Abstract
Summary: Francisella tularensis is a Gram-negative intracellular pathogen and the causative agent of the disease tularemia. Inhalation of as few as 10 bacteria is sufficient to cause severe disease, making F. tularensis one of the most highly virulent bacterial pathogens. The initial stage of infection is characterized by the “silent” replication of bacteria in the absence of a significant inflammatory response. Francisella achieves this difficult task using several strategies: (i) strong integrity of the bacterial surface to resist host killing mechanisms and the release of inflammatory bacterial components (pathogen-associated molecular patterns [PAMPs]), (ii) modification of PAMPs to prevent activation of inflammatory pathways, and (iii) active modulation of the host response by escaping the phagosome and directly suppressing inflammatory pathways. We review the specific mechanisms by which Francisella achieves these goals to subvert host defenses and promote pathogenesis, highlighting as-yet-unanswered questions and important areas for future study.
INTRODUCTION
Francisella tularensis was first identified as the causative agent of a fatal plague-like disease in a population of ground squirrels in Tulare County, CA, in 1911 (147). Originally called Bacterium tularense, it was later renamed Francisella tularensis in honor of Edward Francis, who spent his career extensively studying and characterizing the transmission and growth of this bacterium (209). Although it causes disease in squirrels, rabbits, and numerous other mammals, no animal has been identified as a reservoir. Instead, the reservoir may be freshwater or amoebae living therein. As there is no person-to-person spread, F. tularensis is acquired primarily by humans via arthropod vectors or zoonotic transmission, though it can also be transmitted by inhalation of aerosolized bacteria or ingestion of contaminated food or water (2). Inhalation of F. tularensis causes the most severe infections, and only 10 bacteria can lead to a potentially fatal disease. This high infectivity, along with its ease of aerosolization, have led to its history of weaponization (209).
Francisella species are endemic only in the Northern Hemisphere. F. tularensis subsp. tularensis (F. tularensis) is the most virulent etiologic agent of tularemia in humans and is the primary disease-causing Francisella species in North America. Francisella tularensis subsp. holarctica (F. holarctica) is responsible for the majority of reported cases of tularemia in Europe and Asia. The current vaccine is an attenuated live vaccine strain (LVS) derived from virulent F. holarctica by serial passage. LVS causes a very mild infection in humans but can cause a lethal infection in mice and is therefore commonly used as a model to study Francisella pathogenesis. The closely related Francisella novicida species rarely causes disease in humans, though some cases have been documented (31, 125). However, F. novicida is highly virulent in mice, has over 98% identity to F. tularensis at the DNA level (188), shares many of the same virulence genes (43), and is also used as a model system to study Francisella virulence. Finally, Francisella tularensis subsp. mediasiatica is a species of intermediate virulence in humans and is found in Central Asia, while Francisella philomiragia and Francisella noatunensis can cause infections in aquatic organisms, including wild and farmed fish (57). Throughout this paper we will refer to “Francisella” when discussing general characteristics shared by numerous species and subspecies and will otherwise refer to specific species and subspecies by name. It is important to note that there are significant differences between highly virulent and less pathogenic strains in terms of the requirements of genes for pathogenesis, susceptibilities to host defenses, and the types of immune responses induced. Therefore, caution must be used when interpreting results from experiments using less pathogenic species and drawing conclusions about the characteristics of highly virulent species.
F. tularensis subspecies are the etiological agents of the disease tularemia, also known as rabbit fever. Tularemia is characterized by a 3- to 5-day incubation period (209) during which the bacteria replicate almost “silently” in macrophages and other types of host cells. The eventual release of bacteria from these cells may coincide with the presentation of flu-like symptoms. There are several manifestations of tularemia, each dependent on the route of acquisition (159). The most common form of tularemia is ulceroglandular disease, which can result from insect bites or from contact with infected animal tissues following mechanical damage to the skin. A cutaneous ulcer develops at the site of infection, and bacteria drain to lymph nodes, subsequently causing a systemic infection. Less common forms of the disease include pneumonic, oculoglandular, and oropharyngeal tularemia. Streptomycin or doxycycline is indicated for treatment. Tularemia may be fatal; however, survivors gain robust immunity that has been found to last for up to 30 years (79).
Upon infection, Francisella initially comes into contact with extracellular defenses such as complement, antibody, and cationic antimicrobial peptides (28, 29, 51, 190). Binding of these components to bacteria directly or indirectly leads to lysis and killing (189). Therefore, Francisella uses multiple surface structures and outer membrane modifications (capsule, lipopolysaccharide [LPS] O antigen, modifications that increase surface charge, etc.) to resist these components and block killing. In addition, this prevents structural damage that would release proinflammatory bacterial components capable of initiating a strong immune response. Francisella also enters host cells as an efficient way of evading extracellular defenses.
After engulfment by phagocytic cells, including macrophages, Francisella is taken up into phagosomes that contain an array of toxic antimicrobials aimed at degrading the bacteria (Fig. 1). However, this pathogen has an equally diverse cache of defenses to counteract host antimicrobials. These once again prevent not only killing but also the release of proinflammatory bacterial components that could be recognized by host innate immune receptors (including Toll-like receptors [TLRs]) that stimulate inflammatory responses. Furthermore, similarly to entering host cells to avoid extracellular antimicrobials, Francisella escapes the phagosome to avoid phagosomal antimicrobials and, importantly, reach the cytosol, where it can replicate (Fig. 1). The cytosol is also, however, guarded by innate recognition and defense systems (including the inflammasome) with which the bacteria must contend. In order to replicate in this host compartment, Francisella must also obtain the nutrients required to sustain its rapid cell division and actively counteract host defenses aimed at limiting nutrient availability.
Fig 1.
Stages of Francisella pathogenesis in the macrophage. Francisella can be detected by multiple macrophage receptors (see “Mechanisms of Entry and Fate of Intracellular Francisella” below) and is engulfed by a unique pseudopod loop mechanism. It then traffics to an early phagosome called the Francisella-containing phagosome (FCP). Francisella uses multiple mechanisms to evade host defenses in this harsh environment (inset). Francisella blocks the NADPH oxidase and also detoxifies reactive oxygen species (ROS). It can also resist the action of antimicrobial peptides (AMPs). Francisella does not signal through TLR4 but does activate TLR2 and may induce TLR9 signaling. Francisella then escapes the FCP to replicate within the cytosol. Subsequently, Francisella associates with autophagosomes, although the outcome of this interaction is unknown. Francisella can also induce host cell death.
In addition to subverting extracellular and intracellular defenses to facilitate replication, Francisella suppresses the activation of adaptive immune defenses that have the capability of destroying infected host cells. For example, CD8 T cells can directly lyse infected host cells, clearing these havens for bacterial replication. They and other cells also secrete the cytokine gamma interferon (IFN-γ), which can activate strong defenses in host cells, allowing them to resist Francisella replication. Therefore, Francisella uses several strategies to skew adaptive responses and block IFN-γ signaling to help preserve its replication niche in infected host cells (35, 119, 166, 189). Here we review the multitude of ways in which Francisella subverts host defenses at each of the stages of intracellular infection as well as its effects on adaptive immune responses.
SUBVERSION OF HOST DEFENSES
Following its transmission, Francisella must resist extracellular host defenses prior to entry into host cells. Furthermore, there is evidence that a potentially important extracellular phase exists after the initial stage of infection (87, 136, 239). Up to 75% of viable LVS and F. tularensis SchuS4 bacteria in the blood are present extracellularly at multiple time points following intranasal and intradermal murine infection (87). This finding was confirmed in studies with F. novicida in which ∼80% of viable bacteria in the blood were present extracellularly in serum (239). Of important note, these extracellular Francisella organisms are still infectious, suggesting that this extracellular phase may play an important role in dissemination from the site of infection (239). These data highlight the fact that Francisella must subvert extracellular host defenses to initiate infection and likely to spread and cause disease at later stages as well.
Complement and Antibody
During extracellular phases of infection, Francisella must be able to subvert, resist, or evade killing by a plethora of antimicrobial defenses. One major extracellular defense is the complement system that is present and active in mammalian blood. The complement system consists of an intricately regulated cascade of extracellular signaling proteins that can be triggered by the recognition of a microbe, ultimately leading to its lysis by a pore-forming complex or promoting uptake and killing by phagocytes (190). Throughout the process of complement activation, host signaling molecules which act to promote inflammation and recruit phagocytes and T cells to the site of infection are formed. Furthermore, inhibition of complement activation is necessary to maintain the integrity and viability of the pathogen, as well as to prevent the release of immunostimulatory bacterial components (pathogen-associated molecular patterns [PAMPs]) that would activate an enhanced immune response.
Complement activation is a complex process and is described in detail in a number of reviews (88, 190). Briefly, this process is initiated by the binding of lectins or antibodies to bacteria, which subsequently promote the recruitment and activation of the complement factors C1 through C4. This leads to the generation of the C3 convertase, a central signaling molecule in the complement cascade. When C3 is cleaved to C3a (an anaphylatoxin) and C3b, the canonical complement cascade continues, ultimately leading to bacterial lysis (Fig. 2). Briefly, lysis occurs when C3b, bound to the bacterial envelope, interacts with other complement factors to signal for the formation of the membrane attack complex (MAC). The MAC forms a pore in the bacterial envelope, leading to loss of membrane integrity and osmotic potential, in turn leading to lysis of the bacterium.
Fig 2.
Complement Evasion by Francisella. Antibodies (Abs) can bind to the bacterial surface (left panel), leading to binding of complement component C3 (right panel), triggering activation of the complement cascade and generation of C3a, an inflammatory anaphylatoxin, and C3b. C3b ultimately leads to the formation of the membrane attack complex (MAC) and lysis of the bacterial cell. Francisella counteracts these defenses by binding host plasmin, which inhibits antibody binding (left panel) and thus complement activation. Francisella also binds host factor H, which inhibits C3 cleavage to C3a and C3b and instead acts with host factor I to create the complement-inhibitory molecules C3d and C3bi. This skews the complement cascade from promoting MAC formation and lysis to facilitating phagocytosis. Factor H binding may be promoted by fibrinogen, which is converted to fibrin and is known to bind plasmin, but this has yet to be demonstrated.
Although C3 binds to the bacteria, Francisella blocks complement activation by cleaving this protein into the smaller inhibitory fragments C3bi and C3d (Fig. 2) (28, 29, 51). Generation of these fragments prevents the ability of C3 to signal toward MAC formation. The exact mechanism of C3 inactivation on the Francisella surface is not yet known; however, host factor H readily binds the Francisella cell surface (29) and can inhibit formation of the C3 convertase (Fig. 2). Importantly, factor H can also serve as a cofactor for host factor I-mediated cleavage of C3b to C3bi and C3d. Despite the evidence that factor H binds Francisella, factor I (which is necessary for C3b cleavage) has not yet been shown to interact directly with the bacterial surface. Factor I may interact only transiently, or if it does not bind, a Francisella-encoded factor may be responsible for C3b cleavage to C3bi and C3d.
Generation of C3bi and C3d is not only responsible for inhibiting subsequent MAC formation but is also important in promoting opsonophagocytosis. By binding complement receptor 3 (CR3) on host phagocytes, C3bi and C3d allow Francisella to enter host cells, further promoting escape from extracellular antimicrobials such as complement, antibody, and antimicrobial peptides (20, 28) and facilitating entry of the bacteria into an intracellular replicative niche. It should be noted, however, that uptake mediated by C3bi/C3d and CR3 leads to limited Francisella replication compared to uptake by nonopsonic receptors (see “Mechanisms of Entry and Fate of Intracellular Francisella” below for a more detailed discussion). Therefore, Francisella encounters complement upon infection and can prevent complement-mediated lysis by converting C3 into C3bi and C3d. This routes Francisella to a nonoptimal replicative pathway within host cells, but it is nonetheless an effective way to subvert extracellular antimicrobials and promote intracellular replication.
Francisella's complement-inhibiting activity is highly dependent on the O-antigen oligosaccharide present on LPS on the surface of the bacterial cell (51, 192, 211). Mutants lacking O antigen, either by random selection for rough or gray variants or by targeting wbt locus genes necessary for O-antigen production, are significantly more sensitive to complement than their wild-type counterparts (51, 192, 211). Interestingly, the relative importance of O antigen in complement resistance differs between F. novicida and F. tularensis. F. novicida wbt mutants are more than 2,000-fold more sensitive to serum than wild-type bacteria, while F. tularensis mutants are only 4-fold more sensitive (221). This may be due to the structural differences in O-antigen tetrasaccharide oligomers between these species (221) and/or additional O-antigen-independent mechanisms of complement resistance in F. tularensis that have not yet been elucidated.
The sensitivity of Francisella O-antigen mutants to complement is due to increased binding of complement factors compared to that in the wild type and an inability to catalyze the cleavage of C3 to C3bi (51, 192, 211). This results in the binding of complement factors C5 to C9, which form the MAC and do not bind wild-type Francisella, ultimately causing lysis of the mutant bacteria and decreased survival in serum (29, 51, 192, 211). Therefore, the presence of the O antigen is vital not only for regulating the quantity of complement factors which bind to the Francisella envelope but also for subverting the ability of bound complement components to mediate lysis of the bacteria.
The importance of maintaining resistance to complement lysis is demonstrated by the fact that complement-sensitive mutants are also severely attenuated in vivo. Francisella strains with mutations in the wbt locus are attenuated during infection of both macrophages (130, 135, 221) and mice (129, 153, 204, 221, 232). Furthermore, mutants lacking genes necessary for O-antigen production that are outside the wbt locus are also attenuated for intracellular replication and in vivo virulence compared to the wild type (130).
Additionally, strains with mutations in genes predicted to be important for capsule production are also significantly attenuated in vitro and in vivo (throughout this paper, “in vitro” refers to infection of host cells in culture, while “in vivo” refers to infection of mammalian hosts such as mice) (110, 130, 153, 232). This is important to note, since recent evidence has suggested that the O antigen and capsule of Francisella are structurally similar, with the capsule consisting of a polymer of O-antigen tetrasaccharide subunits (9) as well as a glycoprotein component termed a “capsule-like complex” (18). As such, some mutants deficient in LPS O-antigen production are also impaired in the production of capsule (9, 130), including specific wbt operon mutants (wbtA, wbtC, wbtM, and wbtI). wbtK and lpxL mutants lose the ability to synthesize LPS O antigen but maintain the polysaccharide capsule (9). Conversely, the F. tularensis genes FTT_0673 and FTT_0674 are necessary for capsule production but dispensable for O-antigen production (9). However, to the best of our knowledge, the ability of these mutants to resist complement has not been established. Testing the level of complement resistance in these mutants would allow the elucidation of which structure(s) is required. Thus, the current literature suggests that complement resistance in Francisella depends on O antigen, an O-antigen polysaccharide capsule, or both, as well as other potential factors that have yet to be elucidated.
Interestingly, the Francisella genome is known to contain an operon with similarity to the Bacillus species capBCA genes necessary for the production of a poly-d-glutamic acid capsule (71, 110, 153). However, to date, no poly-d-glutamic acid has been identified in Francisella. Although Francisella capB mutants are attenuated during murine infection, an LVS capB mutant did not exhibit increased sensitivity to serum (110, 213). Furthermore, it is not known whether CapB is required for production of the O-antigen capsule, the glycoprotein capsule-like complex, or a different bacterial envelope structure.
In addition to their sensitivity to complement, recent work has demonstrated that LPS O-antigen and capsule mutants are significantly more proinflammatory than wild-type cells (153, 171). An important, but perhaps overlooked, aspect of O-antigen and capsule function may be the ability of these large surface structures to mask and prevent the release of PAMPs on the Francisella surface (Fig. 3). In support of this hypothesis, LPS O-antigen mutants release more DNA into the cytosol of host cells during infection, leading to an increase in host inflammation and proinflammatory cell death (discussed further in “Cytosolic Defenses” below) (171). However, it is unclear whether this release of bacterial DNA (a PAMP) is due to a general instability of the Francisella envelope or an increased sensitivity to serum components and/or other antimicrobials. Taken together, these data highlight the important role that polysaccharide surface structures play in preventing host recognition of Francisella and virulence.
Fig 3.
Shielding of inflammatory PAMPs in Francisella. Francisella modifies and limits the release of its inflammatory components (PAMPs) that can be recognized by pattern recognition receptors, such as TLRs, expressed by antigen-presenting cells (APC). Francisella bacterial lipoproteins (BLPs) and DNA (shown in red) are PAMPs with inflammatory activity, while the capsule, LPS, free lipid A, and membrane phospholipids do not elicit an inflammatory response. Francisella modifies its LPS and free lipid A such that these PAMPs do not activate host TLR4. It is unclear whether Francisella peptidoglycan (PGN) has inflammatory activity (represented in brown). Membrane proteins that are not BLPs and are not as a class considered PAMPs are shown in purple. The capsule and LPS O antigen provide resistance to antimicrobials such as complement and antimicrobial peptides, preventing damage to the bacterial membranes, which would result in the release of immunostimulatory BLPs and DNA. In addition to providing resistance to the damaging effects of antimicrobials, the capsule and O antigen may also serve as physical barriers to the release of PAMPs into the environment.
In addition to the strategies described above, some bacteria use additional mechanisms to inhibit complement activation. For example, Streptococcus pneumoniae uses its surface M protein to bind host fibrinogen, a protein that can bind the complement inhibitor factor H, thus allowing fibrinogen-coated bacteria to inhibit complement activation (233). To our knowledge, no study to date has demonstrated that Francisella is capable of binding fibrinogen. However, recent work has shown that Francisella binds host plasminogen on its surface, which is in turn converted to plasmin (54), which can bind fibrinogen (137). Thus, although not yet directly demonstrated, this may be the mechanism used by Francisella to bind and sequester factor H, contributing to complement resistance (Fig. 2). Plasmin can also degrade soluble antibody, allowing Francisella to prevent antibody-mediated complement activation (59). Highlighting the potential importance of plasmin binding in pathogenesis, F. tularensis binds plasmin while the attenuated LVS strain does not, and LVS is therefore unable to evade this pathway. Together, Francisella utilizes LPS O antigen, its O-antigen polysaccharide capsule, and plasmin in order to subvert complement activation, promoting extracellular bacterial survival and preventing the release of proinflammatory PAMPs that would subsequently initiate a strong host immune response leading to clearance of the infection.
