Abstract
Arabidopsis cryptochrome 2 (CRY2) is a blue-light receptor mediating blue-light inhibition of hypocotyl elongation and photoperiodic promotion of floral initiation. CRY2 is a constitutive nuclear protein that undergoes blue-light-dependent phosphorylation, ubiquitination, photobody formation, and degradation in the nucleus, but the relationship between these blue-light-dependent events remains unclear. It has been proposed that CRY2 phosphorylation triggers a conformational change responsible for the subsequent ubiquitination and photobody formation, leading to CRY2 function and/or degradation. We tested this hypothesis by a structure-function study, using mutant CRY2–GFP fusion proteins expressed in transgenic Arabidopsis. We show that changes of lysine residues of the NLS (Nuclear Localization Signal) sequence of CRY2 to arginine residues partially impair the nuclear importation of the CRY2K541R and CRY2K554/5R mutant proteins, resulting in reduced phosphorylation, physiological activities, and degradation in response to blue light. In contrast to the wild-type CRY2 protein that forms photobodies exclusively in the nucleus, the CRY2K541R and CRY2K554/5R mutant proteins form protein bodies in both the nucleus and cytosol in response to blue light. These results suggest that photoexcited CRY2 molecules can aggregate to form photobody-like structure without the nucleus-dependent protein modifications or the association with the nuclear CRY2-interacting proteins. Taken together, the observation that CRY2 forms photobodies markedly faster than CRY2 phosphorylation in response to blue light, we hypothesize that the photoexcited cryptochromes form oligomers, preceding other biochemical changes of CRY2, to facilitate photobody formation, signal amplification, and propagation, as well as desensitization by degradation.
Keywords: protein phosphorylation, signal transduction, fluorescence imaging, protein degradation, photobody
INTRODUCTION
Cryptochromes are blue-light receptors that regulate photomorphogenic development and circadian clock (Cashmore, 2003; Lin and Shalitin, 2003; Sancar, 2003; Muller and Ahmad, 2011). The Arabidopsis genome encodes three cryptochromes: CRY1, CRY2, and CRY3. CRY1 and CRY2 primarily regulate de-etiolation and photoperiodic flowering, respectively (Ahmad and Cashmore, 1993; Guo et al., 1998). Besides, these two blue-light receptors mediate many other regulations (Liu et al., 2011c), such as light entrainment of the circadian clock (Somers et al., 1998), light stimulation of guard cell development and stomatal opening (Mao et al., 2005; Kang et al., 2009; Wang et al., 2010a), responses to bacterial and viral pathogens (Jeong et al., 2010; Wu and Yang, 2010), etc. Arabidopsis CRY3 acts as a single-strand DNA photolyase in chloroplast/mitochondria (Kleine et al., 2003; Selby and Sancar, 2006). Although CRY2 primarily regulates photoperiodic promotion of floral initiation (Guo et al., 1998; El-Din El-Assal et al., 2001), its function partially overlaps with CRY1 in that both CRY1 and CRY2 mediate blue-light inhibition of hypocotyl elongation (Lin et al., 1998; Mockler et al., 1999).
CRY2 contains two domains: the N-terminal PHR (Photolyase Homologous Region) domain responsible for photon absorption and the C-terminal CCE (Cryptochrome C-Terminal Extension) effecter domain. The CCE domain of CRY2 contains a putative bipartite NLS (Nuclear Localization Sequence) (Guo et al., 1999; Kleiner et al., 1999). Like other bipartite NLS sequences, the CRY2 NLS (KRVKPEEEEERDMKKSR) contains two clusters of basic amino acids (Lysine or Arginine) separated by a spacer of about 8–10 amino acids rich in the negatively charged residues (Dingwall et al., 1988; Robbins et al., 1991) (Figure 1A). The NLS structural element of a protein can be recognized by importin α and importin β for nuclear transportation across the nuclear pore complexes (Conti and Kuriyan, 2000; Cansizoglu et al., 2007). In addition to nuclear importation, residues of NLS may also be ubiquitinated to serve other biochemical functions. For example, monoubiquitination of a lysine residue of the NLS of the mammalian P53 protein or the Nipah virus matrix (NiV-M) protein prevents nuclear importation of the respective protein (Marchenko et al., 2009; Wang et al., 2010b). Although the nuclear importation is critical to the function and regulation of CRY2, the function of the putative NLS of CRY2 has not been investigated experimentally.
Figure 1.
The Lysine Residues of the Putative NLS of CRY2 Are Important for the Nuclear Importation of CRY2.
(A) The putative NLS of CRY2. The diagram at the bottom shows the structure of CRY2 protein containing the N-terminal PHR domain and C-terminal CCE domain. The putative NLS of CRY2 (residues from K541 to R557) is aligned with the NLSs of Nucleoplasmin and Nipah virus matrix (NiV-M). Two clusters of basic residues are boxed by blue rectangles. The sequences of the K541R and K554/5R mutants are shown in red.
