Abstract
Mucins are high molecular weight glycoproteins that play important roles in diagnostic and prognostic prediction and in carcinogenesis and tumor invasion. Regulation of expression of mucin genes has been studied extensively, and signaling pathways, transcriptional regulators, and epigenetic modification in promoter regions have been described. Detection of the epigenetic status of cancer-related mucin genes is important for early diagnosis of cancer and for monitoring of tumor behavior and response to targeted therapy. Effects of micro-RNAs on mucin gene expression have also started to emerge. In this review, we discuss the current views on epigenetic mechanisms of regulation of mucin genes (MUC1, MUC2, MUC3A, MUC4, MUC5AC, MUC5B, MUC6, MUC16, and MUC17) and the possible clinical applications of this epigenetic information.
Keywords: Mucin, Cancer, Epigenetics, DNA methylation, Histone modification, Micro-RNAs
Introduction
Mucins are high molecular weight glycoproteins with oligosaccharides attached to serine or threonine residues of the mucin core protein backbone by O-glycosidic linkages (Hollingsworth and Swanson 2004). These proteins are produced by various types of epithelial cells. In the past two decades, core proteins for human mucins (MUC1–MUC8, MUC12–13, MUC15–17, MUC19–21) have been identified and categorized as membrane-associated mucins (MUC1, MUC3A, MUC3B, MUC4, MUC12, MUC13, MUC15, MUC16, MUC17, MUC20, and MUC21) and secreted mucins (MUC2, MUC5AC, MUC5B, MUC6, MUC7, MUC8, and MUC19) (Chen et al. 2004; Higuchi et al. 2004; Hollingsworth and Swanson 2004; Itoh et al. 2008; Lehmann et al. 1989; Moniaux et al. 2001). Mucins are responsible for the physical properties of mucus gels and are involved in epithelial cell protection and maintenance of the local molecular microenvironment (Bhaskar et al. 1992; Ho et al. 2006; Linden et al. 2004, 2008). However, transmembrane mucins, in particular, are overexpressed and aberrantly glycosylated in most cases of adenocarcinoma and are also associated with invasive proliferation of tumors and a poor outcome (Hollingsworth and Swanson 2004; Kufe 2009). Immunohistochemical studies of mucin expression in human tumors have demonstrated that expression of MUC1 and MUC4 is a poor prognostic factor, whereas MUC2 expression is associated with a favorable outcome in neoplasms including pancreatic ductal adenocarcinomas. MUC5AC is a common neoplastic marker in pancreatobiliary neoplasms (Nagata et al. 2007; Yonezawa et al. 2008, 2010).
Epigenetic regulation, including DNA methylation, has been a focus of studies of mucin gene expression. CpG methylation in genomic DNA plays an important role in gene regulation and especially in gene silencing, and generally, the promoter region of a transcribed gene is hypomethylated (Bird 1992). There are three primary DNA methyltransferases (DNA methyltransferase 1 (DNMT1), DNMT3A, and DNMT3B). DNMT1 methylates hemi-methylated DNA and is essential for maintaining methylation patterns. DNMT3A and DNMT3B target previously unmethylated CpGs (Oka et al. 2005; Rodriguez-Paredes and Esteller 2011). Aberrant DNA methylation is strongly associated with human cancer and may serve as a good risk marker for future tumor development and as a target for chemotherapy (Hiraki et al. 2010; Lenhard et al. 2005; Matsubayashi et al. 2006).
Nucleosomes are the basic unit of DNA packaging in eukaryotes, and each nucleosome consists of approximately 147 bp DNA and a histone octamer (core histones) (White et al. 2001). The core histones have an N-terminal amino acid tail of about 20–35 residues in length that is rich in basic amino acids (Munshi et al. 2009). Posttranslational modification of histone tails plays a critical role in epigenetic silencing (Kondo et al. 2003; Wolffe and Matzke 1999). Histone tails are subject to many different chemical modifications, including methylation, acetylation, and phosphorylation (Marmorstein 2001). Such modifications affect the access of regulatory factors and alter histone complexes with chromatin, thereby influencing gene expression. Acetylation of lysines 9, 14, and 27 of histone H3 is associated with euchromatin formation (less condensed), and acetylation of promoter-proximal histones is associated with gene expression (Shi et al. 2003). In contrast, methylation of lysine 9 of histone H3 (H3-MeK9) facilitates formation of heterochromatin (highly condensed), and elevated levels of H3-MeK9 at promoter sequences are associated with suppression of gene expression (Mutskov and Felsenfeld 2004; Nguyen et al. 2002). Methylation of lysine 27 of histone H3 is also a marker for constitutive and facultative heterochromatin (Alvarez-Venegas and Avramova 2005). Another histone modification, methylation of K4 of histone H3, localizes to sites of active transcription, and this modification may stimulate transcription. Combinations of histone modifications at different residues may act synergistically or antagonistically to affect gene expression (Liang et al. 2004).
Gene expression may also be regulated by micro-RNAs (miRNAs), which are a class of small non-coding RNAs of approximately 22 nucleotides (He and Hannon 2004). In mammals, miRNAs are incorporated into RNA-induced silencing complexes and bind imperfectly to the 3′ untranslated region (3′-UTR) of target messenger RNAs (mRNAs) and repress translation (Nakahara and Carthew 2004). Epigenetic regulation of small non-coding RNAs plays a critical role in the modulation of mammalian gene expression (Lujambio and Esteller 2007; Zhang et al. 2008). In addition, miRNAs have been reported to regulate both tumor suppressor genes and oncogenes (Medina and Slack 2008; Saito and Jones 2006).
Pathological events such as carcinogenesis can be caused by alteration in established methylation patterns or dysregulation of chromatin remodeling, both of which can result in significant and consequential changes in gene expression (Das and Singal 2004; DeAngelis et al. 2008). Thus, investigation of DNA methylation, chromatin modification, and miRNA expression is important for diagnosis of carcinogenic risk and prediction of outcomes in patients with cancer.
In this review, we discuss current views on epigenetic mechanisms of human mucin family genes, and we examine the possible clinical applications of this epigenetic information (Table 1).
Table 1.
CpGs, histone modifications, and other factors or miRNAs that affect the expression of epigenetically regulated mucin genes
| Mucins | Promoter methylation sites | Histone modification | Factors or miRNA |
|---|---|---|---|
| MUC1 | −70 to +20 | H3-me2K9, aceK9 | miR-125b, miRNA-145, miR-1226 |
| MUC2 | −3,269, −3,199, −2,331, −1,912, −338 to +158 | H3-me2K4, me3K4, meK9, me2K9, aceK9, me2K27, aceK27, H3-ace, H4-ace | DNMT1, HDAC2 |
| MUC3A | −345 to −75 | N.D. | N.D. |
| MUC4 | −121 to −81a | H3-ace, me2K9, aceK9, me3K27 | Sp1, DNMT3A, DNMT3B, HDAC1, HDAC3 |
| −170 to −102b | |||
| MUC5AC | −3,718 to −3,670 | H3-me2K9, aceK9 | N.D. |
| MUC5B | −2,677/−2,163 | H3-me2K4, me3K4, meK9 | DNMT1, HDAC2 |
| −434/−421 | me2K9, aceK9, me2K27, aceK27 | ||
| MUC17 | −179 to +52 | H3-me2K9, aceK9 | N.D. |
ND not determined
aRegion reported by Vincent et al.
bRegion reported by Yamada et al.
Analysis of DNA methylation and histone modification
There are two main methods for genome-wide analysis of DNA methylation patterns, which are referred to as bisulfite sequencing and MassARRAY analysis. In DNA methylation analysis, sodium bisulfite is used to distinguish between methylated and unmethylated cytosine. Bisulfite treatment of DNA converts cytosine to uracil, but does not alter methylated cytosine. In bisulfite sequencing, polymerase chain reaction (PCR) amplification of converted DNA is used to replace the uracil with thymine, and the PCR product is analyzed by Sanger dideoxy terminator sequencing (Zilberman and Henikoff 2007). This method is a standard tool in genome-wide CpG methylation analysis, but is relatively time-consuming and costly. MassARRAY® quantitative methylation analysis (Sequenom Inc.) is based on the bisulfite conversion biochemistry followed by PCR and base-specific cleavage. The cleavage products are analyzed by matrix-assisted laser desorption ionization time-of-flight mass spectrometry (Ehrich et al. 2005). This combination of methods gives a highly accurate, sensitive, and high-throughput approach for quantitative analysis of DNA methylation.
Meanwhile, methylation-specific PCR (MSP), real-time quantitative MSP with SYBR green or probe (QMSP), and pyrosequencing can be used for the detection of methylation status of specific regions involved in the regulation of gene transcription. MSP and QMSP are bisulfite conversion-based PCR techniques, in which two PCR reactions are carried out using unmethylated DNA primers (U primer) and methylated DNA primers (M primer). MSP and QMSP are rapid and inexpensive methods for detection of the CpG methylation status, but only a limited number of CpGs in a primer sequence can be detected using these methods. Pyrosequencing is a DNA sequencing technique that is based on the detection of released pyrophosphate (PPi) during DNA synthesis (Ronaghi 2001). Pyrosequencing is an accurate and fast method for quantification of CpG methylation.
Chromatin immunoprecipitation (ChIP) analysis is a useful technique for investigating histone modifications and/or protein–DNA interactions. In this procedure, formaldehyde-treated chromatin nucleoprotein complexes are sonicated to reduce the sizes of the DNA fragments to 300 to 500 bp. This is followed by immunoprecipitation using specific antibodies against target histone modifications or transcriptional factors, with subsequent amplification of the immunoprecipitated DNA by PCR (Fullwood and Ruan 2009).
