SUMMARY
Mediator, a large (21 polypeptides, MW ~1 MDa) complex conserved throughout eukaryotes, plays an essential role in control of gene expression by conveying regulatory signals that influence the activity of the preinitiation complex (PIC). However, the precise mode of interaction between Mediator and RNA polymerase II (RNAPII), and the mechanism of regulation by Mediator remain elusive. We used cryo-electron microscopy and reconstituted in vitro transcription assays to characterize a transcriptionally-active complex including the Mediator Head module and components of a minimum pre-initiation complex (RNAPII, TFIIF, TFIIB, TBP, and promoter DNA). Our results reveal how the Head interacts with RNAPII, affecting its conformation and function.
INTRODUCTION
The Mediator complex conveys signals from DNA regulatory sequences to RNA polymerase II (RNAPII) and a set of general transcription factors that assemble at a promoter to form a preinitiation complex (PIC) and initiate transcription (Baek et al., 2002; Flanagan et al., 1990; Hahn, 2004; Kornberg, 1999; Naar et al., 1999). Structural analyses of Mediator have shown that the complex is divided into structural modules (Head, Middle, Tail, and Arm) displaying complex relative motions (Cai et al., 2009), and that interaction with activators, repressors, and general transcription factors, can result in different Mediator conformations (Cai et al., 2010; Meyer et al., 2010; Taatjes et al., 2004). Changes in Mediator conformation required for interaction with RNAPII are similar between the yeast and human complexes, suggesting that essential aspects of the Mediator regulation mechanism might be conserved (Asturias et al., 1999; Cai et al., 2009; Sato et al., 2004).
How Mediator conveys regulatory information to RNAPII and the rest of the basal transcription machinery remains elusive. Clearly, a critical first step for addressing this question is to understand how Mediator interacts with RNAPII in the context of the PIC. The Mediator Head module can partially recapitulate Mediator activity at initiation and we have recently reported its X-ray structure (Imasaki et al., 2011). Our previous structural, biochemical, and genetic analysis of the interaction between the Mediator Head module and RNAPII suggested that the Head might have an effect of RNAPII conformation that could have important functional implications (Cai et al., 2010).
Here we describe the structure, calculated by single particle macromolecular electron microscopy (EM), of a complex in which the Head module interacts with RNAPII in the context of a minimal preinitiation complex (mPIC). We used reconstituted in vitro transcription assays, single particle cryo-electron microscopy (cryo-EM), and novel cryo-EM map interpretation tools, to study the Head-RNAPII interaction, and its effect on the transcriptional activity and structure of RNAPII in the context of a minimal preinitiation complex (mPIC). A cryo-EM map of the Head-mPIC complex shows that the Head module interacts extensively with RNAPII subunits Rpb4-Rpb7 and the clamp. Head interaction triggers a rearrangement of the RNAPII structure likely to affect promoter engagement, and bolsters transcriptional activity. These results are consistent with recent studies of bacterial RNA polymerase pointing to modulation of polymerase conformation as a common phenomenon underlying diverse aspects of transcription regulation (Belogurov et al., 2009; Mukhopadhyay et al., 2008; Tagami et al., 2010), and suggest that a straightforward mechanism for Mediator action could involve a transmission of changes in Mediator conformation to RNAPII.
RESULTS
Assembly of the Head-mPIC complex
We assembled the Head-mPIC complex as described previously (Takagi et al., 2006). Briefly, we mixed a pre-assembled TBP-TFIIB-DNA complex including a short (53 bp long) synthetic promoter DNA (Figure 1A) with RNAPII-TFIIF and recombinant Head module. The synthetic promoter DNA was generated by fusing a canonical TATA-box sequence (TATATAA) to a DNA segment previously used in X-ray studies of RNAPII (Westover et al., 2004) The Head-mPIC complex was purified by affinity chromatography by immobilizing it on a calmodulin- sepharose resin through a calmodulin-binding-peptide tag at the C-terminus of TFIIF subunit Tfg2. Characterization by SDS-PAGE revealed bands corresponding to all components of a Head-mPIC complex (Figure 1B).
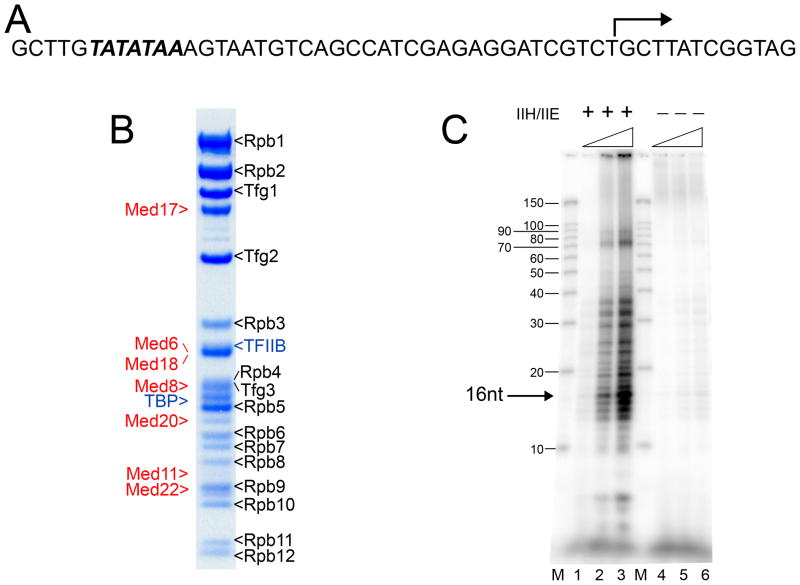
Figure 1. Assembly and functional characterization of the Head-mPIC complex.
(A) Sequence of the 53 base pair promoter DNA used for assembly of the Head-mPIC complex. The TATA box sequence (in italic bold) located about 30 bp upstream of the initiation start site (bent arrow) was fused to the DNA template used previously for RNAPII elongation complex assembly (Westover et al., 2004). (B) SDS-PAGE analysis of the Head-mPIC complex. The subunits of RNAPII (Rpb1-12), TFIIF (Tfg1-3), and the Mediator Head module (Med17, Med6, Med20, Med8, Med18, Med11, and Med22) were identified as indicated. (C) Transcriptional activity of the Head-mPIC complex was tested in a reconstituted transcription assays performed in the presence (lanes 1–3; 0.37, 0.75, and 1.5 pmol Head-mPIC, respectively) or absence (lanes 4–6; same Head-mPIC concentrations as 1–3) of TFIIE and TFIIH. Transcripts were separated in a 15% denaturing gel and imaged with a fluorescent image analyzer. The predominant transcript (16 nucleotides long) is indicated. M: labeled RNA Decade™ marker. See also Figure S1.
Functional characterization of the Head-mPIC complex
We used a reconstituted in vitro transcription assay to test the pre-assembled Head-mPIC complex for promoter-specific basal transcriptional activity from the synthetic promoter DNA. The Head-mPIC showed robust, TFIIE/TFIIH–dependent transcriptional activity (Figure 1C, lanes 1–3, and Figure S1). The predominant transcript products (accounting for ~50% of all observed transcript, Figure S1) were between 13 and 20 nucleotides in length (the major product was 16-nucleotides long) consistent the use of multiple start sites (Hampsey, 1998) selected as RNAPII “scanned” DNA for an optimal initiator sequence (Giardina and Lis, 1993), with initiation from start sites clustered at positions ~25bp downstream of the TATA box (Figure S1). In contrast, in the absence of TFIIE and TFIIH, the pre-assembled Head-mPIC produced virtually no transcripts (Figure 1C, lanes 4–6), indicating TFIIE-TFIIH mediated open promoter complex formation. Very short transcripts (<10 nucleotides) likely resulted from abortive initiation (Yan et al., 1999) and accounted for an additional ~20% of transcript products (Figure S1). The origin of weaker bands corresponding to longer transcripts is harder to explain, but none of these longer products accounts for more than 3% of the observed transcripts (Figure S1). The observation of a variety of transcripts is consistent with the reported behavior of yeast RNAPII, and could also partially relate to the generic nature of the short synthetic template used for assembly of the Head-mPIC. However, the requirement for TFIIH–TFIIE, and the predominance of transcripts whose length corresponds to that expected for promoter–driven initiation, indicates that the Head-mPIC was transcriptionally active.
