor significance was tested using Wald z-tests [42]. Variance homogeneity, residuals normality and residuals independence have been checked. Significant differences were considered at p < 0.05.
Acknowledgements
This work was supported by grants from the French Polar Institute (Institut Paul Emile Victor, Programme 131). C.D. and L.T. were in receipt of a fellowship from the Ministère de l'Enseignement Supérieur et de la Recherche.
References
- 1.Ducharme M. B., Lounsbury D. S. 2007. Self-rescue swimming in cold water: the latest advice. Appl. Physiol. Nutr. Metab. 32, 799–807 10.1139/H07-042 (doi:10.1139/H07-042) [DOI] [PubMed] [Google Scholar]
- 2.Charrassin J., Bost C. 2001. Utilisation of the oceanic habitat by king penguins over the annual cycle. Mar. Ecol. Prog. Ser. 221, 285–298 10.3354/meps221285 (doi:10.3354/meps221285) [DOI] [Google Scholar]
- 3.Pütz K. 2002. Spatial and temporal variability in the foraging areas of breeding king penguins. Condor 104, 528–538 10.1650/0010-5422(2002)104[0528:SATVIT]2.0.CO;2 (doi:10.1650/0010-5422(2002)104[0528:SATVIT]2.0.CO;2) [DOI] [Google Scholar]
- 4.Cherel Y., Durant J. M., Lacroix A. 2004. Plasma thyroid hormone pattern in king penguin chicks: a semi-altricial bird with an extended posthatching developmental period. Gen. Comp. Endocrinol. 136, 398–405 10.1016/j.ygcen.2004.02.003 (doi:10.1016/j.ygcen.2004.02.003) [DOI] [PubMed] [Google Scholar]
- 5.Corbel H., Morlon F., Groscolas R. 2008. Is fledging in king penguin chicks related to changes in metabolic or endocrinal status? Gen. Comp. Endocrinol. 155, 804–813 10.1016/j.ygcen.2007.11.006 (doi:10.1016/j.ygcen.2007.11.006) [DOI] [PubMed] [Google Scholar]
- 6.Dumonteil E., Barré H., Rouanet J. L., Diarra M., Bouvier J. 1994. Dual core and shell temperature regulation during sea acclimatization in Gentoo penguins (Pygoscelis papua). Am. J. Physiol. 266, R1319–R1326 [DOI] [PubMed] [Google Scholar]
- 7.Handrich Y., Bevan R., Charrassin J., Butler P., Pütz K., Woakes A., Lage J., Le Maho Y. 1997. Hypothermia in foraging king penguins. Nature 388, 64–67 10.1038/40392 (doi:10.1038/40392) [DOI] [Google Scholar]
- 8.Williams C. L., Meir J. U., Ponganis P. J. 2011. What triggers the aerobic dive limit? Patterns of muscle oxygen depletion during dives of emperor penguins. J. Exp. Biol. 214, 1802–1812 10.1242/jeb.052233 (doi:10.1242/jeb.052233) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kooyman G. L., Croxall J. P., Costa D. P. 1982. Diving depths and energy requirements of king penguins. Science 217, 726–727 10.1126/science.7100916 (doi:10.1126/science.7100916) [DOI] [PubMed] [Google Scholar]
- 10.Nagy K. A., Kooyman G. L., Ponganis P. J. 2001. Energetic cost of foraging in free-diving emperor penguins. Physiol. Biochem. Zool. 74, 541–547 10.1086/322165 (doi:10.1086/322165) [DOI] [PubMed] [Google Scholar]
- 11.Stahel C. D., Nicol S. C. 1988. Ventilation and oxygen extraction in the little penguin (Eudyptula minor), at different temperatures in air and water. Respir. Physiol. 71, 387–398 10.1016/0034-5687(88)90030-8 (doi:10.1016/0034-5687(88)90030-8) [DOI] [PubMed] [Google Scholar]
- 12.Barré H., Roussel B. 1986. Thermal and metabolic adaptation to first cold-water immersion in juvenile penguins. Am. J. Physiol. 251, R456–R462 [DOI] [PubMed] [Google Scholar]
- 13.Bost C., et al. 1997. Foraging habitat and food intake of satellite-tracked king penguins during the austral summer at Crozet Archipelago. Mar. Ecol. Prog. Ser. 150, 21–33 10.3354/meps150021 (doi:10.3354/meps150021) [DOI] [Google Scholar]
- 14.Vaillancourt E., Prud'Homme S., Haman F., Guglielmo C., Weber J. 2005. Energetics of a long-distance migrant shorebird (Philomachus pugnax) during cold exposure and running. J. Exp. Biol. 208, 317–325 10.1242/jeb.01397 (doi:10.1242/jeb.01397) [DOI] [PubMed] [Google Scholar]
