Abstract
Achieving humoral immunity against human immunodeficiency virus (HIV) is a major obstacle in AIDS vaccine development. Despite eliciting robust humoral responses to HIV, exposed hosts rarely produce broadly neutralizing antibodies. The present study utilizes simian immunodeficiency virus (SIV) to identify viral epitopes that conferred antibody neutralization to clone SIV/17E-CL, an in vivo variant derived from neutralization resistant SIVmac239. Neutralization assays using rhesus macaque monoclonal antibodies were performed on viruses engineered to express single or multiple amino acid mutations. Results identified a single amino acid mutation, P334R, in the carboxy-terminal half of the V3 loop as a critical residue that induced neutralization while retaining normal glycoprotein expression on the surface of the virus. Furthermore, the R334 residue yielded neutralization sensitivity by antibodies recognizing diverse conformational and linear epitopes of gp120, suggesting that neutralization phenotype was a consequence of global structural changes of the envelope protein rather than a specific site epitope.
Introduction
Infection of nonhuman primates with simian immunodeficiency virus (SIV) provides a model for studying immune responses associated with HIV/AIDS in humans (Johnson and Hirsch, 1992). Both cellular and humoral immune responses have been correlated with protective immunity against SIV (Clements et al., 1995; Maecker and Maino, 2003; Paiardini et al., 2008; Sato and Johnson, 2007). Passive protection studies have further demonstrated that antibodies can provide protective immunity when present prior to or immediately preceding HIV or SIV challenge (Haigwood et al., 1996; Lewis et al., 1993; Mascola et al., 1999; Mascola et al., 2003; Mascola et al., 2000; Nishimura et al., 2002; Nishimura et al., 2003; Parren et al., 2001; Van Rompay et al., 1998). Unfortunately, these protective antibodies are infrequently observed in exposed hosts and are predominantly directed to intricate, conformationally dependent epitopes of the SIV envelope (env) proteins in regions with a propensity for mutation and subsequent immune evasion (Cole et al., 2001; Haigwood et al., 1992; Javaherian et al., 1994; Javaherian et al., 1992; Sato and Johnson, 2007). Therefore, identifying viral epitopes that play a role in antibody neutralization are important for our understanding of the immunogenic properties of SIV and HIV and will serve to enhance vaccine development.
Some limited HIV and SIV antibody neutralizing epitopes have been identified within the viral env gene (Pantophlet and Burton, 2006; Sato and Johnson, 2007). The HIV and SIV env gene produces a polyprotein (gp160) that is extensively modified by post-translational polysaccharide addition and is cleaved by host protease (furin) into two separate glycoproteins, gp120 [surface subunit (SU)] and gp41 [transmembrane subunit (TM)] (Luciw, 1996). On the surface of the virus, complete env complexes are comprised of a trimer of noncovalently linked heterodimers of gp120 and gp41. The role of these proteins in virus attachment and entry has been well characterized (Wyatt and Sodroski, 1998). Gp120 is involved with CD4 and co-receptor recognition and binding, while gp41 is responsible for forming the env trimer and mediating cell-virus fusion. Less well known is how neutralizing antibody responses to SIV and HIV are directed against the viral epitopes of the env proteins. Neutralizing antibody responses against SIV have been demonstrated to be associated with the gp41 cytoplasmic tail, ecto and transmembrane domains (Bonavia et al., 2005; Overholser et al., 2005; Puffer et al., 2004). Within gp120, V1/V2 (Johnson et al., 2002; Johnson et al., 2003; Puffer et al., 2004), V3 loop (Means et al., 2001; Palker et al., 1996; Pohlmann et al., 2004) and V4 regions (Choi et al., 1994; Kinsey et al., 1996) have also demonstrated significant roles in affecting virus neutralization by antibodies. Identifying and characterizing these determinants of neutralization in SIV has increased our understanding of the antigenic qualities of envelope proteins. However, more information is required to solve the precise molecular structure and antigenic qualities of these proteins.
The present study was designed to identify the amino acid residues within the env gene that contribute to the antibody neutralization phenotype of SIV/17E-CL, a naturally derived clone of neutralization resistant SIVmac239 that acquired nine amino acid mutations in gp120 (V67M, K141R, T158A, K176N, Q217K, M309I, P334R, K340R and G382R) (Anderson et al., 1993; Regier and Desrosiers, 1990; Sharma et al., 1992). Our earlier investigations using surface plasmon resonance (SPR) determined that differences in association and dissociation kinetic rates of antibody with SIV/17E-CL env proteins were causative for the neutralization phenotype (Steckbeck et al., 2005). However, the region of the viral protein or the epitope(s) involved remained unknown. Here we investigated antibody mediated neutralization in vitro with viruses engineered to express amino acid residues from SIV/17E-CL gp120 within the SIVmac239 backbone. Results from these studies described a novel V3 epitope that conferred neutralization of SIVmac239 by monoclonal antibodies to both linear and conformational epitopes, suggesting that neutralization phenotype was the consequence of global structural changes of the env protein complex.
Materials and Methods
Cells
293T cells and SIV permissive TZM-bl cells (Wei et al., 2002), which express CD4, CCR5, CXCR4 and HIV/SIV inducible firefly luciferase, were maintained in Dulbecco’s modified Eagle’s medium (DMEM) media (Gibco-Invotrogen, Carlsbad, CA) supplemented with 10% fetal bovine serum (FBS) (Atlanta Biologicals, Lawrenceville, GA).
Construction of mutant proviruses
Previously characterized provirus constructs SIVmac239 (Regier and Desrosiers, 1990) and SIVmac17E-CL (Anderson et al., 1993) were digested with Nhe1 and Bpu10I to isolate the encoding region of gp120 and a portion of the ectodomain of gp41 (fig 1). This region was ligated with homologous restriction sites into a CMV-promoter expression vector, TR600 (Bower et al., 2004), using T4 DNA ligase (New England Biolabs, Ipswich, MA) and recombinant DNA was synthesized in DH5α competent cells. The wild-type gp120 env sequences were mutagenized using the QuickChange XL Site Directed Mutagenesis kit (Stratagene, La Jolla, CA) according to manufacturer’s protocol and sequence specific primers. The mutant env constructs were ligated back into the SIV proviral backbone using Nhe1 and Bpu10I restriction sites and T4 DNA ligase and propagated in Stbl2 competent cells (Invitrogen, Carlsbad, CA). Transfection grade plasmids clones were verified by DNA sequencing at the University of Pittsburgh Genomics and Proteomics Core laboratory (Pittsburgh, PA) using an ABI 3730xl DNA analyzer (Applied Biosystems, Foster City, CA).