Functional role and epigenetic regulation mechanisms of mucin genes
MUC1
The MUC1 transmembrane glycoprotein (gene cluster on chromosome 1q21) is expressed at a basal level by normal ductal epithelial cells of secretory organs, including pancreas, breast, lung, and gastrointestinal tract (Lan et al. 1990), and overexpressed and aberrantly glycosylated in most cases of adenocarcinoma (Patton et al. 1995). An elevated level of MUC1 protein plays a role in tumor progression, especially in the process of metastasis (Hollingsworth and Swanson 2004; Kufe 2009). Our clinicopathological studies have demonstrated that MUC1 expression is a poor prognostic factor in various human neoplasms (Higashi et al. 1999; Kitamura et al. 1996; Sagara et al. 1999; Takao et al. 1999; Tamada et al. 2002; Utsunomiya et al. 1998).
The epigenetic mechanisms of MUC1 were not examined for more than a decade after initial methylation analysis of the MUC1 coding region (Zrihan-Licht et al. 1995). However, our recent comprehensive DNA methylation analysis revealed that MUC1 gene expression is regulated by DNA methylation and histone H3 lysine 9 (H3-K9) modification at the MUC1 promoter for the first time (Yamada et al. 2008). In the approximately 3,000 bp of the MUC1 promoter region, there are 184 CpG sites. Analysis of all these sites using the MassARRAY compact system for base-specific cleavage of nucleic acids was performed in MUC1-positive and MUC1-negative breast, pancreas, and colon cancer cell lines. The methylation status of nine CpG sites near the transcriptional start site and histone H3-K9 modification in the vicinity of the start site were both related to MUC1 gene expression. Collectively, a series of epigenetic analyses revealed that DNA demethylation, histone H3-K9 demethylation, and histone H3-K9 acetylation in the 5′-flanking region of MUC1 might all be necessary for MUC1 gene expression (Fig. 1). Most recently, downregulation of MUC1 expression by micro-RNA has been reported by several groups (Jin et al. 2010; Rajabi et al. 2010; Sachdeva and Mo 2010). Sachdeva and Mo showed that miR-145 directly targets MUC1 by interaction with the 3′-UTR and found that miR-145 acts as a tumor suppressor in part by affecting invasion and metastasis by targeting MUC1 (Sachdeva and Mo 2010). Moreover, Suh et al. found that miR-145 is silenced by DNA hypermethylation at the miR-145 promoter region in prostate cancer cells and that 5-azadC treatment induced miR-145 expression in prostate cancer cell lines with miR-145 hypermethylation (Suh et al. 2011). In breast cancer cells, Rajabi et al. indicated that miR-125b suppresses translation of the MUC1 and that miR-125b thereby functions as a tumor suppressor (Rajabi et al. 2010). Jin et al. showed that MUC1 expression is suppressed by miR-1226 and that miR-1226-induced downregulation of MUC1 is correlated with cell death of MUC1-positive breast cancer cells (Jin et al. 2010). These findings suggest the possible functional importance of MUC1 under both physiological and pathological conditions.
Fig. 1.
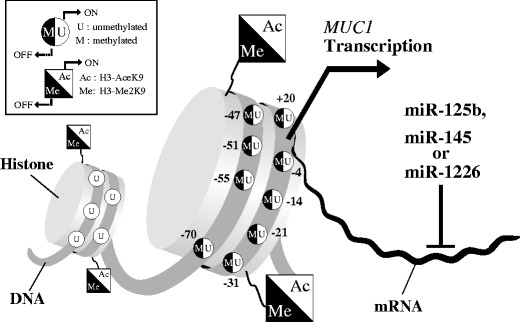
Schematic model of MUC1 gene regulation through epigenetic modifications. Open chromatin is characterized by an unmethylated status of the MUC1 promoter and histones with acetylated H3-K9. This may allow assembly of transcription factors and transcription by RNA polymerase. DNA methylation and histone deacetylation result in condensation of chromatin into a compact state that is inaccessible to transcription factors. The MUC1 gene can also be posttranscriptionally regulated by miRNAs (miR-125b, miR-145, or miR-1226) that target the 3′-UTR of MUC1 mRNA and cause mRNA degradation or translational repression
MUC2
MUC2 is a gel-forming secretory mucin that is expressed in many organs, including the colon, small intestine, and respiratory tract. The corresponding gene has been mapped to human chromosome 11p15 (Chang et al. 1994; Desseyn et al. 1998). Alteration in the protection and lubrication of the intestinal mucosal surface due to a lack of MUC2 might induce an increase of bacterial flora with pro-carcinogenic effects, result in the release of intestinal mucosa-derived factors that are normally depressed by MUC2-stimulating pro-carcinogenic factors, or lead to the destruction of the physical barrier to dietary carcinogens. Loss of MUC2 might also compromise signaling that contributes to epithelial differentiation and proliferation through contacts with membrane-bound mucins or alter the differentiation program of the intestinal mucosa, resulting in an increased probability of tumor formation. Muc2 knockout mice frequently develop adenomas in the small intestine that progress to invasive adenocarcinoma (Velcich et al. 2002). However, high levels of MUC2 expression seen in indolent human pancreatobiliary neoplasms (Higashi et al. 1999; Horinouchi et al. 2003; Nakamura et al. 2002; Yamashita et al. 1993; Yonezawa et al. 1997) are associated with a favorable prognosis (Kitamura et al. 1996; Osako et al. 1993; Tamada et al. 2002; Utsunomiya et al. 1998; Yonezawa et al. 1997, 2008; Yonezawa and Sato 1997).
Epigenetic analysis of the MUC2 gene promoter region has been widely studied (Gratchev et al. 2001; Hanski et al. 1997; Mesquita et al. 2003; Siedow et al. 2002). We have determined the detailed methylation status of a wide area of the MUC2 promoter in breast, lung, pancreas, and colon cancer cell lines using bisulfite genomic sequencing and MassARRAY analysis (Hamada et al. 2005; Yamada et al. 2010). Our results indicated that CpG methylation near the MUC2 transcriptional start site (−338 to +158) plays a critical role in MUC2 gene expression. ChIP assays were also performed using anti-dimethyl-H3-K4/K9/K27, anti-trimethyl-H3-K4/K9, and anti-acetyl-H3-K9/K14/K27 antibodies and revealed that histone H3-K4 methylation, histone H3-K9 dimethylation, and histone H3-K9/K27 acetylation in the 5′-flanking region of the MUC2 may be necessary for gene expression (Yamada et al. 2006). Moreover, treatment with the DNA methylation inhibitor 5-aza-2′-deoxycytidine (5-azadC) and/or a histone deacetylase inhibitor, trichostatin A (TSA) decreased the DNA methylation level in MUC2-negative cells, whereas histone H3-K4/K9 methylation and H3-K9/K27 acetylation were changed to the levels in MUC2-positive cells (Yamada et al. 2006). In the distal region of the MUC2 promoter, the CpG sites at −3,269, −3,199, −2,331, and −1,912 was associated with expression of MUC2 (Vincent et al. 2007). Vincent et al. also showed that MUC2 expression is controlled by HDAC2-enhanced DNMT1 and that methylation of the MUC2 promoter markedly impaired its activation by the methylation-insensitive transcription factor SP1 (Vincent et al. 2007). These studies suggest that epigenetic mechanisms are tightly correlated with MUC2 expression in epithelial cells in various organs. Attempts to apply these findings to clinical samples have begun, and it has been shown that hypomethylation of MUC2 plays an important role in the high level of MUC2 expression in mucinous colorectal cancer (Okudaira et al. 2010).
MUC3A
MUC3A was identified and mapped to a mucin cluster on chromosome 7q22 and categorized as a membrane-associated mucin (Crawley et al. 1999; Gum et al. 1990, 1997; Leroy et al. 2003). An association between MUC3A expression and poor prognosis has been shown in pancreatic, breast, gastric, and renal cancers (Leroy et al. 2003; Park et al. 2003; Rakha et al. 2005; Wang and Fang 2003). However, little is known about the functional role of MUC3A in cancer pathology, and understanding the expression mechanisms of MUC3A may be a key step in developing new strategies for cancer diagnosis and treatment.
In the MUC3A proximal promoter (−620 to +209), the methylation status of 30 CpG sites in breast, lung, pancreas, and colon cancer cells were examined by MassARRAY analysis (Kitamoto et al. 2010). The methylation status showed a good correlation with the level of MUC3A expression. MUC3A-negative/low cell lines were hypermethylated in the vicinity of the transcriptional start site (−345 to −75), whereas MUC3A-positive cell lines showed hypomethylation at the same sites. In contrast, restoration of MUC3A mRNA was observed after TSA treatment, although there was no correlation of MUC3A expression with histone modification at H3K4-Me2/Me3 or H3K9-Me2/Ac. These results suggest that histone H3-K4 and H3-K9 do not play critical roles in MUC3A regulation. Although the roles of MUC3A in cancer development are still unclear, some evidence suggests that MUC3A expression is associated with a poor prognosis in many tumor types. Thus, the methylation status of the MUC3A promoter may be a novel epigenetic marker for the diagnosis of carcinogenic risk and prediction of outcomes for patients with cancer.
MUC4
MUC4 is a large transmembrane mucin with a very long glycosylated extracellular domain that is expressed in various normal tissues. The corresponding gene has been mapped to a gene cluster at chromosome 3q29 (Audie et al. 1993, 1995; Buisine et al. 1999; Gipson et al. 1999). MUC4 is often overexpressed in epithelial cancers, as for MUC1, and is associated with invasive proliferation of tumors and a poor outcome (Saitou et al. 2005; Shibahara et al. 2004b; Tamada et al. 2006; Tsutsumida et al. 2007). MUC4 serves as a intramembrane ligand for the receptor tyrosine kinase ErbB2, which is a transmembrane glycoprotein encoded by the c-ErbB-2 proto-oncogene and has a tyrosine kinase domain that is highly homologous with the epidermal growth factor receptor (Yamamoto et al. 1986).