- 15.McFarlan J. T., Bonen A., Guglielmo C. G. 2009. Seasonal upregulation of fatty acid transporters in flight muscles of migratory white-throated sparrows (Zonotrichia albicollis). J. Exp. Biol. 212, 2934–2940 10.1242/jeb.031682 (doi:10.1242/jeb.031682) [DOI] [PubMed] [Google Scholar]
- 16.Weber J. 2009. The physiology of long-distance migration: extending the limits of endurance metabolism. J. Exp. Biol. 212, 593–597 10.1242/jeb.015024 (doi:10.1242/jeb.015024) [DOI] [PubMed] [Google Scholar]
- 17.Guglielmo C. G. 2010. Move that fatty acid: fuel selection and transport in migratory birds and bats. Integr. Comp. Biol. 50, 336–345 10.1093/icb/icq097 (doi:10.1093/icb/icq097) [DOI] [PubMed] [Google Scholar]
- 18.Ponganis P. J., Kooyman G. L. 2000. Diving physiology of birds: a history of studies on polar species. Comp. Biochem. Physiol. A 126, 143–151 10.1016/S1095-6433(00)00208-7 (doi:10.1016/S1095-6433(00)00208-7) [DOI] [PubMed] [Google Scholar]
- 19.Degletagne C., Keime C., Rey B., de Dinechin M., Forcheron F., Chuchana P., Jouventin P., Gautier C., Duchamp C. 2010. Transcriptome analysis in non-model species: a new method for the analysis of heterologous hybridization on microarrays. BMC Genomics 11, 344. 10.1186/1471-2164-11-344 (doi:10.1186/1471-2164-11-344) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Groscolas R. 1988. The use of body mass loss to estimate metabolic rate in fasting sea birds: a critical examination based on emperor penguins (Aptenodytes forsteri). Comp. Biochem. Physiol. A 90, 361–366 10.1016/0300-9629(88)90203-4 (doi:10.1016/0300-9629(88)90203-4) [DOI] [PubMed] [Google Scholar]
- 21.Cherel Y., Charrassin J. B., Challet E. 1994. Energy and protein requirements for molt in the king penguin Aptenodytes patagonicus. Am. J. Physiol. 266, R1182–R1188 [DOI] [PubMed] [Google Scholar]
- 22.Talbot D. A., Duchamp C., Rey B., Hanuise N., Rouanet J. L., Sibille B., Brand M. D. 2004. Uncoupling protein and ATP/ADP carrier increase mitochondrial proton conductance after cold adaptation of king penguins. J. Physiol. 558, 123–135 10.1113/jphysiol.2004.063768 (doi:10.1113/jphysiol.2004.063768) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Barré H., Nedergaard J., Cannon B. 1986. Increased respiration in skeletal muscle mitochondria from cold-acclimated ducklings: uncoupling effects of free fatty acids. Comp. Biochem. Physiol. B 85, 343–348 10.1016/0305-0491(86)90010-6 (doi:10.1016/0305-0491(86)90010-6) [DOI] [PubMed] [Google Scholar]
- 24.Bézaire V., Seifert E. L., Harper M. E. 2007. Uncoupling protein-3: clues in an ongoing mitochondrial mystery. FASEB J. 21, 312–324 10.1096/fj.06-6966rev (doi:10.1096/fj.06-6966rev) [DOI] [PubMed] [Google Scholar]
- 25.Rey B., Halsey L., Dolmazon V., Rouanet J. L., Roussel D., Handrich Y., Butler P., Duchamp C. 2008. Long-term fasting decreases mitochondrial avian UCP-mediated oxygen consumption in hypometabolic king penguins. Am. J. Physiol. 295, R92–R100 10.1152/ajpcell.00023.2008 (doi:10.1152/ajpcell.00023.2008) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Cheng S. Y., Leonard J. L., Davies P. J. 2010. Molecular aspects of thyroid hormone actions. Endocr. Rev. 31, 139–170 10.1210/er.2009-0007 (doi:10.1210/er.2009-0007) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Taleux N., Guigas B., Dubouchaud H., Moreno M., Weitzel J. M., Goglia F., Favier R., Leverve X. M. 2009. High expression of thyroid hormone receptors and mitochondrial glycerol-3-phosphate dehydrogenase in the liver is linked to enhanced fatty acid oxidation in Lou/C, a rat strain resistant to obesity. J. Biol. Chem. 13, 4308–4316 [DOI] [PubMed] [Google Scholar]
- 28.Dozin B., Magnuson M. A., Nikodem V. M. 1986. Thyroid hormone regulation of malic enzyme synthesis. Dual tissue-specific control. J. Biol. Chem. 261, 10 290–10 292 [PubMed] [Google Scholar]