(B) Immunoblot showing the level of protein expression of the CRY2–GFP and mutant proteins of the indicated transgenic lines. Seven-day-old seedlings were grown in the ½ MS medium in continuous red light; total protein extracts were fractioned by a 10% SDS–PAGE gel, blotted, and probed with the anti-CRY2 (CRY2). The Ponceau S staining of the Rubisco band is included as the loading control.
(C) REU (Relative Expression Unit) is calculated by the formula [CRY2mt/Rubiscomt]/[CRY2wt/Rubiscowt], in which ‘CRY2’ and ‘Rubisco’ designate the digitized band intensity of CRY2 or Rubisco of the respective CRY2 mutants (mt) or the wild-type CRY2–GFP control (wt).
(D) Representative fluorescence images showing the subcellular localization of CRY2–GFP, K541R, and K554/5R in epidermal cells at the top half of the hypocotyls of seedlings grown in dark.
(E) The nuclear/cytoplasmic distribution of the indicated proteins is represented by RIU (Relative Intensity Unit), which is calculated by the formula [nuclear GFP fluorescence intensity]/[cytoplasmic GFP fluorescence intensity], n ≥ 20, bar = 5 μm.
CRY2 completes its posttranslational lifecycle in the nucleus (Yu et al., 2007). Upon photoexcitation, CRY2 is phosphorylated, ubiquitinated, and degraded by the 26S proteasome in the nucleus (Shalitin et al., 2002; Yu et al., 2007; Liu et al., 2011a). Depending on the fluence rate of blue light, CRY2 phosphorylation and degradation occur in the timescale of minutes in etiolated seedlings exposed to blue light, whereas CRY2 degradation requires longer blue-light illumination (Shalitin et al., 2002; Yu et al., 2007). In addition to blue-light-dependent protein modification and degradation, CRY2 also undergoes other light-dependent changes. For example, CRY2 forms nuclear protein bodies in response to blue light (Mas et al., 2000; Yu et al., 2009), which resembles morphologically to the protein bodies of photoexcited phytochromes. The nuclear photoreceptor protein bodies are originally discovered for phytochromes, and they are now referred to as photobodies (Sakamoto and Nagatani, 1996; Kircher et al., 1999; Van Buskirk et al., 2011; Gu et al., 2012). It has been proposed that photobodies are the major site of the function and regulation of photoreceptors. Depending on the fluence rate of blue light, CRY2 forms visually discernable photobodies within the timescale of seconds in etiolated seedlings exposed to blue light, which seems much faster than CRY2 phosphorylation and degradation (Yu et al., 2009). The biochemical and physiological significance of the CRY2 photobodies has been investigated by a comparative study of the endogenous CRY2, GFP–CRY2 (with GFP fused to the N-terminus of CRY2), and CRY2–GFP (with GFP fused to the C-terminus of CRY2) fusion proteins (Yu et al., 2009). The endogenous CRY2 and GFP–CRY2 behave similarly in response to blue light: they undergo blue-light-induced phosphorylation and degradation with the similar kinetics, and they were both physiologically active in a blue-light-specific manner (Yu et al., 2009). It is generally difficult to observe the photobody formation of the endogenous CRY2 and GFP–CRY2, except under conditions that the activity of the 26S proteasome is inhibited, suggesting that the CRY2 photobody is probably associated with ubiquitination and degradation of CRY2. In contrast to the endogenous CRY2 and the GFP–CRY2 fusion protein, the CRY2–GFP fusion protein showed constitutive physiological activity, but much sluggish degradation in response to blue light. Interestingly, the CRY2–GFP photobodies are more readily observed in the absence of the proteasome inhibitor (Yu et al., 2009). Because the endogenous CRY2 and the GFP–CRY2 fusion proteins are rapidly degraded in blue light and their photobodies can only be observed in the presence of the proteasome inhibitor MG132, whereas the CRY2–GFP protein showed more prominent and readily formed photobody and it is degraded slowly, it was hypothesized that CRY2 photobody is associated with the proteins involved in CRY2 degradation. Alternatively, it is also possible that CRY2 form photobodies only when the local concentration of the photoexcited photoreceptor molecules is higher than a certain threshold. These two hypotheses have not been directly tested.
We investigated the structure-function relationship of the putative NLS of CRY2 for its nuclear localization, ubiquitination, degradation, and photobody formation, using the CRY2–GFP fusion proteins mutated at each of the three lysine residues. We found that the lysine residues 541 and 554/555 are critical for CRY2 nuclear localization. Importantly, we found that the cytoplasmic CRY2K541R and CRY2K554/5R mutant proteins, which are apparently not phosphorylated or degraded, formed protein bodies morphologically similar to the CRY2 photobodies. Because the cytosolic protein bodies of the mutant CRY2 fusion proteins are morphologically similar to the nuclear photobody of the endogenous CRY2, and the formation of both cytosolic and nuclear CRY2 protein bodies is strictly dependent on blue light, we hypothesize that photoexcited CRY2 may oligomerize to form photobodies to facilitate the function and regulation of the photoreceptor.
