Abstract
The chloramphenicol producer Streptomyces venezuelae contains an enzyme, SvTrpEG, that has a high degree of amino acid sequence similarity to the phenazine biosynthetic enzyme PhzE of certain species of Pseudomonas. PhzE has the sequence signature of an anthranilate synthase, but recent evidence indicates that it catalyzes the production of 2-amino-2-deoxyisochorismate (ADIC), an intermediate in the two-step anthranilate synthase reaction, not anthranilate. In order to determine if SvTrpEG is likewise an ADIC synthase, we have cloned the gene for SvTrpEG, expressed the recombinant enzyme in Escherichia coli, and purified the enzyme. Analysis of the SvTrpEG-catalyzed reaction mixture using UV-visible spectrophotometry, fluorescence spectrometry, and high-performance liquid chromatography shows that the product of the reaction is anthranilate, not ADIC. Our results therefore reveal that, despite its sequence similarity to PhzE, SvTrpEG is an anthranilate synthase, not an ADIC synthase.
Keywords: Anthranilate Synthase, TrpEG, Streptomyces venezuelae, 2-Amino-2-Deoxyisochorismate (ADIC) Synthase
Introduction
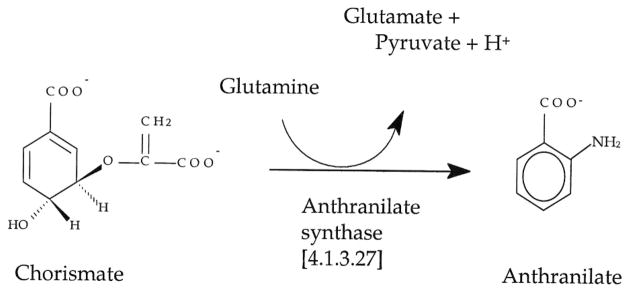
The anthranilate synthase-catalyzed reaction of the tryptophan biosynthetic pathway (Figure 1) is a two-step process.1 The first step involves the transfer of an amido group from glutamine bound to a glutamine amidotransferase (TrpG) subunit to chorismate bound to an anthranilate synthase (TrpE) subunit (and loss of a hydroxyl group from chorismate), forming the intermediate 2-amino-2-deoxyisochorismate (ADIC). The second step utilizes an ADIC lyase activity in TrpE to remove a pyruvate group (and a proton) from ADIC, generating the aromatic compound anthranilate. Anthranilate synthase is feedback inhibited by L-tryptophan. In most cases, the enzyme functions as a heterotetramer, TrpE2TrpG2, made up of separate (non-fused) TrpE and TrpG subunits. Anthranilate synthase enzymes from a number of microorganisms, including Salmonella typhimurium,2,3 Serratia marcescens,4 Solobus solfataricus,5,6 and Archaeoglobus fulgidus,7 have been studied.
Fig 1.
The anthranilate synthase-catalyzed reaction. Anthranilate, the product of the reaction, absorbs at ≈312 nm and fluoresces at 395 nm.
Phenazine compounds that have antibacterial or cytotoxic properties are produced by a number of species of Pseudomonas and Streptomyces. A seven-gene locus (phzABCDEFG) for the synthesis of the compound phenazine-1-carboxylic acid has been identified in the fluorescent soil-dwelling bacterium Pseudomonas fluorescens 2-79.8 Similar genetic loci have been identified in P. aureofaciens9 and in P. aeruginosa,10 an opportunistic pathogen that infects and forms difficult to treat biofilms in cystic fibrosis patients. Within the seven-gene loci is a gene, phzE (or phzB in P. aureofaciens), encoding a protein with amino acid sequence similarity to an anthranilate synthase. The N-terminal two thirds of the PhzE (and PhzB) protein has sequence homology to the TrpE subunit of anthranilate synthase, and the C-terminal one third has homology to TrpG. With its fused TrpE and TrpG components (TrpEG), PhzE most closely resembles a set of similarly-fused anthranilate synthases found in Streptomyces venezuelae,11 Streptomyces coelicolor,12 and some nitrogen-fixing bacteria.13,14
At the time that the fused trpEG gene was discovered in S. venezuelae, it was not clear whether it encoded a bona fide anthranilate synthase or a chorismate-utilizing enzyme that might catalyze the production of a phenazine precursor.15 (Presently, no evidence indicates that S. venezuelae makes a phenazine compound.) Lending support to this second possibility was the fact that S. venezuelae TrpEG (SvTrpEG) had a much higher sequence similarity to PhzE (62%) than to the TrpE and TrpG subunits of the anthranilate synthase from Escherichia coli (≈42%). (See Figure 2 for a sequence alignment of SvTrpEG with PhzE and other homologous enzymes.) A recent report showed that the phenazine precursor is ADIC, the intermediate in the anthranilate synthase reaction,17 and unpublished work (Dr. T-W Yu, personal communication) indicated that PhzE is an ADIC synthase. The question then became whether SvTrpEG might be an ADIC synthase also. The work reported here was designed to answer this question.