Initial EM analysis of the Head-mPIC complex
We prepared cryo-EM samples on perforated carbon support films, which allowed us to use the highest possible protein concentration (about 300 ug protein ml−1) to minimize the likelihood that components of the Head-mPIC complex could dissociate. Initial cryo-EM images were recorded on film (Figure 2A) and analyzed with the SPIDER EM image processing package (Frank et al., 1996). Iterative reference-free alignment and clustering of ~15,000 particle images resulted in a set of class averages which, as expected, were generally reminiscent of projections of the RNAPII structure (RNAPII being the central component of, and accounting for most of the mass in the Head-mPIC complex). In addition, many averages showed a distinct, elongated feature that clearly distinguished them from regular RNAPII projections (Figure 2B and Figure S2). A set of class averages expected to correspond to different projections of the Head-mPIC structure were selected and subjected to common lines analysis (Penczek et al., 1996), resulting in an initial 3D map of the Head-mPIC complex where RNAPII appeared as a large, central globular density with a wide cleft (Figure 2C, top). An elongated feature corresponding in size and shape to our published EM structure of the Head module (Cai et al., 2010) could be seen extending from the Rpb4-Rpb7 side (identified because of its position relative to the active site cleft) of the RNAPII structure, consistent with expected interaction of the Head module with those RNAPII subunits (Cai et al., 2010). A 35 Å resolution molecular model of RNAPII derived from its X-ray structure (Bushnell and Kornberg, 2003), and a previously published ~35 Å resolution EM map of the Head module (Cai et al., 2010) could be fitted into the Head-mPIC common-lines structure (Figure 2C, bottom), and were used to generate a volume that we used for clustering of images in a large (~102,000 images) Head-mPIC cryo-EM dataset recorded with the Leginon automated EM data collection package (Suloway et al., 2005).
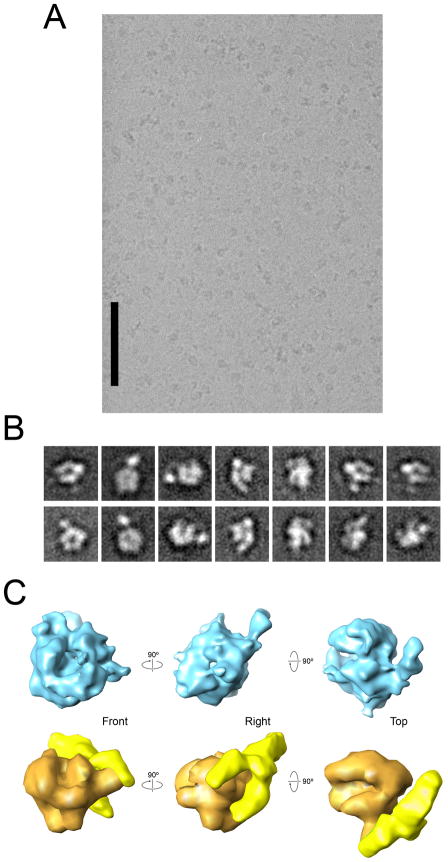
Figure 2. Initial cryo-EM analysis of a Head-mPIC complex.
(A) A raw micrograph showing images of Head-mPIC particles preserved in amorphous ice. The scale bar corresponds to 100 nm. (B) Some initial class averages including clear non-RNAPII features obtained after reference-free alignment and clustering of Head-mPIC cryo-EM images. (C) Initial 3D structure of the Head-mPIC complex (top) obtained by back projection after determining the relative orientations of Head-mPIC class averages, and a distortion-free model of the Head-mPIC structure (bottom) constructed by fitting a ~35 Å resolution molecular map of RNAPII derived from its X-ray structure (Bushnell and Kornberg, 2003) (in orange) and a ~35 Å EM map of the open conformation of the Head module (Cai et al., 2010) (in bright yellow) into the initial Head-mPIC structure. See also Figures S2 and S3.
After using projections of the initial Head-mPIC reference structure to separate cryo-images into groups according to orientation, we discarded all alignment parameters and performed iterative reference-free alignment and clustering within each orientation group (Brignole et al., 2009; Chaban et al., 2008) to minimize any bias related to the initial reference. This resulted in a set of more detailed average projections of the Head-mPIC structure. Examination of these projections suggested that the Head (recognizable in many projections as an elongated feature) might have been missing in some Head-mPIC particles (Figure S3). Therefore, we implemented a competitive projection matching procedure (Craighead et al., 2002; Gao et al., 2004), using the initial Head-mPIC reference structure (Figure 2C) and a low (~35 Å) resolution structure of RNAPII as competing references. This competitive projection matching revealed that about half of the Head-mPIC images showed higher cross-correlation to projections of RNAPII alone, confirming that the Head had dissociated from some Head-mPIC particles. Therefore, we separated the Head-mPIC images into two subsets. About ~50,000 “Head-less” images showing higher correlation to projections of the low-resolution RNAPII reference structure were refined separately and generated a map closely resembling the structure of RNAPII alone (Figure S4A). The remaining ~53,000 images were also refined separately and generated a structure with features clearly reminiscent of RNAPII, but also showing considerable additional density (Figure S4B).
The general agreement of the Head-mPIC cryo-EM map with the X-ray structures of RNAPII and the Head is a strong indication of the overall correctness of the EM map. Nonetheless, to further validate the Head-mPIC map obtained with SPIDER (Figure S4B), we discarded all image alignment parameters and used the SPARX image analysis package and an object (Figure S5A) approximating the general dimensions and shape of the Head-mPIC complex (obtained by low-pass filtering the initial Head-mPIC model to ~50 Å resolution), to select ~27,000 best-behaved Head-mPIC images (those with the most reproducible alignment parameters). We used SPARX to refine a Head-mPIC cryo-EM map with a resolution of ~24 Å from these most stable images (Figure S5B) and found it to be nearly indistinguishable from the Head-mPIC map previously calculated with SPIDER. The consistency between two independently calculated cryo-EM maps of the Head-mPIC complex, one calculated by using SPIDER to refine alignment parameters for ~51,000 Head-mPIC images starting from an initial common-lines volume of the Head-mPIC complex (Figure S4B), and a second one (Figure S5B) calculated by using SPARX to identify and refine alignment parameters for ~27,000 Head-mPIC images starting from an object resembling the overall shape of the initial common-lines volume of the Head-mPIC complex but lacking any features beyond ~50 Å resolution (Figure S5A), provides strong evidence to support the validity of our Head-mPIC cryo-EM 3D map.
In a second test of the validity of the Head-mPIC map, we applied a new iterative clustering algorithm implemented in SPARX (Yang et al., 2012) that reports only clusters whose members can be stably aligned and reproducibly clustered together. This resulted in ~400 completely independent Head-mPIC class averages, which closely resemble reprojections of the final Head-mPIC structure (Figure S6). This, along with the correspondence between independently-derived maps of the Head-mPIC complex, provides strong evidence that the structure we calculated is correct.
A final map of the Head-mPIC complex was calculated by using SPARX to refine alignment parameters for ~51,000 Head-mPIC images, in an iterative procedure that included normalization of the rotationally-averaged power spectrum of intermediate volumes to the rotationally-averaged power spectrum of an EM model derived from the RNAPII X-ray structure. The final Head-mPIC cryo-EM map (Figure 3A) we obtained had a resolution of ~16 Å (Figure S7, A and B).
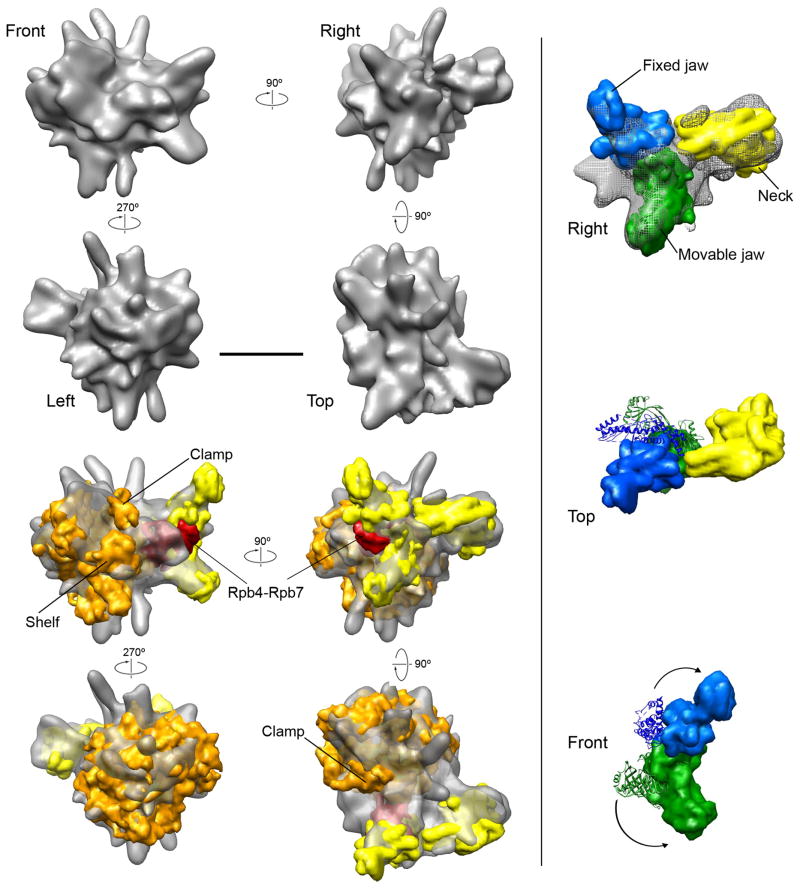
Figure 3. Cryo-EM structure of the Head-mPIC complex and docking of the RNAPII and Head module structures.
