Abstract
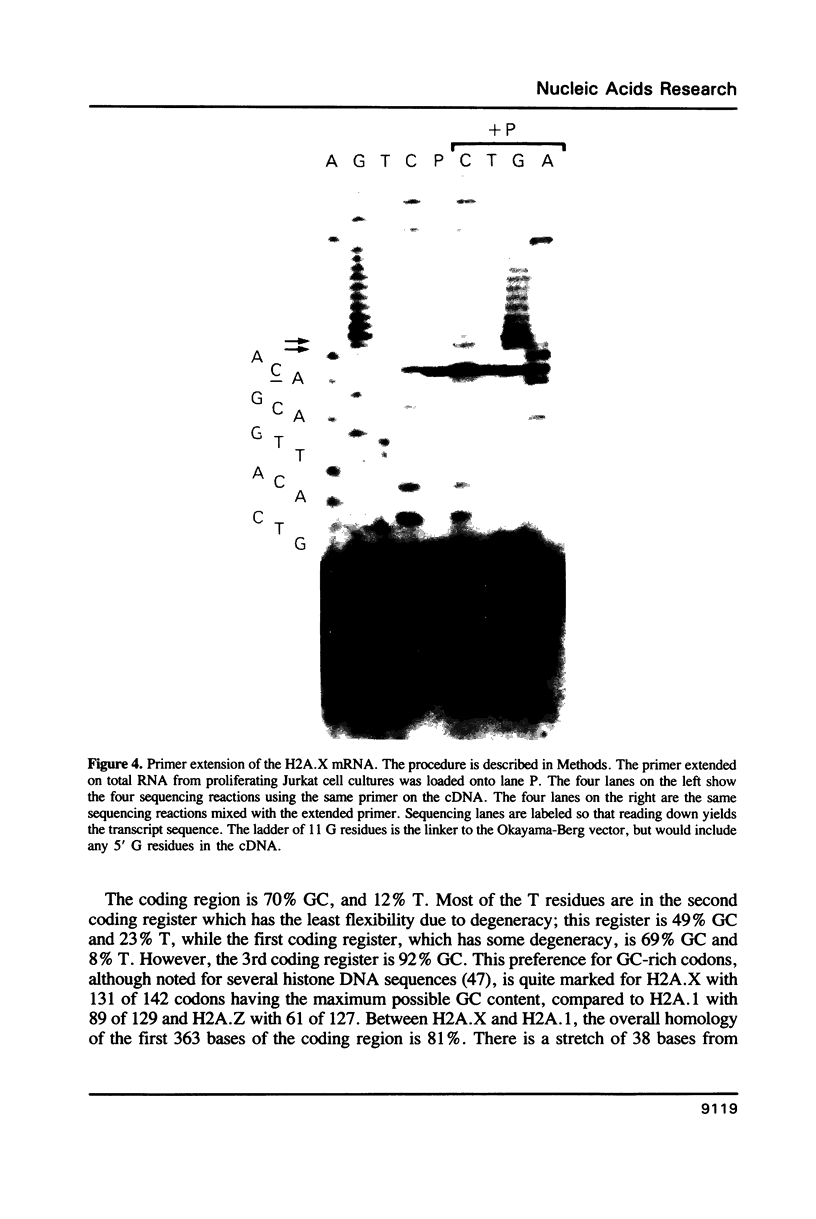
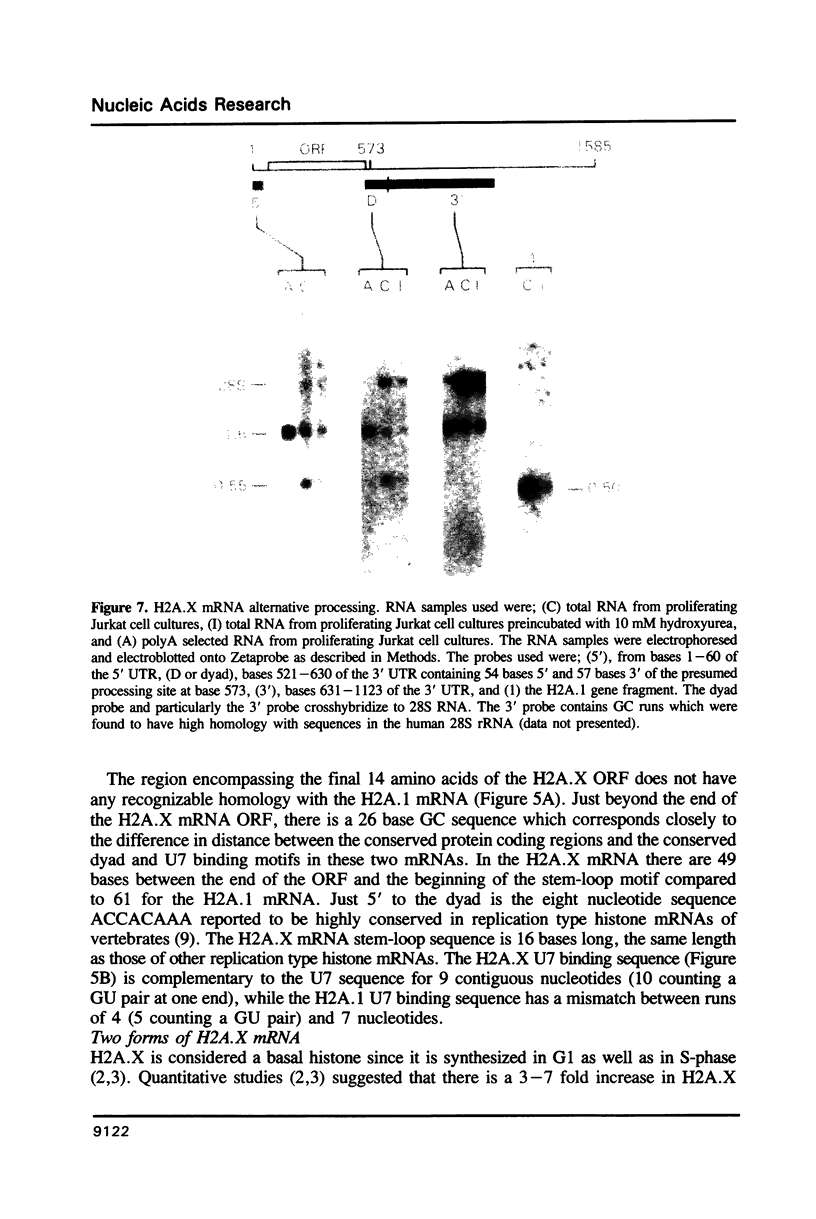
A full length cDNA clone that directs the in vitro synthesis of human histone H2A isoprotein H2A.X has been isolated and sequenced. H2A.X contains 142 amino acid residues, 13 more than human H2A.1. The sequence of the first 120 residues of H2A.X is almost identical to that of human H2A.1. The sequence of the carboxy-terminal 22 residues of H2A.X is unrelated to any known sequence in vertebrate histone H2A; however, it contains a sequence homologous with those of several lower eukaryotes. This homology centers on the carboxy-terminal tetrapeptide which in H2A.X is SerGlnGluTyr. Homologous sequences are found in H2As of three types of yeasts, in Tetrahymena and Drosophila. Seven of the nine carboxy-terminal amino acids of H2A.X are identical with those of S. cerevisiae H2A.1. It is suggested that this H2A carboxy-terminal motif may be present in all eukaryotes. The H2A.X cDNA is 1585 bases long followed by a polyA tail. There are 73 nucleotides in the 5' UTR, 432 in the coding region, and 1080 in the 3' UTR. Even though H2A.X is considered a basal histone, being synthesized in G1 as well as in S-phase, and its mRNA contains polyA addition motifs and a polyA tail, its mRNA also contains the conserved stem-loop and U7 binding sequences involved in the processing and stability of replication type histone mRNAs. Two forms of H2A.X mRNA, consistent with the two sets of processing signals were found in proliferating cell cultures. One, about 1600 bases long, contains polyA; the other, about 575 bases long, lacks polyA. The short form behaves as a replication type histone mRNA, decreasing in amount when cell cultures are incubated with inhibitors of DNA synthesis, while the longer behaves as a basal type histone mRNA.