Jonckheere et al. (2004) found that treatment with TSA restored MUC4 mRNA expression in MUC4-negative pancreatic cancer cells. Singh et al. also showed that the expression level of MUC4 was restored by treatment with 5-azadC and sodium butyrate, an inhibitor of histone deacetylase, in MUC4-negative prostate cancer cells, and suggested that MUC4 is regulated epigenetically (Singh et al. 2006). An investigation of the detailed epigenetic mechanisms of MUC4 expression showed that regulation of MUC4 expression involves both DNA methylation (particularly cytosines at −81, −93, −102, −113, and −121) and histone H3 modification (including H3-K9 and H3-K27 in the MUC4 5′-UTR) mediated by DNA methyltransferases (DNMT3A and DNMT3B) and histone deacetylases (HDAC1 and HDAC3) in pancreatic and gastric epithelial cancer cell lines (Vincent et al. 2008). SP1 binding to the MUC4 promoter (−276/−271 and −166/−156) was also found to participate in the regulation of MUC4 gene expression (Vincent et al. 2008).
We used quantitative MassARRAY methylation analysis to examine the methylation status of 92 CpG sites in the MUC4 promoter (−3,629 to +29) in breast, lung, pancreas, and colon cancer cell lines (Yamada et al. 2009). In 10 cancer cell lines, methylation of 5 CpG sites in the 5′-flanking region of MUC4 (−170 to −102) was associated with the expression of MUC4. Five CpG sites (−121 to −81) had previously been correlated with the expression of MUC4 in these cells (Vincent et al. 2008), but 2 of these CpG sites (−93 and −81) were unrelated to expression of MUC4 in our study. This issue may be resolved by the use of different analytical methods in future studies. Most recently, an examination of the relationships among MUC4 promoter methylation, pancreatic cancer progression, and MUC4 mRNA expression in clinical tissue samples led to the suggestion that aberrant MUC4 promoter hypomethylation may be involved in pancreatic carcinogenesis and malignant development of pancreatic ductal adenocarcinoma (Zhu et al. 2010).
MUC5AC and MUC5B
MUC5AC is a gastric-type secreted gel-forming mucin that is located in the chromosome 11p15 region in a cluster of complex mucin genes (Kim et al. 2002). Although MUC5AC expression in the surface mucous cells in normal gastric mucosa served as a positive control for expression, MUC5AC are never expressed in normal pancreatic tissue. In contrast, MUC5AC has high de novo expression in many types of precancerous lesions of pancreatic ductal adenocarcinoma (PDAC), including pancreatic intraepithelial neoplasia (PanIN) (Nagata et al. 2007), intraductal papillary mucinous neoplasm of the pancreas (Yonezawa et al. 1999), and mucinous cystic neoplasm of the pancreas (Gratchev et al. 2001; Handra-Luca et al. 2005). MUC5AC is also expressed in the lesions of bile duct adenocarcinoma, including biliary intraepithelial neoplasia, mucin-producing bile duct tumor, intraductal papillary neoplasm of the bile duct, and bile duct cystic neoplasm (Shibahara et al. 2004a; Zen et al. 2006). Thus, MUC5AC expression may be a useful marker for early detection of pancreatobiliary neoplasms.
Similar to MUC5AC, MUC5B is also a gel-forming mucin and the corresponding gene has been mapped to chromosome 11p15 (Desseyn et al. 1997). MUC5B is widely expressed in normal tissues in the respiratory and digestive tract (Andrianifahanana et al. 2001). Little is known about MUC5B, but aberrant MUC5B expression has been observed in inflamed or cancerous tissues (Andrianifahanana et al. 2001; Kamio et al. 2005; Pinto-de-Sousa et al. 2004).
Previous studies on the epigenetics of human MUC5AC gene expression have focused on a 1.3-kb promoter region and found little influence of epigenetic regulation, despite restoration of MUC5AC mRNA expression in cancer cells by 5-azadC treatment (Ho et al. 2003; Vincent et al. 2007). In contrast, a recent study has shown that methylation in the “CpG island shore” distal to CpG islands is correlated with silencing of numerous genes (Irizarry et al. 2009). In the MUC5AC promoter region, there is no CpG site from −1.3 to −3.1 kb, but it is of interest that CpG sites reappear from about −3.1 kb onwards. This led us to widen the scope of our study to examine epigenetic mechanisms over about 4.0 kb of the MUC5AC promoter (Yamada et al. 2010). The DNA methylation status of the MUC5AC promoter (−3,855 to +321) in 10 cancer cell lines (breast, lung, pancreas, and colon) was mapped using MassARRAY analysis. The results showed that the CpG methylation status of the MUC5AC promoter from −3,718 to −3,670 is correlated with MUC5AC expression. These results were also consistent with data from pyrosequencing analysis. The modification status of histone H3-K9 in the distal promoter region (−3,718 to −3,670) was also tightly correlated with expression of MUC5AC.
Epigenetic regulation of MUC5B has been shown in several studies (Perrais et al. 2001; Van Seuningen et al. 2001). Detailed epigenetic analysis of the MUC5B promoter region (2.3 kb), including bisulfite sequencing and ChIP assays, in esophageal, gastric, pancreatic, and colon cancer cell lines showed that MUC5B is highly sensitive to DNA methylation and histone modifications in the MUC5B distal promoter (CpG sites at −2,677/−2,163), as well as at cytosines at −434 and −421 (Vincent et al. 2007). This study also indicated that MUC5B expression is regulated by HDAC2-enhanced DNMT1 and that methylation of the MUC5B promoter markedly impaired its activation by SP1 (Vincent et al. 2007). Methylation profiling of the MUC5B distal promoter using bisulfite sequencing and pyrosequencing has also been performed in normal lung and in MUC5B-negative lung cancer tissues (Van Seuningen and Vincent 2009).
MUC6
MUC6 was originally isolated from a gastric cDNA library and mapped to a mucin cluster on chromosome 11p15 (Toribara et al. 1993). High expression of MUC6 is observed in gastric mucosa, duodenal Brunner’s glands, gall bladder, seminal vesicles, pancreatic centroacinar cells and ducts, and periductal glands of the common bile duct. Focal expression is seen in basal endometrial and endocervical glands. MUC6 is also highly expressed in pancreatic cancers and cholangiocarcinomas and focally expressed in endocervical adenocarcinomas (Bartman et al. 1998). In breast tissues, Matsukita et al. have shown a correlation between MUC6 expression and mucinous carcinoma of the breast, suggesting that high expression of MUC6, as well as MUC2, in mucinous carcinoma may act as a barrier to cancerous growth and result in less aggressive biological behavior (Matsukita et al. 2003).
Epigenetic regulation of MUC6 seems to be unlikely, since treatment with 5-aza or TSA did not lead to restoration of MUC6 mRNA in MUC6-negative cell lines, despite the identification of three key methylated cytosines at −282, −271, and −72 by bisulfite sequencing (Vincent et al. 2007). Vincent et al. suggested that repression of MUC6 in cancer cells is rather due to the absence of necessary transcription factors or by an unknown repressive mechanism than to the methylation of its promoter (Vincent et al. 2007).
MUC16 (CA125)
MUC16 (CA125) was identified and mapped to a mucin cluster on chromosome 19p13 and categorized as a transmembrane mucin (O’Brien et al. 2001). Because highly O-glycosylated repeats are the landmark of the mucin family of glycoproteins, CA125 was also named MUC16. MUC16 is overexpressed in many carcinomas (e.g., ovarian cancer, uterine cancer, pancreatic cancer, and biliary cancer) and is known as a tumor marker that is associated with poor prognosis (O’Brien et al. 2001, 2002; Yin and Lloyd 2001).
The structure and function of MUC16 have been well studied, but the mechanisms of regulation of MUC16 expression have yet to be clarified. We have examined the epigenetic status of the MUC16 5′-flanking region in breast, lung, pancreas, and colon cancer cells using MassARRAY analysis and ChIP assays (unpublished data). Treatment with 5-azadC and TSA showed restoration of MUC16 mRNA expression, but MassARRAY and ChIP results showed no correlation between histone H3 modification and MUC16 gene expression. Our findings suggest that epigenetic mechanisms are involved in the regulation of MUC16 expression in cancer cells, but DNA methylation and histone H3-K9 modification in the MUC16 5′-flanking region are unlikely to regulate MUC16 transcription directly. It will be necessary to perform promoter analysis to clarify the details of the expression mechanism of MUC16, which is a poor prognostic factor in patients with cancer.
MUC17
MUC17 glycoprotein was identified and mapped to a mucin cluster at chromosome 7q22 and categorized as a membrane-associated mucin (Gum et al. 2002; Williams et al. 1999). MUC17 is mainly expressed in the digestive tract, including the duodenum, ileum, and transverse colon (Gum et al. 2002; Moehle et al. 2006), but its physiological function is unclear. Aberrant MUC17 expression is observed in PDAC, compared to no expression in the normal pancreas or in pancreatitis (Moniaux et al. 2006), and MUC17 is an independent prognostic factor associated with lymph node metastasis in PDAC (Hirono et al. 2010).