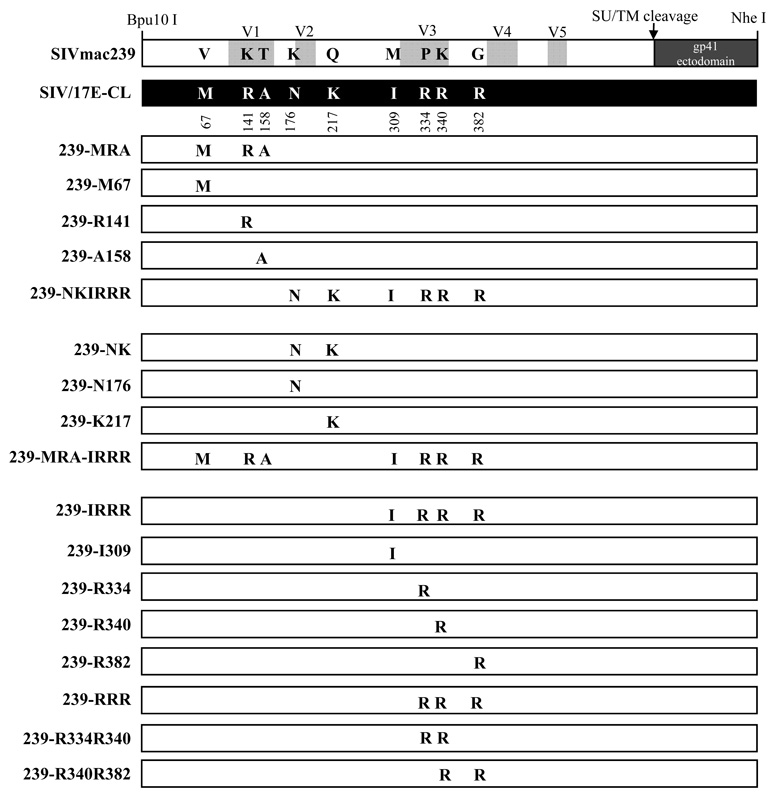
Figure 1.
Design and characterization of SIV mutant viruses. Mutations in SIV/17E-CL are distributed throughout the gp120 env gene and cluster around the V1/V2 and V3 regions. The cloning strategy utilizes the Bpu10I restriction site, 5’ to the start codon of gp120, and the Nhe1 restriction site on the 3’ end of the ectodomain of gp41. All mutant constructs created infectious virus in vitro that activated LTR-luc expression in the TZM-bl. Viral titer stocks ranged from 102–105 TCID50 per ml.
Production of viruses
SIVmac239, SIV/17E-CL and mutant proviral DNA was transiently transfected as previously described (Bower et al., 2004). Briefly, 8µg of proviral DNA was conjugated with 24 µl Lipofectamine 2000 (Invitrogen) in Opti-MEM media (Invitrogen) and incubated with 6–7 × 106 293T cells for 5 hours at 37°C. The media was replaced with 10ml of fresh Opti-MEM and the cultures were incubated an additional 48 hours at 37°C. Cell free supernatants were harvested and frozen at −80°C prior to experimental analysis. Virus titer was determined by serially diluting viral supernatants in DMEM media and culturing with TZM-bl cells (104 cells per well) in 96 well plates (BD-Falcon, San Jose, CA). Following an incubation of 48 hours, supernatants were removed and the cells were lysed with 50 µl cell lysis buffer (Promega, Madison, WI) for 15 minutes at room temperature. Lysates (40 µL) were transferred to white luminescence plates (USA Scientific, Ocala, Fl) and assayed for luciferase production on a Lmax II luminometer (Molecular Devices, Union City, CA) with 25 µl of Luciferase Assay Substrate (Promega) per well. The TCID50 was determined using the Reed and Meunch method (Poli and Fauci, 2004), with a positive cutoff value 2.5 times that of the negative samples.
Virus neutralization determination
MAbs (table 1) were serially diluted 5-fold in DMEM containing 10% FBS in sterile 96 well V-bottom plates (Sarstedt, Newton, NC) for a final assay concentration range of 10 - 0.00013 µg/ml. Virus was diluted in DMEM with 10% FBS to yield an MOI of 0.01, mixed with diluted MAb and pre-incubated 1h at 37°C. The virus-MAb mixture was added to TZM-bl cells (104 cells per well) in 96 well plates (BD-Falcon) and cultured in the presence of 20 µg/ml DEAE-Dextran (Amersham Biosciences, Piscataway, NJ) for 48 hours. SIV infection was assayed by luciferase production as described above. Neutralizing titers are reported as the IC50, the MAb concentration (µg/ml) that neutralized 50% of the virus infection, using the method of Karber (Karber, 1931). All values represent the averages of at least two independent experiments assayed in duplicate.
Table 1.
Monoclonal antibody phenotypes: epitope specificity and SIV neutralization
| MAb | Binding groupa | Putative binding domainb |
Epitope conformationc |
Neutralizing antibody titer (µg/ml)d | |
|---|---|---|---|---|---|
| SIV/17E-CL | SIVmac239 | ||||
| 3.8E | I | N' terminus | conformational | >25 | >25 |
| 3.10A | II | V1 | linear | >25 | >25 |
| 5.5B | III | V2 | linear | >25 | >25 |
| 3.11H | IV | V3-cys loop | linear | 0.070 | >25 |
| 1.11A | V | CCR5 bs | conformational | 0.125 | >25 |
| 4.7A | V | CCR5 bs | conformational | 0.021 | >25 |
| 3.11E | VIa | CCR5 bs | conformational | 0.038 | >25 |
| 3.2C | VIb | V4 | conformational | 0.015 | >25 |
| E31 | VIb | V4 | conformational | 0.039 | >25 |
| 1.10A | VII | V4 | conformational | 0.009 | >25 |
| C26 | VII | V4 | conformational | 0.006 | >25 |
| 3B3 | VIII | Unknown | conformational | >25 | >25 |
Competition groups were previously defined by cross-competition ELISA(Cole et al. 2001)
Putative binding domains are based on previously published data (Cole et al. 2001)
Epitope conformation was previously determined by reducing/nonreducing immunoblot (Cole et al. 2001)
Neutralization antibody titers were previously assayed and reported as the concentration of MAb that inhibited 50% infection (IC50) using the TZM-bl assay (Steckbeck et al. 2005)
Western blot analysis
To quantitate envelope content of each SIV mutant, viral supernatants were pelleted over a 10% glycerol cushion at 19,000 × g for 1.5 h at 4°C, denatured by boiling in reducing buffer and resolved by SDS-PAGE using Tris-HCl 5 and 12.5% gels (Bio-Rad, Hercules, CA). Viral proteins were transferred to polyvinylidine difluoride membrane and probed for gp120 env using a MAb directed against a linear epitope in V3 not affected by the mutations in this study [3.11H, (Cole et. al. 2001)] and p27gag using MAb 2F12 (AIDS Research and Reference Reagent Program). Band densities of gp120 env and p27gag were determined using ImageJ software (National Institutes of Health, Bethesda, MD). Each immunoblot contained a positive control of SIVmac239, thus allowing for calculations of the gp120/p27gag ratio for each mutant virus in relation to the parental strain SIVmac239.