Full text
PDF











Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ballantine J. E., Woodland H. R. Polyadenylation of histone mRNA in Xenopus oocytes and embryos. FEBS Lett. 1985 Jan 28;180(2):224–228. doi: 10.1016/0014-5793(85)81075-9. [DOI] [PubMed] [Google Scholar]
- Barnes W. M., Bevan M., Son P. H. Kilo-sequencing: creation of an ordered nest of asymmetric deletions across a large target sequence carried on phage M13. Methods Enzymol. 1983;101:98–122. doi: 10.1016/0076-6879(83)01008-3. [DOI] [PubMed] [Google Scholar]
- Bird A. P. CpG-rich islands and the function of DNA methylation. Nature. 1986 May 15;321(6067):209–213. doi: 10.1038/321209a0. [DOI] [PubMed] [Google Scholar]
- Bird R. C., Jacobs F. A., Stein G., Stein J., Sells B. H. A unique subspecies of histone H4 mRNA from rat myoblasts contains poly(A). Proc Natl Acad Sci U S A. 1985 Oct;82(20):6760–6764. doi: 10.1073/pnas.82.20.6760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnstiel M. L., Busslinger M., Strub K. Transcription termination and 3' processing: the end is in site! Cell. 1985 Jun;41(2):349–359. doi: 10.1016/s0092-8674(85)80007-6. [DOI] [PubMed] [Google Scholar]
- Bonner W. M., West M. H., Stedman J. D. Two-dimensional gel analysis of histones in acid extracts of nuclei, cells, and tissues. Eur J Biochem. 1980 Aug;109(1):17–23. doi: 10.1111/j.1432-1033.1980.tb04762.x. [DOI] [PubMed] [Google Scholar]
- Busslinger M., Portmann R., Birnsteil M. L. A regulatory sequence near the 3' end of sea urchin histone genes. Nucleic Acids Res. 1979 Jul 11;6(9):2997–3008. doi: 10.1093/nar/6.9.2997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Challoner P. B., Moss S. B., Groudine M. Expression of replication-dependent histone genes in avian spermatids involves an alternate pathway of mRNA 3'-end formation. Mol Cell Biol. 1989 Mar;9(3):902–913. doi: 10.1128/mcb.9.3.902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers A., Old R. RNA 3' cleavage and polyadenylation in oocytes and unfertilized eggs of Xenopus laevis. Dev Biol. 1988 Feb;125(2):237–245. doi: 10.1016/0012-1606(88)90207-2. [DOI] [PubMed] [Google Scholar]
- Choe J., Kolodrubetz D., Grunstein M. The two yeast histone H2A genes encode similar protein subtypes. Proc Natl Acad Sci U S A. 1982 Mar;79(5):1484–1487. doi: 10.1073/pnas.79.5.1484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choe J., Schuster T., Grunstein M. Organization, primary structure, and evolution of histone H2A and H2B genes of the fission yeast Schizosaccharomyces pombe. Mol Cell Biol. 1985 Nov;5(11):3261–3269. doi: 10.1128/mcb.5.11.3261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Connor W., Mezquita J., Winkfein R. J., States J. C., Dixon G. H. Organization of the histone genes in the rainbow trout (Salmo gairdnerii). J Mol Evol. 1984;20(3-4):227–235. doi: 10.1007/BF02104729. [DOI] [PubMed] [Google Scholar]
- Davie J. R., Delcuve G. P., Nickel B. E., Moirier R., Bailey G. Reduced levels of histones H1o and H1b, and unaltered content of methylated DNA in rainbow trout hepatocellular carcinoma chromatin. Cancer Res. 1987 Oct 15;47(20):5407–5410. [PubMed] [Google Scholar]
- DeLisle A. J., Graves R. A., Marzluff W. F., Johnson L. F. Regulation of histone mRNA production and stability in serum-stimulated mouse 3T6 fibroblasts. Mol Cell Biol. 1983 Nov;3(11):1920–1929. doi: 10.1128/mcb.3.11.1920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ernst S. G., Miller H., Brenner C. A., Nocente-McGrath C., Francis S., McIsaac R. Characterization of a cDNA clone coding for a sea urchin histone H2A variant related to the H2A.F/Z histone protein in vertebrates. Nucleic Acids Res. 1987 Jun 11;15(11):4629–4644. doi: 10.1093/nar/15.11.4629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fusauchi Y., Iwai K. Tetrahymena histone H2A. Isolation and two variant sequences. J Biochem. 1983 Jun;93(6):1487–1497. doi: 10.1093/oxfordjournals.jbchem.a134286. [DOI] [PubMed] [Google Scholar]