RESULTS AND DISCUSSION
The NLS of CRY2 Is Required for Its Nuclear Localization and Function
To investigate the putative NLS of CRY2 in the function and regulation of CRY2, we prepared CRY2 mutants, CRY2K541R, CRY2K544R, and CRY2K554/5R, by replacing the lysine residues of NLS with the similar amino acid arginine. The residues K541 and K544 or K554 and K555 are located in the first or the second positively charged segment of the putative bipartite NLS of CRY2 (K541RVK544PEEEEERDMK554K555SR), respectively (Figure 1A). Because the CRY2–GFP fusion protein is physiologically active and it forms easily detectable photobodies, we expressed CRY2 mutants in the form of CRY2–GFP fusion proteins in the cry1cry2 mutant background. The mutation of the residue K544 (CRY2K544R) showed little effect on the function and regulation of CRY2 (data not shown), so we focused our study on the other two mutants: CRY2K541R and CRY2K554/5R. For simplicity and clarity, the CRY2–GFP, CRY2K541R–GFP, and CRY2K554/5R–GFP are referred to as CRY2–GFP, CRY2K541R, and CRY2 K554/5R, respectively in the text, or as CRY2, K541R, and K554/5R, respectively, in the figures of this report. We selected the transgenic lines that express the comparable amount of the total protein of CRY2–GFP, CRY2K541R, and CRY2K554/5R by the immunoblot for further investigation (Figure 1B and 1C). Similar phenotypes were observed in multiple transgenic lines expressing the CRY2K541R or CRY2K554/5R mutants, although most of our experiments were performed in the K541R-1 and K554/5R-1 transgenic lines.
We first investigated the subcellular localization of the CRY2K541R and CRY2K554/5R mutant proteins in the epidermal or cortex cells at the top half of the hypocotyls of etiolated seedlings, which were fixed in formaldehyde and observed under a fluorescence microscope. Both the K541R-1 and K554/5R-1 mutations impaired CRY2 nuclear importation. In contrast to CRY2–GFP control that accumulated exclusively in the nucleoplasm, green fluorescence was observed in the cytoplasm for both CRY2K541R and CRY2K554/5R mutants (Figure 1D). CRY2K541R showed a relatively minor cytoplasmic accumulation, whereas CRY2K554/5R exhibited a near uniform distribution in the nucleoplasm and cytoplasm (Figure 1D and Supplemental Figures 1 and 2). The relative fraction of CRY2–GFP and CRY2 mutant proteins accumulated in the nucleus was estimated by the fluorescence intensity of the nuclear GFP normalized by the cytoplasmic fluorescence intensity (Figure 1E). The result of this analysis showed that relative intensity of nuclear CRY2K541R or CRY2K544/5R is approximately 20 or 10% of that of the CRY2–GFP control, respectively (Figure 1E). This result confirms the important function of the bipartite NLS in the nuclear importation of CRY2.
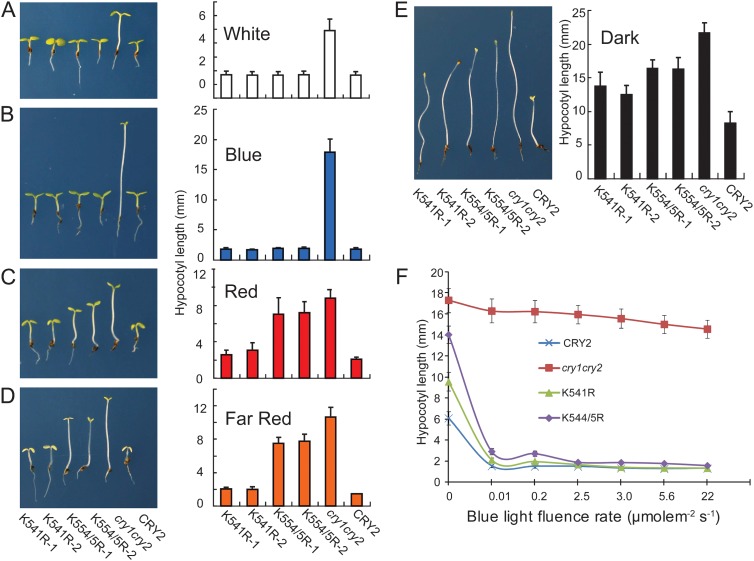
We have previously reported that the CRY2–GFP fusion protein is constitutively active, suppressing hypocotyl elongation, although its activity is also responsive to blue light (Yu et al., 2009). Similarly, the CRY2K541R and CRY2K554/5R mutants showed constitutive activity inhibiting hypocotyl growth (Figure 2). Figure 2 shows that the CRY2K541R and CRY2K554/5R mutants had the highest activity in blue light, suppressing hypocotyl growth, but they apparently also function in the absence of blue light. The CRY2K541R and CRY2K554/5R mutants are still responsive to the increased fluence rates of blue light (Figure 2F), suggesting that, similarly to CRY2–GFP, the CRY2K541R and CRY2K554/5R mutant proteins remain responsive to blue light. The activities of CRY2K541R and CRY2K554/5R mutants are relatively weaker than that of the CRY2–GFP control (Figure 2C–2F). Importantly, there is a clear correlation between the extent of nuclear enrichment and the relative strength of hypocotyl inhibition activity among the three proteins tested. CRY2–GFP accumulated exclusively in the nucleus and it showed strongest activity. Both CRY2K541R and CRY2K554/5R mutant proteins were impaired in both the nuclear importation and hypocotyl inhibition. Among the two mutant proteins, CRY2K541R exhibited better nuclear importation and stronger hypocotyl inhibition activity than CRY2K554/5R (Figure 2). These results are consistent with the notion that CRY2 acts only in the nucleus (Yu et al., 2007).