Antimicrobial Peptides
In addition to complement, the host produces a variety of other antimicrobials, most notably cationic antimicrobial peptides such as defensins and cathelicidins (42). Positively charged antimicrobial peptides, capable of disrupting the negatively charged bacterial membrane, are present extracellularly on mucosal surfaces as well as within macrophages and neutrophils. Upon infection, the bacteria first encounter either the epithelium of the lungs (after inhalation) or the skin (upon arthropod bites or contact with infected animal tissue). Subsequently, macrophages and neutrophils are among the first cell types the bacteria enter. Therefore, resistance to cationic antimicrobial peptides is likely a critical component of early pathogenesis.
Three defensin antimicrobial peptides (hBD-1, hBD-2, and hBD-3) are present in the human airway mucosa (42). The 50% effective concentration (EC50) (the concentration that kills 50% of the bacteria) for hBD-1 and hBD-2 against F. novicida is higher than that against other bacteria such as Pseudomonas, Staphylococcus, and Escherichia coli (roughly 100-fold greater for hBD-1 and 10-fold greater for hBD-2) (108). Accordingly, hBD-1 and hBD-2 are ineffective at killing F. novicida (98), and at sites of infection, it is estimated that hBD-1 and hBD-2 concentrations are below the levels required for killing (198). It is likely that even though in vitro infection of lung epithelial cells with F. novicida induces the expression of hBD-1 and hBD-2 (10- and 40-fold, respectively), these levels are still too low to effectively kill the bacteria. hBD-3, however, is effective at killing F. novicida (in broth culture) (98), but it is present at levels only about 2-fold higher than the EC50 (198), and infection does not further induce its expression (98). Therefore, it is likely that defensins play very little role in host defense against Francisella, due to its high resistance to hBD-1 and hBD-2 as well as its lack of induction of hBD-3 during infection.
Similar to the case for hBD-1 and hBD-2, F. novicida significantly induces the production of the cathelicidin LL-37 in lung epithelial cells in vitro (5). Furthermore, it is estimated that at sites of infection, LL-37 peptide concentrations average approximately 25 μg/ml (198), about 100-fold higher than the EC50 for F. novicida (5). Similar to its evasion of complement, Francisella must likely resist the action of LL-37 and presumably other host antimicrobials (such as hBD-1 and hBD-2) to prevent not only lysis and death but also the release of PAMPs that would subsequently trigger a proinflammatory response.
The literature to date has described at least two different mechanisms by which Francisella can either resist or evade the action of host antimicrobials: (i) by altering the charge of its surface and thus being able to use electrostatic interactions to repel cationic antimicrobial peptides and (ii) by encoding a number of efflux systems that are necessary for resistance to antimicrobials as well as for virulence in vivo.
Francisella modifies its lipid A, an outer membrane glycolipid structure that is a component of LPS but is also present in Francisella in a “free” form which lacks the canonical LPS core and O-antigen sugars (discussed in greater detail in “Toll-Like Receptors” below) (230). The bacteria remove the negatively charged phosphate groups from the 1 and 4′ positions of the lipid A portion of complete LPS (Fig. 4), which serves to increase the overall charge of the bacterial surface and repel cationic antimicrobials (94). Mutants that do not remove the 4′ phosphate are much more sensitive to cationic antimicrobial peptides and are highly attenuated in vivo (229). This attenuation is attributed to the increased susceptibility to cationic antimicrobials, which leads to membrane damage and the release of higher levels of PAMPs, resulting in a greater proinflammatory response, including increased neutrophil recruitment that results in clearance of the bacteria (210, 229).
Fig 4.
E. coli and Francisella lipid A structures. The structures of the E. coli lipid A moiety (A) and free lipid A from Francisella species (B) are shown. For both structures, the sugar backbone is highlighted in blue and acyl chains are represented in black, with numbers denoting length. Phosphate groups at the E. coli lipid A 4′ and 1 positions (absent from lipid A of full-length Francisella LPS) are highlighted in red. Francisella free lipid A contains the phosphate at the 1 position and is modified with galactosamine (GalN) (in green).
Preventing interactions with cationic host antimicrobials is one of Francisella's first lines of defense against the host immune response. However, increased surface charge does not prevent all antimicrobials from acting on the bacterial cell, particularly uncharged or negatively charged antimicrobials that could potentially be attracted more efficiently. Francisella, like many other bacteria, including Neisseria (95), Pseudomonas (179), and Salmonella (23), encodes at least one known multidrug efflux pump that allows resistance to a number of different antimicrobials and detergents, the AcrAB/TolC efflux pump. In addition, other predicted efflux pump proteins have been identified in screens for Francisella virulence genes, including FTN_1066, FTN_1217, FTN_1277, FTN_1654, and FTN_1657 (130, 140, 232). While the exact functions of these predicted efflux proteins are unknown, they may play a role complementary to the known functions of the AcrAB/TolC efflux pump.
The AcrAB inner membrane efflux platform (30), coupled with the TolC outer membrane transporter (91, 178), forms a pump that facilitates the active efflux of toxic compounds and detergents from the bacteria, preventing their antimicrobial action. This multidrug efflux pump is important for Francisella resistance to β-lactams, tetracyclines, aminoglycosides, quinolones, detergents (notably bile salts), and antimicrobial dyes (30, 91). Highlighting the importance of this system in pathogenesis, mutants with mutations in pump components are severely attenuated in vivo (30, 91, 178). While the AcrAB/TolC system is known to provide resistance against host antimicrobial peptides in bacteria (95) and it is assumed that Francisella mutants would be more sensitive to host-derived antimicrobials during infection, to the best of our knowledge this has not been directly tested.
Interestingly, TolC mutants stimulate an increased proinflammatory response compared to that of wild-type Francisella (178). This may be due to increased sensitivity to antimicrobials in vivo leading to increased PAMP release or to a general membrane instability defect eventually leading to PAMP release. Furthermore, since TolC has been shown to be involved in the secretion of virulence factors in other bacteria such as Pseudomonas (100) and E. coli (227), it cannot be ruled out that it may play a similar role in Francisella pathogenesis, accounting for some of the virulence defects exhibited by mutants lacking this protein.
Another important aspect of antimicrobial peptide resistance in Francisella is the potential contribution of the O antigen and/or capsule. The Klebsiella capsule prevents interaction with antimicrobial peptides (39), and the O antigen of Salmonella has been shown to act similarly (212). However, to the best of our knowledge, whether these systems contribute to Francisella resistance is not yet known. Together, the ability of Francisella to resist and evade the action of complement, antibody, and antimicrobial peptides within the host allows it not only to survive but also to maintain the integrity of its envelope, preventing the release of proinflammatory molecules that would induce a host response and promoting its virulence.
Mechanisms of Entry and Fate of Intracellular Francisella
As mentioned above, one mechanism used by bacteria to evade extracellular defenses is simply to enter host cells. Macrophages are sentinels of infection and are one of the first cell types to encounter Francisella (97). Upon initial contact with macrophages, Francisella binds to host receptors and is taken up via spacious, asymmetrical protrusions called pseudopod loops (52). Ultrastructural analyses have shown that pseudopod loops are formed during uptake of unopsonized and opsonized Francisella and are present in multiple cell types (52). This novel phenomenon differs from other described bacterial uptake mechanisms, including coiling phagocytosis seen at the surface of phagocytes during Legionella pneumophila infection and conventional phagocytosis observed during infection with multiple bacterial and viral species (102, 103, 199, 200).
Furthermore, Francisella LVS can recruit host cell membrane cholesterol-rich lipid domains, or “lipid rafts,” with caveolin-1 for successful entry into macrophages (217). Depletion of plasma membrane cholesterol or removal of glycosylphosphatidylinositol (GPI)-anchored proteins results in a severe decrease in Francisella uptake and subsequent intracellular replication (217). Interestingly, in nonphagocytic cells such as hepatocytes, LVS and F. novicida use cholesterol-rich lipid domains dependent on clathrin, not caveolin-1, for entry (123). Thus, Francisella uses multiple mechanisms for uptake into phagocytic and nonphagocytic cells.
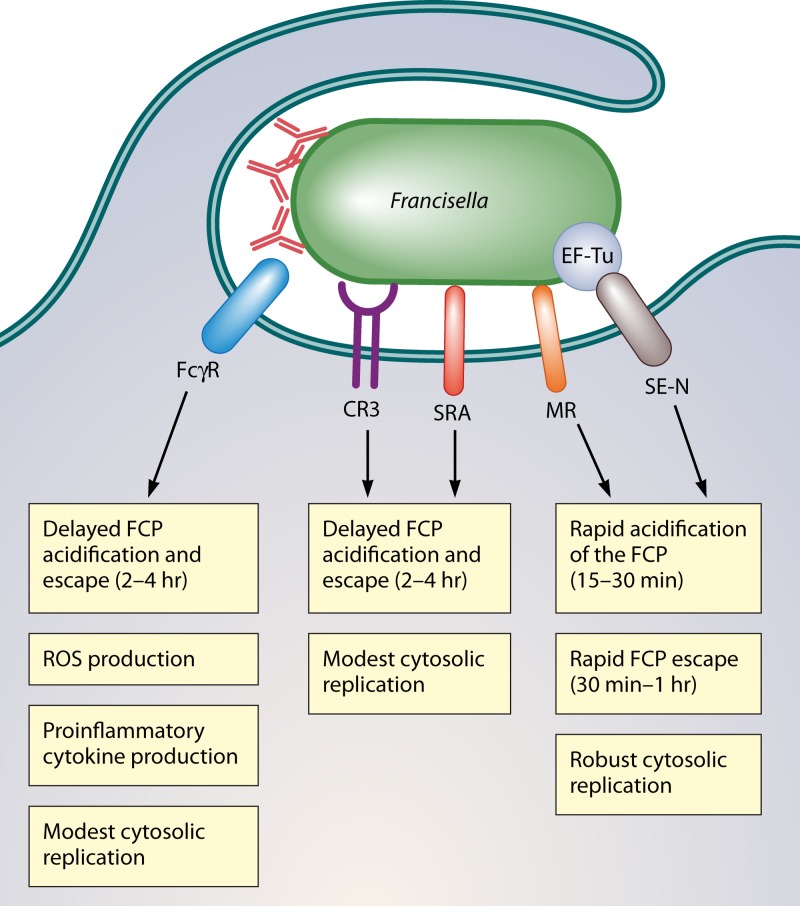
The efficiency of Francisella uptake by macrophages depends in large part on whether the bacteria are opsonized, since serum- or antibody-opsonized bacteria are taken up by macrophages at 10-fold-higher levels than unopsonized bacteria (90). While this increased efficiency of uptake helps the bacteria evade extracellular defenses, it comes at a cost since the intracellular fates of opsonized and unopsonized bacteria are different (Fig. 5). Opsonized bacteria escape the phagosome with delayed kinetics and replicate modestly in the host cell cytosol (28, 90, 176, 202). In contrast, unopsonized Francisella escapes the phagosome rapidly and replicate robustly in the cytosol (17, 202). Therefore, the route of uptake has a profound impact on the outcome of infection, and as discussed below, Francisella uses several strategies to promote entry mechanisms that favor its survival and replication.
Fig 5.
Intracellular fates of Francisella after uptake by different macrophage receptors. Antibody-opsonized Francisella is taken up via the FcγR, leading to increased ROS production and induction of proinflammatory cytokines, delayed FCP acidification and bacterial escape (2 to 4 h) from the phagosome, and only moderate levels of cytosolic replication. Uptake of serum (complement)-opsonized Francisella is mediated mainly by complement receptor 3 (CR3) and scavenger A receptors (SRA), which lead to slowed FCP acidification and phagosomal escape (2 to 4 h) and result in modest cytosolic replication. Lastly, uptake of unopsonized Francisella is mediated by the mannose receptor (MR) and surface-exposed nucleolin (SE-N), leading to rapid acidification (15 to 30 min) and escape from the FCP (30 min to 1 h) and robust cytosolic replication.
Uptake of unopsonized Francisella depends largely on the mannose receptor (MR). When the MR is blocked with antibody or mannan, uptake of unopsonized Francisella decreases by more than 50% (17, 202). Unopsonized Francisella taken up by the MR escapes the phagosome rapidly and replicate to high numbers. In addition, the MR is known to induce relatively weak inflammatory responses compared to those induced by other phagocytic receptors (225). All of these outcomes favor the bacteria, and therefore, MR-mediated uptake makes host cells more permissive for Francisella replication, similar to what is observed during uptake of Mycobacterium tuberculosis (113).
In contrast, serum-opsonized bacteria are bound by complement, leading to uptake mediated in large part by complement receptor 3 (CR3). Ablation of CR3 (CR3−/− macrophages) results in at least a 50% decrease in uptake of serum-opsonized LVS (202). The scavenger A receptor (SRA), although traditionally known to play a role in uptake of unopsonized bacteria, contributes to the uptake of serum-opsonized Francisella, since SRA−/− macrophages exhibit a 20% reduction in the uptake of serum-opsonized LVS (176). Macrophage cell surface-exposed nucleolin has also been implicated in uptake of serum-opsonized Francisella by binding the bacterial membrane protein EF-Tu (19). Finally, MR-mediated uptake also plays a role in uptake of serum-opsonized Francisella (roughly 30% decreased uptake when blocked) (90). As mentioned above, this multireceptor uptake pathway predominated by CR3 results in delayed phagosomal escape and replication and is therefore nonoptimal for Francisella.
Opsonization with antibody, specifically IgG, leads to FcγR-mediated uptake of Francisella. FcγR−/− macrophages exhibit a 90% reduction in uptake of IgG-opsonized Francisella, although they have no defect in the uptake of unopsonized or serum-opsonized bacteria (90). FcγR-mediated uptake leads to increased activation of the NADPH oxidase, a phagosomal enzyme complex that produces toxic reactive oxygen species (ROS) and is discussed in greater detail below. This activation results in a more inhospitable environment for the bacteria in the phagosome, acting to delay phagosomal escape and severely limit subsequent bacterial replication. The efficacy of this host defense is revealed when ROS production is abrogated by genetic deletion of an NADPH oxidase component (using gp91phox−/− macrophages), since IgG-opsonized Francisella bacteria are rescued and can replicate to high levels (90).
Binding of plasmin by Francisella (mentioned in “Complement and Antibody” above) leads to degradation of soluble antibody at the bacterial surface (Fig. 2) (59), which limits uptake by the FcγR pathway and therefore subversion of this pathway, facilitating higher levels of intracellular replication. Francisella also subverts the CR3-mediated pathway using LPS O antigen and capsule, which inhibit binding of complement factors. It should be noted that when complement binding does occur, the bacteria alter the complement pathway to prevent MAC formation and lysis (Fig. 2). However, the alternate complement fragments generated actually promote phagocytosis via CR3. While uptake via this pathway is not optimal, it does still support bacterial replication and is a much better outcome for the bacteria than complement-mediated lysis. The most optimal entry pathway, however, is mediated via the MR. Therefore, Francisella uses the aforementioned subversion mechanisms to promote uptake by the more favorable MR-mediated pathway, facilitating the highest levels of intracellular replication.
Phagosomal Acidification
After uptake by the macrophage, Francisella resides within the Francisella-containing phagosome (FCP) and subsequently escapes into the cytosol, where it can replicate (Fig. 1) (184). The brief time spent in the FCP is a dynamic step in infection during which Francisella must actively evade host antimicrobial defenses, including acidification of the FCP, reactive oxygen species, and antimicrobial peptides. Acidification of the phagosome is a defense mechanism that lowers the pH in this compartment, preventing many types of bacteria from efficiently replicating. Many intracellular pathogens must subvert this host defense mechanism by either blocking it, escaping the phagosome, or maintaining intracellular pH in spite of the lowered pH in the environment (184). It is interesting to note that acidification is actually required for the rapid phagosomal escape of the intracellular pathogen Listeria monocytogenes (27). Studies with Francisella have shown a brief acidification of the FCP that is dependent on the route of uptake, although there has been debate about whether this step is required for rapid escape and about its effect on the outcome of infection, as detailed below (49, 52, 53, 194).
Uptake of unopsonized Francisella leads to transient acidification of the FCP and rapid escape (49, 194). Santic et al. reported acidification of the phagosome at 15 to 30 min postinfection of human monocyte-derived macrophages (MDM) with unopsonized F. novicida, followed by escape at 30 min to 1 h postinfection (194). Interestingly, they showed that inhibition of the vacuolar proton ATPase (vATPase) pump by bafilomycin A (BFA) does not block escape but results in delayed escape (6 to 12 h). These results suggest that acidification is required for rapid escape from the FCP (194). Additionally, Chong et al. observed brief acidification of the FCP in macrophages infected with unopsonized F. tularensis (49). Similarly, BFA did not inhibit phagosomal escape or cytosolic replication but slowed infection kinetics (49). Together these data show that acidification of the FCP ensures rapid escape and robust replication of unopsonized Francisella; however, it is not required for these processes.