We performed MassARRAY quantitative methylation analysis in human cancer cell lines (breast, lung, pancreas, and colon) to determine the methylation status of CpG islands in the MUC17 5′-flanking region (−620 to +209) (Kitamoto et al. 2011). Five CpG sites (−179 to +52) involved in MUC17 expression were identified and these results were consistent with MSP data. To examine the relationship between DNA methylation and histone H3 modification, ChIP assays were performed. Dimethyl-H3-K9 was found in MUC17-negative/low cells, whereas histone H3-K9 was more highly acetylated in MUC17-positive cells. The level of MUC17 mRNA expression was restored by treatment with 5-azadC or TSA. Using MSP analysis, we also investigated the methylation status of normal and cancerous pancreatic tissues from patients with PDAC and found that the methylation status in the five CpG sites of the MUC17 promoter (−179 to +52) was consistent with the expression of MUC17 at both the mRNA and protein levels. Although further studies are needed to clarify these relationships, our findings indicate that the methylation status of the MUC17 promoter could be a novel epigenetic marker for diagnosis of PDAC. Moreover, miRNA microarray analysis in 11 cancer cell lines revealed 5 candidates with a significant relationship with the regulation of MUC17 expression: miR-17, miR-20a, miR-20b, miR-30c, miR-30e, using target prediction based on the miRBase. Of these candidates, it is unclear which are authentic miRNAs in vivo. Additional studies are required, but our data suggest that the MUC17 gene might be regulated posttranscriptionally by miRNAs.
Conclusions and clinical perspectives
The epigenetic mechanisms of human mucin family genes are gradually emerging (Table 1). However, establishment of clinical tests for cancer screening, diagnosis of carcinogenic risk, and tumor staging requires improved understanding of the roles of modifier enzymes (CpG methylation and chromatin) and epigenetic modifications accompanying tumor differentiation. An aberrant increase in both MUC4 mRNA expression and hypomethylation frequency in the PanIN–PDAC progression modality has been shown (Zhu et al. 2010), and increased DNMT1 expression may be involved in multistage pancreatic carcinogenesis from the precancerous stage to malignant progression of ductal carcinomas and may be a predictor of poor prognosis (Peng et al. 2005). Further studies are needed to elucidate the relationships among expression levels of mucins, binding of transcriptional regulatory factors, and recruitment of DNMTs, HDACs, methylated DNA binding domain proteins, and polycomb group proteins.
Mucin expression is tissue- and cell-type specific (Yonezawa et al. 2008), and these specificities may cause different forms of epigenetic regulation at different levels. For MUC1 and MUC4, we were unable to show a relationship between DNA methylation status and expression of MUC1 in the LS174T colon cancer cell line (Yamada et al. 2008, 2009). However, expression of MUC1 in Caco2 cells, which are derived from cancer cells of the same organ, was related to the DNA methylation status. Similarly, BxPC-3 pancreatic cancer cells with low MUC5AC expression had a high level of CpG methylation, whereas PANC1 cells (which have low or no MUC5AC expression) showed hypomethylation. Our results indicate that regulation of expression by DNA methylation is not organ specific, but varies in individual cell lines. Moreover, coordinated changes in DNA methylation and histone modification may be important for epigenetic regulation of mucin genes in cancer cell lines derived from various organs (Kitamoto et al. 2010, 2011; Yamada et al. 2006, 2008, 2009). However, further studies are needed to clarify how these different epigenetic changes are involved in tissue-specific mucin expression in cancer.
Many studies of epigenetic gene regulation have addressed tumor suppressor genes (Garinis et al. 2002). Meanwhile, evidence for promoter hypomethylation of oncogenes has gradually emerged in tumors of several organs (Sato et al. 2003; Sun et al. 2010; Wolff et al. 2010). Currently, epigenetic regulation of oncogenes is a phenomenon of interest. Transmembrane mucin genes associated with a poor prognosis (e.g., MUC1 and MUC4) may also be regulated by epigenetic mechanisms. In cancers like pancreatobiliary and lung carcinomas with a high potential for malignancy showing MUC1 and MUC4 high expression but no MUC2 expression (Yonezawa et al. 2008), promoter CpG sites in transmembrane mucin genes like MUC1 and MUC4 associated with a poor outcome may be hypomethylated, while promoter CpG sites of MUC2 associated with a favorable outcome may be hypermethylated, according to the data based on our studies using human cancer cells (Yamada et al. 2006, 2008, 2009). Improved understanding of the regulatory mechanisms controlling these polar opposite events may enhance the understanding of epigenetic mechanisms associated with other oncogenes and tumor suppressor genes.
Posttranscriptional regulation of gene expression by miRNAs is attracting increased attention (Hartmann et al. 2004; Krol et al. 2010; Rajabi et al. 2010). Among mucin genes, MUC1 expression has been shown to be directly downregulated by miR-145, with the further finding that miR-145 acts as a tumor suppressor in cell invasion and metastasis by targeting MUC1 (Sachdeva and Mo 2010). miR-125b and miR-1226 have also been reported to target MUC1 expression (Jin et al. 2010), and it is likely that these miRNAs could be attractive targets for therapeutic intervention in advanced cancers.
Epigenetic analysis of cancer-related genes in clinical specimens is relatively common, especially using DNA methylation analysis (Kamalakaran et al. 2011; Okudaira et al. 2010). There are several techniques for the evaluation of CpG methylation, but MSP, including quantitative MSP, is a leading strategy for the analysis of clinical samples (Lee et al. 2009; Zhu et al. 2010). However, MSP identifies only regions recognized by the primer sequence, and MassARRAY and pyrosequencing analysis are increasingly being used to overcome this problem (Shinojima et al. 2010; Van Seuningen and Vincent 2009). MassARRAY analysis permits high-throughput identification of methylation sites and semi-quantitative measurement at single or multiple CpG sites. Pyrosequencing is also a reliable technique for quantification of methylation at a single CpG site (Brakensiek et al. 2007). Moreover, methylation analysis by these technologies gives highly reproducible quantification (Shinojima et al. 2010; Van Seuningen and Vincent 2009). In these techniques, however, as clinical samples include various components other than tumor cells, evaluation of the average percentage of methylation of the individual CpG sites is not easy. To permit analysis of discharged fluids, such as pancreatic juice, bile, or sputum, there is a need to develop the technique that allows the detection of individual CpG methylation patterns to be detected.
Acknowledgments
We thank Ms. Izumi Houjou for secretarial assistance. This work was supported by Grants-in-Aid for Scientific Research on Priority Areas 219447 to N. Yamada (JSPS Fellowship), and Scientific Research (C) 20590345 and Scientific Research (B) 23390085 to S. Yonezawa, and Scientific Research (C) 21590399 from the Ministry of Education, Science, Sports, Culture and Technology, Japan and the Kodama Memorial Foundation to M. Higashi.
Competing interest statement
The authors declare that they have no conflicts of interest.
References
- Alvarez-Venegas R, Avramova Z. Methylation patterns of histone H3 Lys 4, Lys 9 and Lys 27 in transcriptionally active and inactive Arabidopsis genes and in atx1 mutants. Nucleic Acids Res. 2005;33:5199–5207. doi: 10.1093/nar/gki830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrianifahanana M, Moniaux N, Schmied BM, Ringel J, Friess H, Hollingsworth MA, et al. Mucin (MUC) gene expression in human pancreatic adenocarcinoma and chronic pancreatitis: a potential role of MUC4 as a tumor marker of diagnostic significance. Clin Cancer Res. 2001;7:4033–4040. [PubMed] [Google Scholar]
- Audie JP, Janin A, Porchet N, Copin MC, Gosselin B, Aubert JP. Expression of human mucin genes in respiratory, digestive, and reproductive tracts ascertained by in situ hybridization. J Histochem Cytochem. 1993;41:1479–1485. doi: 10.1177/41.10.8245407. [DOI] [PubMed] [Google Scholar]