Figure 2.
The Hypocotyl Inhibition Response of Transgenic Seedlings Expressing the CRY2 and Mutant Fusion Proteins.
(A–E) Images (left) and hypocotyl lengths (right, n ≥ 20) of 7-day-old seedlings of indicated genotypes grown in compound soil under continuous white light (90 μmol m−2 s−1) (A), blue light (15 μmol m−2 s−1) (B), red light (10 μmol m−2 s−1) (C), far-red light (3 μmol m−2 s−1) (D), or in darkness (E).
(F) A fluence-rate response of hypocotyl growth of the indicated genotypes grown under continuous blue light for 5 d in ½ MS medium. Hypocotyl lengths and standard deviations (n ≥ 20) are shown. All transgenic lines are in the cry1cry2 background.
The Nuclear Localization of CRY2 Is Important for Its Light-Dependent Phosphorylation and Degradation
Because both CRY2K541R and CRY2K554/5R mutant proteins are defective in the nuclear importation, we expected that these two mutants might also show defects in the blue-light-dependent phosphorylation and degradation. To test this expectation, we analyzed the kinetics of phosphorylation and degradation of CRY2K541R and CRY2K554/5R in the seedlings grown in continuous red light and transferred to blue light (∼22 μmol m−2 s−1). Figure 3 shows that CRY2K541R was degraded at a markedly slower rate than the CRY2–GFP control, whereas very little degradation was detected for the CRY2K554/5R mutant in seedlings exposed to blue light. In contrast to the CRY2–GFP control that reached 80% degradation within 2 h under the fluence rate of blue light tested, the CRY2K541R mutant protein took more than 3 h to reach the 60% degradation (Figure 3B). Under the same fluence rate of blue light, the CRY2K554/5R mutant was degraded even more slowly—less than 20% of CRY2K554/5R was degraded after 8 h of blue-light illumination (Figure 3D).
Figure 3.
Blue-Light-Dependent Phosphorylation and Degradation of the CRY2 Mutant Proteins.
(A–D) Immunoblots showing the level of the CRY2–GFP (A, C), K541R (A), and K554/5R (C) proteins and semi-quantification of degradation (B, D). Seedlings were grown in dark for 7 d, transferred to blue light (∼22 μmol m−2 s−1) for the indicated time (h), and the total protein extracts were analyzed in a 10% SDS–PAGE for immunoblot analyses, which were probed with anti-CRY2 (CRY2); the vPPase detected by vPPase antibody is included as the loading control. Arrows and arrowheads indicate the unphosphorylated or hyperphosphorylated CRY2 fusion proteins (B, E). The relative expression level of CRY2 represented by ‘REU’ (Relative Expression Unit) is calculated by the formula [CRY2t/vPPase t]/[CRY20/vPPase 0], in which ‘CRY2’ and ‘vPPase’ denote digitized band intensities of CRY2 or vPPase of the respective samples collected at time zero (0) or the indicated time after blue-light exposure (t).
(E, F) The immunoblot showing the level of CRY2–GFP and CRY2 mutant proteins (E) and a semi-quantification of phosphorylation (F). Seven-day-old etiolated seedlings were transferred to blue light (∼22 μmol m−2 s−1) for the indicated time (h), and analyzed as in (A) and (C). The relative level of CRY2 phosphorylation is calculated by the formula [CRY2Pi]/[CRY2+CRY2Pi], in which CRY2Pi and CRY2 denote digitized band intensities of phosphorylated and unphosphorylated CRY2 signals, respectively.