Concurrently, it has been reported that uptake of serum-opsonized Francisella results in acidification of a modest 20% to 30% of phagosomes containing serum-opsonized LVS and that it is not required for escape or replication (53). Additionally, the authors found that the maturing FCP does not acquire high levels of the acid hydrolase cathepsin D, which is a cellular marker of acidification, or the endosomal-lysosomal markers CD63, LAMP1, and LAMP2, which are cellular markers of phagosomal maturation (52, 53). In contrast to the previously mentioned work using unopsonized bacteria, BFA did not significantly delay infection kinetics in these experiments using serum-opsonized bacteria (53). These data demonstrate that acidification of the FCP is not required for escape and replication of unopsonized or serum-opsonized Francisella but is required for rapid escape of unopsonized Francisella. The fact that Francisella does not exhibit a significant survival or replication defect in the presence or absence of acidification demonstrates that this pathogen has devised ways to efficiently resist this host defense. This resistance is mediated in part by physically escaping the FCP, where acidification takes place (discussed in “Francisella Escape from the Phagosome” below). It is also likely that Francisella has evolved mechanisms to maintain intracellular pH while in the acidified environment of the FCP.
Inhibition of Reactive Oxygen Species
In addition to subverting the potentially toxic effect of acidification of the FCP, Francisella must contend with reactive oxygen species (ROS) produced in this compartment. ROS are produced by the NADPH oxidase, a membrane-bound multicomponent enzyme system that converts molecular oxygen into toxic superoxide anions (161). In a resting phagocyte, NADPH oxidases are unassembled, with the gp91phox and p22phox (also called flavocytochrome b558) components localizing to the plasma membrane and p47phox, p40phox, p67phox, and Rac2 to the cytosol (145, 161). Upon phagocytosis of a microbe, the cytosolic subunits traffic to the phagosome and assemble with the membrane subunits to create the active NADPH oxidase that then produces ROS. Similar to the case for numerous extracellular and intracellular bacterial species, including Helicobacter pylori and Salmonella spp., multiple Francisella species block NADPH oxidase assembly in neutrophils and macrophages (4, 89, 145, 156, 203).
Francisella species use several approaches to inhibit ROS, including blocking initial assembly of NADPH oxidase components at the phagosomal membrane, blocking ROS production in complexes that have assembled, and detoxifying ROS that are generated. F. novicida requires four putative acid phosphatases (AcpA, AcpB, AcpC, and Hap) for inhibition of NADPH oxidase assembly (155, 186). AcpA colocalizes with the cytosolic NADPH oxidase component p47phox during infection, and purified AcpA dephosphorylates p47phox and p40phox (155). Without phosphorylated membrane-bound components, the cytosolic NADPH oxidase components are not recruited and assembly cannot occur (14). These data suggest that AcpA interacts directly with NADPH oxidase components to block complex assembly.
While AcpA plays a role in limiting the oxidative burst, its relative contribution to virulence is unclear. Two studies found that AcpA is required for replication of F. novicida in human macrophage-like cells, indicating that blocking the NADPH oxidase is an important factor in facilitating replication (11, 155). In contrast, another study using F. novicida determined that AcpA did not play a role in replication in murine macrophages (22). In F. tularensis, deletion of acpA, or even of acpA, acpB, and acpC together, did not influence virulence in murine macrophages or human monocytes (48, 146). These conflicting data indicate that AcpA is not required for Francisella replication under all conditions tested. Its requirement may depend on the species or host cells used and the specific infection conditions.
In addition to blocking assembly, F. tularensis can also inhibit the generation of ROS when NADPH oxidase assembly is induced by exogenous stimuli (146). In spite of the formation of this complex, F. tularensis can inhibit the production of ROS. These data show that F. tularensis can block NADPH oxidase-dependent ROS production postassembly (146), although the mechanism by which this occurs has not yet been elucidated.
Although Francisella uses the mechanisms described above to significantly suppress activation of the NADPH oxidase, low levels of ROS are produced in the phagosome during infection (203). Like many pathogens, Francisella can directly detoxify ROS using proteins, including catalase (134, 151, 214) and superoxide dismutases (15, 16, 151), whose specific mechanisms of action are reviewed extensively elsewhere (135). In addition to these well-known mechanisms of ROS resistance, we identified a previously uncharacterized protein, FTN_1133, that is required for virulence and resistance to organic hydroperoxides (135). FTN_1133 has sequence similarity to Ohr proteins, which are involved in resistance to organic hydroperoxides created during the interaction of ROS with lipids of the bacterial cell membrane (135). The FTN_1133 mutant was attenuated for replication, although this could be restored in gp91phox−/− macrophages and mice (135). These data together indicate that Francisella has numerous overlapping mechanisms with which to subvert the NADPH oxidase and ROS, facilitating pathogenesis.
Francisella Escape from the Phagosome
Phagosomal escape is the last step in Francisella's subversion of the phagocytic pathway, allowing it to escape the toxic phagosome and reach the cytosol, where it can replicate. The timing of escape has been a topic of debate and, as mentioned above, is largely dependent on the route of Francisella uptake by host cells. Unopsonized Francisella escapes the phagosome rapidly (within 1 h) whereas opsonized Francisella exhibits delayed escape (2 to 4 h) (92).
The exact mechanism of escape is not yet known; however, the proteins encoded in the Francisella pathogenicity island (FPI) are absolutely required. The FPI encodes a putative type VI secretion system (T6SS) that is essential for Francisella replication and pathogenesis (21; reviewed in reference 160). The requirement of a specialized secretion system for phagosomal escape would be in line with the situation in many intracellular bacteria, including Shigella flexneri (T3SS), Listeria monocytogenes (Sec pathway), and Burkholderia spp. (T3SS) (70, 168, 177, 193, 197). Briefly, the FPI proteins IglA and IglB share homology with proteins encoded in T6SS clusters in multiple bacterial species, are required for phagosomal escape, and may form the putative outer tube of the T6SS “needle” (64, 65). IglC has been proposed to form the inner tube (64), while IglI and VgrG are secreted and therefore may interact with host proteins during infection (21). In addition, numerous proteins that are not encoded in the FPI have also been implicated in phagosome escape and are reviewed elsewhere (11, 49).
To facilitate phagosomal escape, Francisella must subvert the action of host factors that have evolved to slow or block this process in order to control infection. Activation of the phosphatidylinositol 3-kinase (PI3K)/Akt pathway by bacteria leads to production of proinflammatory cytokines, increased ROS production, and retention of bacteria in the phagosome (61, 104, 181). In accordance, activation of the PI3K/Akt pathway during F. novicida infection blocks phagosomal escape and cytosolic replication (181). Interestingly, F. novicida infection of macrophages activates the SH2 domain-containing inositol phosphatase (SHIP) by an unknown mechanism that is dependent on live bacteria (167). SHIP antagonizes activation of the PI3K/Akt pathway, leading to rapid escape from the phagosome and robust cytosolic replication (167). Additionally, the PI3K/Akt pathway can be activated by cell surface receptors, including FcγR (143) and TLRs (121). Therefore, Francisella mechanisms to evade these receptors (discussed in “Complement and Antibody” above and in “Toll-Like Receptors” below) may also function to block activation of the PI3K/Akt pathway and ensure rapid phagosomal escape.
Toll-Like Receptors
Upon contact with host cells, extracellular and intracellular microbes encounter host pathogen recognition receptors (PRRs) that are capable of detecting conserved microbial components (PAMPs) (109). These receptors can then trigger multiple pathways, including phagocytosis and inflammatory signaling (109) (115). Toll-like receptors (TLRs) are important PRRs that can recognize PAMPs outside the host cell and in the endosome/phagosome (115, 216). TLR signaling is mediated by TIR domain-containing adaptor proteins, including MyD88, TRIF, and TIRAP, that activate transcription factors such as NF-κB and IRF3 (115). These transcriptional regulators induce the expression of inflammatory cytokines and type I interferons, resulting in the activation of innate and adaptive immune cells (109, 115). Numerous TLRs recognize bacterial PAMPs, including TLR2, which senses bacterial lipoproteins (BLPs) and peptidoglycan (PGN) (12, 73), TLR4, which signals in response to LPS from Gram-negative bacteria, TLR5, which recognizes flagellin, and TLR9, which senses bacterial CpG DNA (115).
A central component of Francisella's success as a pathogen is its ability to avoid recognition and subvert the host inflammatory response, particularly in the early stages of infection. Indeed, Francisella can likely evade or suppress inflammatory signaling by all of the aforementioned bacterium-sensing TLRs. For example, Francisella does not encode flagellin, and therefore TLR5 is not activated in response to infection (128). Though TLR recognition of Francisella DNA has been suggested (77), TLR9 (present in the membranes of endosomes and phagosomes) was shown to be unimportant for the host response to Francisella infection in vivo (56). This suggests that Francisella may subvert TLR9 activation, possibly by limiting the release of its DNA through resistance to damaging antimicrobial agents (see “Complement and Antibody” and “Antimicrobial Peptides” above) and/or by directly modulating TLR9 signaling. In support of the idea that Francisella maintains strong structural integrity to prevent DNA release, Peng et al. have shown that some hyperinflammatory F. novicida mutants exhibit increased bacteriolysis and DNA release during in vitro infection (171). Though Francisella rapidly escapes the phagosome (see “Francisella Escape from the Phagosome” above), it is intuitive that bacterial damage or killing and resulting DNA release during the time within this compartment would activate TLR9. It would be interesting to determine the extent of bacterial damage in the phagosome, whether phagosomal bacteria can trigger TLR9 signaling, and whether Francisella DNA is capable of acting as a TLR9 activator.
While TLR4 is considered a primary sensor of Gram-negative bacteria, Francisella LPS does not efficiently activate TLR4 compared to LPS from E. coli and other Gram-negative pathogens (180). Many Gram-negative bacteria that elicit robust TLR4 signaling synthesize a hexa-acylated lipid A portion of LPS with acyl chains of 12 to 14 carbons and phosphate groups at the 1 and 4′ positions (Fig. 4) (180). However, Francisella modifies or removes these important signaling structures. For example, Francisella lipid A acyl chains are two to six carbons longer than those in E. coli LPS (180). In addition, Francisella lipid A is tetra-acylated, as it lacks the canonical 3′ double acyl chain, and both the 1 and 4′ phosphate groups are absent (Fig. 4) (180). These modifications are critical for virulence, since an F. novicida mutant lacking the lpxF gene and producing a penta-acylated lipid A containing the 4′ phosphate group is rapidly cleared in a mouse infection model (229). This mutant exhibits hypersensitivity to the cationic antimicrobial peptide polymyxin B and elicits an increased local cytokine response and increased neutrophil recruitment in vivo. TLR4 is not involved in this response, suggesting that the attenuation is not due to an increase in the signaling capacity of the mutant LPS. Instead, it is likely that the observed attenuation is due to the mutant's increased sensitivity to antimicrobial peptides that lead to outer membrane damage and leakage of other bacterial PAMPs, such as BLPs and DNA, within the phagosome, where they could trigger TLR2- or TLR9-dependent inflammatory responses.
Unlike in other bacteria, between 70% and 90% of total Francisella lipid A exists as “free lipid A” that does not contain core polysaccharides or O antigen (Fig. 4) (230, 243). Similar to the case for the complete LPS, free lipid A is tetra-acylated with elongated acyl chains and lacks the 4′ phosphate group. However, instead of being removed as it is in complete LPS, the phosphate at the 1 position is present and modified with galactosamine (230, 243). This modification to free lipid A alone is critical for pathogenesis, since an flmK mutant that does not add the galactosamine to free lipid A is highly attenuated in vivo (114). To the best of our knowledge, free lipid A is unique to Francisella and could represent a novel mechanism of virulence that might be involved in strengthening the outer membrane. However, it is somewhat counterintuitive that Francisella would produce free lipid A without O antigen, since O antigen is so critical for virulence (see “Complement and Antibody” above). Further studies of the role of Francisella's free lipid A during host interactions will elucidate its contribution to virulence.
Since Francisella LPS (and likely free lipid A) is a poor TLR4 activator, it is expected that TLR4 would not play a critical role in host defense against this pathogen. Most reports support this conclusion, although some disagree as to the magnitude of TLR4 signaling that Francisella LPS can elicit as well as whether TLR4 plays a minor role or no role in host defense. For example, studies have reported that, in vitro, LPS from F. novicida triggers a low level inflammatory response (96, 116) while LVS LPS does not, even at high concentrations (5 μg/ml) (6, 116, 219). However, other studies reported that LVS LPS can induce TLR4 signaling in both transiently transfected HEK293 cells and human monocytes, but only when added at high doses that are likely not biologically relevant (≥2.5 μg/ml) (72). The varied LPS signaling responses may be due to differences in the cell types used or slight structural variations, such as the 6′ glucose addition to F. novicida LPS or the 4′ or 6′ hexose modifications of the LVS LPS (25, 94). Overall, these data show that the LPS from the Francisella species tested is markedly less inflammatory than LPS from other Gram-negative organisms such as E. coli, Salmonella, or Bordetella (6, 72, 96, 219). This finding is in agreement with the general lack of TLR4 activation during in vivo Francisella infections, as discussed below.
Though early reports indicated that TLR4-deficient mice exhibit increased sensitivity to LVS infection (8, 138), the bulk of the current evidence suggests that TLR4 is not essential for the host response to infection. For example, TLR4-deficient mice are not more susceptible than wild-type mice to low-dose F. tularensis or LVS aerosol challenge (46), LVS low-dose intradermal (i.d.) infection (47), or intranasal inoculation with either LVS or F. novicida (1). However, the 50% lethal dose (LD50) for LVS is 1 log lower in TLR4-deficient mice than in wild-type mice after high-dose i.d. infection (106 to 107 CFU) (47). Therefore, TLR4 may play a minor role in the host response to high-dose infections, though doubts remain about whether such a high inoculum is physiologically relevant. Even if TLR4 does play a minor role in host defense, it is not nearly as important as other innate defense proteins (tumor necrosis factor alpha [TNF-α], IFN-γ, or MyD88) that are critically required for resistance to infection as judged by the susceptibility to infection of mice lacking these proteins (1, 126). Overall, Francisella LPS is a poor TLR4 ligand, and TLR4 does not appear to play an important role in the host response to infection in vitro or in vivo.
TLR2 is the primary TLR involved in the inflammatory response to Francisella infection (1). Known Francisella TLR2 agonists include the uncharacterized lipoproteins LpnA/Tul4 and FTT_1103 (86, 220). To our knowledge, TLR2 recognition of Francisella PGN has not been reported, and the status of PGN as a TLR2 ligand is still under debate (12, 73, 222). Several reports have shown that TLR2 is essential for the early inflammatory response to Francisella infection in macrophages in vitro, as well as a critical component of the host response to in vivo Francisella infection as demonstrated by its requirement for control of pulmonary and intradermal infection (1, 16, 55).
However, while Francisella elicits TLR2-dependent signaling, it can also dampen this response. Specific Francisella genes that play a role in this suppression of the host inflammatory response have been identified. For example, a strain with a mutation in the FPI gene iglC not only failed to escape the phagosome or replicate in macrophages but also was unable to suppress TNF-α production and other inflammatory responses (64, 131, 196, 219). In addition, infection of macrophages with wild-type LVS blocked TLR2 and TLR4 activation in response to the addition of E. coli BLP and LPS, respectively, while the iglC mutant could not block this signaling (219). Further supporting the attenuation of TLR2 signaling by Francisella, infection with F. tularensis has been shown to reduce TLR2 expression (38). Mechanisms that the bacteria use to resist damage by antimicrobials, and therefore the release of BLPs, are also an indirect way of evading TLR2 signaling. It has been reported that high-molecular-weight (HMW) carbohydrates from “host-adapted” LVS and F. tularensis capsules impede TLR2-dependent cytokine production in murine macrophages (240), possibly by shielding the bacteria from antimicrobials. Taken together, these data indicate that Francisella is capable of subverting TLR2 signaling, while the host uses this pathway as a mechanism of innate defense.
Preliminary evidence suggests that highly virulent F. tularensis is able to suppress host recognition and inflammatory signaling even further. Melillo et al. recently reported that F. tularensis strain SchuS4 has a greater ability to prevent proinflammatory cytokine production than LVS during infection of murine macrophages (150). Indeed, 2 days after pulmonary infection, F. tularensis replicates to high numbers in the lungs of mice but does not induce inflammatory signaling and in fact triggers an increase in anti-inflammatory signaling (7, 34, 35). In addition, F. tularensis infection of dendritic cells (DC) suppresses the response of both infected and bystander cells to TLR agonists, due at least in part to a heat-stable bacterial component (44). Taken together, these findings suggest that F. tularensis is able to suppress the host inflammatory response to an even greater degree than other Francisella species, correlating with its heightened virulence. Further studies querying the role of TLR signaling in response to F. tularensis infection could shed light on the mechanisms of inflammatory subversion utilized by this pathogen.
Finally, Collazo et al. demonstrated that mice lacking the adaptor MyD88, but not those lacking TLR2, TLR4, or TLR9, had greatly increased susceptibility to LVS infection (56). This indicates that multiple TLRs may play overlapping roles in resistance to infection, such that no one TLR knockout displays highly enhanced susceptibility (56). Alternatively, Medina et al. propose that an unknown MyD88-dependent PRR may be involved in the host response to LVS infection (148). Overall, Francisella species are able to largely evade recognition by TLRs as well as to suppress TLR2 signaling, significantly contributing to virulence.
Cytosolic Defenses
When Francisella reaches the cytosol, it has trafficked past TLRs and phagosomal defenses. It is nonetheless faced with a formidable challenge: replicate to high numbers without triggering an effective immune response. This is all the more challenging since the process of bacterial replication results in the release of PAMPs that can be recognized by cytosolic PRRs. Like the cell surface and the phagosome, the cytosol is equipped with numerous PRRs that recognize an array of bacterial products and elicit an immune response aimed at clearing the invaders. One large family of cytosolic PRRs is the Nod-like receptor (NLR) family, whose 22 members respond to a diverse set of PAMPs, including PGN (Nod1 and Nod2), flagellin (NLRC4, NAIP5, and NAIP6) (118), components of bacterial type III secretion systems (NLRC4), and damage induced by pore-forming toxins (NLRP3) (152). Nod1 and Nod2 are membrane-associated NLRs that detect muropeptide subunits of PGN and induce NF-κB activation and proinflammatory cytokine production (174). A role for Nods during Francisella infection has yet to be described, but these receptors may recognize and respond to PGN from this pathogen. Alternatively, Francisella may subvert Nods by modifying its PGN (similar to the case for Listeria monocytogenes, which N-deacetylates its PGN [33]) or by suppressing the Nod signaling pathway.