- Audie JP, Tetaert D, Pigny P, Buisine MP, Janin A, Aubert JP, et al. Mucin gene expression in the human endocervix. Hum Reprod. 1995;10:98–102. doi: 10.1093/humrep/10.1.98. [DOI] [PubMed] [Google Scholar]
- Bartman AE, Buisine MP, Aubert JP, Niehans GA, Toribara NW, Kim YS, et al. The MUC6 secretory mucin gene is expressed in a wide variety of epithelial tissues. J Pathol. 1998;186:398–405. doi: 10.1002/(SICI)1096-9896(199812)186:4<398::AID-PATH192>3.0.CO;2-X. [DOI] [PubMed] [Google Scholar]
- Bhaskar KR, Garik P, Turner BS, Bradley JD, Bansil R, Stanley HE, et al. Viscous fingering of HCl through gastric mucin. Nature. 1992;360:458–461. doi: 10.1038/360458a0. [DOI] [PubMed] [Google Scholar]
- Bird A. The essentials of DNA methylation. Cell. 1992;70:5–8. doi: 10.1016/0092-8674(92)90526-I. [DOI] [PubMed] [Google Scholar]
- Brakensiek K, Wingen LU, Langer F, Kreipe H, Lehmann U. Quantitative high-resolution CpG island mapping with pyrosequencing reveals disease-specific methylation patterns of the CDKN2B gene in myelodysplastic syndrome and myeloid leukemia. Clin Chem. 2007;53:17–23. doi: 10.1373/clinchem.2007.072629. [DOI] [PubMed] [Google Scholar]
- Buisine MP, Devisme L, Copin MC, Durand-Reville M, Gosselin B, Aubert JP, et al. Developmental mucin gene expression in the human respiratory tract. Am J Respir Cell Mol Biol. 1999;20:209–218. doi: 10.1165/ajrcmb.20.2.3259. [DOI] [PubMed] [Google Scholar]
- Chang SK, Dohrman AF, Basbaum CB, Ho SB, Tsuda T, Toribara NW, et al. Localization of mucin (MUC2 and MUC3) messenger RNA and peptide expression in human normal intestine and colon cancer. Gastroenterology. 1994;107:28–36. doi: 10.1016/0016-5085(94)90057-4. [DOI] [PubMed] [Google Scholar]
- Chen Y, Zhao YH, Kalaslavadi TB, Hamati E, Nehrke K, Le AD, et al. Genome-wide search and identification of a novel gel-forming mucin MUC19/Muc19 in glandular tissues. Am J Respir Cell Mol Biol. 2004;30:155–165. doi: 10.1165/rcmb.2003-0103OC. [DOI] [PubMed] [Google Scholar]
- Crawley SC, Gum JR, Jr, Hicks JW, Pratt WS, Aubert JP, Swallow DM, et al. Genomic organization and structure of the 3′ region of human MUC3: alternative splicing predicts membrane-bound and soluble forms of the mucin. Biochem Biophys Res Commun. 1999;263:728–736. doi: 10.1006/bbrc.1999.1466. [DOI] [PubMed] [Google Scholar]
- Das PM, Singal R. DNA methylation and cancer. J Clin Oncol. 2004;22:4632–4642. doi: 10.1200/JCO.2004.07.151. [DOI] [PubMed] [Google Scholar]
- DeAngelis JT, Farrington WJ, Tollefsbol TO. An overview of epigenetic assays. Mol Biotechnol. 2008;38:179–183. doi: 10.1007/s12033-007-9010-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Desseyn JL, Guyonnet-Duperat V, Porchet N, Aubert JP, Laine A. Human mucin gene MUC5B, the 10.7-kb large central exon encodes various alternate subdomains resulting in a super-repeat. Structural evidence for a 11p15.5 gene family. J Biol Chem. 1997;272:3168–3178. doi: 10.1074/jbc.272.27.16873. [DOI] [PubMed] [Google Scholar]
- Desseyn JL, Buisine MP, Porchet N, Aubert JP, Degand P, Laine A. Evolutionary history of the 11p15 human mucin gene family. J Mol Evol. 1998;46:102–106. doi: 10.1007/PL00006276. [DOI] [PubMed] [Google Scholar]
- Ehrich M, Nelson MR, Stanssens P, Zabeau M, Liloglou T, Xinarianos G, et al. Quantitative high-throughput analysis of DNA methylation patterns by base-specific cleavage and mass spectrometry. Proc Natl Acad Sci USA. 2005;102:15785–15790. doi: 10.1073/pnas.0507816102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fullwood MJ, Ruan Y. ChIP-based methods for the identification of long-range chromatin interactions. J Cell Biochem. 2009;107:30–39. doi: 10.1002/jcb.22116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garinis GA, Patrinos GP, Spanakis NE, Menounos PG. DNA hypermethylation: when tumour suppressor genes go silent. Hum Genet. 2002;111:115–127. doi: 10.1007/s00439-002-0783-6. [DOI] [PubMed] [Google Scholar]
- Gipson IK, Spurr-Michaud S, Moccia R, Zhan Q, Toribara N, Ho SB, et al. MUC4 and MUC5B transcripts are the prevalent mucin messenger ribonucleic acids of the human endocervix. Biol Reprod. 1999;60:58–64. doi: 10.1095/biolreprod60.1.58. [DOI] [PubMed] [Google Scholar]
- Gratchev A, Siedow A, Bumke-Vogt C, Hummel M, Foss HD, Hanski ML, et al. Regulation of the intestinal mucin MUC2 gene expression in vivo: evidence for the role of promoter methylation. Cancer Lett. 2001;168:71–80. doi: 10.1016/S0304-3835(01)00498-0. [DOI] [PubMed] [Google Scholar]
- Gum JR, Hicks JW, Swallow DM, Lagace RL, Byrd JC, Lamport DT, et al. Molecular cloning of cDNAs derived from a novel human intestinal mucin gene. Biochem Biophys Res Commun. 1990;171:407–415. doi: 10.1016/0006-291X(90)91408-K. [DOI] [PubMed] [Google Scholar]
- Gum JR, Jr, Ho JJ, Pratt WS, Hicks JW, Hill AS, Vinall LE, et al. MUC3 human intestinal mucin. Analysis of gene structure, the carboxyl terminus, and a novel upstream repetitive region. J Biol Chem. 1997;272:26678–26686. doi: 10.1074/jbc.272.42.26678. [DOI] [PubMed] [Google Scholar]
- Gum JR, Jr, Crawley SC, Hicks JW, Szymkowski DE, Kim YS. MUC17, a novel membrane-tethered mucin. Biochem Biophys Res Commun. 2002;291:466–475. doi: 10.1006/bbrc.2002.6475. [DOI] [PubMed] [Google Scholar]
- Hamada T, Goto M, Tsutsumida H, Nomoto M, Higashi M, Sugai T, et al. Mapping of the methylation pattern of the MUC2 promoter in pancreatic cancer cell lines, using bisulfite genomic sequencing. Cancer Lett. 2005;227:175–184. doi: 10.1016/j.canlet.2004.11.058. [DOI] [PubMed] [Google Scholar]
- Handra-Luca A, Lamas G, Bertrand JC, Fouret P. MUC1, MUC2, MUC4, and MUC5AC expression in salivary gland mucoepidermoid carcinoma: diagnostic and prognostic implications. Am J Surg Pathol. 2005;29:881–889. doi: 10.1097/01.pas.0000159103.95360.e8. [DOI] [PubMed] [Google Scholar]
- Hanski C, Riede E, Gratchev A, Foss HD, Bohm C, Klussmann E, et al. MUC2 gene suppression in human colorectal carcinomas and their metastases: in vitro evidence of the modulatory role of DNA methylation. Lab Invest. 1997;77:685–695. [PubMed] [Google Scholar]
- Hartmann C, Corre-Menguy F, Boualem A, Jovanovic M, Lelandais-Briere C. MicroRNAs: a new class of gene expression regulators. Med Sci (Paris) 2004;20:894–898. doi: 10.1051/medsci/20042010894. [DOI] [PubMed] [Google Scholar]
- He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–531. doi: 10.1038/nrg1379. [DOI] [PubMed] [Google Scholar]
- Higashi M, Yonezawa S, Ho JJ, Tanaka S, Irimura T, Kim YS, et al. Expression of MUC1 and MUC2 mucin antigens in intrahepatic bile duct tumors: its relationship with a new morphological classification of cholangiocarcinoma. Hepatology. 1999;30:1347–1355. doi: 10.1002/hep.510300609. [DOI] [PubMed] [Google Scholar]
- Higuchi T, Orita T, Katsuya K, Yamasaki Y, Akiyama K, Li H, et al. MUC20 suppresses the hepatocyte growth factor-induced Grb2–Ras pathway by binding to a multifunctional docking site of met. Mol Cell Biol. 2004;24:7456–7468. doi: 10.1128/MCB.24.17.7456-7468.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hiraki M, Kitajima Y, Sato S, Nakamura J, Hashiguchi K, Noshiro H, et al. Aberrant gene methylation in the peritoneal fluid is a risk factor predicting peritoneal recurrence in gastric cancer. World J Gastroenterol. 2010;16:330–338. doi: 10.3748/wjg.v16.i3.330. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirono S, Yamaue H, Hoshikawa Y, Ina S, Tani M, Kawai M, et al. Molecular markers associated with lymph node metastasis in pancreatic ductal adenocarcinoma by genome-wide expression profiling. Cancer Sci. 2010;101:259–266. doi: 10.1111/j.1349-7006.2009.01359.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ho JJ, Han SW, Pan PL, Deng G, Kuan SF, Kim YS. Methylation status of promoters and expression of MUC2 and MUC5AC mucins in pancreatic cancer cells. Int J Oncol. 2003;22:273–279. [PubMed] [Google Scholar]
- Ho SB, Dvorak LA, Moor RE, Jacobson AC, Frey MR, Corredor J, et al. Cysteine-rich domains of muc3 intestinal mucin promote cell migration, inhibit apoptosis, and accelerate wound healing. Gastroenterology. 2006;131:1501–1517. doi: 10.1053/j.gastro.2006.09.006. [DOI] [PubMed] [Google Scholar]
- Hollingsworth MA, Swanson BJ. Mucins in cancer: protection and control of the cell surface. Nat Rev Cancer. 2004;4:45–60. doi: 10.1038/nrc1251. [DOI] [PubMed] [Google Scholar]