Results
Synthesis of mutant viruses
The envelope gene of SIV/17E-CL differs from SIVmac239 by 9 amino acids in gp120 (V67M, K141R, T158A, K176N, Q217K, M309I, P334R, K340R and G382R), see figure 1 (Anderson et al., 1993). Single and multiple rounds of site directed mutagenesis (see materials and methods) were used to generate amino acid mutations in the gp120 env SIVmac239 backbone according to a scheme that represents mutation sets in and around the V1,V2 or V3 regions of SIV/17E-CL (fig 1). Viral stocks were generated from recombinant virus construct in mammalian tissues as described in the materials and methods section. Synthesis of infectious virus was verified from every engineered construct and yielded viral stock titers ranging from 102–105 TCID50 per ml (data not shown).
Envelope expression in SIV mutants
Previous reports suggest that mutations in SIV gp120 that reduce envelope incorporation into viral membranes and/or increase env shedding from the viral membrane result in enhanced sensitivity to antibody mediated neutralization (Klasse and Moore, 1996; Yuste et al., 2005). We previously demonstrated that SIV/17E-CL env expression is similar to SIVmac239 and is therefore not a factor for neutralization (Steckbeck et al., 2005). To determine whether the mutant viruses studied here displayed altered env expression, we assessed the quantity of gp120 present in mutant viruses. Figure 2a demonstrated that with the exception of 239-A158, 239-NK-IRRR, 239-NK and 239-N176, all mutant viruses expressed levels of gp120 env similar to SIVmac239. To measure the level of gp120 env in each mutant clone, band densities from duplicate experiments were determined for gp120 env and p27gag and the ratios were compared to the parental virus, SIVmac239. Using this semi-quantitative analysis, four mutant viruses (239-A158, 239-NKIRRR, 239-NK and 239-N176) displayed significant decreases in gp120 expression (35, 4.9, 17 and 11% respectively) (fig 2b). Interestingly, while 239-A158 displayed reduced gp120 expression, 239-MRA and 239-MRA-IRRR, which also contained the T158A mutation, did not express a reduced gp120 env phenotype. So, the significant reduction of env by T158A only occurred when T158A was expressed alone (239-A158). Figure 2b also demonstrated that mutant viruses with the K176N mutation (239-NKIRRR, 239-NK and 239-N176) expressed significantly lower levels of gp120 env. Thus, the analysis of gp120 expression showed that env content was unperturbed in most mutants and significant variability was limited to few mutants with distinct point mutations.
Figure 2.
Analysis of gp120 expression in mutant viruses. (a) Virus supernatants were pelleted and denatured viral proteins were immunodetected for gp120 env (MAb 3.11H) and structural protein p27gag (MAb 2F12). (b) Band densities were determined and the gp120/ p27gag ratio of each mutant was calculated. The data reported is the percent change in mean gp120/ p27gag ratio (±S.D.) compared to SIVmac239 from two independent experiments.
Lastly, the immunoblot analysis revealed one further observation. Figure 2a showed that gp120 env from mutants with the A158 residue (239-MRA, 239-A158 and 239-MRA-IRRR) resolved at a lower molecular weight than SIVmac239 and mutants with T158. This migration pattern of the env proteins would suggest that the T158A mutation resulted in a loss of an N-linked glycosylation residue, as expected by the change in amino acid sequence, N-C-T →N-C-A.
Analysis of V1/V2 region residues for neutralization sensitivity to monoclonal antibodies
Each mutant virus was examined for neutralization sensitivity using the TZM-bl assay (Steckbeck et al., 2005). Neutralization studies were divided into study groups of mutants according to region, and are summarized in fig 1. Data from V1 region mutants showed that the 239-MRA virus (mutations V67M, K141R and T158A) was not neutralized by MAb at the highest concentration (10µg/ml) (table 2). The 239-M67 (V67M) and 239-R141 (K141R) mutants alone were also not sensitive to neutralization (table 2). However, the 239-A158 mutant (T158A) yielded a clone that was partially sensitive to neutralization by MAbs that recognize epitopes on the carboxy-terminal half of gp120. Four of the eight neutralizing antibodies yielded mean IC50 values (0.64 - 0.037 µg/ml, table 2) within one log10 deviation from the IC50 values observed in SIV/17E-CL (table 1). Interestingly, the 239-NKIRRR mutant that contains the V2 and V3 mutations and none of the V1 mutations demonstrated a neutralization profile more comparable to SIV/17E-CL (tables 1) with IC50 values ranging from 0.39 to 0.003 µg/ml (table 2). Therefore, the single point mutation T158A in V1 conferred partial neutralization sensitivity in only 239-A158, while determinants outside of V1 imparted full neutralization.
Table 2.
Neutralization phenotype resulting from V1 and V2 mutations a
| Monoclonal Antibody |
Binding Domain |
V1 | V2 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Mac239 | 17E-CL | 239-MRA | 239-M67 | 239-R141 | 239-A158 | 239-NKIRRR | 239-NK | 239-N176 | 239-K217 | 239-MRA-IRRR | ||
| 3.8E | N' terminus | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 |
| 3.10A | V1 | >10 | >10 | >10 | >10 | >10 | >10 | 6.43 (0.81) | >10 | >10 | >10 | >10 |
| 5.5B | V2 | >10 | >10 | >10 | >10 | >10 | >10 | 1.18 (0.45) | >10 | >10 | >10 | >10 |
| 3.11H | V3-cys loop | >10 | 0.046 (0.02) | >10 | >10 | 8.30 (2.4) | 0.510 (0.41) | 0.028 (0.02) | 1.01 (0.43) | 1.11 (1.1) | >10 | 0.260 (0.18) |
| 1.11A | CCR5 bs | >10 | 0.075 (0.05) | >10 | >10 | >10 | 0.143 (0.14) | 0.039 (0.01) | 0.215 (0.01) | 0.857 (0.90) | >10 | 0.673 (0.31) |
| 4.7A | CCR5 bs | >10 | 0.016 (0.01) | >10 | >10 | >10 | 0.300 (0.27) | 0.017 (0.01) | 0.530 (0.32) | 0.233 (0.15) | >10 | 0.300 (0.16) |
| 3.11E | CCR5 bs | >10 | 0.024 (0.02) | >10 | >10 | 8.65 (1.9) | 0.640 (0.65) | 0.010 (0.01) | 1.24 (1.1) | 0.154 (0.19) | >10 | 0.220 (0.07) |
| 3.2C | V4 | >10 | 0.022 (0.01) | >10 | >10 | >10 | 0.225 (0.22) | 0.029 (0.01) | 0.273 (0.16) | 0.331 (0.29) | >10 | 0.133 (0.07) |
| E31 | V4 | >10 | 0.029 (0.02) | >10 | >10 | >10 | 0.175 (0.02) | 0.023 (0.01) | 0.195 (0.02) | 0.042 (0.01) | >10 | 0.520 (0.28) |
| 1.10A | V4 | >10 | 0.005 (0.004) | >10 | >10 | 5.25 (3.7) | 0.026 (0.02) | 0.002 (0.001) | 0.084 (0.01) | 0.029 (0.03) | 5.49 (4.1) | 0.033 (0.01) |
| C26 | V4 | >10 | 0.004 (0.003) | >10 | >10 | >10 | 0.037 (0.003) | 0.003 (0.001) | 0.089 (0.03) | 0.031 (0.04) | >10 | 0.026 (0.01) |
| 3B3 | Unknown | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 |
IC50 neutralization titers are reported as the mean concentration of MAb (µg/ml) (±S.D.) from at least two independent experiments. Values in bold type represent mean IC50 values that do not exceed 1 log10 variance from the IC50 of SIV/17E-CL.