Fig 2.
Multiple sequence alignment among P. fluorescens PhzE, P. aureofaciens PhzB, P. aeruginosa PhzE, S. venezuelae TrpEG, and S. coelicolor TrpEG. Catalysis: Residues important for catalysis in Salmonella typhimurium anthranilate synthase (StAS)2 are indicated at their predicted positions (based on alignment) using the upper-case single-letter code for the amino acid; residues implicated based on x-ray data are indicated with a lowercase single-letter code.4 Regulation: Residues important for tryptophan inhibition of StAS are indicated in bold uppercase.16 The polypeptide linker between putative TrpE and TrpG domains is also indicated. Numbering (superscripts) is that of StAS. The C-terminal end of TrpG has been truncated in the figure: all sequences extend for another 70 or so amino acids.
Methods
Polymerase Chain Reaction (PCR) and Cloning
Using nucleotide sequences available in GenBank, PCR primers were designed and used to amplify the gene for SvTrpEG from a plasmid (pDQ187)15 generously provided by Dr. Leo Vining of Dalhousie University. The forward primer had the sequence 5′-GGAACGACGGACCATATGGACCTCTC-3′ and contained an Nde I restriction site (underlined). The reverse primer had the sequence 5′-GGCGTCCGGGATCCTAGACCAGCAC-3′ and contained a BamH1 site (underlined). The resulting 1.9 kb PCR product was cloned by means of its Nde1 and BamH1 sites into the expression plasmid pET-15b (Novagen), which generates a His-tagged fusion protein upon induction of expression. The coding region of svtrpEG within the recombinant plasmid was sequenced to confirm that no mutations were introduced by the PCR. (None were, and the sequence matched the sequence for svtrpEG in GenBank, accession number AF012627).
Protein Expression and Purification
The svtrpEG/pET-15b plasmid was transformed into the E. coli expression strain BL21(DE3)/pLysS (Novagen). Fresh transformants containing the recombinant plasmid were grown at 37°C in overnight LB cultures containing ampicillin (100 μg/mL) and chloramphenicol (25 μg/mL). These were used to inoculate larger LB cultures containing ampicillin (100 μg/mL), chloramphenicol (25 μg/mL), sorbitol (1 M), and betaine (2.5 mM). The cultures were initially grown at 37°C but were shifted to a lower temperature (22°C) and induced with isopropylthio-galactoside (IPTG, 100 μM) when cell density reached OD600 of 0.6. They were grown for an additional 15 hours after induction. The cells were then harvested, resuspended in buffer A (lysis buffer: 100 mM Tris-Cl, pH 8.0; 10% glycerol; 1 mM EDTA; 1 mM dithio-threitol [DTT]; 1 mM phenylmethyl-sulfonyl fluoride [PMSF]) 25 units/mL Benzonase nuclease [Novagen]; and 15 mM imidazole), and broken open by using sonication. Cell debris was removed by centrifugation, and the crude extract was analyzed for expression of SvTrpEG by using sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and by Western blotting with an anti-His tag antibody. The crude extract containing overexpressed protein was loaded onto a Ni-NTA spin column (Qiagen) that had been equilibrated with buffer B (50 mM Tris-HCl, pH 7.5; 300 mM NaCl; 5 mM β-mercaptoethanol; and 5% glycerol) containing 20 mM imidazole. The loaded column was washed sequentially with buffer B containing 30 mM imidazole and 100 mM imidazole. The His-tagged SvTrpEG was eluted with 250 mM imidazole in buffer B, and the eluant was analyzed for purity by using SDS-PAGE. This final enzyme preparation was desalted into buffer C (lysis buffer lacking imidazole, nuclease and EDTA), dialyzed in this same buffer containing 50% glycerol, and stored at −20°C.
Activity Assay
The standard 2-mL activity assay contained chorismate (100 μM), glutamine (7.0 mM), MgCl2 (12.5 mM) and SvTrpEG enzyme (10 μg) in 50 mM Tricine solution buffered at pH 7.5. A Fluoromax-3 photon-counting fluorescence spectrometer (Jobin-Yvonne) was used for measuring production of anthranilate over time. The excitation and emission wavelengths were set at 312 nm and 395 nm, respectively (slit widths 2 nm). Protein concentrations were determined by using the BioRad protein reagent.