- Horinouchi M, Nagata K, Nakamura A, Goto M, Takao S, Sakamoto M, et al. Expression of different glycoforms of membrane mucin (MUC1) and secretory mucin (MUC2, MUC5AC and MUC6) in pancreatic neoplasms. Acta Histochem Cytochem. 2003;36:443–453. doi: 10.1267/ahc.36.443. [DOI] [Google Scholar]
- Irizarry RA, Ladd-Acosta C, Wen B, Wu Z, Montano C, Onyango P, et al. The human colon cancer methylome shows similar hypo- and hypermethylation at conserved tissue-specific CpG island shores. Nat Genet. 2009;41:178–186. doi: 10.1038/ng.298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Itoh Y, Kamata-Sakurai M, Denda-Nagai K, Nagai S, Tsuiji M, Ishii-Schrade K, et al. Identification and expression of human epiglycanin/MUC21: a novel transmembrane mucin. Glycobiology. 2008;18:74–83. doi: 10.1093/glycob/cwm118. [DOI] [PubMed] [Google Scholar]
- Jin C, Rajabi H, Kufe D. miR-1226 targets expression of the mucin 1 oncoprotein and induces cell death. Int J Oncol. 2010;37:61–69. doi: 10.3892/ijo_00000653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jonckheere N, Perrais M, Mariette C, Batra SK, Aubert JP, Pigny P, et al. A role for human MUC4 mucin gene, the ErbB2 ligand, as a target of TGF-beta in pancreatic carcinogenesis. Oncogene. 2004;23:5729–5738. doi: 10.1038/sj.onc.1207769. [DOI] [PubMed] [Google Scholar]
- Kamalakaran S, Varadan V, Giercksky Russnes HE, Levy D, Kendall J, Janevski A, et al. DNA methylation patterns in luminal breast cancers differ from non-luminal subtypes and can identify relapse risk independent of other clinical variables. Mol Oncol. 2011;5(1):77–92. doi: 10.1016/j.molonc.2010.11.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamio K, Matsushita I, Hijikata M, Kobashi Y, Tanaka G, Nakata K, et al. Promoter analysis and aberrant expression of the MUC5B gene in diffuse panbronchiolitis. Am J Respir Crit Care Med. 2005;171:949–957. doi: 10.1164/rccm.200409-1168OC. [DOI] [PubMed] [Google Scholar]
- Kim GE, Bae HI, Park HU, Kuan SF, Crawley SC, Ho JJ, et al. Aberrant expression of MUC5AC and MUC6 gastric mucins and sialyl Tn antigen in intraepithelial neoplasms of the pancreas. Gastroenterology. 2002;123:1052–1060. doi: 10.1053/gast.2002.36018. [DOI] [PubMed] [Google Scholar]
- Kitamoto S, Yamada N, Yokoyama S, Houjou I, Higashi M, Yonezawa S. Promoter hypomethylation contributes to the expression of MUC3A in cancer cells. Biochem Biophys Res Commun. 2010;397:333–339. doi: 10.1016/j.bbrc.2010.05.124. [DOI] [PubMed] [Google Scholar]
- Kitamoto S, Yamada N, Yokoyama S, Houjou I, Higashi M, Goto M, et al. DNA methylation and histone H3-K9 modifications contribute to MUC17 expression. Glycobiology. 2011;21:247–256. doi: 10.1093/glycob/cwq155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitamura H, Yonezawa S, Tanaka S, Kim YS, Sato E. Expression of mucin carbohydrates and core proteins in carcinomas of the ampulla of Vater: their relationship to prognosis. Jpn J Cancer Res. 1996;87:631–640. doi: 10.1111/j.1349-7006.1996.tb00270.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kondo Y, Shen L, Issa JP. Critical role of histone methylation in tumor suppressor gene silencing in colorectal cancer. Mol Cell Biol. 2003;23:206–215. doi: 10.1128/MCB.23.1.206-215.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krol J, Loedige I, Filipowicz W. The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet. 2010;11:597–610. doi: 10.1038/nrg2843. [DOI] [PubMed] [Google Scholar]
- Kufe DW. Mucins in cancer: function, prognosis and therapy. Nat Rev Cancer. 2009;9:874–885. doi: 10.1038/nrc2761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lan MS, Batra SK, Qi WN, Metzgar RS, Hollingsworth MA. Cloning and sequencing of a human pancreatic tumor mucin cDNA. J Biol Chem. 1990;265:15294–15299. [PubMed] [Google Scholar]
- Lee BB, Lee EJ, Jung EH, Chun HK, Chang DK, Song SY, et al. Aberrant methylation of APC, MGMT, RASSF2A, and Wif-1 genes in plasma as a biomarker for early detection of colorectal cancer. Clin Cancer Res. 2009;15:6185–6191. doi: 10.1158/1078-0432.CCR-09-0111. [DOI] [PubMed] [Google Scholar]
- Lehmann JM, Riethmuller G, Johnson JP. MUC18, a marker of tumor progression in human melanoma, shows sequence similarity to the neural cell adhesion molecules of the immunoglobulin superfamily. Proc Natl Acad Sci USA. 1989;86:9891–9895. doi: 10.1073/pnas.86.24.9891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lenhard K, Bommer GT, Asutay S, Schauer R, Brabletz T, Goke B, et al. Analysis of promoter methylation in stool: a novel method for the detection of colorectal cancer. Clin Gastroenterol Hepatol. 2005;3:142–149. doi: 10.1016/S1542-3565(04)00624-X. [DOI] [PubMed] [Google Scholar]
- Leroy X, Gouyer V, Ballereau C, Zerimech F, Huet G, Copin MC, et al. Quantitative RT-PCR assay for MUC3 and VEGF mRNA in renal clear cell carcinoma: relationship with nuclear grade and prognosis. Urology. 2003;62:771–775. doi: 10.1016/S0090-4295(03)00560-0. [DOI] [PubMed] [Google Scholar]
- Liang G, Lin JC, Wei V, Yoo C, Cheng JC, Nguyen CT, et al. Distinct localization of histone H3 acetylation and H3-K4 methylation to the transcription start sites in the human genome. Proc Natl Acad Sci USA. 2004;101:7357–7362. doi: 10.1073/pnas.0401866101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linden S, Mahdavi J, Hedenbro J, Boren T, Carlstedt I. Effects of pH on Helicobacter pylori binding to human gastric mucins: identification of binding to non-MUC5AC mucins. Biochem J. 2004;384:263–270. doi: 10.1042/BJ20040402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linden SK, Sutton P, Karlsson NG, Korolik V, McGuckin MA. Mucins in the mucosal barrier to infection. Mucosal Immunol. 2008;1:183–197. doi: 10.1038/mi.2008.5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lujambio A, Esteller M. CpG island hypermethylation of tumor suppressor microRNAs in human cancer. Cell Cycle. 2007;6:1455–1459. doi: 10.4161/cc.6.12.4408. [DOI] [PubMed] [Google Scholar]
- Marmorstein R. Protein modules that manipulate histone tails for chromatin regulation. Nat Rev Mol Cell Biol. 2001;2:422–432. doi: 10.1038/35073047. [DOI] [PubMed] [Google Scholar]
- Matsubayashi H, Canto M, Sato N, Klein A, Abe T, Yamashita K, et al. DNA methylation alterations in the pancreatic juice of patients with suspected pancreatic disease. Cancer Res. 2006;66:1208–1217. doi: 10.1158/0008-5472.CAN-05-2664. [DOI] [PubMed] [Google Scholar]
- Matsukita S, Nomoto M, Kitajima S, Tanaka S, Goto M, Irimura T, et al. Expression of mucins (MUC1, MUC2, MUC5AC and MUC6) in mucinous carcinoma of the breast: comparison with invasive ductal carcinoma. Histopathology. 2003;42:26–36. doi: 10.1046/j.1365-2559.2003.01530.x. [DOI] [PubMed] [Google Scholar]
- Medina PP, Slack FJ. MicroRNAs and cancer: an overview. Cell Cycle. 2008;7:2485–2492. doi: 10.4161/cc.7.16.6453. [DOI] [PubMed] [Google Scholar]
- Mesquita P, Peixoto AJ, Seruca R, Hanski C, Almeida R, Silva F, et al. Role of site-specific promoter hypomethylation in aberrant MUC2 mucin expression in mucinous gastric carcinomas. Cancer Lett. 2003;189:129–136. doi: 10.1016/S0304-3835(02)00549-9. [DOI] [PubMed] [Google Scholar]
- Moehle C, Ackermann N, Langmann T, Aslanidis C, Kel A, Kel-Margoulis O, et al. Aberrant intestinal expression and allelic variants of mucin genes associated with inflammatory bowel disease. J Mol Med. 2006;84:1055–1066. doi: 10.1007/s00109-006-0100-2. [DOI] [PubMed] [Google Scholar]
- Moniaux N, Escande F, Porchet N, Aubert JP, Batra SK. Structural organization and classification of the human mucin genes. Front Biosci. 2001;6:D1192–D1206. doi: 10.2741/Moniaux. [DOI] [PubMed] [Google Scholar]
- Moniaux N, Junker WM, Singh AP, Jones AM, Batra SK. Characterization of human mucin MUC17. Complete coding sequence and organization. J Biol Chem. 2006;281:23676–23685. doi: 10.1074/jbc.M600302200. [DOI] [PubMed] [Google Scholar]
- Munshi A, Shafi G, Aliya N, Jyothy A. Histone modifications dictate specific biological readouts. J Genet Genomics. 2009;36:75–88. doi: 10.1016/S1673-8527(08)60094-6. [DOI] [PubMed] [Google Scholar]