Next, mutations in the V2 region were examined. The 239-NK virus containing both the K176N and Q217K mutations was partially sensitive to neutralization (table 2), albeit to a lesser degree than SIV/17E-CL (table 1). Only two of the eight neutralizing antibodies displayed IC50 values within a log10 deviation of SIV/17E-CL. Further analysis of the V2 mutations was performed with clones 239-N176 (K176N) and 239-K217 (Q217K). 239-N176 was sensitive to antibody neutralization (IC50 values 1.11 to 0.031 µg/ml), while 239-K217 was not neutralized by the antibodies tested (table 2). Clone 239-MRA-IRRR lacking the V2 region mutations, but containing the V1 and V3 mutations was more sensitive to neutralization than the 239-NK virus (table 2), with IC50 values ranging from 0.67 to 0.026 µg/ml. Therefore, the data of the V2 region analysis showed that the K176N mutation was sufficient to confer partial neutralization in SIV/17E-CL, but that the V3 region contained factors for neutralization as well.
Analysis of V3 region residues for neutralization sensitivity to monoclonal antibodies
Lastly we evaluated mutations in and around the V3 region (M309I, P334R, K340R and G382R) for neutralization sensitivity. Mutant clone 239-IRRR, expressing all four point mutations around V3, showed IC50 values equivalent to SIV/17E-CL, ranging from 0.31 to 0.006 µg/ml (table 3). To elucidate the critical neutralizing mutations in V3, additional single point mutants were generated and assayed for neutralization. The 239-I309 (M309I), 239-R340 (K340R) and 239-R382 (G382R) mutants were resistant to neutralization by all MAbs tested (table 3). Of these clones, only 239-R334 (P334R) demonstrated a neutralization phenotype equivalent to 239-IRRR and SIV/17E-CL (table 3). To determine the contribution of the neighboring mutations that also introduced an arginine (R) in and around V3 (K340R and G382R), combination mutant clones were assayed for neutralization. The data from clones 239-RRR, 239-R334R340 and 239-R340R382 reinforced the observation of the P334R mutation as the neutralizing determinant, as only those clones containing the R334 residue (239-RRR and 239-R334R340) yielded a neutralization phenotype equal to 239-IRRR (table 3). Furthermore, all mutant clones assayed that contained P334R (239-NKIRRR, 239-MRA-IRRR, 239-IRRR, 239-R334, 239-RRR and 239-R334R340) were neutralization sensitive (table 2 and table 3) with IC50 values for MAbs that were equivalent to SIV/17E-CL.
Table 3.
Neutralization phenotype resulting from V3 mutations a
| Monoclonal Antibody |
Binding Domain |
V3 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Mac239 | 17E-CL | 239-IRRR | 239-I309 | 239-R334 | 239-R340 | 239-R382 | 239-RRR | 239-R334R340 | 239-R340R382 | ||
| 3.8E | N' terminus | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 |
| 3.10A | V1 | >10 | >10 | 3.58 (0.39) | >10 | 6.43 (2.8) | >10 | >10 | 7.13 (4.9) | >10 | >10 |
| 5.5B | V2 | >10 | >10 | 9.08 (1.85) | >10 | 5.58 (4.7) | 8.90 (2.2) | >10 | >10 | >10 | >10 |
| 3.11H | V3-cys loop | >10 | 0.046 (0.02) | 0.313 (0.38) | >10 | 0.111 (0.06) | 8.98 (2.0) | 9.36 (1.1) | 0.086 (0.06) | 0.435 (0.50) | >10 |
| 1.11A | CCR5 bs | >10 | 0.075 (0.05) | 0.228 (0.24) | >10 | 0.110 (0.07) | >10 | >10 | 0.069 (0.03) | 0.200 (0.12) | >10 |
| 4.7A | CCR5 bs | >10 | 0.016 (0.01) | 0.056 (0.07) | >10 | 0.051 (0.03) | >10 | >10 | 0.042 (0.02) | 0.075 (0.02) | >10 |
| 3.11E | CCR5 bs | >10 | 0.024 (0.02) | 0.059 (0.07) | >10 | 0.062 (0.04) | 7.85 (4.3) | 7.68 (2.7) | 0.050 (0.02) | 0.072 (0.05) | >10 |
| 3.2C | V4 | >10 | 0.022 (0.01) | 0.014 (0.02) | >10 | 0.048 (0.03) | 7.10 (3.6) | 7.76 (4.4) | 0.016 (0.01) | 0.060 (0.04) | >10 |
| E31 | V4 | >10 | 0.029 (0.02) | 0.010 (0.13) | >10 | 0.101 (0.03) | >10 | >10 | 0.161 (0.15) | 0.050 (0.01) | >10 |
| 1.10A | V4 | >10 | 0.005 (0.004) | 0.005 (0.004) | >10 | 0.004 (0.002) | 6.73 (2.8) | 7.08 (3.6) | 0.004 (0.002) | 0.023 (0.03) | 7.80 (3.8) |
| C26 | V4 | >10 | 0.004 (0.003) | 0.006 (0.007) | >10 | 0.004 (0.004) | 7.85 (4.3) | 9.06 (1.6) | 0.006 (0.002) | 0.010 (0.01) | >10 |
| 3B3 | Unknown | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 | >10 |
IC50 neutralization titers are reported as the mean concentration of MAb (µg/ml) (±S.D.) from at least two independent experiments. Values in bold type represent mean IC50 values that do not exceed 1 log10 variance from the IC50 of SIV/17E-CL.
