Abstract
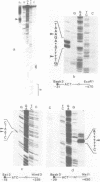
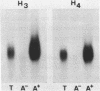
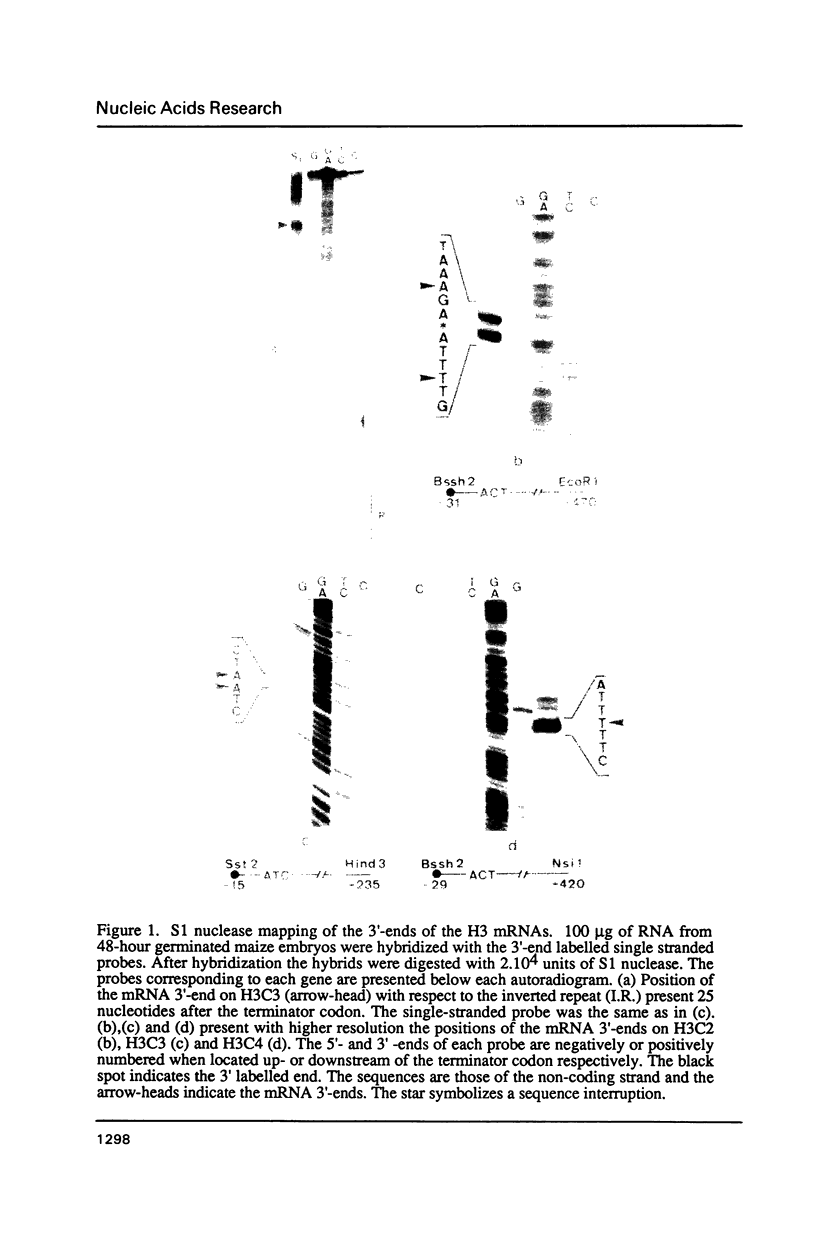
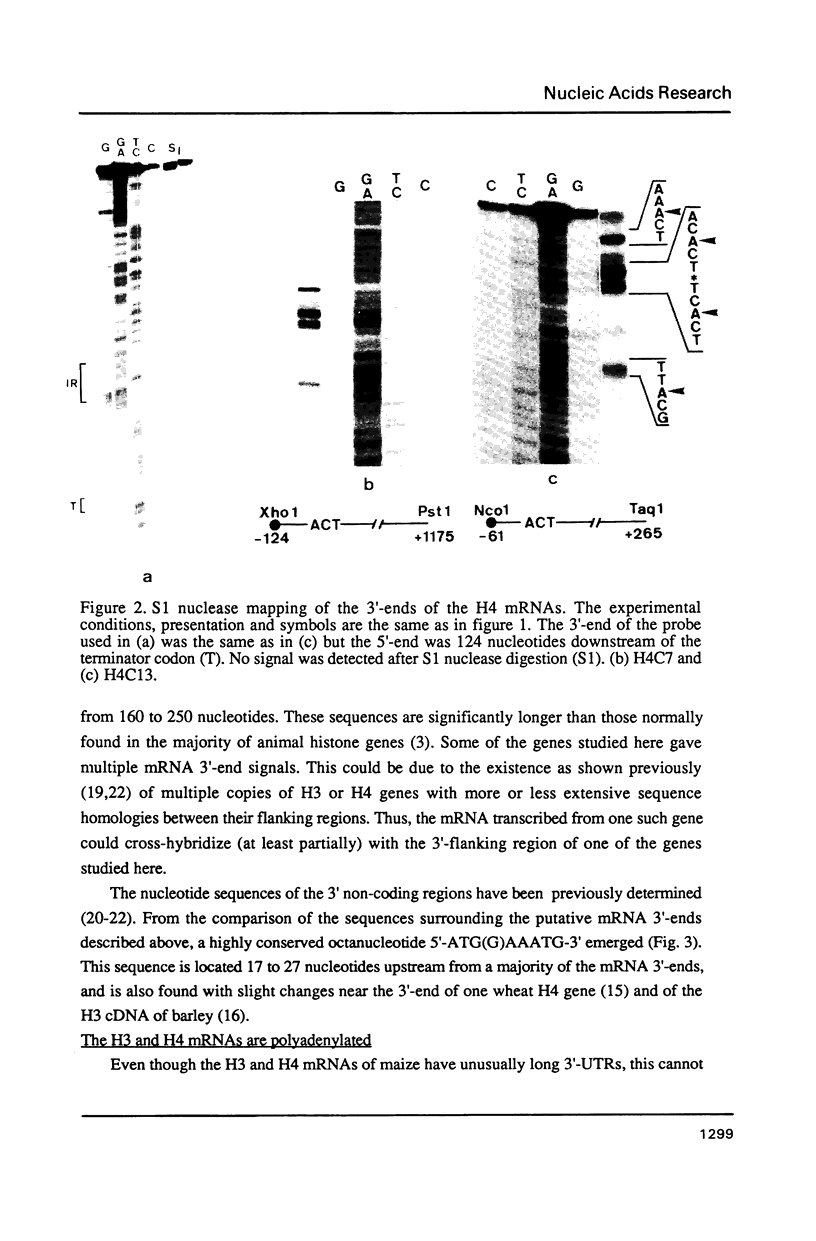
Northern blot analysis revealed that the histone H3 and H4 mRNAs are of unusual large size in germinating maize embryos. S1-mapping experiments show that the 3'-untranslated regions of the mRNAs transcribed from 3 H3 and 2 H4 maize genes previously described are much longer than in the non-polyadenylated histone mRNAs which represent a major class in animals. Moreover, oligo d(T) cellulose fractionation of RNAs isolated at different developmental stages indicates that more than 99% of the maize H3 and H4 mRNAs are polyadenylated. A putative polyadenylation signal is present in all five genes 17 to 27 nucleotides before the 3'-ends of the mRNAs.
Full text
PDF


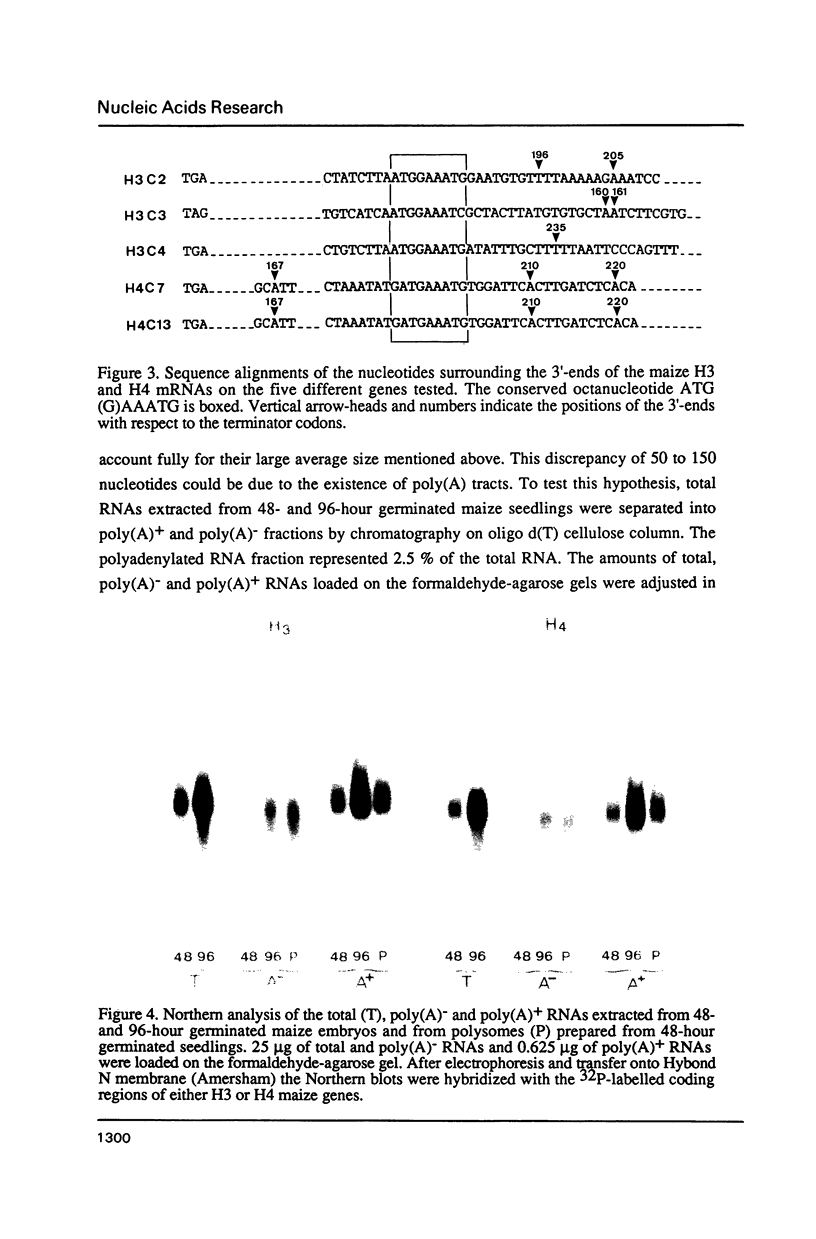




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ahmed C. M., Padayatty J. D. Identification of individual histone mRNAs from rice embryos. Indian J Biochem Biophys. 1982 Jun;19(3):160–166. [PubMed] [Google Scholar]
- Aviv H., Leder P. Purification of biologically active globin messenger RNA by chromatography on oligothymidylic acid-cellulose. Proc Natl Acad Sci U S A. 1972 Jun;69(6):1408–1412. doi: 10.1073/pnas.69.6.1408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bannon G. A., Calzone F. J., Bowen J. K., Allis C. D., Gorovsky M. A. Multiple, independently regulated, polyadenylated messages for histone H3 and H4 in Tetrahymena. Nucleic Acids Res. 1983 Jun 25;11(12):3903–3917. doi: 10.1093/nar/11.12.3903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bird R. C., Jacobs F. A., Stein G., Stein J., Sells B. H. A unique