(A) Different views of a ~16 Å resolution cryo-EM map of the Head-mPIC complex illustrate how the main portion of the structure is strongly reminiscent of RNAPII, and show that a large segment of additional density matches the EM structure of the open conformation of the Head module (Cai et al., 2010) (shown in red in the Front, Right, and Top views). The scale bar corresponds to 10 nm. (B) Molecular models (at 10 Å resolution) of RNAPII (in orange) and the Head module (in bright yellow), derived from their X-ray structures (Cramer et al., 2001; Imasaki et al., 2011) were fitted into the Head-mPIC cryo-EM map. RNAPII could be docked in a single orientation. The Head structure was segmented into its known structural modules (neck, fixed and movable jaws) and these were re-positioned to match the distribution of Head density in the Head-mPIC map. RNAPII subunits Rpb4-Rpb7 are positioned between the Head module jaws. (C) The conformation of the Head module in the Head-mPIC complex (top panel) can be compared to that revealed by X-ray crystallography analysis by matching the position of the neck and considering changes in the positions of the fixed and movable jaws. Upon interaction with RNAPII, both the fixed and movable jaws undergo large rotations (original positions in the X-ray structure are shown as ribbons in the middle and bottom panels; positions in the cryo-EM map of the Head-mPIC complex as solid surfaces), resulting in a conformation that allows the jaws to interact closely with RNAPII. See also Figures S4, S5, S6 and S7.
Head-RNAPII interaction in the Head-mPIC complex
We examined the Head-mPIC cryo-EM map (Figure 3A) and could readily identify RNAPII structural features, as well as the presence of considerable additional density resembling the published EM structure of the Head module (Cai et al., 2010). We found that 10 Å resolution molecular models of RNAPII and the Head module, calculated from their X-ray structures (Bushnell and Kornberg, 2003; Imasaki et al., 2011), could be fitted into the Head-mPIC map in unique, complementary orientations (Figure 3B). We could match the neck (originally named “handle”) and jaws in the Head X-ray structure to corresponding portions of the Head-mPIC EM map, and observed extensive close contact between the Head jaws and RNAPII subunits Rpb4-Rpb7. Two important observations about interaction of the Head module with RNAPII are as follows: First, as suggested by our previous biochemical, structural, and genetic results (Cai et al., 2010), the Head module interacts directly with RNAPII subunits Rpb4-Rpb7, which are positioned between the Head jaws (Figure 3B). This is in agreement with our previous low resolution structure of a Head-Rpb4-Rpb7 complex (Cai et al., 2010), and with the mode of Mediator-RNAPII interaction we proposed based on our most recent interpretation of the Mediator-RNAPII holoenzyme EM structure (Cai et al., 2009). Second, the conformation of the Head module is dramatically altered upon interaction with RNAPII (Figure 3C). Our previous EM studies of the Head module (Cai et al., 2010) revealed that the angle between the Head jaws could vary. Interaction with RNAPII leads to very substantial opening and rotation of the Head jaws with respect to the neck domain, resulting in a Head conformation that differs from those observed in the EM and X-ray structures of the isolated Head (Cai et al., 2010; Imasaki et al., 2011). Matching the neck of the Head X-ray structure to the corresponding portion of the Head-mPIC structure shows that the jaws undergo a large rotation from the position they adopt in the X-ray structure (Figure 3C). The Head’s X-ray structure revealed that the Head jaws and neck come together to form a poorly ordered domain named the central joint, which must facilitate large changes in the relative position of the jaws and neck (Imasaki et al., 2011). The structure of the Head-mPIC complex now reveals that conformational changes accommodated by the Head’s central joint are essential for interaction of the Head with RNAPII.
Conformation of RNAPII in the Head-mPIC complex
Comparing features in the Head-mPIC structure to those of RNAPII alone suggests that interaction with the Head module leads to changes in RNAPII conformation. In particular, the clamp domain of the RNAPII X-ray structure model is entirely outside the Head-mPIC map (Figure 3B, Front view), suggesting that this domain, which usually forms one side of a narrow cleft surrounding the RNAPII active site, is altered upon interaction with the Head. Also, additional density not attributable to RNAPII or the Head module is apparent throughout the Head-mPIC structure. To interpret the non-Head portion of the Head-mPIC cryo-EM map, we segmented the RNAPII X-ray structure into modules (core, jaw-lobe, shelf, clamp, and Rpb4-Rpb7) previously identified by X-ray crystallography studies (Cramer et al., 2001) (Figure 4A, top left).
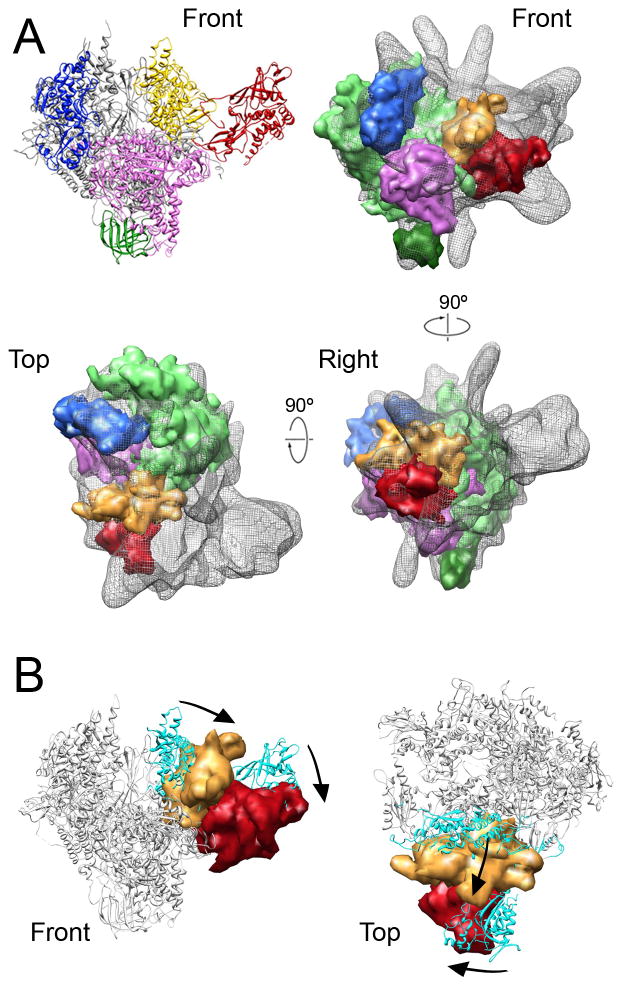
Figure 4. Conformation of RNAPII in the Head-mPIC complex.
(A) Molecular maps (at 18 Å resolution) derived from RNAPII structural modules (top left; core in light green, Rpb8 in dark green, jaw-lobe in blue, shelf in magenta, clamp in yellow, and Rpb4-Rpb7 in red) defined by X-ray crystallography studies of RNAPII (Cramer et al., 2001) docked into the Head-mPIC EM map (shown as a semi-transparent gray mesh). All modules were docked independently and computed docking positions were minimally adjusted to preserve inter-module crystal contacts. (B) Comparing the positions of the clamp and the Rpb4-Rpb7 subunits in the Head-mPIC cryo-EM map (solid surfaces; clamp in gold and Rpb4-Rpb7 in red) with those in the X-ray structure of RNAPII (the RNAPII crystal structure is shown as a gray ribbon model with the clamp and Rpb4-Rpb7 colored in teal) reveals that Head interaction results in a large rearrangement of the clamp, likely facilitated by repositioning of Rpb4-Rpb7 (movement of the clamp and Rpb4-Rpb7 also indicated by the arrows). See also Figures S8 and S9.
Monte Carlo Docking Analysis: Mean docking positions
Because all RNAPII structural modules are relatively large and have sizeable exposed surfaces, we could assign an approximate initial position for each module by matching it to features in the Head-mPIC EM map. However, too arrive at an objective interpretation of the RNAPII conformation in the Head-mPIC cryo-EM map, we used a Monte Carlo-based docking protocol (Supplemental Methods). Individual X-ray segments were independently docked into many noise-corrupted versions of an original EM map, and the resulting positions were used to calculate a mean docking position for each X-ray segment, as well as a 3D “cloud” enclosing a region that encompasses all docking positions below a certain probability threshold. This allowed us to quantitatively evaluate the reliability of the RNAPII module positions and arrive at a molecular interpretation of the RNAPII consistent with the intrinsic limitations of the Head-mPIC cryo-EM map.
Examining the mean docking positions from Monte Carlo docking analysis (Figure S8A, left column) shows that the RNAPII core (the largest module) was matched to the side of the Head-mPIC structure opposite to the Head. The jaw-lobe module rotated slightly about an axis roughly parallel to the RNAPII active site cleft and fitting of the shelf module required a relatively small rotation toward the back end of the active site cleft. In contrast, the computed position for the Rpb4-Rpb7 polymerase subunits and the clamp module were markedly different from those in the X-ray conformation. Polymerase subunits Rpb4-Rpb7, positioned between the Head jaws (Figure 3B), move down and forward toward the downstream end of the active site cleft. The repositioning of Rpb4-Rpb7 likely allows the clamp to swing outwards by as much as 45° (Figure S8A, left column), generating a wide-open RNAPII active site cleft. We performed slight adjustment of the computed module mean positions to preserve inter-module crystal contacts, and repositioned subunit Rpb8 to avoid overlap with core and shelf module densities (see below), to arrive at a final interpretation of the RNAPII conformation in the Head-mPIC complex (Figure 4A).