Other cytosolic PRRs, such as RIG-I and MDA-5, bind bacterial or viral nucleic acids and induce the production of type I interferons (IFNs) (157). The type I IFN family includes numerous IFN-α proteins, a single IFN-β protein, and other IFNs (66). These secreted cytokines have a well-established role in interfering with viral replication but can also be induced in response to bacterial infection (157). F. novicida induces type I IFN production independently of TLRs, Nod1/2, RIG-I, or MDA-5 (99). The cytosolic PRR that induces type I IFN production in response to F. novicida is currently unknown. However, the adaptor protein STING (stimulator of interferon genes) is required for type I IFN production during F. novicida infection and is speculated to act downstream of the yet-to-be-identified cytosolic PRR (112). STING is known to induce type I IFN production in response to cytosolic double-stranded DNA (dsDNA) derived from transfected plasmids, viruses, and some bacteria (107). Therefore, it is likely that Francisella DNA, which has been observed in the host cytosol by confocal microscopy (112), is the ligand that activates this cytosolic defense system.
It is not clear how Francisella DNA reaches the cytosol to trigger type I IFN, but bacterial escape from the phagosome is required for induction of this host response (99, 112). Perhaps damage incurred by Francisella in phagosomes allows for the release of DNA from ruptured phagosomes following escape. Interestingly, an auxotrophic mutant of LVS that escapes the phagosome but cannot replicate in the cytosol failed to trigger cytosolic defense pathways (169), suggesting that bacterial replication might be required to increase the amount of DNA to a threshold level to which the host responds. In this context, mechanisms of maintaining structural integrity (LPS modifications, O antigen, and capsule; see “Complement and Antibody” and “Toll-Like Receptors” above) may prevent damage to the bacteria and the release of DNA in the cytosol and therefore would be hypothesized to promote subversion of these cytosolic defenses.
In addition to triggering the type I IFN pathway, cytosolic DNA released during Francisella infection can also be recognized by the PRR absent in melanoma 2 (AIM2) (82, 183), whose expression is upregulated by IFN-β (112). AIM2 is a member of the PYHIN (pyrin and HIN-200) family of proteins that binds dsDNA through a HIN-200 domain (81, 101, 187). AIM2 contributes to host defense by initiating the formation of a multiprotein complex called the inflammasome that is comprised of a PRR (from the NLR or PYHIN families), the scaffolding protein ASC, and the cysteine protease caspase-1 (144). Inflammasome activation causes infected cells to undergo an inflammatory form of programmed cell death called pyroptosis (84). This cell death may release bacteria into the extracellular environment, where they can easily be taken up by cells such as neutrophils that are not permissive for replication (210). Additionally, pyroptosis is accompanied by the release of the proinflammatory cytokines interleukin-1β (IL-1β) and IL-18 from dying cells, serving to recruit and activate other immune cells and further promote bacterial clearance. AIM2 inflammasome activation is essential for controlling F. novicida infection, since mice lacking components of this defense system succumb to infection much more rapidly than their wild-type counterparts (81, 141). The murine NLRC4 and NLRP3 inflammasomes do not play an obvious role in combating F. novicida during infection of macrophages or mice (141, 142). However, the NLRP3 inflammasome, along with the AIM2 inflammasome, is activated during Francisella infection of human epithelial cells and monocytes/macrophages (13). The mechanism underlying NLRP3 activation in human cells is not clear, since Francisella does not express known NLRP3 ligands.
Unlike the less virulent F. novicida, highly virulent F. tularensis fails to efficiently activate the inflammasome (234). Macrophages and dendritic cells infected with F. tularensis secrete very low levels of the inflammasome-dependent cytokine IL-18 in vitro, and there is very little caspase-1 activation induced in the spleens and livers of infected mice (234). As mentioned, microarray analysis of human monocytes infected with F. tularensis revealed that this pathogen downregulates the expression of several genes belonging to the TLR and type I IFN pathways (38, 60). TLR2 signaling is necessary for the expression of IL-1β and increases the rate of inflammasome activation during F. novicida infection (111), while type I IFN is essential for inflammasome activation (99). Therefore, hampering TLR2 and IFN signaling, two major host defense pathways that contribute to inflammasome activation, could lead to a lack of activation of this complex during F. tularensis infection. It is likely that highly virulent F. tularensis also has additional ways by which to limit inflammasome activation.
Several Francisella genes have been implicated in modulating inflammasome activation (82, 106, 224, 232). However, Peng et al. recently showed that these genes were not important for actively modulating the inflammasome (171). Instead, the increased induction of macrophage death triggered by a panel of mutants lacking genes encoding membrane-associated proteins was due to increased bacteriolysis in the cytosol that allowed for the leakage of DNA and increased inflammasome activation. This study suggests that maintenance of membrane integrity is critical for Francisella to prevent the release of PAMPs and induction of the inflammasome. F. tularensis may also directly suppress inflammasome activation; however, genes important for direct suppression of the inflammasome complex by F. tularensis have not been identified. The presence of such genes in F. tularensis but not less virulent Francisella species could explain the divergence in activation of this complex.
During the latter stages of F. tularensis infection, infected cells undergo caspase-3-dependent programmed cell death or apoptosis. Unlike pyroptosis, this form of cell death is noninflammatory (120). Wickstrum et al. observed a significant increase in apoptosis in the livers and spleens of F. tularensis-infected mice between days 3 and 4 postinfection. This spike in apoptosis was preceded by an exponential increase in bacteria and antigen distribution in the infected organs (234). F. tularensis may direct host cells to undergo apoptosis instead of pyroptosis following cytosolic replication, facilitating dissemination to neighboring cells without triggering a strong inflammatory response. Additionally, phagocytosis of apoptotic bodies by activated macrophages impairs the ability of these cells to produce proinflammatory cytokines (80). In the context of an F. tularensis infection, these macrophages that have been rendered immunologically suppressed by taking up bacterium-containing apoptotic bodies could serve as reservoirs for further replication. Interestingly, it is not clear how Francisella egresses from infected cells and disseminates throughout the host. These impaired macrophages could also serve as Trojan horses, trafficking the bacteria systemically.
Another strategy commonly used by viruses and some pathogenic bacteria to exit host cells is to hijack the autophagic pathway (69). Autophagy is a process carried out by eukaryotic cells in which cytoplasmic material is engulfed into double-membrane-bound vacuoles called autophagosomes that subsequently fuse with lysosomes for degradation (127). This pathway was originally described as being important for maintaining cellular homeostasis, but it is now evident that it is also important for host defense. Following cytosolic replication, some Francisella organisms reside inside autophagosomes (45), although it is not clear whether this is a host-induced response to control infection or a pathogen-induced mechanism to promote virulence. Francisella autophagosomes contain major histocompatibility complex (MHC) class II (105), suggesting that the host might induce autophagy to degrade bacteria and present their antigens to activate the immune system and control bacterial replication. However, F. tularensis modulates the expression of atg5, beclin 1, and several other autophagy-related genes (60) during macrophage infection, supporting the notion that Francisella may act to subvert this host defense system. Furthermore, autophagy was recently shown to regulate the level of AIM2 inflammasome activation by targeting these complexes for lysosomal degradation (206). Therefore, hijacking the autophagic pathway could serve as a critical step in F. tularensis pathogenesis by hampering inflammasome-mediated bacterial clearance and providing this pathogen with a method by which to escape from host cells.
Replicating in the cytosol of host cells without inducing an inflammatory response is one of the most challenging yet critical immune subversion tactics employed by F. tularensis during infection. Although many of the virulence determinants that aid in evasion of cytosolic defenses are not currently known, it is likely that there are numerous genes necessary to subvert recognition by multiple PRRs, activation of cell death pathways, and killing via autophagy.
Nutritional Defenses
Although subverting host defenses in the cytosol is essential to facilitate intracellular replication and virulence, Francisella must also ensure that it can acquire sufficient nutrients in this host cell compartment to sustain its high growth rate. The host, however, has developed mechanisms that limit the availability of specific nutrients, providing a “nutritional” defense against pathogens. The fight over nutrients highlights another critical aspect of host-pathogen interactions.
Many Francisella genes annotated as being involved in metabolism, for both energy production and biosynthesis of macromolecules, have been identified in screens for genes important in intracellular replication and in vivo virulence (130, 135, 140, 214, 232). These screens identified genes required for growth during infection but not in rich media, and they therefore suggest that Francisella must utilize distinct metabolic pathways in order to successfully survive and replicate within the host, compared to those required for growth in broth. Mapping the genes identified in these screens onto predicted pathways has highlighted specific metabolic pathways critical to Francisella replication in the host, such as the biosynthesis of purines and pyrimidines as well as gluconeogenesis and glycolysis. The importance of these pathways in Francisella intracellular replication has recently been reviewed by Meibom and Charbit (149). We will focus on the mechanisms utilized by Francisella to directly access nutrients, such as iron and tryptophan, which are actively limited by the host during infection.
Iron, one of the most thoroughly studied nutrients in terms of host-pathogen interactions, is necessary for a variety of enzymatic functions in bacteria and is a vital component of various redox reactions that take place during growth. Within the host, nearly 99.9% of total iron is intracellular (37). Extracellular iron is highly insoluble, which thus represents a barrier to its uptake (36). Therefore, the little iron present outside host cells is associated with iron storage molecules such as heme, transferrin, and lactoferrin (36, 228). Francisella can overcome this first barrier to iron acquisition by entering host cells, where the iron content is substantially higher. However, even within host cells, iron is often sequestered by iron-containing redox enzymes and iron storage proteins such as ferritin and lactoferrin, making its acquisition by the bacteria challenging (36, 228).
Furthermore, the host has additional mechanisms to try to sequester iron away from pathogens as it responds to infection. Past work has demonstrated that when macrophages sense LPS or are stimulated with IFN-γ, the host cells respond by decreasing expression of the transferrin receptor (TfR) that shuttles iron-bound transferrin into the cell (158). These signs of infection also signal the host to increase production of ferritin, which binds and sequesters free iron within host cells (185). Together, these host strategies decrease the availability of free iron both extracellularly and intracellularly.
Due in part to these mechanisms, serum iron concentrations significantly decrease in otherwise healthy human volunteers during F. tularensis SchuS4 inhalational infection (170), as occurs during other microbial infections (231). This can be correlated with the fact that instead of the decrease in TfR expression typically observed in response to infection, TfR is significantly upregulated during LVS infection (165) and even colocalizes with the early Francisella-containing phagosome (FCP) (165). Interestingly, TfR localization does not occur with killed bacteria, indicating that LVS actively alters host TfR localization through an unknown mechanism (165). Increases in TfR would lead to increased intracellular pools of iron, while decreasing serum iron levels. Also, unlike stimulation with IFN-γ and/or LPS, LVS infection does not induce expression of ferritin (165). Therefore, LVS may also actively prevent the host from sequestering the intracellular iron brought in by TfR, allowing the bacteria to utilize the increased iron pool. Together, these changes in host expression of TfR and ferritin, as well as TfR localization to the FCP, cause a significant increase in usable intracellular iron that can enhance Francisella replication (Fig. 6).
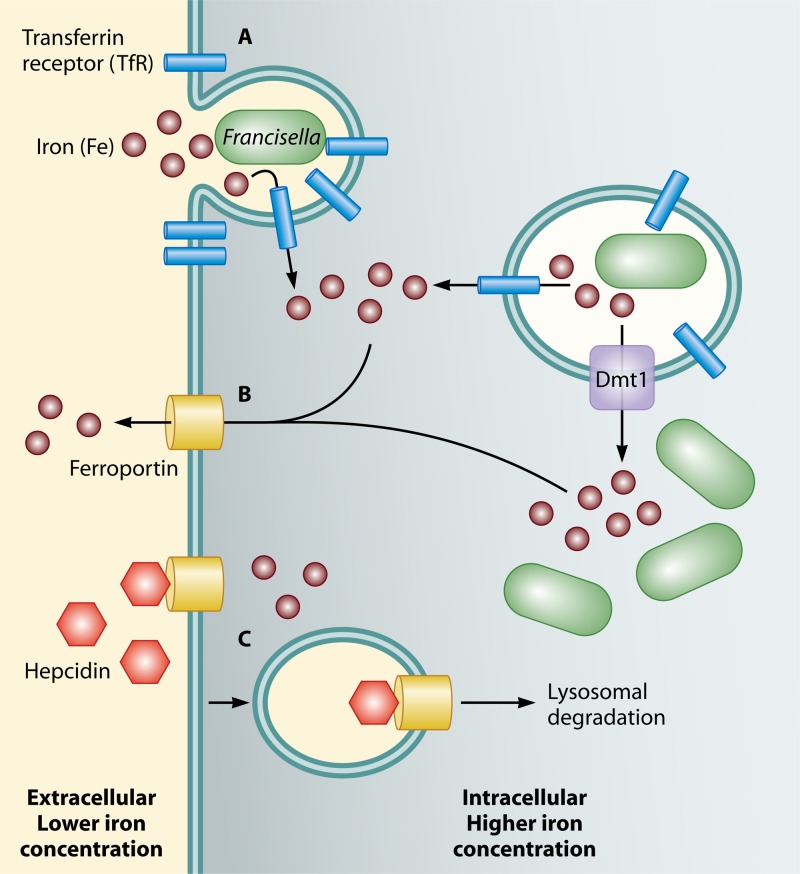
Fig 6.
Competition for iron during Francisella infection. (A) Upon infection of macrophages, Francisella induces the upregulation and relocalization of the transferrin receptor (TfR) to the forming phagosome, facilitating an influx of iron into the host cell. (B) The host counteracts this by upregulating Dmt1 and ferroportin, which export iron from the phagosome to the cytosol and from the cytosol to the extracellular space, respectively. (C) Francisella, however, counters this by inducing host hepcidin, which binds ferroportin and causes it to be degraded. In total, this leads to an increase of intracellular iron available in the cytosol for Francisella replication.
Further highlighting the tug-of-war between host and pathogen, the host counters the influx of iron due to increased TfR expression during LVS infection by increasing expression of Dmt1, a phagosomal membrane protein which transports iron from phagosomes into the cytosol, as well as ferroportin, which shuttles iron from the cytosol out of the cell (Fig. 6) (165). The combined action of these proteins would limit the amount of intracellular iron available to replicating Francisella in the cytosol. Countering this restriction, LVS infection induces the production of hepcidin, an antimicrobial peptide secreted by hepatic cells (165). Hepcidin binds to ferroportin on the host cell surface, causing it to become internalized and subsequently be degraded (162). Thus, hepcidin induction and subsequent inhibition of efflux of cytosolic iron appear to be one strategy employed by LVS to maintain intracellular iron pools at levels permissive for replication. However, whether LVS actively induces hepcidin production to specifically inhibit ferroportin or whether hepcidin's antiferroportin action is an unintentional consequence to the host and secondary to its antimicrobial activity is not known. The exact mechanisms by which Francisella promotes the expression of hepcidin and TfR, while simultaneously preventing ferritin expression, are unknown.
Further contributing to its ability to capture and acquire iron, Francisella utilizes a rhizoferrin-like siderophore (215). Siderophores are small molecules that selectively bind iron at a significantly higher affinity than host iron-sequestering compounds, creating a biochemical flux in favor of the siderophore-producing organism (154). This allows Francisella to outcompete the host for available iron. The Francisella locus necessary for production and utilization of its siderophore is fslABCDE (also termed figABCDE) (117, 215). fslABCD are directly involved in the production and/or export of the Francisella siderophore (117, 215). fslE, on the other hand, is not necessary for production or export, but instead is required for the utilization of the siderophore, through either siderophore binding or import (117, 182). Francisella lacks genes with significant similarity to the known (and typically highly conserved) siderophore receptor and importer genes tonB, exbB, and exbD (122). It is therefore hypothesized that fslE, or one of the two fslE orthologs present in the genome, may act as a novel siderophore importer (182). The two fslE-like genes, termed fupAB (encoded in LVS by a single protein fusion of these two paralogs, named fupA/B) (132, 205), are important for siderophore uptake, particularly in LVS compared to F. tularensis, which relies more heavily on fslE for siderophore import (132, 182, 205).
In the context of an infection, siderophore production and iron acquisition proteins are a vital component of Francisella's ability to maintain iron homeostasis and ultimately survive and replicate in the host. fslABC and feoB, encoding a ferrous iron importer utilized by other bacteria to directly import free iron (41, 122, 215), have each been identified in in vivo virulence screens (214, 232). fslABC have also been shown to be necessary for replication in macrophages (135). Furthermore, fupAB are required for full virulence in mice during LVS (205) and F. tularensis (223) infection and for LVS replication in macrophages (140, 205). Interestingly, while unable to produce siderophores, an fslA mutant of F. tularensis SchuS4 is not attenuated during infection of mice, suggesting that other iron acquisition pathways, such as the feoB importer or the fupAB system, may be playing a parallel role. On the other hand, while not identified in previous screens for virulence factors, we have identified fslE in a negative selection screen in F. tularensis for genes necessary for survival in mice (D. S. Weiss et al., unpublished observation).
Although iron acquisition may play an important role in pathogenesis, recent work has demonstrated that Francisella must also limit its total intracellular iron concentration (133). Increased intracellular iron concentrations can lead to increased susceptibility to oxygen radical damage within the bacteria. In fact, more virulent strains, such as F. tularensis SchuS4, have lower levels of intracellular iron than the less virulent LVS, which correlates with increased resistance to oxygen radical-mediated stresses (133). It is important to note that these studies were performed using bacteria grown on agar plates. It would be interesting to measure the iron content of each strain during infection to see if the differences mentioned above are maintained. Overall, these data suggest that Francisella maintains homeostasis of intracellular iron, importing enough to promote survival while preventing excessively high and toxic concentrations.