subspecies of histone H4 mRNA from rat myoblasts contains poly(A). Proc Natl Acad Sci U S A. 1985 Oct;82(20):6760–6764. doi: 10.1073/pnas.82.20.6760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnstiel M. L., Busslinger M., Strub K. Transcription termination and 3' processing: the end is in site! Cell. 1985 Jun;41(2):349–359. doi: 10.1016/s0092-8674(85)80007-6. [DOI] [PubMed] [Google Scholar]
- Dean C., Tamaki S., Dunsmuir P., Favreau M., Katayama C., Dooner H., Bedbrook J. mRNA transcripts of several plant genes are polyadenylated at multiple sites in vivo. Nucleic Acids Res. 1986 Mar 11;14(5):2229–2240. doi: 10.1093/nar/14.5.2229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dhaese P., De Greve H., Gielen J., Seurinck L., Van Montagu M., Schell J. Identification of sequences involved in the polyadenylation of higher plant nuclear transcripts using Agrobacterium T-DNA genes as models. EMBO J. 1983;2(3):419–426. doi: 10.1002/j.1460-2075.1983.tb01439.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engel J. D., Sugarman B. J., Dodgson J. B. A chicken histone H3 gene contains intervening sequences. Nature. 1982 Jun 3;297(5865):434–436. doi: 10.1038/297434a0. [DOI] [PubMed] [Google Scholar]
- Ernst S. G., Miller H., Brenner C. A., Nocente-McGrath C., Francis S., McIsaac R. Characterization of a cDNA clone coding for a sea urchin histone H2A variant related to the H2A.F/Z histone protein in vertebrates. Nucleic Acids Res. 1987 Jun 11;15(11):4629–4644. doi: 10.1093/nar/15.11.4629. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fahrner K., Yarger J., Hereford L. Yeast histone mRNA is polyadenylated. Nucleic Acids Res. 1980 Dec 11;8(23):5725–5737. doi: 10.1093/nar/8.23.5725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gantt J. S., Key J. L. Molecular cloning of a pea H1 histone cDNA. Eur J Biochem. 1987 Jul 1;166(1):119–125. doi: 10.1111/j.1432-1033.1987.tb13490.x. [DOI] [PubMed] [Google Scholar]