Monte Carlo Docking Analysis: Docking probability clouds
In general, 1σ probability clouds, including ~70% of all possible docking positions determined by the Monte Carlo analysis, resemble the overall shape of the corresponding RNAPII X-ray segments (Figure S8A, right column) and are roughly centered at the mean segment positions, extending uniformly by 5–10 Â. However, the shelf module, which is adjacent to the clamp, shows a large 1σ probability cloud, asymmetrically-distributed around the shelf’s mean position (Figure S8B). This suggests that rearrangement of Rpb4-Rpb7 and the clamp results in considerable conformational variability of the adjacent shelf. The computed placement of the shelf would require a downward displacement of Rpb8, which could explain the relatively elongated appearance of the EM map around Rpb8.
An important issue is whether the Head-mPIC cryo-EM map contains enough information to conclusively establish that Head interaction triggers a change in RNAPII conformation. A straightforward way of approaching this question is to determine if the RNAPII X-ray model can be fitted into the union of all 1σ clouds obtained from the Monte Carlo docking analysis. If the mean positions for the docked RNAPII segments represent the locations most consistent with the Head-mPIC map, other locations are consistent with the EM map only at a comparatively lower probability level. If the core module of the X-ray structure is matched to the core module docked into the EM map, the X-ray position of the core, jaw-lobe, shelf, and Rpb8, are largely within the respective 1σ clouds from docking analysis. In contrast, the clamp and Rpb4-Rpb7 portions of the X-ray structure are positioned completely outside the respective 1σ clouds from docking analysis (Figure S8C). This indicates that our EM map represents a conformation of the polymerase that differs significantly from the X-ray model due to a large outward swing of the clamp module, likely facilitated by re-positioning of Rpb4-Rpb7 triggered by interaction with the Head (Figure 4B).
Interestingly, the motions required to place the jaw-lobe, shelf, and clamp modules in the positions they adopt in the Head-mPIC complex are analogous to changes in position observed between different crystallographic conformations of RNAPII (Cramer et al., 2001) but larger, especially for Rpb4-Rpb7 and the clamp (Figure S9A). This suggests that the rearrangement of RNAPII structure triggered by Head interaction is facilitated by the normal flexibility of the polymerase. Outward rotation of the clamp results in a wide-open active site cleft that could facilitate promoter access to the RNAPII active site. X-ray crystallographic analysis of RNAPII has documented clamp position changes of up to 25°, but the change induced by Head binding could be nearly twice as large (Figure S9B). Other changes in the polymerase structure appear to be geared toward facilitating motion of the clamp. In particular, downward and forward movement of Rpb4-Rpb7, would facilitate opening of the clamp, as X-ray structures of the 12-subunit form of RNAPII show that a portion of Rpb4 extends below the clamp and would restrict it from swinging outwards (Armache et al., 2003; Bushnell and Kornberg, 2003). To gain a more detailed understanding of the effect of Head module interaction on RNAPII conformation, we investigated possible conformational variability in the Head-mPIC structure.
Codimensional Principal Component Analysis of the Head-mPIC complex
The Monte Carlo docking analysis just described established that interaction of the Head with RNAPII changes the structure of the polymerase and results in clamp opening. Next we wanted to investigate whether the change in clamp conformation observed in the Head-mPIC structure reflected the conformation of all Head-mPIC particles, or resulted from averaging of molecules in different conformations. To answer to this question, we performed Codimensional Principal Component Analysis (CD-PCA) (Penczek et al., 2011) to obtain a quantitative description of variability in the Head-mPIC structure. A total of 12,000 resampled Head-mPIC volumes were calculated by applying a stratified EM projection data resampling approach to ~27K Head-mPIC images with highly stable alignment parameters. CD-PCA of these resampled volumes resulted in 5 eigenvolumes, and reprojections of these eigenvolumes were used to compute factorial coordinates for all 2D projection images. These factorial coordinates were then clustered using a K-means algorithm to obtain 5 groups of projection images. Three-dimensional reconstruction from images in each K-means clustering group yielded 5 initial CD-PCA maps corresponding to plausible conformers of the Head-mPIC complex (Figure 5A and Figure S10A).
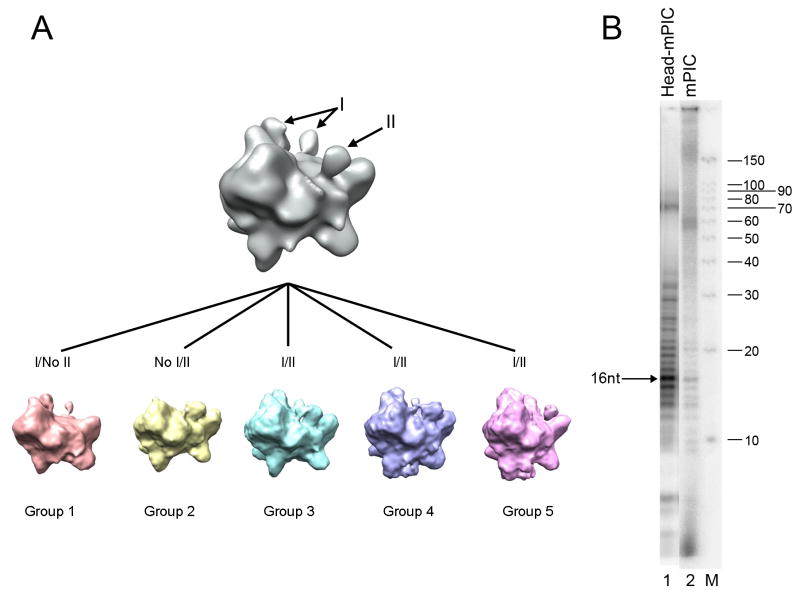
Figure 5. Conformational variability and transcriptional activity of the Head-mPIC complex.
(A) Five Head-mPIC maps generated through Codimensional Principal Component Analysis (CD-PCA) provide a detailed description of variability in the conformation of the Head-mPIC complex. Domains at the upstream (I) and downstream (II) ends of the active site cleft are variably present/ordered in the different conformations of the Head-mPIC complex revealed by CD-PCA. (B) Comparison between transcriptional activity of the Head-mPIC and mPIC complexes in the presence of TFIIH and TFIIE. Assays were performed with the Head–mPIC (1.5 pmol, lane 1), and mPIC (1.5 pmol, lane 2) and results were analyzed as before. The amounts of 16-nucleotide long transcript generated were 2.2 fmol for the Head-mPIC, and 0.12 fmol for mPIC alone. M: labeled RNA Decade™ marker. See also Figure S10.
The validity of the structures obtained from CD-PCA analysis was ascertained by recalculating alignment parameters for images in each group: the original alignment parameters were discarded and new parameters were determined through 3D refinement within each K-means image group by using a single very low-resolution template (shown in Figure S6A) as initial reference. The resulting maps (Figure S10B) faithfully reproduced features in the original CD-PCA maps. In a final test, the initial CD-PCA maps were used to initialize multi-reference refinement of the image data. Considering the localized nature of differences between CD-PCA maps, changes in projection image assignments were limited (~40% agreement with the original PCA assignments) and the resulting maps again resembled closely the original CD-PCA maps (Figure S10C), providing strong evidence for the accuracy of the initial clustering.
Examination of the CD-PCA maps (Figure 5A) highlights both within-group consistencies and inter-group differences in Head-mPIC particles. For example, except for the open clamp, the conformation of RNAPII in the map arising from the fifth group seems closest to that of the RNAPII X-ray structure (e.g., the bottom (foot) portion of the shelf module is clearly visible). On the other hand, the first and second CD-PCA maps represent much larger changes in RNAPII conformation and are closer to the Head-mPIC structure. Another noticeable difference is variability in the presence and/or appearance of domains with comparatively low density (marked I and II in Figure S10) extending from the RNAPII portion of the Head-mPIC map. The map calculated from Head-less images in the Head-mPIC dataset unequivocally shows a polymerase structure with a closed clamp (Figure S4A). In contrast, the Head module and an open clamp are clearly apparent in all 5 CD-PCA structures calculated from Head-mPIC images. Therefore, we conclude that interaction with the Head module directly results in a shift to RNAPII conformations that represent variations on an open-clamp state of RNAPII.
Effect of clamp opening on mPIC transcriptional activity
Crystal structures of RNAPII show the enzyme’s active site at the bottom of a cleft that is too narrow to accommodate double-stranded promoter DNA (Bushnell and Kornberg, 2003; Cramer et al., 2000; Cramer et al., 2001). Therefore, a change in the structure of either promoter DNA (unwinding of the template and non-template strands) or RNAPII (opening of the active site cleft) would seem essential for promoter engagement and initiation of transcription. We anticipated that opening of the RNAPII clamp by the Head might facilitate promoter DNA engagement and lead to an increase transcription activity. To test this idea, we compared the transcriptional activity of the Head-mPIC to that of an otherwise identical Head-less mPIC. Like the Head-mPIC, the mPIC alone showed TFIIE/TFIIH-dependent transcriptional activity. However, the amount of the prevalent (16-nucleotide long) specific transcript from the Head-mPIC was approximately 18-fold higher than that from the mPIC alone (Figure 5B, lanes 1 and 2, respectively). The magnitude of this stimulatory effect of the Head on basal transcription is comparable to that reported for the entire Mediator complex (Kim et al., 1994).