- Gardiner-Garden M., Frommer M. CpG islands in vertebrate genomes. J Mol Biol. 1987 Jul 20;196(2):261–282. doi: 10.1016/0022-2836(87)90689-9. [DOI] [PubMed] [Google Scholar]
- Gick O., Krämer A., Vasserot A., Birnstiel M. L. Heat-labile regulatory factor is required for 3' processing of histone precursor mRNAs. Proc Natl Acad Sci U S A. 1987 Dec;84(24):8937–8940. doi: 10.1073/pnas.84.24.8937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldknopf I. L., Busch H. Isopeptide linkage between nonhistone and histone 2A polypeptides of chromosomal conjugate-protein A24. Proc Natl Acad Sci U S A. 1977 Mar;74(3):864–868. doi: 10.1073/pnas.74.3.864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harvey R. P., Whiting J. A., Coles L. S., Krieg P. A., Wells J. R. H2A.F: an extremely variant histone H2A sequence expressed in the chicken embryo. Proc Natl Acad Sci U S A. 1983 May;80(10):2819–2823. doi: 10.1073/pnas.80.10.2819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hatch C. L., Bonner W. M. Sequence of cDNAs for mammalian H2A.Z, an evolutionarily diverged but highly conserved basal histone H2A isoprotein species. Nucleic Acids Res. 1988 Feb 11;16(3):1113–1124. doi: 10.1093/nar/16.3.1113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayashi T., Ohe Y., Hayashi H., Iwai K. Human spleen histone H2A. Isolation and four variant sequences. J Biochem. 1980 Jul;88(1):27–34. [PubMed] [Google Scholar]
- Heintz N., Sive H. L., Roeder R. G. Regulation of human histone gene expression: kinetics of accumulation and changes in the rate of synthesis and in the half-lives of individual histone mRNAs during the HeLa cell cycle. Mol Cell Biol. 1983 Apr;3(4):539–550. doi: 10.1128/mcb.3.4.539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hentschel C. C., Birnstiel M. L. The organization and expression of histone gene families. Cell. 1981 Aug;25(2):301–313. doi: 10.1016/0092-8674(81)90048-9. [DOI] [PubMed] [Google Scholar]
- Huang S. Y., Barnard M. B., Xu M., Matsui S., Rose S. M., Garrard W. T. The active immunoglobulin kappa chain gene is packaged by non-ubiquitin-conjugated nucleosomes. Proc Natl Acad Sci U S A. 1986 Jun;83(11):3738–3742. doi: 10.1073/pnas.83.11.3738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. An analysis of 5'-noncoding sequences from 699 vertebrate messenger RNAs. Nucleic Acids Res. 1987 Oct 26;15(20):8125–8148. doi: 10.1093/nar/15.20.8125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marzluff W. F., Pandey N. B. Multiple regulatory steps control histone mRNA concentrations. Trends Biochem Sci. 1988 Feb;13(2):49–52. doi: 10.1016/0968-0004(88)90027-8. [DOI] [PubMed] [Google Scholar]
- Matsumoto S., Yanagida M. Histone gene organization of fission yeast: a common upstream sequence. EMBO J. 1985 Dec 16;4(13A):3531–3538. doi: 10.1002/j.1460-2075.1985.tb04113.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- May G. S., Morris N. R. The unique histone H2A gene of Aspergillus nidulans contains three introns. Gene. 1987;58(1):59–66. doi: 10.1016/0378-1119(87)90029-1. [DOI] [PubMed] [Google Scholar]