Discussion
This study analyzed the antibody neutralization phenotypes resulting from single and multiple point mutations in SIVmac239 gp120 representing mutations derived from SIV/17E-CL. Using a broad panel of rhesus MAbs with linear and conformational epitopes to multiple domains throughout SIV gp120, we sought to identify the mutation(s) responsible for inducing antibody mediated neutralization seen in SIV/17E-CL. The first mutation identified, T158A, only influenced neutralization when expressed alone, as seen in virus 239-A158 (table 2). Interestingly, the mutation of threonine 158 to alanine removes a potential N-linked glycosylation site from SIVmac239 (g6) by disrupting the N-linked glycosylation signal (Asn-X-Ser/Thr), N-C-T →N-C-A. Previous research suggests that the carbohydrates around V1/V2 and V3 shield epitopes on env because removal of glycosylation sites in SIV often enhances antibody neutralization (Chackerian, Rudensey, and Overbaugh, 1997; Cole et al., 2004; Reitter and Desrosiers, 1998; Reitter, Means, and Desrosiers, 1998). In our study, the T158A Δg6 mutation induced a neutralizing phenotype when expressed alone (239-A158), but showed no effect when other mutations were expressed in cis, 239-MRA (V67M, K141R and T158A). A more significant observation of the 239-A158 (T158A) mutant was a significant reduction in gp120 env/p27gag ratio compared to SIVmac239 (figure 2b). Reduced envelope content in the virion has been correlated with neutralization sensitivity of HIV and SIV (Klasse and Moore, 1996; Yuste et al., 2005). Therefore, reduced env protein expression would explain the neutralization sensitivity of the 239-A158 clone, yet does not dictate the neutralization phenotype of SIV/17E-CL which does not have lowered env expression.
Next, a K176N mutation amino-terminal to V2 was observed to induce monoclonal antibody mediated neutralization in vitro, albeit to a lesser degree than SIV/17E-CL (table 1 and table 2). As reported previously, V1/V2 plays a role in neutralization, given that mutations and deletions of V1/V2 induce a neutralizing phenotype (Johnson et al., 2002; Johnson et al., 2003; Puffer et al., 2004). Neutralization sensitivity has been reported for SIVmac316 that expresses a mutation at amino acid 176 in gp120 (Johnson et al., 2003; Means et al., 2001). Furthermore, the MERT cluster of mutations (V67M, K176E, G382R K573T) are implicated in the macrophage tropic phenotype associated with SIVmac316 (Mori et al., 1992). Macrophage tropism is often associated with antibody neutralization sensitivity in SIV (Means et al., 2001; Puffer et al., 2002; Rudensey et al., 1998). Indeed, SIV/17E-CL is macrophage tropic (Mankowski et al., 1997), however the V67M and G382R mutations had no effect on neutralization sensitivity in this study (table 2 and table 3). More importantly the K176N mutation yielded significant reductions in envelope expression (fig 2). As may be the case for the T158A mutation, the atypically lowered expression of the glycoproteins on the viral membrane of the SIV K176N mutant induced the observed antibody neutralization phenotype.
In this study, the most significant changes in antibody mediated neutralization were observed in the carboxy-terminal portion of the V3 loop at residue 334 (table 3). All mutants assayed that contained R334 (239-NKIRRR, 239-MRA-IRRR, 239-IRRR, 239-R334, 239-RRR and 239-R334R340) were neutralized by MAbs at concentrations equivalent to SIV/17E-CL while retaining normal protein expression of env (fig. 2). Although impressive, it is not so surprising given the documented importance that HIV and SIV V3 has in regard to viral tropism and neutralization phenotypes (Hartley et al., 2005). Evidence thus far suggests that the V3 loop may be split into two domains each with a different role in determining phenotype. The amino-terminal portion is suggested to directly interact with neutralizing antibodies, while the carboxy-terminal portion is weakly or non-immunogenic and may interact with other domains of the gp120 monomer (Benichou et al., 1993; Huang et al., 2005; McBride et al., 1993; Miller, Murphey-Corb, and Montelaro, 1992; Palker et al., 1996; Pohlmann et al., 2004; Rosen et al., 2005; Rosen, Samson, and Anglister, 2008). Changes in the carboxy-terminal domain of V3 are suspected to influence phenotype via structural changes of the individual monomers or the complete trimer (Huang et al., 2005; Rosen, Samson, and Anglister, 2008; Rosen et al., 2006). Our data supports this theory in part because the neutralizing phenotype obtained by the P334R mutation was apparent not only for a V3 binding antibody (3.11H), but for the entire panel of MAbs recognizing both linear and conformational epitopes clustered throughout the gp120 molecule (table 3).
One particular model in HIV could explain structure changes associated with R334. The “11/25” rule states that in HIV-1 negative/neutral charges at residues 11 or 25 in the V3 loop confer CCR5 co-receptor usage and macrophage tropism, while positive charges yield CXCR4 co-receptor usage and T-lymphocyte or dual tropism phenotype (De Jong et al., 1992; Fouchier et al., 1992; Resch, Hoffman, and Swanstrom, 2001). The corresponding amino acids in T-lymphotrophic SIVmac239 V3 are 11 (P306) and 25 (P334). Interestingly, Rosen et al. (Rosen, Samson, and Anglister, 2008) recently demonstrated that the 25th amino acid in HIV-1 V3, residue 322, displays electrostatic interaction with residue 440 in the C4 domain. The 440 residue in C4 was mapped as the second amino acid following a conserved YAPP domain, which is also present in SIVmac as a YLPP domain. Interestingly, the second amino acid carboxy-terminal to this domain in SIV is a negatively charge glutamic acid E454. If SIV structure is homologous to HIV-1, the R334 and E454 would theoretically interact with electrostatic attraction and yield structure and phenotype of macrophage tropic viruses. Indeed, studies of SIV V3 mutations found that changing P334 to uncharged amino acids leucine or glutamine did not confer macrophage tropism in SIVmac239 (Kirchhoff, Mori, and Desrosiers, 1994; Pohlmann et al., 2004), yet SIV/17E-CL (R334) is a macrophage tropic virus (Mankowski et al., 1997). Therefore, it is feasible that the 11/25 rule for positivity applies to HIV-1, but could more broadly reflect the necessity for position specific electrostatic interactions between V3 and other regions of SIV gp120, such as V4.
This study indentified the P334R mutation in the C’-terminal portion of the V3 loop to be the main determinant of neutralization in SIV/17E-CL. Given that this residue affected neutralization by MAbs recognizing diverse conformational and linear epitopes in gp120 and that the V3 region has been observed to be associated with viral tropism and neutralization phenotypes, it is likely that the neutralization sensitive phenotype of SIV/17E-CL is a result of the structural changes of the envelope protein elicited by the P334R mutation. Given the currently limited data describing V3 structure within the context of gp41/gp120 trimers, this report supports the importance of further studies on the V3 region to assist in developing effective therapeutics and preventative strategies against HIV AIDS.
Acknowledgments
This work was supported by the University of Pittsburgh Center for Vaccine Research and funding from a National Institutes of Health grant (AI 52058) to KC.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Competing Interests
The authors declare that they have no competing interests.
Contributor Information
Seth A. Faith, Email: saf31@pitt.edu.
Yingyun Wu, Email: ywu@novavax.com.
David Kuhrt, Email: dmk26@pitt.edu.
Jonathan D. Steckbeck, Email: jds170@pitt.edu.
Jodi K. Craigo, Email: craigoj@pitt.edu.
Janice E. Clements, Email: jclement@bs.jhmi.edu.
Kelly Stefano Cole, Email: stefcole@pitt.edu.