- Mutskov V, Felsenfeld G. Silencing of transgene transcription precedes methylation of promoter DNA and histone H3 lysine 9. EMBO J. 2004;23:138–149. doi: 10.1038/sj.emboj.7600013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagata K, Horinouchi M, Saitou M, Higashi M, Nomoto M, Goto M, et al. Mucin expression profile in pancreatic cancer and the precursor lesions. J Hepatobiliary Pancreat Surg. 2007;14:243–254. doi: 10.1007/s00534-006-1169-2. [DOI] [PubMed] [Google Scholar]
- Nakahara K, Carthew RW. Expanding roles for miRNAs and siRNAs in cell regulation. Curr Opin Cell Biol. 2004;16:127–133. doi: 10.1016/j.ceb.2004.02.006. [DOI] [PubMed] [Google Scholar]
- Nakamura A, Horinouchi M, Goto M, Nagata K, Sakoda K, Takao S, et al. New classification of pancreatic intraductal papillary-mucinous tumour by mucin expression: its relationship with potential for malignancy. J Pathol. 2002;197:201–210. doi: 10.1002/path.1109. [DOI] [PubMed] [Google Scholar]
- Nguyen CT, Weisenberger DJ, Velicescu M, Gonzales FA, Lin JC, Liang G, et al. Histone H3-lysine 9 methylation is associated with aberrant gene silencing in cancer cells and is rapidly reversed by 5-aza-2′-deoxycytidine. Cancer Res. 2002;62:6456–6461. [PubMed] [Google Scholar]
- O’Brien TJ, Beard JB, Underwood LJ, Dennis RA, Santin AD, York L. The CA 125 gene: an extracellular superstructure dominated by repeat sequences. Tumour Biol. 2001;22:348–366. doi: 10.1159/000050638. [DOI] [PubMed] [Google Scholar]
- O’Brien TJ, Beard JB, Underwood LJ, Shigemasa K. The CA 125 gene: a newly discovered extension of the glycosylated N-terminal domain doubles the size of this extracellular superstructure. Tumour Biol. 2002;23:154–169. doi: 10.1159/000064032. [DOI] [PubMed] [Google Scholar]
- Oka M, Meacham AM, Hamazaki T, Rodic N, Chang LJ, Terada N. De novo DNA methyltransferases Dnmt3a and Dnmt3b primarily mediate the cytotoxic effect of 5-aza-2′-deoxycytidine. Oncogene. 2005;24:3091–3099. doi: 10.1038/sj.onc.1208540. [DOI] [PubMed] [Google Scholar]
- Okudaira K, Kakar S, Cun L, Choi E, Wu Decamillis R, Miura S, et al. MUC2 gene promoter methylation in mucinous and non-mucinous colorectal cancer tissues. Int J Oncol. 2010;36:765–775. doi: 10.3892/ijo_00000552. [DOI] [PubMed] [Google Scholar]
- Osako M, Yonezawa S, Siddiki B, Huang J, Ho JJ, Kim YS, et al. Immunohistochemical study of mucin carbohydrates and core proteins in human pancreatic tumors. Cancer. 1993;71:2191–2199. doi: 10.1002/1097-0142(19930401)71:7<2191::AID-CNCR2820710705>3.0.CO;2-X. [DOI] [PubMed] [Google Scholar]
- Park HU, Kim JW, Kim GE, Bae HI, Crawley SC, Yang SC, et al. Aberrant expression of MUC3 and MUC4 membrane-associated mucins and sialyl Le(x) antigen in pancreatic intraepithelial neoplasia. Pancreas. 2003;26:e48–e54. doi: 10.1097/00006676-200304000-00022. [DOI] [PubMed] [Google Scholar]
- Patton S, Gendler SJ, Spicer AP. The epithelial mucin, MUC1, of milk, mammary gland and other tissues. Biochim Biophys Acta. 1995;1241:407–423. doi: 10.1016/0304-4157(95)00014-3. [DOI] [PubMed] [Google Scholar]
- Peng DF, Kanai Y, Sawada M, Ushijima S, Hiraoka N, Kosuge T, et al. Increased DNA methyltransferase 1 (DNMT1) protein expression in precancerous conditions and ductal carcinomas of the pancreas. Cancer Sci. 2005;96:403–408. doi: 10.1111/j.1349-7006.2005.00071.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perrais M, Pigny P, Buisine MP, Porchet N, Aubert JP, Seuningen-Lempire I. Aberrant expression of human mucin gene MUC5B in gastric carcinoma and cancer cells. Identification and regulation of a distal promoter. J Biol Chem. 2001;276:15386–15396. doi: 10.1074/jbc.M010534200. [DOI] [PubMed] [Google Scholar]
- Pinto-de-Sousa J, Reis CA, David L, Pimenta A, Cardoso-de-Oliveira M. MUC5B expression in gastric carcinoma: relationship with clinico-pathological parameters and with expression of mucins MUC1, MUC2, MUC5AC and MUC6. Virchows Arch. 2004;444:224–230. doi: 10.1007/s00428-003-0968-y. [DOI] [PubMed] [Google Scholar]
- Rajabi H, Jin C, Ahmad R, McClary C, Joshi MD, Kufe D. Mucin 1 oncoprotein expression is suppressed by the miR-125b Oncomir. Genes Cancer. 2010;1:62–68. doi: 10.1177/1947601909357933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rakha EA, Boyce RW, Abd El-Rehim D, Kurien T, Green AR, Paish EC, et al. Expression of mucins (MUC1, MUC2, MUC3, MUC4, MUC5AC and MUC6) and their prognostic significance in human breast cancer. Mod Pathol. 2005;18:1295–1304. doi: 10.1038/modpathol.3800445. [DOI] [PubMed] [Google Scholar]
- Rodriguez-Paredes M, Esteller M. Cancer epigenetics reaches mainstream oncology. Nat Med. 2011;17:330–339. doi: 10.1038/nm.2305. [DOI] [PubMed] [Google Scholar]
- Ronaghi M. Pyrosequencing sheds light on DNA sequencing. Genome Res. 2001;11:3–11. doi: 10.1101/gr.11.1.3. [DOI] [PubMed] [Google Scholar]
- Sachdeva M, Mo YY. MicroRNA-145 suppresses cell invasion and metastasis by directly targeting mucin 1. Cancer Res. 2010;70:378–387. doi: 10.1158/0008-5472.CAN-09-2021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sagara M, Yonezawa S, Nagata K, Tezuka Y, Natsugoe S, Xing PX, et al. Expression of mucin 1 (MUC1) in esophageal squamous-cell carcinoma: its relationship with prognosis. Int J Cancer. 1999;84:251–257. doi: 10.1002/(SICI)1097-0215(19990621)84:3<251::AID-IJC9>3.0.CO;2-7. [DOI] [PubMed] [Google Scholar]
- Saito Y, Jones PA. Epigenetic activation of tumor suppressor microRNAs in human cancer cells. Cell Cycle. 2006;5:2220–2222. doi: 10.4161/cc.5.19.3340. [DOI] [PubMed] [Google Scholar]
- Saitou M, Goto M, Horinouchi M, Tamada S, Nagata K, Hamada T, et al. MUC4 expression is a novel prognostic factor in patients with invasive ductal carcinoma of the pancreas. J Clin Pathol. 2005;58:845–852. doi: 10.1136/jcp.2004.023572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato N, Maitra A, Fukushima N, Heek NT, Matsubayashi H, Iacobuzio-Donahue CA, et al. Frequent hypomethylation of multiple genes overexpressed in pancreatic ductal adenocarcinoma. Cancer Res. 2003;63:4158–4166. [PubMed] [Google Scholar]
- Shi H, Wei SH, Leu YW, Rahmatpanah F, Liu JC, Yan PS, et al. Triple analysis of the cancer epigenome: an integrated microarray system for assessing gene expression, DNA methylation, and histone acetylation. Cancer Res. 2003;63:2164–2171. [PubMed] [Google Scholar]
- Shibahara H, Tamada S, Goto M, Oda K, Nagino M, Nagasaka T, et al. Pathologic features of mucin-producing bile duct tumors: two histopathologic categories as counterparts of pancreatic intraductal papillary-mucinous neoplasms. Am J Surg Pathol. 2004;28:327–338. doi: 10.1097/00000478-200403000-00005. [DOI] [PubMed] [Google Scholar]
- Shibahara H, Tamada S, Higashi M, Goto M, Batra SK, Hollingsworth MA, et al. MUC4 is a novel prognostic factor of intrahepatic cholangiocarcinoma-mass forming type. Hepatology. 2004;39:220–229. doi: 10.1002/hep.20031. [DOI] [PubMed] [Google Scholar]
- Shinojima Y, Terui T, Hara H, Kimura M, Igarashi J, Wang X, et al. Identification and analysis of an early diagnostic marker for malignant melanoma: ZAR1 intra-genic differential methylation. J Dermatol Sci. 2010;59:98–106. doi: 10.1016/j.jdermsci.2010.04.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siedow A, Szyf M, Gratchev A, Kobalz U, Hanski ML, Bumke-Vogt C, et al. De novo expression of the Muc2 gene in pancreas carcinoma cells is triggered by promoter demethylation. Tumour Biol. 2002;23:54–60. doi: 10.1159/000048689. [DOI] [PubMed] [Google Scholar]
- Singh AP, Chauhan SC, Bafna S, Johansson SL, Smith LM, Moniaux N, et al. Aberrant expression of transmembrane mucins, MUC1 and MUC4, in human prostate carcinomas. Prostate. 2006;66:421–429. doi: 10.1002/pros.20372. [DOI] [PubMed] [Google Scholar]
- Suh SO, Chen Y, Zaman MS, Hirata H, Yamamura S, Shahryari V et al. (2011) MicroRNA-145 is regulated by DNA methylation and p53 gene mutation in prostate cancer. Carcinogenesis, published online 23 Feb [DOI] [PMC free article] [PubMed]
- Sun W, Liu Y, Glazer CA, Shao C, Bhan S, Demokan S, et al. TKTL1 is activated by promoter hypomethylation and contributes to head and neck squamous cell carcinoma carcinogenesis through increased aerobic glycolysis and HIF1alpha stabilization. Clin Cancer Res. 2010;16:857–866. doi: 10.1158/1078-0432.CCR-09-2604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takao S, Uchikura K, Yonezawa S, Shinchi H, Aikou T. Mucin core protein expression in extrahepatic bile duct carcinoma is associated with metastases to the liver and poor prognosis. Cancer. 1999;86:1966–1975. doi: 10.1002/(SICI)1097-0142(19991115)86:10<1966::AID-CNCR13>3.0.CO;2-M. [DOI] [PubMed] [Google Scholar]