We next analyzed the blue-light-dependent phosphorylation of the mutant proteins. Although it has been previously reported that CRY2–GFP is constitutively phosphorylated (Yu et al., 2009), the result shown in Figure 3 clearly demonstrates that the phosphorylation of CRY2–GFP is dependent on blue light. This discrepancy is most likely caused by different sensitivities of the different methods used in the two studies. The CRY2–GFP phosphorylation was detected by the radioactive 32[P] labeling in our previous study (Yu et al., 2009), or by the band migration shift immunoblot test used in the present study (Figure 3). It is conceivable that a small fraction of the CRY2–GFP phosphorylation sites or a small fraction of the cellular CRY2–GFP is phosphorylated in the absence of blue light to cause constitutive activity of CRY2–GFP, and such a small fraction of phosphorylated CRY2–GFP can be detected by the radioactive labeling method but not by the band shift immunoblot method. Nevertheless, it is clear now that the majority of the phosphorylatable residues or the cellular CRY2–GFP molecules are fully phosphorylated in response to blue light.
In contrast to the highly phosphorylatable CRY2–GFP control, less than 50% of the CRY2K541R mutant protein was phosphorylated or degraded at any time point tested (Figure 3A). Interestingly, only a very low level of phosphorylated band of CRY2K541R was detected after the seedlings were illuminated with blue light for 2 h (Figure 3A). Because only the nuclear CRY2 is phosphorylated and only phosphorylated CRY2 is ubiquitinated and degraded (Shalitin et al., 2002), it is possible that most CRY2K541R protein molecules located in the nucleus were phosphorylated and degraded after seedlings were exposed to blue light for 2 h, whereas the CRY2K541R protein molecules located in the cytoplasm were never phosphorylated or degraded. The CRY2K554/5R mutant protein showed more severe defects in phosphorylation and degradation (Figure 3C and 3D). Only an unphosphorylated band of CRY2K554/5R was detected with blue-light treatment and the unphosphorylated CRY2K554/5R was apparently not degraded at any time point tested (Figure 3C and 3D). The more profound defects in phosphorylation and degradation of CRY2K554/5R in comparison to CRY2K541R are consistent with relatively lower abundance of CRY2K554/5R accumulated in the nucleus (Figure 1D and 1E). In other words, the relatively smaller fraction of CRY2K554/5R imported into the nucleus was phosphorylated (Figure 3C and 3D), physiologically active (Figure 2), and degraded (Figure 3C and 3D), whereas the relatively larger fraction of CRY2K554/5R left in the cytoplasm was not phosphorylated or degraded in response to light (Figure 3C and 3D). To further test this possibility, we compared phosphorylation of CRY2, CRY2K541R, and CRY2K554/5R under an experimental condition that favors the detection of CRY2 phosphorylation over CRY2 degradation (Figure 3E and 3F). In this experiment, etiolated seedlings were exposed to blue light (∼22 μmol m−2 s−1) for a shorter time period, and the immunoblot was overexposed to favor the detection of the relatively minor band of the phosphorylated CRY2K554/5R (Figure 3E). As shown in Figure 3E, the phosphorylated form of CRY2K554/5R was clearly detected under this condition. In contrast to CRY2–GFP that was almost completely phosphorylated within 10–30 min of light treatment, only about 40–50% of CRY2K541R or ∼20% of CRY2K554/5R was phosphorylated (Figure 3F). These results support the hypothesis that phosphorylation of CRY2 is required for its degradation and both biochemical changes of CRY2 occur only in the nucleus.
The Cytoplasmic CRY2 Form Photobody-Like Structure in Response to Blue Light
As reported previously (Yu et al., 2009), CRY2–GFP forms photobodies in a blue-light-dependent manner (Figure 4). Photobodies are defined as nuclear protein bodies formed by photoreceptors and proteins associated with the photoreceptors (Van Buskirk et al., 2011). CRY2 photobodies are associated with not only the function, but also the degradation of CRY2 (Yu et al., 2009). We estimated that the CRY2–GFP photobodies could be detected in over 90% of the hypocotyl epidermal cells examined within 1 min of blue-light treatment (Figure 5A). The percentage of nuclei that contain the CRY2–GFP photobodies decreases after a few minutes of blue-light treatment, but the size of remaining photobodies increases (Yu et al., 2009) (Figure 4). The nuclear photobodies were also detected in seedlings expressing CRY2K541R and CRY2K554/5R within 1 min of blue-light treatment (Figure 4). However, the photobodies of CRY2K541 or CRY2K554/5R seem to decrease faster than that of CRY2–GFP: the photobodies in seedlings expressing the CRY2K541 or CRY2K554/5R decreased to about 50% of that expressing CRY2–GFP within 30 min of blue-light treatment (Figure 5A). This is probably because lower levels of CRY2K541R or CRY2K554/5R accumulate in the nucleus, and they may be removed by the blue-light-dependent degradation at a relatively faster rate. Interestingly, at the same time as the CRY2K541R or CRY2K554/5R photobodies disappeared in the nucleus, these mutant proteins formed photobody-like protein bodies in the cytoplasm (Figure 4, 30', 60'). Similarly to photobodies in the nucleus, formation of the cytoplasmic photobody-like protein bodies of CRY2K541R and CRY2K554/5R is strictly dependent on blue light (Figure 4). This suggests that only the photoexcited CRY2K541R and CRY2K554/5R proteins can form the photobody-like protein bodies. Also analogous to photobodies, the cytoplasmic photobody-like protein bodies of CRY2K541R and CRY2K554/5R decreased in number (Figure 5B) but increased in size (Figure 5C). This observation indicates that the cytoplasmic photobody-like protein bodies of CRY2K541R and CRY2K554/5R behave very similarly to CRY2–GFP in that they may ‘grow’ by oligomerization in response to blue light. Taken together, we propose that photoexcited CRY2 oligomerize to form photobodies, which are primarily composed of the photoexcited photoreceptor molecules. Although the nuclear environment may accelerate photobody formation, the photobody-like structure may form in the absence of the nuclear environment. For example, certain nuclear proteins may facilitate oligomerization of CRY2 or CRY2–GFP located in the nucleus, but the photoexcited CRY2K541R and CRY2K554/5R located in the cytoplasm may still undergo oligomerization albeit at a relatively slower rate. Because the rate of formation of the CRY2 photobody is significantly faster than not only the biochemical changes of CRY2, such as phosphorylation and degradation (Figure 3 and 4) (Shalitin et al., 2002; Yu et al., 2007, 2009), but also physiological activities of CRY2, such as blue-light-induced gene expression changes (Zuo et al., 2011), we hypothesize that photoexcited CRY2 may form photobodies to increase the local concentration of the photoreceptors and facilitate interaction of CRY2 with other proteins.