In a previous study, we had tested the overall effect of the Head on transcription and found only a much smaller (~2-fold) stimulation (Takagi et al., 2006). We believe that this disparity relates directly to an important difference in the way in which the present and previous transcription assays were carried out. In the “standard” assay described in the earlier study, all components (DNA, TBP, TFIIB, TFIIF, TFIIE, TFIIH, RNAPII with or without the Head) were mixed and then tested for transcription activity. This standard assay measures overall transcription activity, which reflects the compounded effect of several steps including assembly of the PIC, activation of the PIC, and transcript synthesis. The assay we used in the present work to compare the transcriptional activity of the mPIC and Head-mPIC complexes differs from a standard assay in that we tested pre-formed complexes that had already undergone the steps required for assembly of the mPIC or Head-mPIC. Therefore, our results show that although the effect of the Head on overall transcription (including PIC assembly and activation) is only ~2-fold, if the Head-mPIC is already assembled, the effect of the Head becomes much larger (~18-fold). This observation suggests that the effect of the Head module on transcriptional activity is largely focused on a post-assembly step, namely, activation of the complete, yet inactive PIC. This idea is consistent with our structural results, which would suggest that the Head activates the PIC by triggering opening of the RNAPII clamp.
Finally, it is important to consider how stimulation of RNAPII CTD phosphorylation by the Head module might affect our interpretation of the transcription assay results. CTD phosphorylation by TFIIH is associated with the transition from initiation to elongation of transcription (Payne et al., 1989), and both Mediator (Kim et al., 1994) and the Head (Imasaki et al., 2011) stimulate phosphorylation of the CTD. Conceivably, stimulation of transcription by the Head in a pre-assembled complex could be partially attributed to stimulation of CTD phosphorylation by the Head. However, we do not think this is the case because in a pure reconstituted system like the one discussed here, CTD phosphorylation is dispensable for basal transcription activity (Serizawa et al., 1993).
Further analysis of the Head-mPIC structure
Consistent with the presence of additional components (TFIIF, TFIIB, TBP, and DNA), several features in the Head-mPIC EM structure cannot be accounted for by fitting of the Head and RNAPII X-ray structures (see Figure 4A). These unidentified densities can be better appreciated in a difference map between the Head-mPIC cryo-EM structure and the fitted Head and RNAPII modules (Figure 6A, light blue surfaces, and Figure S11, light blue mesh). We based our interpretation of these additional features in the Head-mPIC structure on information about the interaction of RNAPII with TFIIF (Chen et al., 2007; Chen et al., 2010; Eichner et al., 2010), and on the structure of the RNAPII-TFIIB complex from X-ray crystallographic studies (Armache et al., 2003; Bushnell and Kornberg, 2003; Kostrewa et al., 2009; Liu et al., 2010).
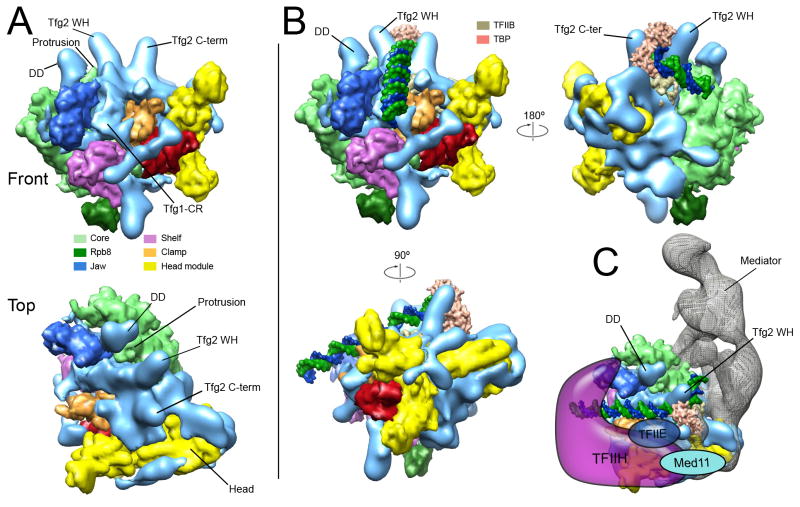
Figure 6. Implications for PIC organization and RNAPII-promoter interaction.
(A) Distribution of additional (non-RNAPII and non-Head) density (depicted as a light blue surface) along the RNAPII surface. Individual features are positioned adjacent to the locations of specific TFIIF domains determined through biochemical studies (see Figure S6) (Chen et al., 2007; Chen et al., 2010; Eichner et al., 2010). Densities that likely correspond to the Tfg1-Tfg2 dimerization domain (DD) and the Tfg2 wing-helix domain (Tfg2 WH) flank the RNAPII protrusion. Density matching the expected location of the Tfg1 charged region (Tfg1-CR) and Tfg2 C-terminus (Tfg2 C-term) are also observed. (B) Fitting of a model for organization of the RNAPII-TFIIB-TBP-DNA complex into the Head-mPIC cryo-EM map. Matching of the core module in the RNAPII-TFIIB and Head-mPIC structures results in TFIIB and TBP being nestled between the Tfg2 WH and Tfg2 C-term densities above the upstream end of the polymerase active site cleft, without any steric clashes with possible TFIIF density. The Tfg2 wing-helix domain (Tfg2 WH) would be positioned to interact with promoter DNA, as expected (Eichner et al., 2010). Promoter DNA would travel along the RNAPII active site cleft and have unimpeded access to the polymerase active site after opening of the clamp. (C) A model for interaction of additional preinitiation complex components with the Head-mPIC complex. The position of the Head would place Mediator subunit Med11 (partially located in the Head’s fixed jaw (Imasaki et al., 2011)) in a position where it could interact with TFIIH, as reported (Esnault et al., 2008). Also, opening of the active site cleft upon outward rotation of the clamp would expose surfaces for interaction with TFIIE and TFIIH. See also Figure S11.
The close physical proximity of most sizeable unassigned features in the Head-mPIC structure to patches of the RNAPII surface pinpointed as TFIIF-interacting areas by a number of studies that have used a combination of site-directed proteolysis, crosslinking, and mass spectrometry (MS) analysis to delineate TFIIF-RNAPII interactions (Chen et al., 2007; Chen et al., 2010; Eichner et al., 2010), suggests that these unidentified features could correspond to TFIIF domains (Figure S11). For example, we observe densities matching the expected positions of the Tfg1-Tfg2 dimerization domain, the Tfg2 wing-helix, and the Tfg2 C-terminus (Figure 6A). More extended density observed along the RNAPII active site cleft could also be at least partially related to TFIIF. The Tfg1-Tfg2 dimerization domain (DD) and the Tfg2 wing-helix domain (Tfg2 WH) would be positioned downstream and upstream of the RNAPII protrusion, as expected (Chen et al., 2010; Eichner et al., 2010).
Little density remains unidentified after tentative assignment of possible TFIIF domains as described above. Although biochemical and functional evidence (Figure 1) indicate that TFIIB, TBP, and promoter DNA must be present, we were unable to conclusively identify density corresponding to these factors. It has been reported that in vivo, the RNAPII PIC might be considerably dynamic (Fan and Struhl, 2009; Sprouse et al., 2008; Wade and Struhl, 2008), and a possible explanation for the absence of TFIIB-TBP-DNA density in the Head-mPIC cryo-EM map is that these components might have been poorly ordered. The results from our in vitro transcription assays (Figure 1C) demonstrate that TFIIB, TBP, and promoter DNA were present in the Head-mPIC complex at concentrations that were over an order of magnitude lower than those used for EM sample preparation, but it must be noted that the efficiency of the Head-mPIC reconstituted transcription assay (Figure 1C), measured on the basis of template utilization, is very low, on the order of 1%. This efficiency is on par with that observed in previous studies (Takagi et al., 2006; Takagi and Kornberg, 2006), but the possibility that only a small fraction of Head-mPIC complexes are organized as functional assemblies cannot be ruled out.
Implications for PIC organization and RNAPII-promoter interaction
Although we could not visualize density corresponding to TBP, TFIIB, or promoter DNA, our cryo-EM map of the Head-mPIC complex has, at a minimum, revealed the structure of the RNAPII-TFIIF-Head complex with which TFIIB, TBP, and promoter DNA would presumably interact. Matching the position of the core RNAPII module in the most current model of the minimal PIC (Kostrewa et al., 2009; Liu et al., 2010) with the position of the core module in the Head-mPIC cryo-EM map, results in a complementary fit between presumed TFIIF densities above the upstream end of the RNAPII active site cleft and the TFIIB-TBP-DNA complex. TFIIB and TBP would be nestled between the Tfg2 wing-helix domain and additional TFIIF density possibly corresponding to the Tfg2 C-terminus (Figure 6B).