- Harvey R. P., Whiting J. A., Coles L. S., Krieg P. A., Wells J. R. H2A.F: an extremely variant histone H2A sequence expressed in the chicken embryo. Proc Natl Acad Sci U S A. 1983 May;80(10):2819–2823. doi: 10.1073/pnas.80.10.2819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hentschel C. C., Birnstiel M. L. The organization and expression of histone gene families. Cell. 1981 Aug;25(2):301–313. doi: 10.1016/0092-8674(81)90048-9. [DOI] [PubMed] [Google Scholar]
- Jackson A. O., Larkins B. A. Influence of Ionic Strength, pH, and Chelation of Divalent Metals on Isolation of Polyribosomes from Tobacco Leaves. Plant Physiol. 1976 Jan;57(1):5–10. doi: 10.1104/pp.57.1.5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kedes L. H. Histone genes and histone messengers. Annu Rev Biochem. 1979;48:837–870. doi: 10.1146/annurev.bi.48.070179.004201. [DOI] [PubMed] [Google Scholar]
- Krieg P. A., Robins A. J., Colman A., Wells J. R. Chicken histone H5 mRNA: the polyadenylated RNA lacks the conserved histone 3' terminator sequence. Nucleic Acids Res. 1982 Nov 11;10(21):6777–6785. doi: 10.1093/nar/10.21.6777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehrach H., Diamond D., Wozney J. M., Boedtker H. RNA molecular weight determinations by gel electrophoresis under denaturing conditions, a critical reexamination. Biochemistry. 1977 Oct 18;16(21):4743–4751. doi: 10.1021/bi00640a033. [DOI] [PubMed] [Google Scholar]