- 29.Collin A., Taouis M., Buyse J., Ifuta N. B., Darras V. M., Van As P., Malheiros R. D., Moraes V. M., Decuypere E. 2003. Thyroid status, but not insulin status, affects expression of avian uncoupling protein mRNA in chicken. Am. J. Physiol. 284, E771–E777 [DOI] [PubMed] [Google Scholar]
- 30.Rey B., Roussel D., Romestaing C., Belouze M., Rouanet J. L., Desplanches D., Sibille B., Servais S., Duchamp C. 2010. Up-regulation of avian uncoupling protein in cold-acclimated and hyperthyroid ducklings prevents reactive oxygen species production by skeletal muscle mitochondria. BMC Physiol. 10, 5. 10.1186/1472-6793-10-5 (doi:10.1186/1472-6793-10-5) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Chen W., Zhang X., Birsoy K., Roeder R. G. 2010. A muscle-specific knockout implicates nuclear receptor coactivator MED1 in the regulation of glucose and energy metabolism. Proc. Natl Acad. Sci. USA 107, 10 196–10 201 10.1073/pnas.1005626107 (doi:10.1073/pnas.1005626107) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Duteil D., Chambon C., Ali F., Malivindi R., Zoll J., Kato S., Geny B., Chambon P., Metzger D. 2010. The transcriptional coregulators TIF2 and SRC-1 regulate energy homeostasis by modulating mitochondrial respiration in skeletal muscles. Cell Metab. 12, 496–508 10.1016/j.cmet.2010.09.016 (doi:10.1016/j.cmet.2010.09.016) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Chen W., Yang Q., Roeder R. G. 2009. Dynamic interactions and cooperative functions of PGC-1a and MED1 in TRα-mediated activation of the brown fat specific UCP-1 gene. Mol. Cell 35, 755–768 10.1016/j.molcel.2009.09.015 (doi:10.1016/j.molcel.2009.09.015) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Otto T., Fandrey J. 2008. Thyroid hormone induces hypoxia-inducible factor 1α gene expression through thyroid hormone receptor β/retinoid X receptor α-dependent activation of hepatic leukemia factor. Endocrinology 149, 2241–2250 10.1210/en.2007-1238 (doi:10.1210/en.2007-1238) [DOI] [PubMed] [Google Scholar]
- 35.Bracken C. P., Whitelax M. L., Peet D. J. 2003. The hypoxia-inducible factors: key transcriptional regulators of hypoxic responses. Cell. Mol. Life Sci. 60, 1376–1393 10.1007/s00018-003-2370-y (doi:10.1007/s00018-003-2370-y) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ma Y., Freitag P., Zhou J., Brüne B., Frede S., Fandrey J. 2004. Thyroid hormone induces erythropoietin gene expression through augmented accumulation of hypoxia-inducible factor-1. Am. J. Physiol. 287, R600–R607 [DOI] [PubMed] [Google Scholar]
- 37.Lowry O., Passonneau J. A. 1972. Flexible system of enzymatic analysis. New York, NY: Academic Press [Google Scholar]
- 38.Froget G., Handrich Y., Le Maho Y., Rouanet J. L., Woakes A. J., Butler P. J. 2002. The heart rate/oxygen consumption relationship during cold exposure of the king penguin: a comparison with that during exercise. J. Exp. Biol. 205, 2511–2517 [DOI] [PubMed] [Google Scholar]
- 39.Barre H., Rouanet J. L. 1983. Calorigenic effect of glucagon and catecholamines in king penguin chicks. Am. J. Physiol. 244, 758–763 [DOI] [PubMed] [Google Scholar]
- 40.Bolstad B. M., Irizarry R. A., Astrand M., Speed T. P. 2003. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–193 10.1093/bioinformatics/19.2.185 (doi:10.1093/bioinformatics/19.2.185) [DOI] [PubMed] [Google Scholar]
- 41.Smyth G. K. 2004. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 1027. [DOI] [PubMed] [Google Scholar]
- 42.Bolker B. M., Brooks M. E., Clark C. J., Geange S. W., Poulsen J. R., Stevens M. H. H., White J. S. S. 2009. Generalized linear mixed models: a practical guide for ecology and evolution. Trends Ecol. Evol. 24, 127–135 10.1016/j.tree.2008.10.008 (doi:10.1016/j.tree.2008.10.008) [DOI] [PubMed] [Google Scholar]