- Tamada S, Goto M, Nomoto M, Nagata K, Shimizu T, Tanaka S, et al. Expression of MUC1 and MUC2 mucins in extrahepatic bile duct carcinomas: its relationship with tumor progression and prognosis. Pathol Int. 2002;52:713–723. doi: 10.1046/j.1440-1827.2002.01414.x. [DOI] [PubMed] [Google Scholar]
- Tamada S, Shibahara H, Higashi M, Goto M, Batra SK, Imai K, et al. MUC4 is a novel prognostic factor of extrahepatic bile duct carcinoma. Clin Cancer Res. 2006;12:4257–4264. doi: 10.1158/1078-0432.CCR-05-2814. [DOI] [PubMed] [Google Scholar]
- Toribara NW, Roberton AM, Ho SB, Kuo WL, Gum E, Hicks JW, et al. Human gastric mucin. Identification of a unique species by expression cloning. J Biol Chem. 1993;268:5879–5885. [PubMed] [Google Scholar]
- Tsutsumida H, Goto M, Kitajima S, Kubota I, Hirotsu Y, Wakimoto J, et al. MUC4 expression correlates with poor prognosis in small-sized lung adenocarcinoma. Lung Cancer. 2007;55:195–203. doi: 10.1016/j.lungcan.2006.10.013. [DOI] [PubMed] [Google Scholar]
- Utsunomiya T, Yonezawa S, Sakamoto H, Kitamura H, Hokita S, Aiko T, et al. Expression of MUC1 and MUC2 mucins in gastric carcinomas: its relationship with the prognosis of the patients. Clin Cancer Res. 1998;4:2605–2614. [PubMed] [Google Scholar]
- Seuningen I, Vincent A. Mucins: a new family of epigenetic biomarkers in epithelial cancers. Expert Opinion. 2009;3:411–427. doi: 10.1517/17530050902852697. [DOI] [PubMed] [Google Scholar]
- Seuningen I, Pigny P, Perrais M, Porchet N, Aubert JP. Transcriptional regulation of the 11p15 mucin genes. Towards new biological tools in human therapy, in inflammatory diseases and cancer? Front Biosci. 2001;6:D1216–D1234. doi: 10.2741/Seuning. [DOI] [PubMed] [Google Scholar]
- Velcich A, Yang W, Heyer J, Fragale A, Nicholas C, Viani S, et al. Colorectal cancer in mice genetically deficient in the mucin Muc2. Science. 2002;295:1726–1729. doi: 10.1126/science.1069094. [DOI] [PubMed] [Google Scholar]
- Vincent A, Perrais M, Desseyn JL, Aubert JP, Pigny P, Seuningen I. Epigenetic regulation (DNA methylation, histone modifications) of the 11p15 mucin genes (MUC2, MUC5AC, MUC5B, MUC6) in epithelial cancer cells. Oncogene. 2007;26:6566–6576. doi: 10.1038/sj.onc.1210479. [DOI] [PubMed] [Google Scholar]
- Vincent A, Ducourouble MP, Seuningen I. Epigenetic regulation of the human mucin gene MUC4 in epithelial cancer cell lines involves both DNA methylation and histone modifications mediated by DNA methyltransferases and histone deacetylases. FASEB J. 2008;22:3035–3045. doi: 10.1096/fj.07-103390. [DOI] [PubMed] [Google Scholar]
- Wang RQ, Fang DC. Alterations of MUC1 and MUC3 expression in gastric carcinoma: relevance to patient clinicopathological features. J Clin Pathol. 2003;56:378–384. doi: 10.1136/jcp.56.5.378. [DOI] [PMC free article] [PubMed] [Google Scholar]
- White CL, Suto RK, Luger K. Structure of the yeast nucleosome core particle reveals fundamental changes in internucleosome interactions. EMBO J. 2001;20:5207–5218. doi: 10.1093/emboj/20.18.5207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams SJ, McGuckin MA, Gotley DC, Eyre HJ, Sutherland GR, Antalis TM. Two novel mucin genes down-regulated in colorectal cancer identified by differential display. Cancer Res. 1999;59:4083–4089. [PubMed] [Google Scholar]
- Wolff EM, Byun HM, Han HF, Sharma S, Nichols PW, Siegmund KD, et al. Hypomethylation of a LINE-1 promoter activates an alternate transcript of the MET oncogene in bladders with cancer. PLoS Genet. 2010;6:e1000917. doi: 10.1371/journal.pgen.1000917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolffe AP, Matzke MA. Epigenetics: regulation through repression. Science. 1999;286:481–486. doi: 10.1126/science.286.5439.481. [DOI] [PubMed] [Google Scholar]
- Yamada N, Hamada T, Goto M, Tsutsumida H, Higashi M, Nomoto M, et al. MUC2 expression is regulated by histone H3 modification and DNA methylation in pancreatic cancer. Int J Cancer. 2006;119:1850–1857. doi: 10.1002/ijc.22047. [DOI] [PubMed] [Google Scholar]
- Yamada N, Nishida Y, Tsutsumida H, Hamada T, Goto M, Higashi M, et al. MUC1 expression is regulated by DNA methylation and histone H3 lysine 9 modification in cancer cells. Cancer Res. 2008;68:2708–2716. doi: 10.1158/0008-5472.CAN-07-6844. [DOI] [PubMed] [Google Scholar]
- Yamada N, Nishida Y, Tsutsumida H, Goto M, Higashi M, Nomoto M, et al. Promoter CpG methylation in cancer cells contributes to the regulation of MUC4. Br J Cancer. 2009;100:344–351. doi: 10.1038/sj.bjc.6604845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamada N, Nishida Y, Yokoyama S, Tsutsumida H, Houjou I, Kitamoto S, et al. Expression of MUC5AC, an early marker of pancreatobiliary cancer, is regulated by DNA methylation in the distal promoter region in cancer cells. J Hepatobiliary Pancreat Sci. 2010;17:844–854. doi: 10.1007/s00534-010-0278-0. [DOI] [PubMed] [Google Scholar]
- Yamamoto T, Ikawa S, Akiyama T, Semba K, Nomura N, Miyajima N, et al. Similarity of protein encoded by the human c-erb-B-2 gene to epidermal growth factor receptor. Nature. 1986;319:230–234. doi: 10.1038/319230a0. [DOI] [PubMed] [Google Scholar]
- Yamashita K, Yonezawa S, Tanaka S, Shirahama H, Sakoda K, Imai K, et al. Immunohistochemical study of mucin carbohydrates and core proteins in hepatolithiasis and cholangiocarcinoma. Int J Cancer. 1993;55:82–91. doi: 10.1002/ijc.2910550116. [DOI] [PubMed] [Google Scholar]
- Yin BW, Lloyd KO. Molecular cloning of the CA125 ovarian cancer antigen: identification as a new mucin, MUC16. J Biol Chem. 2001;276:27371–27375. doi: 10.1074/jbc.M103554200. [DOI] [PubMed] [Google Scholar]
- Yonezawa S, Sato E. Expression of mucin antigens in human cancers and its relationship with malignancy potential. Pathol Int. 1997;47:813–830. doi: 10.1111/j.1440-1827.1997.tb03713.x. [DOI] [PubMed] [Google Scholar]
- Yonezawa S, Sueyoshi K, Nomoto M, Kitamura H, Nagata K, Arimura Y, et al. MUC2 gene expression is found in noninvasive tumors but not in invasive tumors of the pancreas and liver: its close relationship with prognosis of the patients. Hum Pathol. 1997;28:344–352. doi: 10.1016/S0046-8177(97)90134-9. [DOI] [PubMed] [Google Scholar]
- Yonezawa S, Horinouchi M, Osako M, Kubo M, Takao S, Arimura Y, et al. Gene expression of gastric type mucin (MUC5AC) in pancreatic tumors: its relationship with the biological behavior of the tumor. Pathol Int. 1999;49:45–54. doi: 10.1046/j.1440-1827.1999.00823.x. [DOI] [PubMed] [Google Scholar]
- Yonezawa S, Goto M, Yamada N, Higashi M, Nomoto M. Expression profiles of MUC1, MUC2, and MUC4 mucins in human neoplasms and their relationship with biological behavior. Proteomics. 2008;8:3329–3341. doi: 10.1002/pmic.200800040. [DOI] [PubMed] [Google Scholar]
- Yonezawa S, Higashi M, Yamada N, Yokoyama S, Goto M. Significance of mucin expression in pancreatobiliary neoplasms. J Hepatobiliary Pancreat Sci. 2010;17:108–124. doi: 10.1007/s00534-009-0174-7. [DOI] [PubMed] [Google Scholar]
- Zen Y, Sasaki M, Fujii T, Chen TC, Chen MF, Yeh TS, et al. Different expression patterns of mucin core proteins and cytokeratins during intrahepatic cholangiocarcinogenesis from biliary intraepithelial neoplasia and intraductal papillary neoplasm of the bile duct—an immunohistochemical study of 110 cases of hepatolithiasis. J Hepatol. 2006;44:350–358. doi: 10.1016/j.jhep.2005.09.025. [DOI] [PubMed] [Google Scholar]
- Zhang L, Volinia S, Bonome T, Calin GA, Greshock J, Yang N, et al. Genomic and epigenetic alterations deregulate microRNA expression in human epithelial ovarian cancer. Proc Natl Acad Sci USA. 2008;105:7004–7009. doi: 10.1073/pnas.0801615105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu Y, Zhang JJ, Zhu R, Liang WB, Gao WT, Yu JB et al. (2010) The increase in the expression and hypomethylation of MUC4 gene with the progression of pancreatic ductal adenocarcinoma. Med Oncol, published online 5 Oct [DOI] [PubMed]
- Zilberman D, Henikoff S. Genome-wide analysis of DNA methylation patterns. Development. 2007;134:3959–3965. doi: 10.1242/dev.001131. [DOI] [PubMed] [Google Scholar]
- Zrihan-Licht S, Weiss M, Keydar I, Wreschner DH. DNA methylation status of the MUC1 gene coding for a breast-cancer-associated protein. Int J Cancer. 1995;62:245–251. doi: 10.1002/ijc.2910620303. [DOI] [PubMed] [Google Scholar]