Determination of Chorismate Concentration in Assay
Since chorismate degrades slowly over time, and since the commercially-available preparation of chorismate (Sigma) is not 100% pure, we sought to determine accurately the concentration of the chorismate in the activity assay. To do this, we determined (in triplicate), based on the area of the 395 nm peak in the fluorescence spectrum of the reaction product and using an anthranilate standard curve, the amount of anthranilate formed in a three-hour reaction at 35°C. Then, using the 1:1 stoichiometric relationship between anthranilate and chorismate, we calculated the amount of chorismate that had been present in the assay. This calculation allowed us to correct the concentration of the stock chorismate solution that had been used. We were confident that all of the chorismate originally present in the assay had been converted to anthranilate because the equilibrium for the reaction lies far to the right (K′eq≈5×1030),18 and the reaction was essentially complete after three hours (this study).
Results
His-tagged recombinant SvTrpEG was expressed in E. coli and purified by using Ni-NTA affinity chromatography. As can be seen from the SDS-PAGE analysis in Figure 3, the purified recombinant SvTrpEG enzyme preparation gave a single band corresponding to a protein of molecular weight 67 kDa. This matches the molecular weight calculated from the predicted amino acid sequence of the recombinant protein (including the His tag). We sought to determine if SvTrpEG was an ADIC synthase, as expected based on its sequence similarity to PhzE, or an anthranilate synthase. As can be seen in Figure 4, the SvTrpEG-catalyzed reaction produces anthranilate, not ADIC. There are several lines of evidence supporting this conclusion. First, the product of the SvTrpEG-catalyzed reaction gave an absorbance spectrum similar to that of a standard solution of anthranilate. Like anthranilate, it has an absorbance maximum at ≈312 nm (Figure 4A). Notably, ADIC absorbs at a different wavelength, 278 nm.1 Second, a compound that fluoresces at 395 nm after being excited at 312 nm accumulated over time in the reaction mixture (Figure 4B, inset). The fluorescence spectrum of this compound is identical to that of an anthranilate standard (Figure 4B). Whereas anthranilate is known to fluoresce at 395 nm, ADIC is not a fluorescent compound.1 Third, when the SvTrpEG-catalyzed reaction mixture was analyzed by using reversed-phase high-performance liquid chromatography, the fluorescent product that formed had a retention time that was identical to that of a standard solution of anthranilate, but not ADIC (data not shown). Finally, the anthranilate-producing activity of SvTrpEG was found to be directly proportional to the amount of enzyme added in the activity assay (not shown); this result would be the expected one for an enzyme-mediated catalytic process. Altogether, these results demonstrate that SvTrpEG catalyzes a reaction that produces anthranilate. Therefore, SvTrpEG is an anthranilate synthase, not an ADIC synthase.
Fig 3.
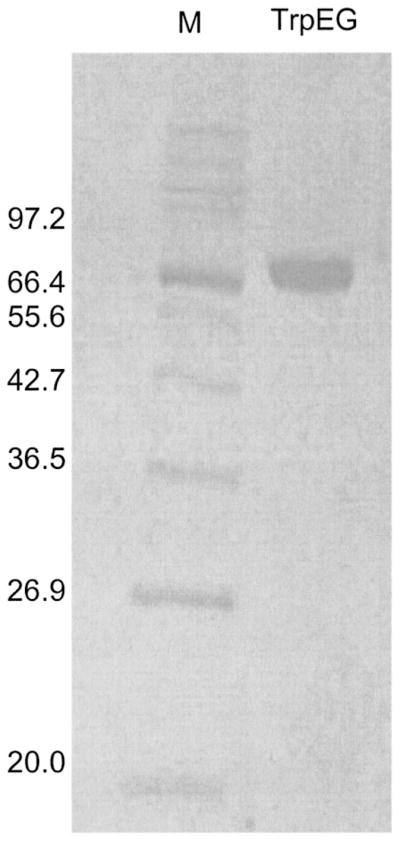
SDS-PAGE analysis of the purified SvTrpEG enzyme. M = molecular mass standards. TrpEG = 20 μg of purified SvTrpEG. Molecular masses in kDa are indicated on the left. SvTrpEG migrates at ≈67 kDa.
Fig 4.
Analysis of the SvTrpEG-catalyzed reaction: A) by UV-visible spectrophotometry. Top line, UV-visible spectrum of the SvTrpEG-catalyzed reaction mixture after 1 hour of reaction at 35°C. Bottom line, anthranilate standard. Both have an absorbance maximum near 312 nm. Note that ADIC absorbs maximally at 278 nm;1 B by fluorescence spectroscopy (excitation at 312 nm, emission at 395 nm). Time course showing a linear increase in fluorescence intensity at 395 nm (in counter per second) over time (in seconds). The slope of the line represents the initial velocity under the conditions of the assay.