Figure 4.
Blue-Light-Dependent Formation of Photobodies and Cytoplasmic Photobody-Like Protein Bodies of the K541R and K554/5R Mutant Proteins.Upper half of hypocotyls of 5-day-old etiolated seedlings exposed to blue light (36 μmol m−2 s−1) for the time (0, 1, 5, 30, and 60 min) indicated. Samples were fixed in 4% paraformaldehyde and examined by a fluorescence microscope. The white arrowheads indicate representative cytoplasmic photobody-like protein bodies.
Figure 5.
Formation of the Cytoplasmic CRY2K541R or CRY2K554/5R Photobody-Like Structures.
(A) The percentage of nucleus-containing photobodies decreases over time. PCN (Photobody Containing Nucleus) was measured as described in Figure 4, with the standard deviations (n ≥ 200) shown.
(B) The number of CPLPB (Cytoplasmic Photobody-Like Protein Bodies) decreases over time. CPLPB per cell is calculated by the formula [total number of CPLPBs in a field of view]/[the number of cells in the same field view]. The standard deviation is derived from three measurements.
(C) The size of CPLPB increases over time.
METHODS
Plant Materials
The Arabidopsis lines used in this report are in the cry1cry2 background derived from the Columbia accession. The CRY2–GFP transgenic line was as described previously (Yu et al., 2009). Amino-acid substitutions of lysine to arginine were introduced into the CRY2 coding region at positions of K541 and K554K555, respectively, using the QuickChange Site-Directed Mutagenesis system according to the manufacturer’s instruction (Stratagene, CA) and the mutant CRY2 fused with GFP were constructed into the binary vector pKYLX as previously described (Yu et al., 2009). The two constructs were introduced into the cry1cry2 mutants by Agrobacterium tumefaciens using the floral dip transformation method and the transgenic lines were examined by Western blot analysis for selection of independent lines that expressed CRY2–GFP proteins at the similar level. Light sources and measurements of hypocotyl elongation are as previously described (Liu et al., 2011b).
Fluorescence Microscopy
For fluorescence microscopy analyses, seedlings were grown on ½ MS agar medium without sucrose for 5 d in the dark. Etiolated seedlings exposed to blue light over time as indicated were collected and fixed in 4% paraformaldehyde in Tris buffer (10 mM Tris-HCl, pH 7.5, 10 mM Na EDTA, 100 mM NaCl) for 20 min, washed twice by the same Tris buffer without paraformaldehyde for 10 min. The whole seedling was placed on the object slide, immersed with one drop of VECTASHIELD® Mounting Medium with DAPI (Vector Laboratories, Catalog No. HT-1200), and covered with a glass coverslip. The slides were pressed to dry by binder clips overnight. The fluorescence images were captured using an inverted fluorescence microscope (BX-71, Olympus). Image analyses were performed using MacBiophotonics ImageJ software and processed using Adobe Photoshop (Adobe Systems). The definition of the nuclear and cytoplasm region is as described previously (Wang et al., 2010b). The average fluorescence intensity in nucleus and cytoplasm were measured with background correction by ImageJ software and the nuclear/cytoplasmic fluorescence intensity ratios were calculated by the formula [average fluorescence intensity in nucleus]/[average fluorescence intensity in cytoplasm]. The number and size of cytoplasmic photobody-like proteins were measured using the ‘color threshold’ and ‘analyze particle’ features of ImageJ software with the nuclear photobody manually excluded by inspection.
SUPPLEMENTARY DATA
Supplementary Data are available at Molecular Plant Online.