DISCUSSION
The overall structure of Mediator, as well as large-scale changes in its conformation required for interaction with RNAPII, appear to be conserved from yeast to humans. This suggests a likely conservation of the basic mechanism for transcription regulation by Mediator (Cai et al., 2009). Because of its limited resolution and quality, our previous EM map of the Mediator-RNAPII complex (Davis et al., 2002) could not be used to unequivocally establish the precise mode of Mediator-RNAPII interaction, and therefore failed to provide information about the possible nature of the regulation mechanism. It had been reported that the Head module alone has a clear effect on basal transcription (Takagi et al., 2006; Takagi and Kornberg, 2006), but the basis of this effect remained elusive. Now, the structure of the Head-mPIC complex demonstrates that extensive interaction of the clamp and Rpb4-Rpb7 polymerase subunits with the Mediator Head module causes a considerable change in the conformation of RNAPII itself, which renders the enzyme’s active site accessible to promoter DNA. Our interpretation of in vitro transcription assay results is that the increase in accessibility to promoter DNA caused by Head interaction is directly responsible for a nearly 20–fold increase in transcriptional activity. The large positive effect of the Head on basal transcription is even more remarkable if one considers that a pre-assembled mPIC prepared on the short DNA template included in the Head-mPIC complex was largely inactive (Figure 5B). In vitro transcription assays with purified yeast proteins typically require the use of circular templates in which supercoiling facilitates promoter opening (Parvin and Sharp, 1993), and we believe that, without the conformational change triggered by the Head module, RNAPII cannot effectively engage the short double-stranded DNA template used for Head-mPIC assembly.
The increase in basal transcriptional activity triggered by the Head is comparable to that elicited by the complete Mediator, suggesting that clamp opening constitutes a fundamental aspect of the Mediator mechanism, and that the full effect of Mediator on basal transcription can be largely recapitulated by the Head module in the Head-mPIC complex. This appears reasonable, given that the interaction of the Head module and RNAPII in the Head-mPIC complex is in complete agreement with the interaction between the entire Mediator and RNAPII. Our initial assessment of the Mediator–RNAPII interaction (Asturias et al., 1999) was based on images of a Mediator–RNAPII complex formed with RNAPII lacking subunits Rpb4/Rpb7, and relied on an incomplete understanding of the RNAPII structure because an X-ray structure of the enzyme was not available at the time. This was followed by our EM analysis of a Mediator–RNAPII holoenzyme (including Rpb4/Rpb7 and TFIIF) purified from yeast as an entire complex (Davis et al., 2002). The 3D EM map of the holoenzyme showed some stain-induced deformation that hampered its interpretation, so we tried to determine the orientation of RNAPII based on cross-correlation analysis of 2D images. A more definitive analysis of the Mediator–RNAPII interaction became possible only after we determined a more faithful and detailed cryo-EM map of Mediator (Cai et al., 2009), and resulted in an interpretation of the Mediator–RNAPII interaction that is consistent with biochemical, genetic, and structural analysis of the Head–RNAPII (Cai et al., 2010), and Head–mPIC interactions. Interestingly, our interpretation of the Mediator–RNAPII interaction does not agree with that reported in the human system (Bernecky et al., 2011). Although the mode of interaction between Mediator and RNAPII could be conserved from yeast to humans, there is no direct evidence that this is indeed the case. Alternatively, it is conceivable that the yeast and human Mediator–RNAPII structures do not correspond to functionally equivalent “states” of Mediator–RNAPII interaction. Finally, it should be noted that in-depth analysis of our 16 Å resolution Head-mPIC structure was necessary to convincingly demonstrate a conformational change in RNAPII, while the model for Mediator–RNAPII interaction in the human system was derived from a structure at 36 Å resolution.
The new RNAPII conformation identified in our study is also a likely platform for assembly of additional components of the PIC. An open cleft may provide access to surfaces important for TFIIE and TFIIH interaction, as both factors are expected to bind around the downstream end of the RNAPII active site cleft (Chen et al., 2007; Drapkin et al., 1994) (Figure 6C). Although the mechanism of initiation regulation by Mediator must involve many aspects, modulation of RNAPII conformation in a way that directly influences promoter engagement is likely a fundamental component. Flexibility of the clamp domain has been documented in both bacterial and eukaryotic RNA polymerases (Cramer et al., 2001; Darst et al., 2002), and the clamp of archaeal polymerase has been recently identified as an important target for regulation of RNAPII initiation and processivity (Grohmann et al., 2011). Clamp opening appears to be critical for promoter engagement in bacterial RNA polymerase (Belogurov et al., 2009; Mukhopadhyay et al., 2008) and recent studies suggest that changes in the conformation of bacterial polymerase similar to the ones described here are involved in initiation regulation (Tagami et al., 2010). Finely regulated changes in Mediator structure would modulate interaction with RNAPII, with opening of the clamp by the Head functioning as a final effect. Determining how activator-induced changes in Mediator conformation might ultimately modulate the effect of Head binding on RNAPII conformation and PIC assembly is an important next challenge in understanding the Mediator mechanism.
METHODS
Assembly and purification of the Head-mPIC complex
Assembly of the Head-mPIC was described previously (Takagi et al., 2006). Briefly, we started by purifying a RNAPII-TFIIF complex from yeast cells through a TAP tag (Puig et al., 2001) placed at the C-terminus of the Tfg2 subunit of TFIIF (Chung et al., 2003). RNAPII-TFIIF was then incubated with a pre-assembled, recombinant TBP-TFIIB-DNA complex (promoter DNA was 53 residues in length, with a TATA box 30 residues upstream of the transcription initiation site, designed after a DNA segment used in previous structural studies (Westover et al., 2004)), to form an mPIC, the minimal assembly capable of promoter-specific initiation. This mPIC was incubated with a 1.5-fold molar excess of recombinant Head module (Takagi et al., 2006). The complex that resulted after incubation of all these components was purified by affinity chromatography on calmodulin-Sepharose, after immobilizing it through a calmodulin-binding peptide tag left at the Tfg2 C-terminus after TAP tagging. For SDS-PAGE of the Head-mPIC complex, approximately 6 μg of the calmodulin column peak fraction was electrophoresed in a 4–12% NuPAGE, followed by staining with Coomassie brilliant blue.
Transcription assays
In vitro transcription assays with pre-assembled Head-mPIC and mPIC complexes were performed essentially as described (Takagi and Kornberg, 2006), with the following modifications: only TFIIE and TFIIH were supplemented; 800 μM ATP, 800 μM GTP, 800 μM CTP and no cold UTP were used; reactions were incubated for 20 min at room temperature; and 15% (w/v) acrylamide, 3% (w/v) bisacrylamide, 7 M urea gels were used. Decade Marker RNA (Decade™ Marker System, Ambion) was labeled with [γ-32P] ATP according to the manufacture’s instruction. Quantification of transcripts on an absolute scale was performed with a FLA-5100 fluorescent image analyzer and the Multi-Gauge software package (FUJIFILM Life Science) after addition of 1 nCi of [α-32P] UTP to the gel 10 min before the end of the run.
Cryo-EM sample preparation
About 3 μl of Head-mPIC sample (~300 ug protein mL−1) were applied to 400 mesh Cu/Rh EM grids (Ted Pella) coated with a freshly glow-discharged (in the presence of amyl amine) perforated carbon film support and preserved by flash freezing in liquid ethane (Dubochet et al., 1988).
Initial cryo-EM analysis of the Head-mPIC complex
Images of Head-mPIC particles were collected under low-dose conditions with a CM200 FEG (Philips/FEI) electron microscope equipped with a field emission gun and operating at an accelerating voltage of 120 kV. Images were recorded on Kodak SO-163 film, at a magnification of 66,000X with underfocus values between 2.3 and 3.7 μm. A total of 19 micrographs were digitized on a Zeiss/SCAI flat bed densitometer (ZI/Zeiss) with a step size of 7 μm. Digitized images were two-fold pixel-averaged, resulting in a final pixel size corresponding to 2.05 Å. A total of ~15,000 particle images were divided into 12 groups according to defocus values calculated independently for 12 distinct sections of every micrograph. We used SPIDER (Frank et al., 1996) to carry out iterative reference-free alignment and clustering to separate images into groups, and tested the homogeneity of the groups by comparing the outcome from multiple rounds of reference-free alignment (Chaban et al., 2008). Class averages showing a large, elongated density expected to correspond to the Head module were assigned relative orientations by using the common lines routine implemented in the SPARX image processing package (Hohn et al., 2007), and assembled into an initial Head-mPIC structure. Next, we constructed a distortion-free model of the Head-mPIC complex by fitting a molecular model of RNAPII derived from its X-ray structure (Bushnell and Kornberg, 2003), and a Random Conical Tilt reconstruction of the recombinant Head module (Cai et al., 2010) into the initial Head-mPIC structure.