Discussion
Lin et al,15 who studied the expression of the fused TrpEG from S. venezuelae in an E. coli TrpE auxotroph, were not able to conclusively establish that the enzyme was an anthranilate synthase. Our results demonstrate that it is and therefore show that a fused anthranilate synthase is present in S. venezuelae. At the time, it was also uncertain whether or not there were two anthranilate synthase-like enzymes in S. venezuelae, one involved in tryptophan biosynthesis and the other involved somehow in the biosynthesis of a secondary metabolite.15 This issue is still not settled.
The question remains, then, as to what might be the physiological role of the fused anthranilate synthase in S. venezuelae. Is it involved in the production of a precursor of a secondary metabolite derived from chorismate? Or, does it play a role in primary metabolism, catalyzing a step in the biosynthesis of tryptophan? In trying to elucidate the role of the enzyme, it will be important to determine whether or not it is feedback inhibited by tryptophan. Generally, enzymes involved in primary metabolism (tryptophan biosynthesis) are inhibited, while those of secondary metabolism are not. It is also noteworthy that, although a chorismate-utilizing enzyme involved in the biosynthesis of chloramphenicol in S. venezuelae has been identified, this enzyme was not SvTrpEG. It was rather a 4-Amino-4-deoxychorismate (ADC) synthase that is encoded by a pabAB gene present within the chloramphenicol biosynthetic gene cluster recently identified.19 Thus, the role of TrpEG in S. venezuelae primary and secondary metabolism remains to be established.
Future work from our laboratory will focus on the characterization of SvTrpEG in terms of its steady-state kinetic properties, including possible feedback inhibition by tryptophan, and its oligomerization state. Since SvTrpEG, although similar in amino acid sequence to PhzE, is an anthranilate synthase and therefore has ADIC lyase activity whereas PhzE does not, it will be of interest to identify which residues in SvTrpEG confer this additional lyase activity. (See Figure 2 for the residues involved in catalysis and feedback inhibition of S. typhimurium anthranilate synthase). Site-directed mutagenesis and the construction of chimeric TrpEG/PhzE enzymes will be used to address this question.
In conclusion, the streptomycetes are the primary producers of currently available antibiotics used to treat bacterial infections. The model streptomycete Streptomyces coelicolor, whose genome was sequenced recently,12 contains a fused TrpEG similar to the one present in S. venezuelae. For this reason, an understanding of the structure and function of SvTrpEG and its role in metabolism will indirectly give insights into the microbial physiology of S. coelicolor and of streptomycetes in general. A deeper understanding of metabolism in streptomycetes will, in turn, aid in the discovery of new antibiotics, which will benefit all people, but especially poor and minority populations, who often are most adversely affected by infectious disease.
Acknowledgments
This work was supported by grants G12 RR003048-18 and S06 GM08016-34 funded through the NIH RCMI and MBRS-SCORE programs, respectively.
References
- 1.Morollo AA, Bauerle R. Characterization of composite aminodeoxyisochorismate synthase and aminodeoxyisochorismate lyase activities of anthranilate synthase. Proc Natl Acad Sci U S A. 1993;90(21):9983–9987. doi: 10.1073/pnas.90.21.9983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bauerle R, Hess J, French S. Anthranilate synthase-anthranilate posphoribosyltransferase complex and subunits of Salmonella typhimurium. Methods Enzymol. 1987;142:366–386. doi: 10.1016/s0076-6879(87)42049-1. [DOI] [PubMed] [Google Scholar]
- 3.Morollo AA, Eck MJ. Structure of the cooperative allosteric anthranilate synthase from Salmonella typhimurium. Nat Struct Biol. 2001;8(3):243–247. doi: 10.1038/84988. [DOI] [PubMed] [Google Scholar]