FUNDING
This work is supported in part by the National Institute of Health (Grant No. GM56265 to C.L.), the National Natural Science Foundation of China (Grant No. 31171352 to B.L.), and National Transgenic Crop Initiative (Grant No. 2010ZX08010–002). No conflict of interest declared.
Supplementary Material
References
- Ahmad M, Cashmore AR. HY4 gene of A. thaliana encodes a protein with characteristics of a blue-light photoreceptor. Nature. 1993;366:162–166. doi: 10.1038/366162a0. [DOI] [PubMed] [Google Scholar]
- Cansizoglu AE, Lee BJ, Zhang ZC, Fontoura BM, Chook YM. Structure-based design of a pathway-specific nuclear import inhibitor. Nat. Struct. Mol. Biol. 2007;14:452–454. doi: 10.1038/nsmb1229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cashmore AR. Cryptochromes: enabling plants and animals to determine circadian time. Cell. 2003;114:537–543. [PubMed] [Google Scholar]
- Conti E, Kuriyan J. Crystallographic analysis of the specific yet versatile recognition of distinct nuclear localization signals by karyopherin alpha. Structure. 2000;8:329–338. doi: 10.1016/s0969-2126(00)00107-6. [DOI] [PubMed] [Google Scholar]
- Dingwall C, Robbins J, Dilworth SM, Roberts B, Richardson WD. The nucleoplasmin nuclear location sequence is larger and more complex than that of SV-40 large T antigen. J. Cell Biol. 1988;107:841–849. doi: 10.1083/jcb.107.3.841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- El-Din El-Assal S, Alonso-Blanco C, Peeters AJ, Raz V, Koornneef M. A QTL for flowering time in Arabidopsis reveals a novel allele of CRY2. Nat. Genet. 2001;29:435–440. doi: 10.1038/ng767. [DOI] [PubMed] [Google Scholar]
- Gu NN, Zhang YC, Yang HQ. Substitution of a conserved glycine in the PHR domain of Arabidopsis CRYPTOCHROME 1 confers a constitutive light response. Mol. Plant. 2012;5:85–97. doi: 10.1093/mp/ssr052. [DOI] [PubMed] [Google Scholar]
- Guo H, Duong H, Ma N, Lin C. The Arabidopsis blue light receptor cryptochrome 2 is a nuclear protein regulated by a blue light-dependent post-transcriptional mechanism. Plant J. 1999;19:279–287. doi: 10.1046/j.1365-313x.1999.00525.x. [DOI] [PubMed] [Google Scholar]
- Guo H, Yang H, Mockler TC, Lin C. Regulation of flowering time by Arabidopsis photoreceptors. Science. 1998;279:1360–1363. doi: 10.1126/science.279.5355.1360. [DOI] [PubMed] [Google Scholar]
- Jeong RD, et al. Cryptochrome 2 and phototropin 2 regulate resistance protein-mediated viral defense by negatively regulating an E3 ubiquitin ligase. Proc. Natl Acad. Sci. U S A. 2010;107:13538–13543. doi: 10.1073/pnas.1004529107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang CY, Lian HL, Wang FF, Huang JR, Yang HQ. Cryptochromes, phytochromes, and COP1 regulate light-controlled stomatal development in Arabidopsis. Plant Cell. 2009;21:2624–2641. doi: 10.1105/tpc.109.069765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kircher S, et al. Light quality-dependent nuclear import of the plant photoreceptors phytochrome A and B. Plant Cell. 1999;11:1445–1456. doi: 10.1105/tpc.11.8.1445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kleine T, Lockhart P, Batschauer A. An Arabidopsis protein closely related to Synechocystis cryptochrome is targeted to organelles. Plant J. 2003;35:93–103. doi: 10.1046/j.1365-313x.2003.01787.x. [DOI] [PubMed] [Google Scholar]
- Kleiner O, Kircher S, Harter K, Batschauer A. Nuclear localization of the Arabidopsis blue light receptor cryptochrome 2. Plant J. 1999;19:289–296. doi: 10.1046/j.1365-313x.1999.00535.x. [DOI] [PubMed] [Google Scholar]
- Lin C, Shalitin D. Cryptochrome structure and signal transduction. Annu. Rev. Plant Biol. 2003;54:469–496. doi: 10.1146/annurev.arplant.54.110901.160901. [DOI] [PubMed] [Google Scholar]