- Mizusawa S., Nishimura S., Seela F. Improvement of the dideoxy chain termination method of DNA sequencing by use of deoxy-7-deazaguanosine triphosphate in place of dGTP. Nucleic Acids Res. 1986 Feb 11;14(3):1319–1324. doi: 10.1093/nar/14.3.1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss S. B., Challoner P. B., Groudine M. Expression of a novel histone 2B during mouse spermiogenesis. Dev Biol. 1989 May;133(1):83–92. doi: 10.1016/0012-1606(89)90299-6. [DOI] [PubMed] [Google Scholar]
- Moss T., Cary P. D., Abercrombie B. D., Crane-Robinson C., Bradbury E. M. A pH-dependent interaction between histones H2A and H2B involving secondary and tertiary folding. Eur J Biochem. 1976 Dec 11;71(2):337–350. doi: 10.1111/j.1432-1033.1976.tb11120.x. [DOI] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. Identification of the human U7 snRNP as one of several factors involved in the 3' end maturation of histone premessenger RNA's. Science. 1987 Dec 18;238(4834):1682–1687. doi: 10.1126/science.2825355. [DOI] [PubMed] [Google Scholar]
- Mueller R. D., Yasuda H., Hatch C. L., Bonner W. M., Bradbury E. M. Identification of ubiquitinated histones 2A and 2B in Physarum polycephalum. Disappearance of these proteins at metaphase and reappearance at anaphase. J Biol Chem. 1985 Apr 25;260(8):5147–5153. [PubMed] [Google Scholar]
- Nickel B. E., Davie J. R. Structure of polyubiquitinated histone H2A. Biochemistry. 1989 Feb 7;28(3):964–968. doi: 10.1021/bi00429a007. [DOI] [PubMed] [Google Scholar]
- Okayama H., Berg P. A cDNA cloning vector that permits expression of cDNA inserts in mammalian cells. Mol Cell Biol. 1983 Feb;3(2):280–289. doi: 10.1128/mcb.3.2.280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandey N. B., Marzluff W. F. The stem-loop structure at the 3' end of histone mRNA is necessary and sufficient for regulation of histone mRNA stability. Mol Cell Biol. 1987 Dec;7(12):4557–4559. doi: 10.1128/mcb.7.12.4557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rodrigues J. de A., Brandt W. F., von Holt C. The amino acid sequence of wheat histone H2A(1). A core histone with a C-terminal extension. Eur J Biochem. 1985 Aug 1;150(3):499–505. doi: 10.1111/j.1432-1033.1985.tb09050.x. [DOI] [PubMed] [Google Scholar]
- Ross J., Kobs G. H4 histone messenger RNA decay in cell-free extracts initiates at or near the 3' terminus and proceeds 3' to 5'. J Mol Biol. 1986 Apr 20;188(4):579–593. doi: 10.1016/s0022-2836(86)80008-0. [DOI] [PubMed] [Google Scholar]
- Ross J., Peltz S. W., Kobs G., Brewer G. Histone mRNA degradation in vivo: the first detectable step occurs at or near the 3' terminus. Mol Cell Biol. 1986 Dec;6(12):4362–4371. doi: 10.1128/mcb.6.12.4362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salser W. Globin mRNA sequences: analysis of base pairing and evolutionary implications. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 2):985–1002. doi: 10.1101/sqb.1978.042.01.099. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schümperli D. Multilevel regulation of replication-dependent histone genes. Trends Genet. 1988 Jul;4(7):187–191. doi: 10.1016/0168-9525(88)90074-1. [DOI] [PubMed] [Google Scholar]