- 4.Spraggon G, Kim C, Nhuyen-Huu X, Yee MC, Yanofsky C, Mills SE. The structure of anthranilate synthase of Serratia marcescens crystallized in the presence of (i) its substrates, chorismate and glutamine, and a product, glutamate, and (ii) its end-product inhibitor, L-tryptophan. Proc Natl Acad Sci U S A. 2001;98(11):6021–6026. doi: 10.1073/pnas.111150298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Tutino ML, Tosco A, Marino G, Sannia G. Expression of Sulfolobus solfataricus trpE and trpG genes in E. coli. Biochem Biophys Res Commun. 1997;230(2):306–310. doi: 10.1006/bbrc.1996.5951. [DOI] [PubMed] [Google Scholar]
- 6.Knöchel T, Ivens A, Hester G, et al. The crystal structure of anthranilate synthase from Sulfolobus solfataricus: functional implications. Proc Natl Acad Sci U S A. 1999;96(17):9479–9484. doi: 10.1073/pnas.96.17.9479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Byrnes WM, Vilker VL. Extrinsic factors potassium chloride and glycerol induce thermostability in recombinant anthranilate synthase from Archaeoglobus fulgidus. Extremophiles. 2004;8(6):455–462. doi: 10.1007/s00792-004-0406-3. [DOI] [PubMed] [Google Scholar]
- 8.Mavrodi DV, Ksenzenko VN, Bonsall RF, Cook RJ, Boronin AM, Thomashow LS. A seven-gene locus for synthesis of phenazine-1-carboxylic acid by Pseudomonas fluorescens 2–79. J Bacteriol. 1998;180(9):2541–2548. doi: 10.1128/jb.180.9.2541-2548.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Pierson LS, Gaffney T, Lam S, Gong F. Molecular analysis of genes encoding phenazine biosynthesis in the biological control bacterium Pseudomonas aureofaciens 30–84. FEMS Microbiol Lett. 1995;134(1–2):299–307. doi: 10.1111/j.1574-6968.1995.tb07954.x. [DOI] [PubMed] [Google Scholar]
- 10.Mavrodi DV, Bonsall RF, Delaney SM, Soule MJ, Phillips G, Thomashow LS. Functional analysis of genes for biosynthesis of pyocyanin and phenazine-1-carboxamide from Pseudomonas aeruginosa PAO1. J Bacteriol. 2001;183(21):6454–6465. doi: 10.1128/JB.183.21.6454-6465.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Paradkar AS, Stuttard C, Vining LC. Molecular cloning of the genes for anthranilate synthetase from Streptomyces venezuelae ISP 5230. FEMS Microbiol Lett. 1991;62(2–3):177–181. doi: 10.1016/0378-1097(91)90154-3. [DOI] [PubMed] [Google Scholar]
- 12.Bentley SD, Chater KF, Cerdeno-Tarraga AM, et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2) Nature. 2002;417(6885):141–147. doi: 10.1038/417141a. [DOI] [PubMed] [Google Scholar]
- 13.Bae YM, Holmgren E, Crawford IP. Rhizobium meliloti anthranilate synthase gene: cloning, sequence, and expression in Escherichia coli. J Bacteriol. 1989;171(6):3471–3478. doi: 10.1128/jb.171.6.3471-3478.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.DeTroch P, Dosselaere F, Keijers V, deWilde P, Vanderleyden J. Isolation and characterization of the Azospirillum brasilense trpE(G) gene, encoding anthranilate synthase. Curr Microbiol. 1997;34(1):27–32. doi: 10.1007/s002849900139. [DOI] [PubMed] [Google Scholar]
- 15.Lin C, Paradkar AS, Vining LC. Regulation of an anthranilate synthase gene in Streptomyces venezuelae by a trp attenuator. Microbiology. 1998;144:1971–1980. doi: 10.1099/00221287-144-7-1971. [DOI] [PubMed] [Google Scholar]
- 16.Caligiuri MG, Bauerle R. Identification of amino acid residues in feedback regulation of the anthranilate synthase complex from Salmonella typhimurium. Evidence for an amino-terminal regulatory site. J Biol Chem. 1991;266(13):8328–8335. [PubMed] [Google Scholar]
- 17.McDonald M, Mavrodi KV, Thomashow LS, Floss HG. Phenazine biosynthesis in Pseudomonas fluorescens: branchpoint from the primary shikimate biosynthetic pathway and role of phenzaine-1,6-dicarboxylic acid. J Am Chem Soc. 2001;123:9459–9460. doi: 10.1021/ja011243+. [DOI] [PubMed] [Google Scholar]
- 18.Byrnes WM, Goldberg RN, Holden MJ, Mayhew MP, Tewari YB. Thermodynamics of reactions catalyzed by anthranilate synthase. Biophys Chem. 2000;84:45–64. doi: 10.1016/s0301-4622(99)00145-3. [DOI] [PubMed] [Google Scholar]
- 19.He J, Magarvey N, Piraee M, Vining LC. The gene cluster for chloramphenicol biosynthesis in Streptomyces venezuelae ISP5230 includes novel shikimate pathway homologues and a monomodular non-ribosomal peptide synthetase gene. Microbiology. 2001;147:2817–2829. doi: 10.1099/00221287-147-10-2817. [DOI] [PubMed] [Google Scholar]