- Lin C, Yang H, Guo H, Mockler T, Chen J, Cashmore AR. Enhancement of blue-light sensitivity of Arabidopsis seedlings by a blue light receptor cryptochrome 2. Proc. Natl Acad. Sci. U S A. 1998;95:2686–2690. doi: 10.1073/pnas.95.5.2686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu B, Liu H, Zhong D, Lin C. Searching for a photocycle of the cryptochrome photoreceptors. Curr. Opin. Plant Biol. 2011a;13:578–586. doi: 10.1016/j.pbi.2010.09.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu B, Zuo Z, Liu H, Liu X, Lin C. Arabidopsis cryptochrome 1 interacts with SPA1 to suppress COP1 activity in response to blue light. Genes Dev. 2011b;25:1029–1034. doi: 10.1101/gad.2025011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu H, Liu B, Zhao C, Pepper M, Lin C. The action mechanisms of plant cryptochromes. Trends Plant Sci. 2011c;16:684–691. doi: 10.1016/j.tplants.2011.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mao J, Zhang YC, Sang Y, Li QH, Yang HQ. From the cover: a role for Arabidopsis cryptochromes and COP1 in the regulation of stomatal opening. Proc. Natl Acad. Sci. U S A. 2005;102:12270–12275. doi: 10.1073/pnas.0501011102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marchenko ND, Hanel W, Li D, Becker K, Reich N, Moll UM. Stress-mediated nuclear stabilization of p53 is regulated by ubiquitination and importin-alpha3 binding. Cell Death Differ. 2009;17:255–267. doi: 10.1038/cdd.2009.173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mas P, Devlin PF, Panda S, Kay SA. Functional interaction of phytochrome B and cryptochrome 2. Nature. 2000;408:207–211. doi: 10.1038/35041583. [DOI] [PubMed] [Google Scholar]
- Mockler TC, Guo H, Yang H, Duong H, Lin C. Antagonistic actions of Arabidopsis cryptochromes and phytochrome B in the regulation of floral induction. Development. 1999;126:2073–2082. doi: 10.1242/dev.126.10.2073. [DOI] [PubMed] [Google Scholar]
- Muller P, Ahmad M. Light-activated cryptochrome reacts with molecular oxygen to form a flavin-superoxide radical pair consistent with magnetoreception. J. Biol. Chem. 2011;286:21033–21040. doi: 10.1074/jbc.M111.228940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robbins J, Dilworth SM, Laskey RA, Dingwall C. Two interdependent basic domains in nucleoplasmin nuclear targeting sequence: identification of a class of bipartite nuclear targeting sequence. Cell. 1991;64:615–623. doi: 10.1016/0092-8674(91)90245-t. [DOI] [PubMed] [Google Scholar]
- Sakamoto K, Nagatani A. Nuclear localization activity of phytochrome B. Plant J. 1996;10:859–868. doi: 10.1046/j.1365-313x.1996.10050859.x. [DOI] [PubMed] [Google Scholar]
- Sancar A. Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors. Chem. Rev. 2003;103:2203–2237. doi: 10.1021/cr0204348. [DOI] [PubMed] [Google Scholar]
- Selby CP, Sancar A. A cryptochrome/photolyase class of enzymes with single-stranded DNA-specific photolyase activity. Proc. Natl Acad. Sci. U S A. 2006;103:17696–17700. doi: 10.1073/pnas.0607993103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shalitin D, et al. Regulation of Arabidopsis cryptochrome 2 by blue-light-dependent phosphorylation. Nature. 2002;417:763–767. doi: 10.1038/nature00815. [DOI] [PubMed] [Google Scholar]
- Somers DE, Devlin PF, Kay SA. Phytochromes and cryptochromes in the entrainment of the Arabidopsis circadian clock. Science. 1998;282:1488–1490. doi: 10.1126/science.282.5393.1488. [DOI] [PubMed] [Google Scholar]
- Van Buskirk EK, Decker PV, Chen M. Photobodies in light signaling. Plant Physiol. 2011 doi: 10.1104/pp.111.186411. ? [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang FF, Lian HL, Kang CY, Yang HQ. Phytochrome B is involved in mediating red light-induced stomatal opening in Arabidopsis thaliana. Mol. Plant. 2010a;3:246–259. doi: 10.1093/mp/ssp097. [DOI] [PubMed] [Google Scholar]
- Wang YE, et al. Ubiquitin-regulated nuclear-cytoplasmic trafficking of the Nipah virus matrix protein is important for viral budding. PLoS Pathog. 2010b;6:e1001186. doi: 10.1371/journal.ppat.1001186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu L, Yang HQ. CRYPTOCHROME 1 is implicated in promoting R protein-mediated plant resistance to Pseudomonas syringae in Arabidopsis. Mol. Plant. 2010;3:539–548. doi: 10.1093/mp/ssp107. [DOI] [PubMed] [Google Scholar]
- Yu X, et al. Arabidopsis cryptochrome 2 completes its posttranslational life cycle in the nucleus. Plant Cell. 2007;19:3146–3156. doi: 10.1105/tpc.107.053017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu X, et al. Formation of nuclear bodies of Arabidopsis CRY2 in response to blue light is associated with its blue light-dependent degradation. Plant Cell. 2009;21:118–130. doi: 10.1105/tpc.108.061663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuo Z, Liu H, Liu B, Liu X, Lin C. Blue light-dependent interaction of CRY2 with SPA1 regulates COP1 activity and floral initiation in Arabidopsis. Curr. Biol. 2011;21:841–847. doi: 10.1016/j.cub.2011.03.048. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.