Although it is the most thoroughly studied, iron is not the only nutrient whose availability is actively limited by the host. The intracellular concentration of the amino acid tryptophan is significantly decreased during infection (218) due to an increase in the expression of indoleamine 2,3-dioxygenase (IDO), which initiates the tryptophan catabolic pathway by converting tryptophan to kynurenine (218). IDO is expressed in nearly every cell type, excluding hepatic cells, and is particularly highly expressed in the lungs and placenta (218). Infection by Chlamydia (26) or Streptococcus (172), as well as stimulation with IFN-γ (238) or LPS (237), induces IDO expression and results in decreased tryptophan levels. This is an important defensive mechanism, as it creates an innate nutritional barrier that limits the ability of microbes to replicate.
Upon intranasal infection with Francisella, IDO is significantly upregulated in the lung (172). Francisella is capable of overcoming this host defense by maintaining the ability to synthesize tryptophan de novo, although this process is very energy intensive (241). The importance of tryptophan prototrophy is exemplified by the fact that strains with mutations in trpB, which is necessary for the final step of tryptophan biosynthesis, are significantly attenuated during intranasal infection (50, 172) and that this attenuation can be overcome in IDO−/− mice (172). trpB mutants are capable of limited intracellular replication in macrophages and show decreased survival in IFN-γ-stimulated macrophages compared to wild-type Francisella (50). As IDO expression is induced by IFN-γ, the attenuation of the trpB mutant is also rescued upon infection of IFN-γ receptor (IFNGR)-deficient mice (50). These data indicate that tryptophan prototrophy is a necessary attribute of Francisella pathogenesis, allowing the bacterium to overcome an aspect of the nutritional barrier induced by IFN-γ and IDO upon infection of the lung.
While it can produce tryptophan, Francisella is auxotrophic for 13 amino acids which it cannot synthesize de novo, likely because it does not encode some or all of the enzymes required for their biosynthetic pathways (122, 149, 188). Therefore, it is intuitive that being able to acquire these essential amino acids within the host is a necessary component of Francisella pathogenesis. Francisella is predicted to lack all of the biosynthetic enzymes for arginine, histidine, lysine, methionine, and tyrosine (122). Interestingly, a putative methionine uptake transporter (FTN_1107/FTT1125) (140, 214), lysine importer (FTN_0296/FTT1633c) (232), and tyrosine permease (FTN_1711/FTT1732c) (11) have all been identified in virulence screens, suggesting that they are necessary for pathogenesis. Furthermore, amino acid importers with no predicted substrate (FTN_0097/FTT1688 and FTN_0848/FTT0968c) have been identified as well (130, 135, 140, 214, 232). This suggests that Francisella may have developed strategies to scavenge from the host those amino acids which it cannot synthesize de novo.
Other than tryptophan restriction, to the best of our knowledge, we do not know of another example of active host depletion of an amino acid that is important during Francisella infection. While numerous biosynthetic and metabolic pathways have been lost during the reductive evolution of the more recently evolving F. tularensis and F. holarctica subspecies, Francisella species are capable of de novo biosynthesis of alanine, glutamate, glutamine, asparagine, glycine, phenylalanine, and tryptophan (122, 149, 188). In fact, pathways for the synthesis of glutamate, glycine, and phenylalanine are redundantly represented. This suggests that the ability to synthesize these amino acids is essential for pathogenesis and could potentially counter active host restriction mechanisms, similar to the way Francisella counters tryptophan depletion as described above. In further support of this idea, asparagine synthase (FTN_1421/FTT1456c) which is necessary for asparagine prototrophy (130, 135, 232), and a number of enzymes utilized for the creation of aromatic amino acid precursors have been identified in virulence screens (149).
Another well-known example of Francisella auxotrophy is the requirement for cysteine, which serves as a necessary source of sulfate. This requirement is also necessary in broth and thus does not necessarily represent an active host defense process to deplete cysteine. However, in the host, cysteine is present within glutathione (GSH). The ability to utilize GSH is thus a required component of Francisella intracellular growth. Indeed, F. tularensis mutants lacking a transpeptidase (ggt) necessary for cleavage of GSH are severely attenuated during infection of macrophages and mice (3).
The requirement for other biosynthetic pathways during host infection, such as purine and pyrimidine biosynthesis, is well established for Francisella and other bacterial pathogens (10, 149). Furthermore, the importance of central metabolism (e.g., glycolysis) has also been shown to play a role during infection (149). Additionally, there is evidence that nutritional status may play a role in allowing Francisella to sense the host phagosome compared to being in the host cytosol (40) and to induce expression of virulence genes, including those required for phagosomal escape (40, 67, 191). Mutants that are unable to recognize particular metabolites, such as polyamines present in the host cytosol, are significantly attenuated and unable to alter temporal expression of virulence factors (191). Taken together, these data highlight the importance in the outcome of infection of host and pathogen proteins involved in this nutritional tug-of-war.
Modulation of Adaptive Immune Responses
We have focused a great deal of attention on the ways in which Francisella modulates the immune responsiveness of the cells that it directly infects. We now discuss the effect that these interactions have on the ability of other immune cells to fight infection and the development of an effective adaptive immune response. As noted previously, pulmonary infection caused by F. tularensis in humans is characterized by an initial delay in inflammatory responses during the first 72 h (58). Although it is not entirely clear how F. tularensis evades early immune responses, modulating the activity of innate immune cells that contribute to early cell-mediated immunity likely plays a significant role. Antigen-presenting cells (APCs), such as macrophages and dendritic cells, play an important role in early innate defense against Francisella infection. The ability of these cells to present microbial antigens and activate T cells enables them to serve as a bridge between the innate and adaptive immune systems. During the earliest stages of infection, Francisella resides primarily in macrophages (97). Therefore, it is not surprising that this pathogen uses multiple subversion mechanisms (e.g., facilitating cell entry via nonactivating macrophage receptors, escaping phagosomal killing, and modulating cytosolic defense pathways) to systematically disarm macrophage defenses. However, when activated by the proinflammatory cytokine IFN-γ, these phagocytes are capable of overcoming the modulatory effects exerted by Francisella, preventing its replication in the cytosol (68, 126). Primarily produced by NK cells and dendritic cells during infection, IFN-γ regulates over 200 genes, many of which are involved in host defense pathways important for enhancing nitric oxide production, inducing autophagy, and increasing major histocompatibility complex (MHC) class I and II antigen presentation (32, 226). Activation of these pathways could collectively enhance Francisella clearance and promote host survival. Moreover, a lack of IFN-γ in vivo, through either genetic disruption (76) or anticytokine treatment (126), increased the susceptibility of mice to Francisella infection, indicating that this cytokine is an important mediator of host defense in vivo.
The exact mechanism of IFN-γ-mediated clearance of Francisella from activated macrophages is unclear. IFN-γ can hinder F. novicida and LVS escape from the phagosomes of human monocyte-derived macrophages and murine peritoneal exudate cells, respectively (131, 195). This prolonged retention of Francisella in the phagosome would expose bacteria to phagosomal defenses such as the NADPH oxidase, nitric oxide, and lysosomal enzymes if phagolysosomal fusion occurred (201), all of which are upregulated by IFN-γ, leading to bacterial clearance. In contrast, escape of highly virulent F. tularensis from the phagosome is not affected by IFN-γ activation of macrophages, although its cytosolic replication is nonetheless restricted (74). Therefore, IFN-γ-mediated activation of macrophages leads to both phagosomal and cytosolic defenses that control the virulence of multiple Francisella species.
Since IFN-γ has such a pleiotropic effect on macrophage activation, any mechanism of suppressing the action of this cytokine would likely have broad effects on promoting Francisella pathogenesis. Francisella directly impairs the ability of macrophages to respond to IFN-γ stimulation during infection by downregulating the expression of the alpha subunit of the IFN-γ receptor (IFNGR), which is critical for signaling (189). Nallaparaju et al. found that in the event that signaling through the IFNGR does occur, Francisella uses FopC to indirectly block the phosphorylation and subsequent activation of Stat1 (a transcriptional regulator that is required for IFNGR signaling) (159a, 166). The IFN-γ signaling pathway is further decommissioned during infection as the bacteria induce an increase in the expression of SOCS3, a protein that negatively regulates this pathway (38). Activation of the TLR signaling pathway enhances IFN-γ-dependent cytokine production in macrophages (242). Therefore, suppression of TLR signaling by mechanisms discussed earlier in this review (prevention of PAMP release, modification of LPS/lipid A structure to prevent TLR4 activation, and downregulation of TLR expression) could also dampen activation of the IFN-γ pathway.
In addition to suppressing IFN-γ signaling, F. tularensis induces macrophages to produce the immunomodulatory lipid prostaglandin E2 (PGE2) (236). PGE2 inhibits T cells from producing IFN-γ, favoring bacterial replication (Fig. 7) (175). PGE2 also induces the expression of a >10-kDa protease-resistant host factor that promotes ubiquitin-dependent lysosomal degradation of MHC class II molecules, resulting in a nearly complete absence of MHC class II on the surface of macrophages (235). Therefore, by inducing PGE2, Francisella inhibits bacterial killing mediated by IFN-γ and also dampens the induction of adaptive immune responses by limiting antigen presentation through MHC class II.
Fig 7.
Subversion of adaptive immune responses by Francisella. Francisella infects antigen-presenting cells (APCs) and dampens their ability to produce proinflammatory cytokines. IL-12 is blocked via the induction of beta interferon (IFN-β). IFN-γ is produced by dendritic cells and T cells during Francisella infection, and it induces macrophages to kill the bacteria. Francisella blocks signal transduction through the IFN-γ receptor (IFNGR) by inhibiting Stat1 phosphorylation and through the induction of prostaglandin E2 (PGE2), which inhibits IFN-γ production by T cells. Francisella also hampers T cell activation by inducing the degradation of major histocompatibility complex class II (MHC II) (indicated by its absence from the cell surface, tagging with ubiquitin [Ub], and trafficking toward the lysosome) and directing APCs to produce the anti-inflammatory cytokines TGF-β and IL-10, which promote the development of regulatory T cells (Treg) that are able to suppress inflammation and cell-mediated immune responses.
Dendritic cells play a critical role in directing cell-mediated immunity by producing cytokines that control the way in which T cells are activated and the immune responses that they subsequently promote. In accordance with this pathogen's ability to dampen cell-mediated immune responses, F. tularensis suppresses proinflammatory cytokine production from these cells (35). Among these cytokines is IL-12, which is important for the development of IFN-γ-producing T cells (75). Surprisingly, inhibition of IL-12 by F. tularensis is mediated by IFN-β production (Fig. 7) (24). As described previously in this review, type I IFNs contribute to inflammasome activation by Francisella. However, there was no correlation between IFN-β production and inflammasome activation during F. tularensis infection. Considering that F. tularensis dampens the expression of genes in the type I IFN pathway, it is possible that this pathogen suppresses the expression of type I IFNs to a level that is below the threshold for inflammasome activation but adequate for IL-12 suppression.
F. tularensis also redirects macrophages to become alternatively activated (207). Alternatively activated macrophages exhibit a skewed inflammatory profile that is less antimicrobial, with decreased levels of nitric oxide production upon IFN-γ stimulation and an impaired ability to kill bacteria (78, 93). Additionally, F. tularensis directs dendritic cells to produce the anti-inflammatory cytokines transforming growth factor β (TGF-β) and IL-10 (35, 236). This greatly contributes to the absence of an inflammatory response early in infection because IL-10 inhibits macrophage proliferation and proinflammatory cytokine production (164). In addition, it can lead to reduced MHC class II expression, which would suppress adaptive immune responses (139).
IL-10 or TGF-β can direct dendritic cells to become tolerogenic, inhibiting their ability to activate T cells that induce an inflammatory response (Fig. 7) (139). These cytokines also promote the development of regulatory T cells that can suppress the inflammatory activity of other T cells (85). In fact, pulmonary regulatory T cells develop in the lungs during F. tularensis infection, and this development correlates with an increase in bacterial burden (173). Though it is currently unclear how F. tularensis redirects APCs toward an anti-inflammatory response, inducing immune tolerance through the induction of IL-10 or TGF-β is a survival strategy used by pathogens such as Yersinia pestis, Coxiella burnetii, and Chlamydia pneumoniae (63). In the case of Yersinia pestis, IL-10 production is induced by the type III secretion system effector protein LcrV (124, 208). Further characterization of Francisella secretion systems and effector proteins may shed light on similar mechanisms of immune modulation.
Cell-mediated immunity is generally accepted as being essential for controlling intracellular bacterial infection. In accordance with this dogma, antibodies have been shown to provide little protection against Francisella infection, but antibody-independent B cell responses are important for early protection (62). However, Francisella, like some other intracellular bacterial pathogens (83), is also capable of infecting and replicating inside B cells (119), although it is not known whether it can modulate B cell activity in order to evade immune detection. Salmonella enterica serovar Typhimurium has been shown to induce MyD88-dependent IL-10 production in splenic B cells during systemic infection, promoting its survival in vivo (163). As discussed above, IL-10 induces a tolerogenic state in immune cells, and it is possible that Francisella-induced IL-10 production in APCs could also alter the function of B cells during infection.
Francisella uses multiple strategies to modulate APCs in order to subvert subsequent activation of the adaptive immune system. This modulation not only contributes to the failure in immune activation during the initial stages of infection but also helps provide a suitable environment for replication in subsequent phases of infection by dampening adaptive responses such as the activation of CD8 T cells, which could kill infected host cells harboring Francisella.
CONCLUDING REMARKS
Francisella uses a vast arsenal of virulence mechanisms to subvert host defenses and promote intracellular replication. The main overarching strategy is to prevent the initiation of host inflammatory responses. Since the inflammatory response operates in a positive-feedback loop, it is powerful once started and very difficult, if not impossible, to reign in. If initiated, host defenses encountered by the bacteria (extracellular, phagosomal, and cytosolic defenses) are heightened at each step of infection, taking a toll on replication levels. This is highlighted by comparing the intracellular fates of Francisella in unstimulated or IFN-γ-stimulated macrophages. While Francisella is able to grow rapidly in unstimulated macrophages, IFN-γ-stimulated macrophages are not permissive for replication and are instead able to kill the bacteria. Therefore, if inflammatory processes leading to IFN-γ production are initiated, this drastically alters the course of infection in favor of the host.
Since PAMPs activate inflammatory pathways during infection, a critical aspect of subverting the inflammatory response is preventing PAMP recognition by the host. Francisella achieves this in large part by devoting considerable resources to resisting damage (capsule, LPS O antigen, lipid A modifications, efflux pumps, detoxifying ROS, etc.) induced by antimicrobials that would lead to the release of PAMPs. When PAMPs are inevitably liberated and recognized by the host at some level, Francisella suppresses the signaling of host PRRs, although the mechanisms by which this occurs are not defined and will be a critical area of future research. The importance of subverting early inflammatory responses can be seen when studying numerous mutant strains that lack virulence traits required in the first hours of infection (resistance to complement and phagosomal defenses, defects in phagosome escape, etc), which are often hyperinflammatory and severely attenuated for virulence.
Remaining immunologically “silent” allows the bacteria to reach the host cytosol, where they can replicate. However, an often overlooked virulence trait is the ability to acquire sufficient nutrients to facilitate rapid intracellular replication. Francisella has several strategies to acquire iron and synthesize tryptophan de novo to subvert specific host defense pathways. As mentioned, numerous metabolic genes have been identified in virulence screens and may lead to the elucidation of novel mechanisms by which the bacteria subvert “nutritional defenses.” This is a subject of great interest not just for Francisella pathogenesis but also in the field of bacterial pathogenesis as a whole. As it is one of the most virulent bacterial pathogens known, continued research into how Francisella subverts innate and adaptive host responses will lead to novel discoveries that will enhance our understanding of host-pathogen interactions.
ACKNOWLEDGMENTS
We thank Thomas Kawula for critical reading of the manuscript.
We attempted to mention all of the relevant studies and have undoubtedly omitted some. We apologize in advance to those authors whose work we did not cite.
This work was supported by NIH grants U54-AI057157 from the Southeastern Regional Center of Excellence for Emerging Infections and Biodefense and R56-AI087673 (D.S.W.), the National Science Foundation Graduate Research Fellowship (C.L.J. and T.R.S.), and the ARCS Foundation (T.R.S.).
The contents of this article are solely the responsibility of the authors and do not necessarily represent the official views of the NIH.
Biographies

Crystal L. Jones obtained her B.S. in biology with honors from Georgia State University in 2005. There she studied the molecular mechanisms of iron uptake in Bartonella quintana. She is currently a Ph.D. candidate in the Microbiology and Molecular Genetics Program at Emory University, conducting research in the laboratory of Dr. David Weiss. Her research focuses on understanding the role of TLR2 in host defense against Francisella infection. She was awarded the National Science Foundation Graduate Research Fellowship.

Brooke Napier obtained her undergraduate degree in neuroscience from Agnes Scott College in 2007. She then worked with Dr. Monica Farley's group in the Division of Infectious Diseases, Emory University School of Medicine, examining capsule-related sequences in the nonencapsulated human pathogen Haemophilus influenzae. She is currently working on a Ph.D. in microbiology and molecular genetics at Emory University in the lab of Dr. David Weiss. Her dissertation work focuses on elucidating novel mechanisms that pathogenic bacteria employ to evade the host immune response.

Tim Sampson received his B.S. in microbiology from the University of Pittsburgh, where he studied mycobacteriophage genetics with Dr. Graham Hatfull. There, he helped identify the first transposons within the mycobacteriophage population. He is currently a Ph.D. student in microbiology at Emory University, studying under Dr. David Weiss. He is a recipient of the NSF Graduate Research Fellowship and was named an ARCS Scholar. His current work focuses on the genetic mechanisms utilized by Francisella to evade recognition by the innate immune system, and he hopes to apply this knowledge to the development of novel vaccination strategies.