References
- Anderson MG, Hauer D, Sharma DP, Joag SV, Narayan O, Zink MC, Clements JE. Analysis of envelope changes acquired by SIVmac239 during neuroadaption in rhesus macaques. Virology. 1993;195(2):616–626. doi: 10.1006/viro.1993.1413. [DOI] [PubMed] [Google Scholar]
- Benichou S, Venet A, Beyer C, Tiollais P, Madaule P. Characterization of B-cell epitopes in the envelope glycoproteins of simian immunodeficiency virus. Virology. 1993;194(2):870–874. doi: 10.1006/viro.1993.1333. [DOI] [PubMed] [Google Scholar]
- Bonavia A, Bullock BT, Gisselman KM, Margulies BJ, Clements JE. A single amino acid change and truncated TM are sufficient for simian immunodeficiency virus to enter cells using CCR5 in a CD4-independent pathway. Virology. 2005;341(1):12–23. doi: 10.1016/j.virol.2005.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bower JF, Yang X, Sodroski J, Ross TM. Elicitation of neutralizing antibodies with DNA vaccines expressing soluble stabilized human immunodeficiency virus type 1 envelope glycoprotein trimers conjugated to C3d. J Virol. 2004;78(9):4710–4719. doi: 10.1128/JVI.78.9.4710-4719.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chackerian B, Rudensey LM, Overbaugh J. Specific N-linked and O-linked glycosylation modifications in the envelope V1 domain of simian immunodeficiency virus variants that evolve in the host alter recognition by neutralizing antibodies. J Virol. 1997;71(10):7719–7727. doi: 10.1128/jvi.71.10.7719-7727.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi WS, Collignon C, Thiriart C, Burns DP, Stott EJ, Kent KA, Desrosiers RC. Effects of natural sequence variation on recognition by monoclonal antibodies neutralize simian immunodeficiency virus infectivity. J Virol. 1994;68(9):5395–5402. doi: 10.1128/jvi.68.9.5395-5402.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clements JE, Montelaro RC, Zink MC, Amedee AM, Miller S, Trichel AM, Jagerski B, Hauer D, Martin LN, Bohm RP, et al. Cross-protective immune responses induced in rhesus macaques by immunization with attenuated macrophage-tropic simian immunodeficiency virus. J Virol. 1995;69(5):2737–2744. doi: 10.1128/jvi.69.5.2737-2744.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cole KS, Alvarez M, Elliott DH, Lam H, Martin E, Chau T, Micken K, Rowles JL, Clements JE, Murphey-Corb M, Montelaro RC, Robinson JE. Characterization of neutralization epitopes of simian immunodeficiency virus (SIV) recognized by rhesus monoclonal antibodies derived from monkeys infected with an attenuated SIV strain. Virology. 2001;290(1):59–73. doi: 10.1006/viro.2001.1144. [DOI] [PubMed] [Google Scholar]
- Cole KS, Steckbeck JD, Rowles JL, Desrosiers RC, Montelaro RC. Removal of N-linked glycosylation sites in the V1 region of simian immunodeficiency virus gp120 results in redirection of B-cell responses to V3. J Virol. 2004;78(3):1525–1539. doi: 10.1128/JVI.78.3.1525-1539.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Jong JJ, De Ronde A, Keulen W, Tersmette M, Goudsmit J. Minimal requirements for the human immunodeficiency virus type 1 V3 domain to support the syncytium-inducing phenotype: analysis by single amino acid substitution. J Virol. 1992;66(11):6777–6780. doi: 10.1128/jvi.66.11.6777-6780.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fouchier RA, Groenink M, Kootstra NA, Tersmette M, Huisman HG, Miedema F, Schuitemaker H. Phenotype-associated sequence variation in the third variable domain of the human immunodeficiency virus type 1 gp120 molecule. J Virol. 1992;66(5):3183–3187. doi: 10.1128/jvi.66.5.3183-3187.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haigwood NL, Misher L, Chin SM, Blair M, Planelles V, Scandella CJ, Steimer KS, Gardner MB, Yilma T, Hirsch VM, et al. Characterization of group specific antibodies in primates: studies with SIV envelope in macaques. J Med Primatol. 1992;21(2–3):82–90. [PubMed] [Google Scholar]
- Haigwood NL, Watson A, Sutton WF, McClure J, Lewis A, Ranchalis J, Travis B, Voss G, Letvin NL, Hu SL, Hirsch VM, Johnson PR. Passive immune globulin therapy in the SIV/macaque model: early intervention can alter disease profile. Immunol Lett. 1996;51(1–2):107–114. doi: 10.1016/0165-2478(96)02563-1. [DOI] [PubMed] [Google Scholar]
- Hartley O, Klasse PJ, Sattentau QJ, Moore JP. V3: HIV's switch-hitter. AIDS Res Hum Retroviruses. 2005;21(2):171–189. doi: 10.1089/aid.2005.21.171. [DOI] [PubMed] [Google Scholar]
- Huang CC, Tang M, Zhang MY, Majeed S, Montabana E, Stanfield RL, Dimitrov DS, Korber B, Sodroski J, Wilson IA, Wyatt R, Kwong PD. Structure of a V3-containing HIV-1 gp120 core. Science. 2005;310(5750):1025–1028. doi: 10.1126/science.1118398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Javaherian K, Langlois AJ, Montefiori DC, Kent KA, Ryan KA, Wyman PD, Stott J, Bolognesi DP, Murphey-Corb M, Larosa GJ. Studies of the conformation-dependent neutralizing epitopes of simian immunodeficiency virus envelope protein. J Virol. 1994;68(4):2624–2631. doi: 10.1128/jvi.68.4.2624-2631.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Javaherian K, Langlois AJ, Schmidt S, Kaufmann M, Cates N, Langedijk JP, Meloen RH, Desrosiers RC, Burns DP, Bolognesi DP, et al. The principal neutralization determinant of simian immunodeficiency virus differs from that of human immunodeficiency virus type 1. Proc Natl Acad Sci U S A. 1992;89(4):1418–1422. doi: 10.1073/pnas.89.4.1418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson PR, Hirsch VM. SIV infection of macaques as a model for AIDS pathogenesis. Int Rev Immunol. 1992;8(1):55–63. doi: 10.3109/08830189209056641. [DOI] [PubMed] [Google Scholar]
- Johnson WE, Morgan J, Reitter J, Puffer BA, Czajak S, Doms RW, Desrosiers RC. A replication-competent, neutralization-sensitive variant of simian immunodeficiency virus lacking 100 amino acids of envelope. J Virol. 2002;76(5):2075–2086. doi: 10.1128/jvi.76.5.2075-2086.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson WE, Sanford H, Schwall L, Burton DR, Parren PW, Robinson JE, Desrosiers RC. Assorted mutations in the envelope gene of simian immunodeficiency virus lead to loss of neutralization resistance against antibodies representing a broad spectrum of specificities. J Virol. 2003;77(18):9993–10003. doi: 10.1128/JVI.77.18.9993-10003.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karber G. Beitrag zue kollektiven Behandlung pharmakologischer Reinhenversuche. Arch Exp Pathol Pharmakol. 1931;162:480–487. [Google Scholar]