Automated collection of a larger Head-mPIC EM dataset
A larger Head-mPIC EM image dataset was collected with the Leginon automated data collection system (Suloway et al., 2005) on an FEI Tecnai F20 electron microscope operating at 120 kV. Images were recorded with a Tietz F415 4X4 K pixel CCD camera (15 μm pixel−1), at a dose of <10 e − Å−2 with underfocus values between 0.8 and 3.8 μm; about 500 CCD frames were recorded at a nominal magnification of 50,000X and a pixel size of 1.79 Å pixel−1.
EM image processing and structure refinement
Initial processing of the raw CCD frames was carried out with the Appion software package, which interfaces with the Leginon database infrastructure (Lander et al., 2009). The contrast transfer function (CTF) for each micrograph was estimated with the Automated CTF Estimation (ACE) package (Mallick et al., 2005); a total of 402 images whose CTF estimation had an ACE confidence of 0.8 or better were processed. About 120,000 particles images were picked with the DoG picker (Voss et al., 2009) routine available in Appion, and were extracted using a box size of 150×150 pixels. This dataset was carefully screened to discard questionable images to obtain a final image stack that contained 102,741 particle images. Images were two-fold binned to a final pixel size of 3.58 Å pixel−1. Further image processing was carried with the SPARX and SPIDER EM image processing packages (Frank et al., 1996; Hohn et al., 2007).
Supplementary Material
HIGHLIGHTS.
The Head module interacts with the RNA polymerase II clamp and subunits Rpb4-Rpb7
This interaction causes a structural rearrangement of polymerase and the Head
Clamp opening by the Head increases DNA accessibility to the polymerase active site
Increased promoter engagement results in a large increase in transcription activity
Acknowledgments
This work was supported by NIH grant R01 67167 (F.J.A.), NSF grant MCB 0843026 (Y.T.), American Heart Association Scientist Development Award 0735395N (Y.T.), the Showalter Trust Fund (Y.T.), and NIH grant R01 60635 (P.A.P.). T.I. was supported by an HFSP postdoctoral fellowship. We thank Roger Kornberg for providing materials for this study, and Johnathan Chittuluru for help with some details of EM image analysis.
Footnotes
Accession codes. The cryo-EM map of the Head-mPIC complex was deposited in the Electron Microscopy Data Bank with accession code EMD-5407.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Armache KJ, Kettenberger H, Cramer P. Architecture of initiation-competent 12-subunit RNA polymerase II. Proc Natl Acad Sci U S A. 2003;100:6964–6968. doi: 10.1073/pnas.1030608100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asturias FJ, Jiang YW, Myers LC, Gustafsson CM, Kornberg RD. Conserved structures of mediator and RNA polymerase II holoenzyme. Science. 1999;283:985–987. doi: 10.1126/science.283.5404.985. [DOI] [PubMed] [Google Scholar]
- Baek HJ, Malik S, Qin J, Roeder RG. Requirement of TRAP/mediator for both activator-independent and activator-dependent transcription in conjunction with TFIID-associated TAF(II)s. Mol Cell Biol. 2002;22:2842–2852. doi: 10.1128/MCB.22.8.2842-2852.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belogurov GA, Vassylyeva MN, Sevostyanova A, Appleman JR, Xiang AX, Lira R, Webber SE, Klyuyev S, Nudler E, Artsimovitch I, et al. Transcription inactivation through local refolding of the RNA polymerase structure. Nature. 2009;457:332–335. doi: 10.1038/nature07510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bernecky C, Grob P, Ebmeier CC, Nogales E, Taatjes DJ. Molecular architecture of the human Mediator-RNA polymerase II-TFIIF assembly. PLoS Biol. 2011;9:e1000603. doi: 10.1371/journal.pbio.1000603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brignole EJ, Smith S, Asturias FJ. Conformational flexibility of metazoan fatty acid synthase enables catalysis. Nat Struct Mol Biol. 2009;16:190–197. doi: 10.1038/nsmb.1532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bushnell DA, Kornberg RD. Complete, 12-subunit RNA polymerase II at 4.1-A resolution: implications for the initiation of transcription. Proc Natl Acad Sci U S A. 2003;100:6969–6973. doi: 10.1073/pnas.1130601100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai G, Imasaki T, Takagi Y, Asturias FJ. Mediator structural conservation and implications for the regulation mechanism. Structure. 2009;17:559–567. doi: 10.1016/j.str.2009.01.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai G, Imasaki T, Yamada K, Cardelli F, Takagi Y, Asturias FJ. Mediator head module structure and functional interactions. Nat Struct Mol Biol. 2010;17:273–279. doi: 10.1038/nsmb.1757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaban Y, Ezeokonkwo C, Chung WH, Zhang F, Kornberg RD, Maier-Davis B, Lorch Y, Asturias FJ. Structure of a RSC-nucleosome complex and insights into chromatin remodeling. Nat Struct Mol Biol. 2008;15:1272–1277. doi: 10.1038/nsmb.1524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen HT, Warfield L, Hahn S. The positions of TFIIF and TFIIE in the RNA polymerase II transcription preinitiation complex. Nat Struct Mol Biol. 2007;14:696–703. doi: 10.1038/nsmb1272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen ZA, Jawhari A, Fischer L, Buchen C, Tahir S, Kamenski T, Rasmussen M, Lariviere L, Bukowski-Wills JC, Nilges M, et al. Architecture of the RNA polymerase II-TFIIF complex revealed by cross-linking and mass spectrometry. EMBO J. 2010;29:717–726. doi: 10.1038/emboj.2009.401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung WH, Craighead JL, Chang WH, Ezeokonkwo C, Bareket-Samish A, Kornberg RD, Asturias FJ. RNA polymerase II/TFIIF structure and conserved organization of the initiation complex. Mol Cell. 2003;12:1003–1013. doi: 10.1016/s1097-2765(03)00387-3. [DOI] [PubMed] [Google Scholar]
- Craighead J, Chang W, Asturias F. Structure of Yeast RNA Polymerase II in Solution. Implications for Enzyme Regulation and Interaction with Promoter DNA. Structure (Camb) 2002;10:1117–1125. doi: 10.1016/s0969-2126(02)00813-4. [DOI] [PubMed] [Google Scholar]
- Cramer P, Bushnell DA, Fu J, Gnatt AL, Maier-Davis B, Thompson NE, Burgess RR, Edwards AM, David PR, Kornberg RD. Architecture of RNA polymerase II and implications for the transcription mechanism. Science. 2000;288:640–649. doi: 10.1126/science.288.5466.640. [DOI] [PubMed] [Google Scholar]
- Cramer P, Bushnell DA, Kornberg RD. Structural basis of transcription: RNA polymerase II at 2.8 angstrom resolution. Science. 2001;292:1863–1876. doi: 10.1126/science.1059493. [DOI] [PubMed] [Google Scholar]
- Darst SA, Opalka N, Chacon P, Polyakov A, Richter C, Zhang G, Wriggers W. Conformational flexibility of bacterial RNA polymerase. Proc Natl Acad Sci U S A. 2002;99:4296–4301. doi: 10.1073/pnas.052054099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis JA, Takagi Y, Kornberg RD, Asturias FA. Structure of the yeast RNA polymerase II holoenzyme: Mediator conformation and polymerase interaction. Mol Cell. 2002;10:409–415. doi: 10.1016/s1097-2765(02)00598-1. [DOI] [PubMed] [Google Scholar]
- Drapkin R, Reardon JT, Ansari A, Huang JC, Zawel L, Ahn K, Sancar A, Reinberg D. Dual role of TFIIH in DNA excision repair and in transcription by RNA polymerase II. Nature. 1994;368:769–772. doi: 10.1038/368769a0. [DOI] [PubMed] [Google Scholar]
- Dubochet J, Adrian M, Chang JJ, Homo JC, Lepault J, McDowall AW, Schultz P. Cryo-electron microscopy of vitrified specimens. Q Rev Biophys. 1988;21:129–228. doi: 10.1017/s0033583500004297. [DOI] [PubMed] [Google Scholar]
- Eichner J, Chen HT, Warfield L, Hahn S. Position of the general transcription factor TFIIF within the RNA polymerase II transcription preinitiation complex. EMBO J. 2010;29:706–716. doi: 10.1038/emboj.2009.386. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esnault C, Ghavi-Helm Y, Brun S, Soutourina J, Van Berkum N, Boschiero C, Holstege F, Werner M. Mediator-dependent recruitment of TFIIH modules in preinitiation complex. Mol Cell. 2008;31:337–346. doi: 10.1016/j.molcel.2008.06.021. [DOI] [PubMed] [Google Scholar]
- Fan X, Struhl K. Where does mediator bind in vivo? PLoS One. 2009;4:e5029. doi: 10.1371/journal.pone.0005029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flanagan PM, Kelleher RJ, Feaver WJ, Lue NF, LaPointe JW, Kornberg RD. Resolution of factors required for the initiation of transcription by yeast RNA polymerase II. J Biol Chem. 1990;265:11105–11107. [PubMed] [Google Scholar]