Anna Llewellyn completed her B.S. at Georgia State University in 2006. Her undergraduate research project focused on quorum sensing and interspecies signaling. After a three-month appointment in a Peruvian multidrug-resistant tuberculosis clinical laboratory, she began working on her Ph.D. in microbiology and molecular genetics at Emory University in 2007. As a member of Dr. David Weiss' laboratory, her dissertation work has concentrated on elucidation of Francisella virulence mechanisms and intracellular host-pathogen interactions.

Max R. Schroeder is a graduate student at Emory University. He obtained his B.A. in microbiology from Ohio Wesleyan University, Delaware, OH. There, he studied the evolution of avian plumage coloration and bacterial feather degradation. His research interests are in molecular pathogenesis, and he is currently working on the genetic determinants of Streptococcus pneumoniae pathogenesis.

David S. Weiss is an Assistant Professor of Infectious Diseases at the Emory University School of Medicine and Emory Vaccine Center. He completed his Ph.D. on Toll-like receptors at the New York University School of Medicine under the guidance of Dr. Arturo Zychlinsky. His postdoctoral work in Dr. Denise Monack's lab at Stanford University focused on Francisella tularensis pathogenesis and inflammasome activation. His lab currently studies mechanisms of bacterial pathogenesis and host defense using Francisella, as well as mechanisms of antibiotic resistance using the nosocomial pathogen Acinetobacter baumannii.
REFERENCES
- 1. Abplanalp AL, Morris IR, Parida BK, Teale JM, Berton MT. 2009. TLR-dependent control of Francisella tularensis infection and host inflammatory responses. PLoS One 4:e7920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Akimana C, Kwaik Y. 2011. Francisella-arthropod vector interaction and its role in patho-adaptation to infect mammals. Front. Microbiol. 2:34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Alkhuder K, Meibom KL, Dubail I, Dupuis M, Charbit A. 2009. Glutathione provides a source of cysteine essential for intracellular multiplication of Francisella tularensis. PLoS Pathog. 5:e1000284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Allen LA, Beecher BR, Lynch JT, Rohner OV, Wittine LM. 2005. Helicobacter pylori disrupts NADPH oxidase targeting in human neutrophils to induce extracellular superoxide release. J. Immunol. 174:3658–3667 [DOI] [PubMed] [Google Scholar]
- 5. Amer LS, Bishop BM, van Hoek ML. 2010. Antimicrobial and antibiofilm activity of cathelicidins and short, synthetic peptides against Francisella. Biochem. Biophys. Res. Commun. 396:246–251 [DOI] [PubMed] [Google Scholar]
- 6. Ancuta P, Pedron T, Girard R, Sandstrom G, Chaby R. 1996. Inability of the Francisella tularensis lipopolysaccharide to mimic or to antagonize the induction of cell activation by endotoxins. Infect. Immun. 64:2041–2046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Andersson H, et al. 2006. Transcriptional profiling of host responses in mouse lungs following aerosol infection with type A Francisella tularensis. J. Med. Microbiol. 55:263–271 [DOI] [PubMed] [Google Scholar]
- 8. Anthony LS, Skamene E, Kongshavn PA. 1988. Influence of genetic background on host resistance to experimental murine tularemia. Infect. Immun. 56:2089–2093 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Apicella MA, et al. 2010. Identification, characterization and immunogenicity of an O-antigen capsular polysaccharide of Francisella tularensis. PLoS One 5:e11060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Appelberg R. 2006. Macrophage nutriprive antimicrobial mechanisms. J. Leukoc. Biol. 79:1117–1128 [DOI] [PubMed] [Google Scholar]
- 11. Asare R, Abu Kwaik Y. 2010. Molecular complexity orchestrates modulation of phagosome biogenesis and escape to the cytosol of macrophages by Francisella tularensis. Environ. Microbiol. 12:2559–2586 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Asong J, Wolfert MA, Maiti KK, Miller D, Boons GJ. 2009. Binding and cellular activation studies reveal that Toll-like receptor 2 can differentially recognize peptidoglycan from Gram-positive and Gram-negative bacteria. J. Biol. Chem. 284:8643–8653 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Atianand MK, et al. 2011. Francisella tularensis reveals a disparity between human and mouse NLRP3 inflammasome activation. J. Biol. Chem. 286:39033–39042 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Babior BM. 2002. The leukocyte NADPH oxidase. Isr. Med. Assoc. J. 4:1023–1024 [PubMed] [Google Scholar]
- 15. Bakshi CS, et al. 2008. An improved vaccine for prevention of respiratory tularemia caused by Francisella tularensis SchuS4 strain. Vaccine 26:5276–5288 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Bakshi CS, et al. 2006. Superoxide dismutase B gene (sodB)-deficient mutants of Francisella tularensis demonstrate hypersensitivity to oxidative stress and attenuated virulence. J. Bacteriol. 188:6443–6448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Balagopal A, et al. 2006. Characterization of the receptor-ligand pathways important for entry and survival of Francisella tularensis in human macrophages. Infect. Immun. 74:5114–5125 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Bandara AB, et al. 2011. Isolation and mutagenesis of a capsule-like complex (CLC) from Francisella tularensis, and contribution of the CLC to F. tularensis virulence in mice. PLoS One 6:e19003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Barel M, et al. 2008. A novel receptor-ligand pathway for entry of Francisella tularensis in monocyte-like THP-1 cells: interaction between surface nucleolin and bacterial elongation factor Tu. BMC Microbiol. 8:145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Barker JH, et al. 2009. The role of complement opsonization in interactions between F. tularensis subsp. novicida and human neutrophils. Microbes Infect. 11:762–769 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Barker JR, et al. 2009. The Francisella tularensis pathogenicity island encodes a secretion system that is required for phagosome escape and virulence. Mol. Microbiol. 74:1459–1470 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Baron GS, Reilly TJ, Nano FE. 1999. The respiratory burst-inhibiting acid phosphatase AcpA is not essential for the intramacrophage growth or virulence of Francisella novicida. FEMS Microbiol. Lett. 176:85–90 [DOI] [PubMed] [Google Scholar]
- 23. Baucheron S, et al. 2004. AcrAB-TolC directs efflux-mediated multidrug resistance in Salmonella enterica serovar Typhimurium DT104. Antimicrob. Agents Chemother. 48:3729–3735 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Bauler TJ, Chase JC, Bosio CM. 2011. IFN-beta mediates suppression of IL-12p40 in human dendritic cells following infection with virulent Francisella tularensis. J. Immunol. 187:1845–1855 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Beasley AS, et al. 2012. A variety of novel lipid a structures obtained from Francisella tularensis live obtained from Francisella tularensis live. Innate Immun. 18:268–278 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Beatty WL, Belanger TA, Desai AA, Morrison RP, Byrne GI. 1994. Tryptophan depletion as a mechanism of gamma interferon-mediated chlamydial persistence. Infect. Immun. 62:3705–3711 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Beauregard KE, Lee KD, Collier RJ, Swanson JA. 1997. pH-dependent perforation of macrophage phagosomes by listeriolysin O from Listeria monocytogenes. J. Exp. Med. 186:1159–1163 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Ben Nasr A, et al. 2006. Critical role for serum opsonins and complement receptors CR3 (CD11b/CD18) and CR4 (CD11c/CD18) in phagocytosis of Francisella tularensis by human dendritic cells (DC): uptake of Francisella leads to activation of immature DC and intracellular survival of the bacteria. J. Leukoc. Biol. 80:774–786 [DOI] [PubMed] [Google Scholar]
- 29. Ben Nasr A, Klimpel GR. 2008. Subversion of complement activation at the bacterial surface promotes serum resistance and opsonophagocytosis of Francisella tularensis. J. Leukoc. Biol. 84:77–85 [DOI] [PubMed] [Google Scholar]
- 30. Bina XR, Lavine CL, Miller MA, Bina JE. 2008. The AcrAB RND efflux system from the live vaccine strain of Francisella tularensis is a multiple drug efflux system that is required for virulence in mice. FEMS Microbiol. Lett. 279:226–233 [DOI] [PubMed] [Google Scholar]
- 31. Birdsell D, et al. 2009. Francisella tularensis subsp. novicida isolated from a human in Arizona. BMC Res. Notes 2:223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Boehm U, Klamp T, Groot M, Howard JC. 1997. Cellular responses to interferon-gamma. Annu. Rev. Immunol. 15:749–795 [DOI] [PubMed] [Google Scholar]
- 33. Boneca IG, et al. 2007. A critical role for peptidoglycan N-deacetylation in Listeria evasion from the host innate immune system. Proc. Natl. Acad. Sci. U. S. A. 104:997–1002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Bosio CM, Bielefeldt-Ohmann H, Belisle JT. 2007. Active suppression of the pulmonary immune response by Francisella tularensis Schu4. J. Immunol. 178:4538–4547 [DOI] [PubMed] [Google Scholar]
- 35. Bosio CM, Dow SW. 2005. Francisella tularensis induces aberrant activation of pulmonary dendritic cells. J. Immunol. 175:6792–6801 [DOI] [PubMed] [Google Scholar]
- 36. Braun V. 2001. Iron uptake mechanisms and their regulation in pathogenic bacteria. Int. J. Med. Microbiol. 291:67–79 [DOI] [PubMed] [Google Scholar]
- 37. Bridges K, Seligman P. 1995. Disorders of iron metabolism. JB Lippincott Company, London, United Kingdom [Google Scholar]
- 38. Butchar JP, et al. 2008. Microarray analysis of human monocytes infected with Francisella tularensis identifies new targets of host response subversion. PLoS One 3:e2924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Campos M, et al. 2004. Capsule polysaccharide mediates bacterial resistance. Infect. Immun. 72:7107–7114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Carlson PE, Jr, et al. 2009. Global transcriptional response to spermine, a component of the intramacrophage environment, reveals regulation of Francisella gene expression through insertion sequence elements. J. Bacteriol. 191:6855–6864 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Cartron ML, Maddocks S, Gillingham P, Craven CJ, Andrews SC. 2006. Feo-transport of ferrous iron into bacteria. Biometals 19:143–157 [DOI] [PubMed] [Google Scholar]
- 42. Cederlund A, Gudmundsson G, Agerberth B. 2011. Antimicrobial peptides important in innate immunity. FEBS J. 278:3942–3951 [DOI] [PubMed] [Google Scholar]
- 43. Champion MD, et al. 2009. Comparative genomic characterization of Francisella tularensis strains belonging to low and high virulence subspecies. PLoS Pathog. 5:e1000459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Chase JC, Celli J, Bosio CM. 2009. Direct and indirect impairment of human dendritic cell function by virulent Francisella tularensis Schu S4. Infect. Immun. 77:180–195 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Checroun C, Wehrly TD, Fischer ER, Hayes SF, Celli J. 2006. Autophagy-mediated reentry of Francisella tularensis into the endocytic compartment after cytoplasmic replication. Proc. Natl. Acad. Sci. U. S. A. 103:14578–14583 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Chen W, KuoLee R, Shen H, Busa M, Conlan JW. 2004. Toll-like receptor 4 (TLR4) does not confer a resistance advantage on mice against low-dose aerosol infection with virulent type A Francisella tularensis. Microb. Pathog. 37:185–191 [DOI] [PubMed] [Google Scholar]
- 47. Chen W, Kuolee R, Shen H, Busa M, Conlan JW. 2005. Toll-like receptor 4 (TLR4) plays a relatively minor role in murine defense against primary intradermal infection with Francisella tularensis LVS. Immunol. Lett. 97:151–154 [DOI] [PubMed] [Google Scholar]
- 48. Child R, Wehrly TD, Rockx-Brouwer D, Dorward DW, Celli J. 2010. Acid phosphatases do not contribute to the pathogenesis of type A Francisella tularensis. Infect. Immun. 78:59–67 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Chong A, et al. 2008. The early phagosomal stage of Francisella tularensis determines optimal phagosomal escape and Francisella pathogenicity island protein expression. Infect. Immun. 76:5488–5499 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Chu P, Rodriguez AR, Arulanandam BP, Klose KE. 2011. Tryptophan prototrophy contributes to Francisella tularensis evasion of gamma interferon-mediated host defense. Infect. Immun. 79:2356–2361 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Clay CD, Soni S, Gunn JS, Schlesinger LS. 2008. Evasion of complement-mediated lysis and complement C3 deposition are regulated by Francisella tularensis lipopolysaccharide O antigen. J. Immunol. 181:5568–5578 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Clemens DL, Lee BY, Horwitz MA. 2005. Francisella tularensis enters macrophages via a novel process involving pseudopod loops. Infect. Immun. 73:5892–5902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Clemens DL, Lee BY, Horwitz MA. 2009. Francisella tularensis phagosomal escape does not require acidification of the phagosome. Infect. Immun. 77:1757–1773 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Clinton SR, Bina JE, Hatch TP, Whitt MA, Miller MA. 2010. Binding and activation of host plasminogen on the surface of Francisella tularensis. BMC Microbiol. 10:76. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Cole LE, et al. 2007. Toll-like receptor 2-mediated signaling requirements for Francisella tularensis live vaccine strain infection of murine macrophages. Infect. Immun. 75:4127–4137 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Collazo CM, Sher A, Meierovics AI, Elkins KL. 2006. Myeloid differentiation factor-88 (MyD88) is essential for control of primary in vivo Francisella tularensis LVS infection, but not for control of intra-macrophage bacterial replication. Microbes Infect. 8:779–790 [DOI] [PubMed] [Google Scholar]
- 57. Colquhoun D, Duobu S. 2011. Francisella infections in farmed and wild aquatic organisms. Vet. Res. 42:47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Conlan JW, Chen W, Shen H, Webb A, KuoLee R. 2003. Experimental tularemia in mice challenged by aerosol or intradermally with virulent strains of Francisella tularensis: bacteriologic and histopathologic studies. Microb. Pathog. 34:239–248 [DOI] [PubMed] [Google Scholar]
- 59. Crane DD, Warner SL, Bosio CM. 2009. A novel role for plasmin-mediated degradation of opsonizing antibody in the evasion of host immunity by virulent, but not attenuated, Francisella tularensis. J. Immunol. 183:4593–4600 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Cremer TJ, Amer A, Tridandapani S, Butchar JP. 2009. Francisella tularensis regulates autophagy-related host cell signaling pathways. Autophagy 5:125–128 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Cremer TJ, et al. 2011. Akt-mediated proinflammatory response of mononuclear phagocytes infected with Burkholderia cenocepacia occurs by a novel GSK3β-dependent, IκB kinase-independent mechanism. J. Immunol. 187:635–643 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Culkin SJ, Rhinehart-Jones T, Elkins KL. 1997. A novel role for B cells in early protective immunity to an intracellular pathogen, Francisella tularensis strain LVS. J. Immunol. 158:3277–3284 [PubMed] [Google Scholar]
- 63. Cyktor JC, Turner J. 2011. Interleukin-10 and immunity against prokaryotic and eukaryotic intracellular pathogens. Infect. Immun. 79:2964–2973 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. de Bruin OM, et al. 2011. The biochemical properties of the Francisella pathogenicity island (FPI)-encoded proteins IglA, IglB, IglC, PdpB and DotU suggest roles in type VI secretion. Microbiology 157:3483–3491 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. de Bruin OM, Ludu JS, Nano FE. 2007. The Francisella pathogenicity island protein IglA localizes to the bacterial cytoplasm and is needed for intracellular growth. BMC Microbiol. 7:1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Decker T, Muller M, Stockinger S. 2005. The yin and yang of type I interferon activity in bacterial infection. Nat. Rev. Immunol. 5:675–687 [DOI] [PubMed] [Google Scholar]
- 67. Deng K, Blick R, Liu W, Hansen E. 2006. Identification of Francisella tularensis genes affected by iron limitation. Infect. Immun. 74:4224–4236 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. De Pascalis R, Taylor BC, Elkins KL. 2008. Diverse myeloid and lymphoid cell subpopulations produce gamma interferon during early innate immune responses to Francisella tularensis live vaccine strain. Infect. Immun. 76:4311–4321 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Deretic V, Levine B. 2009. Autophagy, immunity, and microbial adaptations. Cell Host Microbe 5:527–549 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Desvaux M, Hébraud M. 2006. The protein secretion systems in Listeria: inside out bacterial virulence. FEMS Microbiol. Rev. 30:774–805 [DOI] [PubMed] [Google Scholar]
- 71. Drysdale M, et al. 2005. Capsule synthesis by Bacillus anthracis is required for dissemination in murine inhalation anthrax. EMBO J. 24:221–227 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Duenas AI, et al. 2006. Francisella tularensis LPS induces the production of cytokines in human monocytes and signals via Toll-like receptor 4 with much lower potency than E. coli LPS. Int. Immunol. 18:785–795 [DOI] [PubMed] [Google Scholar]
- 73. Dziarski R, Gupta D. 2005. Staphylococcus aureus peptidoglycan is a Toll-like receptor 2 activator: a reevaluation. Infect. Immun. 73:5212–5216 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Edwards JA, Rockx-Brouwer D, Nair V, Celli J. 2010. Restricted cytosolic growth of Francisella tularensis subsp. tularensis by IFN-gamma activation of macrophages. Microbiology 156:327–339 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Elkins KL, Cooper A, Colombini SM, Cowley SC, Kieffer TL. 2002. In vivo clearance of an intracellular bacterium, Francisella tularensis LVS, is dependent on the p40 subunit of interleukin-12 (IL-12) but not on IL-12 p70. Infect. Immun. 70:1936–1948 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Elkins KL, Rhinehart-Jones TR, Culkin SJ, Yee D, Winegar RK. 1996. Minimal requirements for murine resistance to infection with Francisella tularensis LVS. Infect. Immun. 64:3288–3293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Elkins KL, Rhinehart-Jones TR, Stibitz S, Conover JS, Klinman DM. 1999. Bacterial DNA containing CpG motifs stimulates lymphocyte-dependent protection of mice against lethal infection with intracellular bacteria. J. Immunol. 162:2291–2298 [PubMed] [Google Scholar]