- Kinsey NE, Anderson MG, Unangst TJ, Joag SV, Narayan O, Zink MC, Clements JE. Antigenic variation of SIV: mutations in V4 alter the neutralization profile. Virology. 1996;221(1):14–21. doi: 10.1006/viro.1996.0348. [DOI] [PubMed] [Google Scholar]
- Kirchhoff F, Mori K, Desrosiers RC. The "V3" domain is a determinant of simian immunodeficiency virus cell tropism. J Virol. 1994;68(6):3682–3692. doi: 10.1128/jvi.68.6.3682-3692.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klasse PJ, Moore JP. Quantitative model of antibody- and soluble CD4-mediated neutralization of primary isolates and T-cell line-adapted strains of human immunodeficiency virus type 1. J Virol. 1996;70(6):3668–3677. doi: 10.1128/jvi.70.6.3668-3677.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis MG, Elkins WR, McCutchan FE, Benveniste RE, Lai CY, Montefiori DC, Burke DS, Eddy GA, Shafferman A. Passively transferred antibodies directed against conserved regions of SIV envelope protect macaques from SIV infection. Vaccine. 1993;11(13):1347–1355. doi: 10.1016/0264-410x(93)90106-8. [DOI] [PubMed] [Google Scholar]
- Luciw PA. Human Immunodeficiency Viruses and Their Replication. In: Fields BN, Knipe DM, Howley PM, Chanock RM, Melinick JL, Monath TP, Roizman B, Straus SE, editors. Fields Virology. Philadelphia: Lippincott - Raven Publishers; 1996. p. 1881. [Google Scholar]
- Maecker HT, Maino VC. T cell immunity to HIV: defining parameters of protection. Curr HIV Res. 2003;1(2):249–259. doi: 10.2174/1570162033485294. [DOI] [PubMed] [Google Scholar]
- Mankowski JL, Flaherty MT, Spelman JP, Hauer DA, Didier PJ, Amedee AM, Murphey-Corb M, Kirstein LM, Munoz A, Clements JE, Zink MC. Pathogenesis of simian immunodeficiency virus encephalitis: viral determinants of neurovirulence. J Virol. 1997;71(8):6055–6060. doi: 10.1128/jvi.71.8.6055-6060.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mascola JR, Lewis MG, Stiegler G, Harris D, VanCott TC, Hayes D, Louder MK, Brown CR, Sapan CV, Frankel SS, Lu Y, Robb ML, Katinger H, Birx DL. Protection of Macaques against pathogenic simian/human immunodeficiency virus 89.6PD by passive transfer of neutralizing antibodies. J Virol. 1999;73(5):4009–4018. doi: 10.1128/jvi.73.5.4009-4018.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mascola JR, Lewis MG, VanCott TC, Stiegler G, Katinger H, Seaman M, Beaudry K, Barouch DH, Korioth-Schmitz B, Krivulka G, Sambor A, Welcher B, Douek DC, Montefiori DC, Shiver JW, Poignard P, Burton DR, Letvin NL. Cellular immunity elicited by human immunodeficiency virus type 1/ simian immunodeficiency virus DNA vaccination does not augment the sterile protection afforded by passive infusion of neutralizing antibodies. J Virol. 2003;77(19):10348–10356. doi: 10.1128/JVI.77.19.10348-10356.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mascola JR, Stiegler G, VanCott TC, Katinger H, Carpenter CB, Hanson CE, Beary H, Hayes D, Frankel SS, Birx DL, Lewis MG. Protection of macaques against vaginal transmission of a pathogenic HIV-1/SIV chimeric virus by passive infusion of neutralizing antibodies. Nat Med. 2000;6(2):207–210. doi: 10.1038/72318. [DOI] [PubMed] [Google Scholar]
- McBride BW, Corthals G, Rud E, Kent K, Webster S, Cook N, Cranage MP. Comparison of serum antibody reactivities to a conformational and to linear antigenic sites in the external envelope glycoprotein of simian immunodeficiency virus (SIVmac) induced by infection and vaccination. J Gen Virol. 1993;74(Pt 6):1033–1041. doi: 10.1099/0022-1317-74-6-1033. [DOI] [PubMed] [Google Scholar]
- Means RE, Matthews T, Hoxie JA, Malim MH, Kodama T, Desrosiers RC. Ability of the V3 loop of simian immunodeficiency virus to serve as a target for antibody-mediated neutralization: correlation of neutralization sensitivity, growth in macrophages, and decreased dependence on CD4. J Virol. 2001;75(8):3903–3915. doi: 10.1128/JVI.75.8.3903-3915.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller MA, Murphey-Corb M, Montelaro RC. Identification of broadly reactive continuous antigenic determinants of simian immunodeficiency virus glycoproteins. AIDS Res Hum Retroviruses. 1992;8(6):1153–1164. doi: 10.1089/aid.1992.8.1153. [DOI] [PubMed] [Google Scholar]
- Mori K, Ringler DJ, Kodama T, Desrosiers RC. Complex determinants of macrophage tropism in env of simian immunodeficiency virus. J Virol. 1992;66(4):2067–2075. doi: 10.1128/jvi.66.4.2067-2075.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishimura Y, Igarashi T, Haigwood N, Sadjadpour R, Plishka RJ, Buckler-White A, Shibata R, Martin MA. Determination of a statistically valid neutralization titer in plasma that confers protection against simian-human immunodeficiency virus challenge following passive transfer of high-titered neutralizing antibodies. J Virol. 2002;76(5):2123–2130. doi: 10.1128/jvi.76.5.2123-2130.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishimura Y, Igarashi T, Haigwood NL, Sadjadpour R, Donau OK, Buckler C, Plishka RJ, Buckler-White A, Martin MA. Transfer of neutralizing IgG to macaques 6 h but not 24 h after SHIV infection confers sterilizing protection: implications for HIV-1 vaccine development. Proc Natl Acad Sci U S A. 2003;100(25):15131–15136. doi: 10.1073/pnas.2436476100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Overholser ED, Babas T, Zink MC, Barber SA, Clements JE. CD4-independent entry and replication of simian immunodeficiency virus in primary rhesus macaque astrocytes are regulated by the transmembrane protein. J Virol. 2005;79(8):4944–4951. doi: 10.1128/JVI.79.8.4944-4951.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paiardini M, Frank I, Pandrea I, Apetrei C, Silvestri G. Mucosal immune dysfunction in AIDS pathogenesis. AIDS Rev. 2008;10(1):36–46. [PubMed] [Google Scholar]
- Palker TJ, Muir AJ, Spragion DE, Staats HF, Langlois A, Montefiori DC. The V3 domain of SIVmac251 gp120 contains a linear neutralizing epitope. Virology. 1996;224(2):415–426. doi: 10.1006/viro.1996.0548. [DOI] [PubMed] [Google Scholar]