- Frank J, Radermacher M, Penczek P, Zhu J, Li Y, Ladjadj M, Leith A. SPIDER and WEB: Processing and visualization of images in 3D electron microscopy and related fields. J Struct Biol. 1996;116:190–199. doi: 10.1006/jsbi.1996.0030. [DOI] [PubMed] [Google Scholar]
- Gao H, Valle M, Ehrenberg M, Frank J. Dynamics of EF-G interaction with the ribosome explored by classification of a heterogeneous cryo-EM dataset. J Struct Biol. 2004;147:283–290. doi: 10.1016/j.jsb.2004.02.008. [DOI] [PubMed] [Google Scholar]
- Giardina C, Lis JT. Polymerase processivity and termination on Drosophila heat shock genes. J Biol Chem. 1993;268:23806–23811. [PubMed] [Google Scholar]
- Grohmann D, Nagy J, Chakraborty A, Klose D, Fielden D, Ebright RH, Michaelis J, Werner F. The Initiation Factor TFE and the Elongation Factor Spt4/5 Compete for the RNAP Clamp during Transcription Initiation and Elongation. Mol Cell. 2011;43:263–274. doi: 10.1016/j.molcel.2011.05.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hahn S. Structure and mechanism of the RNA polymerase II transcription machinery. Nat Struct Mol Biol. 2004;11:394–403. doi: 10.1038/nsmb763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hampsey M. Molecular genetics of the RNA polymerase II general transcriptional machinery. Microbiol Mol Biol Rev. 1998;62:465–503. doi: 10.1128/mmbr.62.2.465-503.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hohn M, Tang G, Goodyear G, Baldwin PR, Huang Z, Penczek PA, Yang C, Glaeser RM, Adams PD, Ludtke SJ. SPARX, a new environment for Cryo-EM image processing. J Struct Biol. 2007;157:47–55. doi: 10.1016/j.jsb.2006.07.003. [DOI] [PubMed] [Google Scholar]
- Imasaki T, Calero G, Cai G, Tsai KL, Yamada K, Cardelli F, Erdjument-Bromage H, Tempst P, Berger I, Kornberg GL, et al. Architecture of the Mediator head module. Nature. 2011;475:240–243. doi: 10.1038/nature10162. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim YJ, Bjorklund S, Li Y, Sayre MH, Kornberg RD. A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II. Cell. 1994;77:599–608. doi: 10.1016/0092-8674(94)90221-6. [DOI] [PubMed] [Google Scholar]
- Kornberg RD. Eukaryotic transcriptional control. Trends Cell Biol. 1999;9:M46–49. [PubMed] [Google Scholar]
- Kostrewa D, Zeller ME, Armache KJ, Seizl M, Leike K, Thomm M, Cramer P. RNA polymerase II-TFIIB structure and mechanism of transcription initiation. Nature. 2009;462:323–330. doi: 10.1038/nature08548. [DOI] [PubMed] [Google Scholar]
- Lander GC, Stagg SM, Voss NR, Cheng A, Fellmann D, Pulokas J, Yoshioka C, Irving C, Mulder A, Lau PW, et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J Struct Biol. 2009;166:95–102. doi: 10.1016/j.jsb.2009.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu X, Bushnell DA, Wang D, Calero G, Kornberg RD. Structure of an RNA polymerase II-TFIIB complex and the transcription initiation mechanism. Science. 2010;327:206–209. doi: 10.1126/science.1182015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mallick SP, Carragher B, Potter CS, Kriegman DJ. ACE: automated CTF estimation. Ultramicroscopy. 2005;104:8–29. doi: 10.1016/j.ultramic.2005.02.004. [DOI] [PubMed] [Google Scholar]
- Meyer KD, Lin SC, Bernecky C, Gao Y, Taatjes DJ. p53 activates transcription by directing structural shifts in Mediator. Nat Struct Mol Biol. 2010;17:753–760. doi: 10.1038/nsmb.1816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mukhopadhyay J, Das K, Ismail S, Koppstein D, Jang M, Hudson B, Sarafianos S, Tuske S, Patel J, Jansen R, et al. The RNA polymerase “switch region” is a target for inhibitors. Cell. 2008;135:295–307. doi: 10.1016/j.cell.2008.09.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naar AM, Beaurang PA, Zhou S, Abraham S, Solomon W, Tjian R. Composite co-activator ARC mediates chromatin-directed transcriptional activation. Nature. 1999;398:828–832. doi: 10.1038/19789. [DOI] [PubMed] [Google Scholar]
- Parvin JD, Sharp PA. DNA topology and a minimal set of basal factors for transcription by RNA polymerase II. Cell. 1993;73:533–540. doi: 10.1016/0092-8674(93)90140-l. [DOI] [PubMed] [Google Scholar]
- Payne JM, Laybourn PJ, Dahmus ME. The transition of RNA polymerase II from initiation to elongation is associated with phosphorylation of the carboxyl-terminal domain of subunit IIa. J Biol Chem. 1989;264:19621–19629. [PubMed] [Google Scholar]
- Penczek PA, Kimmel M, Spahn CM. Identifying conformational states of macromolecules by eigen-analysis of resampled cryo-EM images. Structure. 2011;19:1582–1590. doi: 10.1016/j.str.2011.10.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Penczek PA, Zhu J, Frank J. A common-lines based method for determining orientations for N > 3 particle projections simultaneously. Ultramicroscopy. 1996;63:205–218. doi: 10.1016/0304-3991(96)00037-x. [DOI] [PubMed] [Google Scholar]
- Puig O, Caspary F, Rigaut G, Rutz B, Bouveret E, Bragado-Nilsson E, Wilm M, Seraphin B. The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods. 2001;24:218–229. doi: 10.1006/meth.2001.1183. [DOI] [PubMed] [Google Scholar]
- Sato S, Tomomori-Sato C, Parmely TJ, Florens L, Zybailov B, Swanson SK, Banks CA, Jin J, Cai Y, Washburn MP, et al. A set of consensus mammalian mediator subunits identified by multidimensional protein identification technology. Mol Cell. 2004;14:685–691. doi: 10.1016/j.molcel.2004.05.006. [DOI] [PubMed] [Google Scholar]
- Serizawa H, Conaway JW, Conaway RC. Phosphorylation of C-terminal domain of RNA polymerase II is not required in basal transcription. Nature. 1993;363:371–374. doi: 10.1038/363371a0. [DOI] [PubMed] [Google Scholar]
- Sprouse RO, Karpova TS, Mueller F, Dasgupta A, McNally JG, Auble DT. Regulation of TATA-binding protein dynamics in living yeast cells. Proc Natl Acad Sci U S A. 2008;105:13304–13308. doi: 10.1073/pnas.0801901105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suloway C, Pulokas J, Fellmann D, Cheng A, Guerra F, Quispe J, Stagg S, Potter CS, Carragher B. Automated molecular microscopy: the new Leginon system. J Struct Biol. 2005;151:41–60. doi: 10.1016/j.jsb.2005.03.010. [DOI] [PubMed] [Google Scholar]
- Taatjes DJ, Schneider-Poetsch T, Tjian R. Distinct conformational states of nuclear receptor-bound CRSP-Med complexes. Nat Struct Mol Biol. 2004;11:664–671. doi: 10.1038/nsmb789. [DOI] [PubMed] [Google Scholar]
- Tagami S, Sekine S, Kumarevel T, Hino N, Murayama Y, Kamegamori S, Yamamoto M, Sakamoto K, Yokoyama S. Crystal structure of bacterial RNA polymerase bound with a transcription inhibitor protein. Nature. 2010;468:978–982. doi: 10.1038/nature09573. [DOI] [PubMed] [Google Scholar]
- Takagi Y, Calero G, Komori H, Brown JA, Ehrensberger AH, Hudmon A, Asturias F, Kornberg RD. Head module control of mediator interactions. Mol Cell. 2006;23:355–364. doi: 10.1016/j.molcel.2006.06.007. [DOI] [PubMed] [Google Scholar]
- Takagi Y, Kornberg RD. Mediator as a general transcription factor. J Biol Chem. 2006;281:80–89. doi: 10.1074/jbc.M508253200. [DOI] [PubMed] [Google Scholar]
- Voss NR, Yoshioka CK, Radermacher M, Potter CS, Carragher B. DoG Picker and TiltPicker: software tools to facilitate particle selection in single particle electron microscopy. J Struct Biol. 2009;166:205–213. doi: 10.1016/j.jsb.2009.01.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wade JT, Struhl K. The transition from transcriptional initiation to elongation. Curr Opin Genet Dev. 2008;18:130–136. doi: 10.1016/j.gde.2007.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Westover KD, Bushnell DA, Kornberg RD. Structural basis of transcription: separation of RNA from DNA by RNA polymerase II. Science. 2004;303:1014–1016. doi: 10.1126/science.1090839. [DOI] [PubMed] [Google Scholar]
- Yan Q, Moreland RJ, Conaway JW, Conaway RC. Dual roles for transcription factor IIF in promoter escape by RNA polymerase II. J Biol Chem. 1999;274:35668–35675. doi: 10.1074/jbc.274.50.35668. [DOI] [PubMed] [Google Scholar]
- Yang Z, Fang J, Chittuluru J, Asturias FJ, Penczek PA. Iterative Stable Alignment and Clustering of 2D Transmission Electron Microscope Images. Structure. 2012;20:237–247. doi: 10.1016/j.str.2011.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.