- 78. Elser B, et al. 2002. IFN-gamma represses IL-4 expression via IRF-1 and IRF-2. Immunity 17:703–712 [DOI] [PubMed] [Google Scholar]
- 79. Eneslatt K, et al. 2011. Persistence of cell-mediated immunity three decades after vaccination with the live vaccine strain of Francisella tularensis. Eur. J. Immunol. 41:974–980 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Fadok VA, et al. 1998. Macrophages that have ingested apoptotic cells in vitro inhibit proinflammatory cytokine production through autocrine/paracrine mechanisms involving TGF-beta, PGE2, and PAF. J. Clin. Invest. 101:890–898 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Fernandes-Alnemri T, Yu JW, Datta P, Wu J, Alnemri ES. 2009. AIM2 activates the inflammasome and cell death in response to cytoplasmic DNA. Nature 458:509–513 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Fernandes-Alnemri T, et al. 2010. The AIM2 inflammasome is critical for innate immunity to Francisella tularensis. Nat. Immunol. 11:385–393 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Fillatreau S. 2011. Novel regulatory functions for Toll-like receptor-activated B cells during intracellular bacterial infection. Immunol. Rev. 240:52–71 [DOI] [PubMed] [Google Scholar]
- 84. Fink SL, Cookson BT. 2005. Apoptosis, pyroptosis, and necrosis: mechanistic description of dead and dying eukaryotic cells. Infect. Immun. 73:1907–1916 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Fontenot JD, Rudensky AY. 2005. A well adapted regulatory contrivance: regulatory T cell development and the forkhead family transcription factor Foxp3. Nat. Immunol. 6:331–337 [DOI] [PubMed] [Google Scholar]
- 86. Forestal CA, et al. 2008. A conserved and immunodominant lipoprotein of Francisella tularensis is proinflammatory but not essential for virulence. Microb. Pathog. 44:512–523 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87. Forestal CA, et al. 2007. Francisella tularensis has a significant extracellular phase in infected mice. J. Infect. Dis. 196:134–137 [DOI] [PubMed] [Google Scholar]
- 88. Fujita T, Matsushita M, Endo Y. 2004. The lectin-complement pathway—its role in innate immunity and evolution. Immunol. Rev. 198:185–202 [DOI] [PubMed] [Google Scholar]
- 89. Gallois A, Klein JR, Allen LA, Jones BD, Nauseef WM. 2001. Salmonella pathogenicity island 2-encoded type III secretion system mediates exclusion of NADPH oxidase assembly from the phagosomal membrane. J. Immunol. 166:5741–5748 [DOI] [PubMed] [Google Scholar]
- 90. Geier H, Celli J. 2011. Phagocytic receptors dictate phagosomal escape and intracellular proliferation of Francisella tularensis. Infect. Immun. 79:2204–2214 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Gil H, et al. 2006. Deletion of TolC orthologs in Francisella tularensis identifies roles in multidrug resistance and virulence. Proc. Natl. Acad. Sci. U. S. A. 103:12897–12902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Golovliov I, Baranov V, Krocova Z, Kovarova H, Sjöstedt A. 2003. An attenuated strain of the facultative intracellular bacterium Francisella tularensis can escape the phagosome of monocytic cells. Infect. Immun. 71:5940–5950 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Gordon S, Martinez FO. 2010. Alternative activation of macrophages: mechanism and functions. Immunity 32:593–604 [DOI] [PubMed] [Google Scholar]
- 94. Gunn JS, Ernst RK. 2007. The structure and function of Francisella lipopolysaccharide. Ann. N. Y. Acad. Sci. 1105:202–218 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Hagman KE, et al. 1995. Resistance of Neisseria gonorrhoeae to antimicrobial hydrophobic agents is modulated by the mtrRCDE efflux system. Microbiology 141:611–622 [DOI] [PubMed] [Google Scholar]
- 96. Hajjar AM, et al. 2006. Lack of in vitro and in vivo recognition of Francisella tularensis subspecies lipopolysaccharide by Toll-like receptors. Infect. Immun. 74:6730–6738 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Hall JD, et al. 2008. Infected-host-cell repertoire and cellular response in the lung following inhalation of Francisella tularensis Schu S4, LVS, or U112. Infect. Immun. 76:5843–5852 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98. Han S, Bishop BM, van Hoek ML. 2008. Antimicrobial activity of human beta-defensins and induction by Francisella. Biochem. Biophys. Res. Commun. 371:670–674 [DOI] [PubMed] [Google Scholar]
- 99. Henry T, Brotcke A, Weiss DS, Thompson LJ, Monack DM. 2007. Type I interferon signaling is required for activation of the inflammasome during Francisella infection. J. Exp. Med. 204:987–994 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100. Hirakata Y, et al. 2002. Multidrug efflux systems play an important role in the invasiveness of Pseudomonas aeruginosa. J. Exp. Med. 196:109–118 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101. Hornung V, et al. 2009. AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nature 458:514–518 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102. Horwitz MA. 1982. Phagocytosis of microorganisms. Rev. Infect. Dis. 4:104–123 [DOI] [PubMed] [Google Scholar]
- 103. Horwitz MA. 1984. Phagocytosis of the Legionnaires' disease bacterium (Legionella pneumophila) occurs by a novel mechanism: engulfment within a pseudopod coil. Cell 36:27–33 [DOI] [PubMed] [Google Scholar]
- 104. Hoyal CR, et al. 2003. Modulation of p47PHOX activity by site-specific phosphorylation: Akt-dependent activation of the NADPH oxidase. Proc. Natl. Acad. Sci. U. S. A. 100:5130–5135 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105. Hrstka R, et al. 2007. Francisella tularensis strain LVS resides in MHC II-positive autophagic vacuoles in macrophages. Folia Microbiol. (Praha) 52:631–636 [DOI] [PubMed] [Google Scholar]
- 106. Huang MT, et al. 2010. Deletion of ripA alleviates suppression of the inflammasome and MAPK by Francisella tularensis. J. Immunol. 185:5476–5485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107. Ishikawa H, Ma Z, Barber GN. 2009. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461:788–792 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108. Ishimoto H, et al. 2006. Identification of hBD-3 in respiratory tract and serum: the increase in pneumonia. Eur. Respir. J. 27:253–260 [DOI] [PubMed] [Google Scholar]
- 109. Janeway CA, Jr, Medzhitov R. 2002. Innate immune recognition. Annu. Rev. Immunol. 20:197–216 [DOI] [PubMed] [Google Scholar]
- 110. Jia Q, et al. 2010. A Francisella tularensis live vaccine strain (LVS) mutant with a deletion in capB, encoding a putative capsular biosynthesis protein, is significantly more attenuated than LVS yet induces potent protective immunity in mice against F. tularensis challenge. Infect. Immun. 78:4341–4355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111. Jones CL, Weiss DS. 2011. TLR2 signaling contributes to rapid inflammasome activation during F. novicida infection. PLoS One 6:e20609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112. Jones JW, et al. 2010. Absent in melanoma 2 is required for innate immune recognition of Francisella tularensis. Proc. Natl. Acad. Sci. U. S. A. 107:9771–9776 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113. Kang PB, et al. 2005. The human macrophage mannose receptor directs Mycobacterium tuberculosis lipoarabinomannan-mediated phagosome biogenesis. J. Exp. Med. 202:987–999 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114. Kanistanon D, et al. 2008. A Francisella mutant in lipid A carbohydrate modification elicits protective immunity. PLoS Pathog. 4:e24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115. Kawai T, Akira S. 2010. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11:373–384 [DOI] [PubMed] [Google Scholar]
- 116. Kieffer TL, Cowley S, Nano FE, Elkins KL. 2003. Francisella novicida LPS has greater immunobiological activity in mice than F. tularensis LPS, and contributes to F. novicida murine pathogenesis. Microbes Infect. 5:397–403 [DOI] [PubMed] [Google Scholar]
- 117. Kiss K, Liu W, Huntley JF, Norgard MV, Hansen EJ. 2008. Characterization of fig operon mutants of Francisella novicida U112. FEMS Microbiol. Lett. 285:270–277 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118. Kofoed EM, Vance RE. 2011. Innate immune recognition of bacterial ligands by NAIPs determines inflammasome specificity. Nature 477:592–595 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119. Krocova Z, et al. 2008. Interaction of B cells with intracellular pathogen Francisella tularensis. Microb. Pathog. 45:79–85 [DOI] [PubMed] [Google Scholar]
- 120. Lai XH, Golovliov I, Sjostedt A. 2001. Francisella tularensis induces cytopathogenicity and apoptosis in murine macrophages via a mechanism that requires intracellular bacterial multiplication. Infect. Immun. 69:4691–4694 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121. Laird MH, et al. 2009. TLR4/MyD88/PI3K interactions regulate TLR4 signaling. J. Leukoc. Biol. 85:966–977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122. Larsson P, et al. 2005. The complete genome sequence of Francisella tularensis, the causative agent of tularemia. Nat. Genet. 37:153–159 [DOI] [PubMed] [Google Scholar]
- 123. Law HT, et al. 2011. Francisella tularensis uses cholesterol and clathrin-based endocytic mechanisms to invade hepatocytes. Sci. Rep. 1:192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124. Lee VT, Tam C, Schneewind O. 2000. LcrV, a substrate for Yersinia enterocolitica type III secretion, is required for toxin targeting into the cytosol of HeLa cells. J. Biol. Chem. 275:36869–36875 [DOI] [PubMed] [Google Scholar]
- 125. Leelaporn A, Yongyod S, Limsrivanichakorn S, Yungyuen T, Kiratisin P. 2008. Francisella novicida bacteremia, Thailand. Emerg. Infect. Dis. 14:1935–1937 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126. Leiby DA, Fortier AH, Crawford RM, Schreiber RD, Nacy CA. 1992. In vivo modulation of the murine immune response to Francisella tularensis LVS by administration of anticytokine antibodies. Infect. Immun. 60:84–89 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127. Levine B, Mizushima N, Virgin HW. 2011. Autophagy in immunity and inflammation. Nature 469:323–335 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128. Li H, Nookala S, Bina XR, Bina JE, Re F. 2006. Innate immune response to Francisella tularensis is mediated by TLR2 and caspase-1 activation. J. Leukoc. Biol. 80:766–773 [DOI] [PubMed] [Google Scholar]
- 129. Li J, et al. 2007. Attenuation and protective efficacy of an O-antigen-deficient mutant of Francisella tularensis LVS. Microbiology 153:3141–3153 [DOI] [PubMed] [Google Scholar]
- 130. Lindemann SR, et al. 2011. Francisella tularensis Schu S4 O-antigen and capsule biosynthesis gene mutants induce early cell death in human macrophages. Infect. Immun. 79:581–594 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131. Lindgren H, et al. 2004. Factors affecting the escape of Francisella tularensis from the phagolysosome. J. Med. Microbiol. 53:953–958 [DOI] [PubMed] [Google Scholar]
- 132. Lindgren H, et al. 2009. The 58-kilodalton major virulence factor of Francisella tularensis is required for efficient utilization of iron. Infect. Immun. 77:4429–4436 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133. Lindgren H, et al. 2011. Iron content differs between Francisella tularensis subspecies tularensis and subspecies holarctica strains and correlates to their susceptibility to H2O2-induced killing. Infect. Immun. 79:1218–1224 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134. Lindgren H, et al. 2007. Resistance of Francisella tularensis strains against reactive nitrogen and oxygen species with special reference to the role of KatG. Infect. Immun. 75:1303–1309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135. Llewellyn AC, Jones CL, Napier BA, Bina JE, Weiss DS. 2011. Macrophage replication screen identifies a novel Francisella hydroperoxide resistance protein involved in virulence. PLoS One 6:e24201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136. Long GW, et al. 1993. Detection of Francisella tularensis in blood by polymerase chain reaction. J. Clin. Microbiol. 31:152–154 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137. Lucas MA, Fretto LJ, McKee PA. 1983. The binding of human plasminogen to fibrin and fibrinogen. J. Biol. Chem. 258:4249–4256 [PubMed] [Google Scholar]
- 138. Macela A, et al. 1996. The immune response against Francisella tularensis live vaccine strain in Lps(n) and Lps(d) mice. FEMS Immunol. Med. Microbiol. 13:235–238 [DOI] [PubMed] [Google Scholar]
- 139. Mahnke K, Schmitt E, Bonifaz L, Enk AH, Jonuleit H. 2002. Immature, but not inactive: the tolerogenic function of immature dendritic cells. Immunol. Cell Biol. 80:477–483 [DOI] [PubMed] [Google Scholar]
- 140. Maier TM, et al. 2007. Identification of Francisella tularensis Himar1-based transposon mutants defective for replication in macrophages. Infect. Immun. 75:5376–5389 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141. Mariathasan S, Weiss DS, Dixit VM, Monack DM. 2005. Innate immunity against Francisella tularensis is dependent on the ASC/caspase-1 axis. J. Exp. Med. 202:1043–1049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142. Mariathasan S, et al. 2006. Cryopyrin activates the inflammasome in response to toxins and ATP. Nature 440:228–232 [DOI] [PubMed] [Google Scholar]
- 143. Marshall JG, et al. 2001. Restricted accumulation of phosphatidylinositol 3-kinase products in a plasmalemmal subdomain during Fc gamma receptor-mediated phagocytosis. J. Cell Biol. 153:1369–1380 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144. Martinon F, Mayor A, Tschopp J. 2009. The inflammasomes: guardians of the body. Annu. Rev. Immunol. 27:229–265 [DOI] [PubMed] [Google Scholar]
- 145. McCaffrey RL, Allen LA. 2006. Francisella tularensis LVS evades killing by human neutrophils via inhibition of the respiratory burst and phagosome escape. J. Leukoc. Biol. 80:1224–1230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146. McCaffrey RL, et al. 2010. Multiple mechanisms of NADPH oxidase inhibition by type A and type B Francisella tularensis. J. Leukoc. Biol. 88:791–805 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147. McCoy GW. 1911. Some features of the squirrel plague problem. Cal. State J. Med. 9:105–109 [PMC free article] [PubMed] [Google Scholar]
- 148. Medina EA, Morris IR, Berton MT. 2010. Phosphatidylinositol 3-kinase activation attenuates the TLR2-mediated macrophage proinflammatory cytokine response to Francisella tularensis live vaccine strain. J. Immunol. 185:7562–7572 [DOI] [PubMed] [Google Scholar]
- 149. Meibom KL, Charbit A. 2010. Francisella tularensis metabolism and its relation to virulence. Front. Microbiol. 1:140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150. Melillo AA, Bakshi CS, Melendez JA. 2010. Francisella tularensis antioxidants harness reactive oxygen species to restrict macrophage signaling and cytokine production. J. Biol. Chem. 285:27553–27560 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151. Melillo AA, et al. 2009. Identification of Francisella tularensis live vaccine strain CuZn superoxide dismutase as critical for resistance to extracellularly generated reactive oxygen species. J. Bacteriol. 191:6447–6456 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 152. Miao EA, et al. 2010. Innate immune detection of the type III secretion apparatus through the NLRC4 inflammasome. Proc. Natl. Acad. Sci. U. S. A. 107:3076–3080 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 153. Michell S, et al. 2010. Deletion of the Bacillus anthracis capB homologue in Francisella tularensis subspecies tularensis generates an attenuated strain that protects mice against virulent tularaemia. J. Med. Microbiol. 59:1275–1284 [DOI] [PubMed] [Google Scholar]
- 154. Miethke M, Marahiel MA. 2007. Siderophore-based iron acquisition and pathogen control. Microbiol. Mol. Biol. Rev. 71:413–451 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155. Mohapatra NP, et al. 2010. Francisella acid phosphatases inactivate the NADPH oxidase in human phagocytes. J. Immunol. 184:5141–5150 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156. Mohapatra NP, et al. 2008. Combined deletion of four Francisella novicida acid phosphatases attenuates virulence and macrophage vacuolar escape. Infect. Immun. 76:3690–3699 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157. Monroe KM, McWhirter SM, Vance RE. 2010. Induction of type I interferons by bacteria. Cell. Microbiol. 12:881–890 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158. Mulero V, Brock JH. 1999. Regulation of iron metabolism in murine J774 macrophages: role of nitric oxide-dependent and -independent pathways following activation with gamma interferon and lipopolysaccharide. Blood 94:2383–2389 [PubMed] [Google Scholar]
- 159. Murray P, Rosentthal K, Pfaller M. 2005. Medical microbiology, 5th ed Elsevier Mosby, Philadelphia, PA [Google Scholar]
- 159a. Nallaparaju KC, et al. 31 March 2011. Evasion of IFN-γ signaling by Francisella novicida is dependent upon Francisella outer membrane protein C. PLoS One 6:e18201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 160. Nano FE, Schmerk C. 2007. The Francisella pathogenicity island. Ann. N. Y. Acad. Sci. 1105:122–137 [DOI] [PubMed] [Google Scholar]
- 161. Nauseef WM. 2004. Assembly of the phagocyte NADPH oxidase. Histochem. Cell Biol. 122:277–291 [DOI] [PubMed] [Google Scholar]
- 162. Nemeth E, et al. 2004. Hepcidin regulates cellular iron efflux by binding to ferroportin and inducing its internalization. Science 306:2090–2093 [DOI] [PubMed] [Google Scholar]