- Pantophlet R, Burton DR. GP120: target for neutralizing HIV-1 antibodies. Annu Rev Immunol. 2006;24:739–769. doi: 10.1146/annurev.immunol.24.021605.090557. [DOI] [PubMed] [Google Scholar]
- Parren PW, Marx PA, Hessell AJ, Luckay A, Harouse J, Cheng-Mayer C, Moore JP, Burton DR. Antibody protects macaques against vaginal challenge with a pathogenic R5 simian/human immunodeficiency virus at serum levels giving complete neutralization in vitro. J Virol. 2001;75(17):8340–8347. doi: 10.1128/JVI.75.17.8340-8347.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pohlmann S, Davis C, Meister S, Leslie GJ, Otto C, Reeves JD, Puffer BA, Papkalla A, Krumbiegel M, Marzi A, Lorenz S, Munch J, Doms RW, Kirchhoff F. Amino acid 324 in the simian immunodeficiency virus SIVmac V3 loop can confer CD4 independence and modulate the interaction with CCR5 and alternative coreceptors. J Virol. 2004;78(7):3223–3232. doi: 10.1128/JVI.78.7.3223-3232.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poli GF, Fauci AS. Isolation and quanitation of HIV in peripheral blood. In: Coligan JE, Kruisbeek AM, Margulies DH, Shevack EM, S W, editors. Current protocols in immunology. Vol. 3. New York, N.Y: John Wiley & Sons, Inc; 2004. pp. 12.2.1–12.2.11. (ed.) [Google Scholar]
- Puffer BA, Altamura LA, Pierson TC, Doms RW. Determinants within gp120 and gp41 contribute to CD4 independence of SIV Envs. Virology. 2004;327(1):16–25. doi: 10.1016/j.virol.2004.03.016. [DOI] [PubMed] [Google Scholar]
- Puffer BA, Pohlmann S, Edinger AL, Carlin D, Sanchez MD, Reitter J, Watry DD, Fox HS, Desrosiers RC, Doms RW. CD4 independence of simian immunodeficiency virus Envs is associated with macrophage tropism, neutralization sensitivity, and attenuated pathogenicity. J Virol. 2002;76(6):2595–2605. doi: 10.1128/JVI.76.6.2595-2605.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Regier DA, Desrosiers RC. The complete nucleotide sequence of a pathogenic molecular clone of simian immunodeficiency virus. AIDS Res Hum Retroviruses. 1990;6(11):1221–1231. doi: 10.1089/aid.1990.6.1221. [DOI] [PubMed] [Google Scholar]
- Reitter JN, Desrosiers RC. Identification of replication-competent strains of simian immunodeficiency virus lacking multiple attachment sites for N-linked carbohydrates in variable regions 1 and 2 of the surface envelope protein. J Virol. 1998;72(7):5399–5407. doi: 10.1128/jvi.72.7.5399-5407.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reitter JN, Means RE, Desrosiers RC. A role for carbohydrates in immune evasion in AIDS. Nat Med. 1998;4(6):679–684. doi: 10.1038/nm0698-679. [DOI] [PubMed] [Google Scholar]
- Resch W, Hoffman N, Swanstrom R. Improved success of phenotype prediction of the human immunodeficiency virus type 1 from envelope variable loop 3 sequence using neural networks. Virology. 2001;288(1):51–62. doi: 10.1006/viro.2001.1087. [DOI] [PubMed] [Google Scholar]
- Rosen O, Chill J, Sharon M, Kessler N, Mester B, Zolla-Pazner S, Anglister J. Induced fit in HIV-neutralizing antibody complexes: evidence for alternative conformations of the gp120 V3 loop and the molecular basis for broad neutralization. Biochemistry. 2005;44(19):7250–7258. doi: 10.1021/bi047387t. [DOI] [PubMed] [Google Scholar]
- Rosen O, Samson AO, Anglister J. Correlated mutations at gp120 positions 322 and 440: implications for gp120 structure. Proteins. 2008;71(3):1066–1070. doi: 10.1002/prot.21982. [DOI] [PubMed] [Google Scholar]
- Rosen O, Sharon M, Quadt-Akabayov SR, Anglister J. Molecular switch for alternative conformations of the HIV-1 V3 region: implications for phenotype conversion. Proc Natl Acad Sci U S A. 2006;103(38):13950–13955. doi: 10.1073/pnas.0606312103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rudensey LM, Kimata JT, Long EM, Chackerian B, Overbaugh J. Changes in the extracellular envelope glycoprotein of variants that evolve during the course of simian immunodeficiency virus SIVMne infection affect neutralizing antibody recognition, syncytium formation, and macrophage tropism but not replication, cytopathicity, or CCR-5 coreceptor recognition. J Virol. 1998;72(1):209–217. doi: 10.1128/jvi.72.1.209-217.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato S, Johnson W. Antibody-mediated neutralization and simian immunodeficiency virus models of HIV/AIDS. Curr HIV Res. 2007;5(6):594–607. doi: 10.2174/157016207782418515. [DOI] [PubMed] [Google Scholar]
- Sharma DP, Zink MC, Anderson M, Adams R, Clements JE, Joag SV, Narayan O. Derivation of neurotropic simian immunodeficiency virus from exclusively lymphocytetropic parental virus: pathogenesis of infection in macaques. J Virol. 1992;66(6):3550–3556. doi: 10.1128/jvi.66.6.3550-3556.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steckbeck JD, Orlov I, Chow A, Grieser H, Miller K, Bruno J, Robinson JE, Montelaro RC, Cole KS. Kinetic rates of antibody binding correlate with neutralization sensitivity of variant simian immunodeficiency virus strains. J Virol. 2005;79(19):12311–12320. doi: 10.1128/JVI.79.19.12311-12320.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Rompay KK, Berardi CJ, Dillard-Telm S, Tarara RP, Canfield DR, Valverde CR, Montefiori DC, Cole KS, Montelaro RC, Miller CJ, Marthas ML. Passive immunization of newborn rhesus macaques prevents oral simian immunodeficiency virus infection. J Infect Dis. 1998;177(5):1247–1259. doi: 10.1086/515270. [DOI] [PubMed] [Google Scholar]
- Wei X, Decker JM, Liu H, Zhang Z, Arani RB, Kilby JM, Saag MS, Wu X, Shaw GM, Kappes JC. Emergence of resistant human immunodeficiency virus type 1 in patients receiving fusion inhibitor (T-20) monotherapy. Antimicrob Agents Chemother. 2002;46(6):1896–1905. doi: 10.1128/AAC.46.6.1896-1905.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wyatt R, Sodroski J. The HIV-1 envelope glycoproteins: fusogens, antigens, and immunogens. Science. 1998;280(5371):1884–1888. doi: 10.1126/science.280.5371.1884. [DOI] [PubMed] [Google Scholar]
- Yuste E, Johnson W, Pavlakis GN, Desrosiers RC. Virion envelope content, infectivity, and neutralization sensitivity of simian immunodeficiency virus. J Virol. 2005;79(19):12455–12463. doi: 10.1128/JVI.79.19.12455-12463.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]