- Seiler-Tuyns A., Paterson B. M. A chimeric mouse histone H4 gene containing either an intron or poly(A) addition signal behaves like a basal histone. Nucleic Acids Res. 1986 Nov 25;14(22):8845–8862. doi: 10.1093/nar/14.22.8845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sittman D. B., Graves R. A., Marzluff W. F. Histone mRNA concentrations are regulated at the level of transcription and mRNA degradation. Proc Natl Acad Sci U S A. 1983 Apr;80(7):1849–1853. doi: 10.1073/pnas.80.7.1849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sittman D. B., Graves R. A., Marzluff W. F. Structure of a cluster of mouse histone genes. Nucleic Acids Res. 1983 Oct 11;11(19):6679–6697. doi: 10.1093/nar/11.19.6679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stauber C., Lüscher B., Eckner R., Lötscher E., Schümperli D. A signal regulating mouse histone H4 mRNA levels in a mammalian cell cycle mutant and sequences controlling RNA 3' processing are both contained within the same 80-bp fragment. EMBO J. 1986 Dec 1;5(12):3297–3303. doi: 10.1002/j.1460-2075.1986.tb04643.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sures I., Lowry J., Kedes L. H. The DNA sequence of sea urchin (S. purpuratus) H2A, H2B and H3 histone coding and spacer regions. Cell. 1978 Nov;15(3):1033–1044. doi: 10.1016/0092-8674(78)90287-8. [DOI] [PubMed] [Google Scholar]
- Wells D., Kedes L. Structure of a human histone cDNA: evidence that basally expressed histone genes have intervening sequences and encode polyadenylylated mRNAs. Proc Natl Acad Sci U S A. 1985 May;82(9):2834–2838. doi: 10.1073/pnas.82.9.2834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wells D., McBride C. A comprehensive compilation and alignment of histones and histone genes. Nucleic Acids Res. 1989;17 (Suppl):r311–r346. doi: 10.1093/nar/17.suppl.r311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- West M. H., Bonner W. M. Histone 2A, a heteromorphous family of eight protein species. Biochemistry. 1980 Jul 8;19(14):3238–3245. doi: 10.1021/bi00555a022. [DOI] [PubMed] [Google Scholar]
- West M. H., Bonner W. M. Structural comparisons of mouse histones 2A.X and 2A.Z with 2A.1 and 2A.2. Comp Biochem Physiol B. 1983;76(3):455–464. doi: 10.1016/0305-0491(83)90275-4. [DOI] [PubMed] [Google Scholar]
- White E. M., Shapiro D. L., Allis C. D., Gorovsky M. A. Sequence and properties of the message encoding Tetrahymena hv1, a highly evolutionarily conserved histone H2A variant that is associated with active genes. Nucleic Acids Res. 1988 Jan 11;16(1):179–198. doi: 10.1093/nar/16.1.179. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu R. S., Nishioka D., Bonner W. M. Differential conservation of histone 2A variants between mammals and sea urchins. J Cell Biol. 1982 May;93(2):426–431. doi: 10.1083/jcb.93.2.426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu R. S., Panusz H. T., Hatch C. L., Bonner W. M. Histones and their modifications. CRC Crit Rev Biochem. 1986;20(2):201–263. doi: 10.3109/10409238609083735. [DOI] [PubMed] [Google Scholar]
- Wu R. S., Tsai S., Bonner W. M. Patterns of histone variant synthesis can distinguish G0 from G1 cells. Cell. 1982 Dec;31(2 Pt 1):367–374. doi: 10.1016/0092-8674(82)90130-1. [DOI] [PubMed] [Google Scholar]
- Zhong R., Roeder R. G., Heintz N. The primary structure and expression of four cloned human histone genes. Nucleic Acids Res. 1983 Nov 11;11(21):7409–7425. doi: 10.1093/nar/11.21.7409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Andrade Rodrigues J., Brandt W. F., Von Holt C. The primary structure of the histone H2A(2) type from wheat germ. A core histone type with both, N-terminal and C-terminal extensions. Eur J Biochem. 1988 May 2;173(3):555–560. doi: 10.1111/j.1432-1033.1988.tb14035.x. [DOI] [PubMed] [Google Scholar]
- van Daal A., White E. M., Gorovsky M. A., Elgin S. C. Drosophila has a single copy of the gene encoding a highly conserved histone H2A variant of the H2A.F/Z type. Nucleic Acids Res. 1988 Aug 11;16(15):7487–7497. doi: 10.1093/nar/16.15.7487. [DOI] [PMC free article] [PubMed] [Google Scholar]