- 163. Neves P, et al. 2010. Signaling via the MyD88 adaptor protein in B cells suppresses protective immunity during Salmonella typhimurium infection. Immunity 33:777–790 [DOI] [PubMed] [Google Scholar]
- 164. O'Farrell AM, Liu Y, Moore KW, Mui AL. 1998. IL-10 inhibits macrophage activation and proliferation by distinct signaling mechanisms: evidence for Stat3-dependent and -independent pathways. EMBO J. 17:1006–1018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165. Pan X, Tamilselvam B, Hansen EJ, Daefler S. 2010. Modulation of iron homeostasis in macrophages by bacterial intracellular pathogens. BMC Microbiol. 10:64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166. Parsa KV, et al. 2008. Francisella gains a survival advantage within mononuclear phagocytes by suppressing the host IFNgamma response. Mol. Immunol. 45:3428–3437 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167. Parsa KV, et al. 2006. Macrophage pro-inflammatory response to Francisella novicida infection is regulated by SHIP. PLoS Pathog. 2:e71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168. Parsot C. 1994. Shigella flexneri: genetics of entry and intercellular dissemination in epithelial cells. Curr. Top. Microbiol. Immunol. 192:217–241 [DOI] [PubMed] [Google Scholar]
- 169. Pechous R, et al. 2006. Construction and characterization of an attenuated purine auxotroph in a Francisella tularensis live vaccine strain. Infect. Immun. 74:4452–4461 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 170. Pekarek RS, Bostian KA, Bartelloni PJ, Calia FM, Beisel WR. 1969. The effects of Francisella tularensis infection on iron metabolism in man. Am. J. Med. Sci. 258:14–25 [DOI] [PubMed] [Google Scholar]
- 171. Peng K, Broz P, Jones J, Joubert LM, Monack D. 2011. Elevated AIM2-mediated pyroptosis triggered by hypercytotoxic Francisella mutant strains is attributed to increased intracellular bacteriolysis. Cell Microbiol. 13:1586–1600 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172. Peng K, Monack DM. 2010. Indoleamine 2,3-dioxygenase 1 is a lung-specific innate immune defense mechanism that inhibits growth of Francisella tularensis tryptophan auxotrophs. Infect. Immun. 78:2723–2733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 173. Periasamy S, et al. 2011. Development of tolerogenic dendritic cells and regulatory T cells favors exponential bacterial growth and survival during early respiratory tularemia. J. Leukoc. Biol. 90:493–507 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174. Philpott DJ, Girardin SE. 2010. Nod-like receptors: sentinels at host membranes. Curr. Opin. Immunol. 22:428–434 [DOI] [PubMed] [Google Scholar]
- 175. Phipps RP, Stein SH, Roper RL. 1991. A new view of prostaglandin E regulation of the immune response. Immunol. Today 12:349–352 [DOI] [PubMed] [Google Scholar]
- 176. Pierini LM. 2006. Uptake of serum-opsonized Francisella tularensis by macrophages can be mediated by class A scavenger receptors. Cell. Microbiol. 8:1361–1370 [DOI] [PubMed] [Google Scholar]
- 177. Pilatz S, et al. 2006. Identification of Burkholderia pseudomallei genes required for the intracellular life cycle and in vivo virulence. Infect. Immun. 74:3576–3586 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 178. Platz GJ, et al. 2010. A tolC mutant of Francisella tularensis is hypercytotoxic compared to the wild type and elicits increased proinflammatory responses from host cells. Infect. Immun. 78:1022–1031 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 179. Poole K, Krebes K, McNally C, Neshat S. 1993. Multiple antibiotic resistance in Pseudomonas aeruginosa: evidence for involvement of an efflux operon. J. Bacteriol. 175:7363–7372 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 180. Raetz CR, et al. 2009. Discovery of new biosynthetic pathways: the lipid A story. J. Lipid Res. 50(Suppl.):S103–S108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 181. Rajaram MV, et al. 2009. Akt and SHIP modulate Francisella escape from the phagosome and induction of the Fas-mediated death pathway. PLoS One 4:e7919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 182. Ramakrishnan G, Meeker A, Dragulev B. 2008. fslE is necessary for siderophore-mediated iron acquisition in Francisella tularensis Schu S4. J. Bacteriol. 190:5353–5361 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 183. Rathinam VA, et al. 2010. The AIM2 inflammasome is essential for host defense against cytosolic bacteria and DNA viruses. Nat. Immunol. 11:395–402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 184. Ray K, Marteyn B, Sansonetti PJ, Tang CM. 2009. Life on the inside: the intracellular lifestyle of cytosolic bacteria. Nat. Rev. Microbiol. 7:333–340 [DOI] [PubMed] [Google Scholar]
- 185. Recalcati S, Taramelli D, Conte D, Cairo G. 1998. Nitric oxide-mediated induction of ferritin synthesis in J774 macrophages by inflammatory cytokines: role of selective iron regulatory protein-2 downregulation. Blood 91:1059–1066 [PubMed] [Google Scholar]
- 186. Reilly TJ, Baron GS, Nano FE, Kuhlenschmidt MS. 1996. Characterization and sequencing of a respiratory burst-inhibiting acid phosphatase from Francisella tularensis. J. Biol. Chem. 271:10973–10983 [DOI] [PubMed] [Google Scholar]
- 187. Roberts TL, et al. 2009. HIN-200 proteins regulate caspase activation in response to foreign cytoplasmic DNA. Science 323:1057–1060 [DOI] [PubMed] [Google Scholar]
- 188. Rohmer L, et al. 2007. Comparison of Francisella tularensis genomes reveals evolutionary events associated with the emergence of human pathogenic strains. Genome Biol. 8:R102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 189. Roth KM, Gunn JS, Lafuse W, Satoskar AR. 2009. Francisella inhibits STAT1-mediated signaling in macrophages and prevents activation of antigen-specific T cells. Int. Immunol. 21:19–28 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 190. Rus H, Cudrici C, Niculescu F. 2005. The role of the complement system in innate immunity. Immunol. Res. 33:103–112 [DOI] [PubMed] [Google Scholar]
- 191. Russo BC, et al. 2011. A Francisella tularensis locus required for spermine responsiveness is necessary for virulence. Infect. Immun. 79:3665–3676 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 192. Sandstrom G, Lofgren S, Tarnvik A. 1988. A capsule-deficient mutant of Francisella tularensis LVS exhibits enhanced sensitivity to killing by serum but diminished sensitivity to killing by polymorphonuclear leukocytes. Infect. Immun. 56:1194–1202 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 193. Sansonetti PJ, Ryter A, Clerc P, Maurelli AT, Mounier J. 1986. Multiplication of Shigella flexneri within HeLa cells: lysis of the phagocytic vacuole and plasmid-mediated contact hemolysis. Infect. Immun. 51:461–469 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 194. Santic M, Asare R, Skrobonja I, Jones S, Abu Kwaik Y. 2008. Acquisition of the vacuolar ATPase proton pump and phagosome acidification are essential for escape of Francisella tularensis into the macrophage cytosol. Infect. Immun. 76:2671–2677 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 195. Santic M, Molmeret M, Abu Kwaik Y. 2005. Modulation of biogenesis of the Francisella tularensis subsp. novicida-containing phagosome in quiescent human macrophages and its maturation into a phagolysosome upon activation by IFN-gamma. Cell. Microbiol. 7:957–967 [DOI] [PubMed] [Google Scholar]
- 196. Santic M, Molmeret M, Klose KE, Jones S, Kwaik YA. 2005. The Francisella tularensis pathogenicity island protein IglC and its regulator MglA are essential for modulating phagosome biogenesis and subsequent bacterial escape into the cytoplasm. Cell. Microbiol. 7:969–979 [DOI] [PubMed] [Google Scholar]
- 197. Sasakawa C, et al. 1989. Functional organization and nucleotide sequence of virulence region-2 on the large virulence plasmid in Shigella flexneri 2a. Mol. Microbiol. 3:1191–1201 [DOI] [PubMed] [Google Scholar]
- 198. Schaller-Bals S, Schulze A, Bals R. 2002. Increased levels of antimicrobial peptides in tracheal aspirates of newborn infants during infection. Am. J. Respir. Crit. Care Med. 165:992–995 [DOI] [PubMed] [Google Scholar]
- 199. Schlesinger LS, Bellinger-Kawahara CG, Payne NR, Horwitz MA. 1990. Phagocytosis of Mycobacterium tuberculosis is mediated by human monocyte complement receptors and complement component C3. J. Immunol. 144:2771–2780 [PubMed] [Google Scholar]
- 200. Schlesinger LS, Horwitz MA. 1990. Phagocytosis of leprosy bacilli is mediated by complement receptors CR1 and CR3 on human monocytes and complement component C3 in serum. J. Clin. Invest. 85:1304–1314 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 201. Schroder K, Hertzog PJ, Ravasi T, Hume DA. 2004. Interferon-gamma: an overview of signals, mechanisms and functions. J. Leukoc. Biol. 75:163–189 [DOI] [PubMed] [Google Scholar]
- 202. Schulert GS, Allen LA. 2006. Differential infection of mononuclear phagocytes by Francisella tularensis: role of the macrophage mannose receptor. J. Leukoc. Biol. 80:563–571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 203. Schulert GS, et al. 2009. Francisella tularensis genes required for inhibition of the neutrophil respiratory burst and intramacrophage growth identified by random transposon mutagenesis of strain LVS. Infect. Immun. 77:1324–1336 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 204. Sebastian S, et al. 2007. A defined O-antigen polysaccharide mutant of Francisella tularensis live vaccine strain has attenuated virulence while retaining its protective capacity. Infect. Immun. 75:2591–2602 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 205. Sen B, Meeker A, Ramakrishnan G. 2010. The fslE homolog, FTL_0439 (fupA/B), mediates siderophore-dependent iron uptake in Francisella tularensis LVS. Infect. Immun. 78:4276–4285 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 206. Shi CS, et al. 2012. Activation of autophagy by inflammatory signals limits IL-1beta production by targeting ubiquitinated inflammasomes for destruction. Nat. Immunol. 13:255–263 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 207. Shirey KA, Cole LE, Keegan AD, Vogel SN. 2008. Francisella tularensis live vaccine strain induces macrophage alternative activation as a survival mechanism. J. Immunol. 181:4159–4167 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 208. Sing A, et al. 2002. Yersinia V-antigen exploits Toll-like receptor 2 and CD14 for interleukin 10-mediated immunosuppression. J. Exp. Med. 196:1017–1024 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 209. Sjöstedt A. 2007. Tularemia: history, epidemiology, pathogen physiology, and clinical manifestations. Ann. N. Y. Acad. Sci. 1105:1–29 [DOI] [PubMed] [Google Scholar]
- 210. Sjostedt A, Conlan JW, North RJ. 1994. Neutrophils are critical for host defense against primary infection with the facultative intracellular bacterium Francisella tularensis in mice and participate in defense against reinfection. Infect. Immun. 62:2779–2783 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 211. Sorokin VM, Pavlovich NV, Prozorova LA. 1996. Francisella tularensis resistance to bactericidal action of normal human serum. FEMS Immunol. Med. Microbiol. 13:249–252 [DOI] [PubMed] [Google Scholar]
- 212. Stinavage P, Martin L, Spitznagel J. 1989. O antigen and lipid A phosphoryl groups in resistance of Salmonella typhimurium LT-2 to nonoxidative killing in human polymorphonuclear neutrophils. Infect. Immun. 57:3894–3900 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 213. Su J, et al. 2011. The capBCA locus is required for intracellular growth of Francisella tularensis LVS. Front. Microbiol. 2:83. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 214. Su J, et al. 2007. Genome-wide identification of Francisella tularensis virulence determinants. Infect. Immun. 75:3089–3101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 215. Sullivan JT, Jeffery EF, Shannon JD, Ramakrishnan G. 2006. Characterization of the siderophore of Francisella tularensis and role of fslA in siderophore production. J. Bacteriol. 188:3785–3795 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 216. Takeda K, Akira S. 2004. TLR signaling pathways. Semin. Immunol. 16:3–9 [DOI] [PubMed] [Google Scholar]
- 217. Tamilselvam B, Daefler S. 2008. Francisella targets cholesterol-rich host cell membrane domains for entry into macrophages. J. Immunol. 180:8262–8271 [DOI] [PubMed] [Google Scholar]
- 218. Taylor MW, Feng GS. 1991. Relationship between interferon-gamma, indoleamine 2,3-dioxygenase, and tryptophan catabolism. FASEB J. 5:2516–2522 [PubMed] [Google Scholar]
- 219. Telepnev M, Golovliov I, Grundstrom T, Tarnvik A, Sjostedt A. 2003. Francisella tularensis inhibits Toll-like receptor-mediated activation of intracellular signalling and secretion of TNF-alpha and IL-1 from murine macrophages. Cell. Microbiol. 5:41–51 [DOI] [PubMed] [Google Scholar]
- 220. Thakran S, et al. 2008. Identification of Francisella tularensis lipoproteins that stimulate the Toll-like receptor (TLR) 2/TLR1 heterodimer. J. Biol. Chem. 283:3751–3760 [DOI] [PubMed] [Google Scholar]
- 221. Thomas RM, et al. 2007. The immunologically distinct O antigens from Francisella tularensis subspecies tularensis and Francisella novicida are both virulence determinants and protective antigens. Infect. Immun. 75:371–378 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 222. Travassos LH, et al. 2004. Toll-like receptor 2-dependent bacterial sensing does not occur via peptidoglycan recognition. EMBO Rep. 5:1000–1006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 223. Twine S, et al. 2005. A mutant of Francisella tularensis strain SCHU S4 lacking the ability to express a 58-kilodalton protein is attenuated for virulence and is an effective live vaccine. Infect. Immun. 73:8345–8352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 224. Ulland TK, et al. 2010. Mutation of Francisella tularensis mviN leads to increased macrophage absent in melanoma 2 inflammasome activation and a loss of virulence. J. Immunol. 185:2670–2674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 225. Underhill DM, Ozinsky A. 2002. Phagocytosis of microbes: complexity in action. Annu. Rev. Immunol. 20:825–852 [DOI] [PubMed] [Google Scholar]
- 226. Virgin HW, Levine B. 2009. Autophagy genes in immunity. Nat. Immunol. 10:461–470 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 227. Wandersman C, Delepelaire P. 1990. TolC, an Escherichia coli outer membrane protein required for hemolysin secretion. Proc. Natl. Acad. Sci. U. S. A. 87:4776–4780 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 228. Wandersman C, Stojiljkovic I. 2000. Bacterial heme sources: the role of heme, hemoprotein receptors and hemophores. Curr. Opin. Microbiol. 3:215–220 [DOI] [PubMed] [Google Scholar]
- 229. Wang X, Ribeiro AA, Guan Z, Abraham SN, Raetz CR. 2007. Attenuated virulence of a Francisella mutant lacking the lipid A 4′-phosphatase. Proc. Natl. Acad. Sci. U. S. A. 104:4136–4141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 230. Wang X, et al. 2006. Structure and biosynthesis of free lipid A molecules that replace lipopolysaccharide in Francisella tularensis subsp. novicida. Biochemistry 45:14427–14440 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 231. Weinberg ED. 1974. Iron and susceptibility to infectious disease. Science 184:952–956 [DOI] [PubMed] [Google Scholar]
- 232. Weiss DS, et al. 2007. In vivo negative selection screen identifies genes required for Francisella virulence. Proc. Natl. Acad. Sci. U. S. A. 104:6037–6042 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 233. Whitnack E, Beachey EH. 1985. Inhibition of complement-mediated opsonization and phagocytosis of Streptococcus pyogenes by D fragments of fibrinogen and fibrin bound to cell surface M protein. J. Exp. Med. 162:1983–1997 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 234. Wickstrum JR, et al. 2009. Francisella tularensis induces extensive caspase-3 activation and apoptotic cell death in the tissues of infected mice. Infect. Immun. 77:4827–4836 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 235. Wilson JE, Katkere B, Drake JR. 2009. Francisella tularensis induces ubiquitin-dependent major histocompatibility complex class II degradation in activated macrophages. Infect. Immun. 77:4953–4965 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 236. Woolard MD, et al. 2007. Francisella tularensis-infected macrophages release prostaglandin E2 that blocks T cell proliferation and promotes a Th2-like response. J. Immunol. 178:2065–2074 [DOI] [PubMed] [Google Scholar]
- 237. Yoshida R, Hayaishi O. 1978. Induction of pulmonary indoleamine 2,3-dioxygenase by intraperitoneal injection of bacterial lipopolysaccharide. Proc. Natl. Acad. Sci. U. S. A. 75:3998–4000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 238. Yoshida R, Oku T, Imanishi J, Kishida T, Hayaishi O. 1986. Interferon: a mediator of indoleamine 2,3-dioxygenase induction by lipopolysaccharide, poly(I) X poly(C), and pokeweed mitogen in mouse lung. Arch. Biochem. Biophys. 249:596–604 [DOI] [PubMed] [Google Scholar]
- 239. Yu JJ, et al. 2008. The presence of infectious extracellular Francisella tularensis subsp. novicida in murine plasma after pulmonary challenge. Eur. J. Clin. Microbiol. Infect. Dis. 27:323–325 [DOI] [PubMed] [Google Scholar]
- 240. Zarrella TM, et al. 2011. Host-adaptation of Francisella tularensis alters the bacterium's surface-carbohydrates to hinder effectors of innate and adaptive immunity. PLoS One 6:e22335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 241. Zegarra-Moran O, et al. 2004. Double mechanism for apical tryptophan depletion in polarized human bronchial epithelium. J. Immunol. 173:542–549 [DOI] [PubMed] [Google Scholar]
- 242. Zhao J, et al. 2006. IRF-8/interferon (IFN) consensus sequence-binding protein is involved in Toll-like receptor (TLR) signaling and contributes to the cross-talk between TLR and IFN-gamma signaling pathways. J. Biol. Chem. 281:10073–10080 [DOI] [PubMed] [Google Scholar]
- 243. Zhao J, Raetz CR. 2010. A two-component Kdo hydrolase in the inner membrane of Francisella novicida. Mol. Microbiol. 78:820–836 [DOI] [PMC free article] [PubMed] [Google Scholar]