- Maxson R., Cohn R., Kedes L., Mohun T. Expression and organization of histone genes. Annu Rev Genet. 1983;17:239–277. doi: 10.1146/annurev.ge.17.120183.001323. [DOI] [PubMed] [Google Scholar]
- Molgaard H. V., Perucho M., Ruiz-Carrillo A. Histone H5 messenger RNA is polyadenylated. Nature. 1980 Jan 31;283(5746):502–504. doi: 10.1038/283502a0. [DOI] [PubMed] [Google Scholar]
- Old R. W., Woodland H. R. Histone genes: not so simple after all. Cell. 1984 Oct;38(3):624–626. doi: 10.1016/0092-8674(84)90256-3. [DOI] [PubMed] [Google Scholar]
- Philipps G., Chaubet N., Chaboute M. E., Ehling M., Gigot C. Genomic organization and nucleotide sequences of two corn histone H4 genes. Gene. 1986;42(2):225–229. doi: 10.1016/0378-1119(86)90301-x. [DOI] [PubMed] [Google Scholar]
- Ruderman J. V., Pardue M. L. A portion of all major classes of histone messenger RNA in amphibian oocytes is polyadenylated. J Biol Chem. 1978 Mar 25;253(6):2018–2025. [PubMed] [Google Scholar]
- Wells D., Bains W., Kedes L. Codon usage in histone gene families of higher eukaryotes reflects functional rather than phylogenetic relationships. J Mol Evol. 1986;23(3):224–241. doi: 10.1007/BF02115579. [DOI] [PubMed] [Google Scholar]
- Wells D., Hoffman D., Kedes L. Unusual structure, evolutionary conservation of non-coding sequences and numerous pseudogenes characterize the human H3.3 histone multigene family. Nucleic Acids Res. 1987 Apr 10;15(7):2871–2889. doi: 10.1093/nar/15.7.2871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wells D., Kedes L. Structure of a human histone cDNA: evidence that basally expressed histone genes have intervening sequences and encode polyadenylylated mRNAs. Proc Natl Acad Sci U S A. 1985 May;82(9):2834–2838. doi: 10.1073/pnas.82.9.2834. [DOI] [PMC free article] [PubMed] [Google Scholar]