Abstract
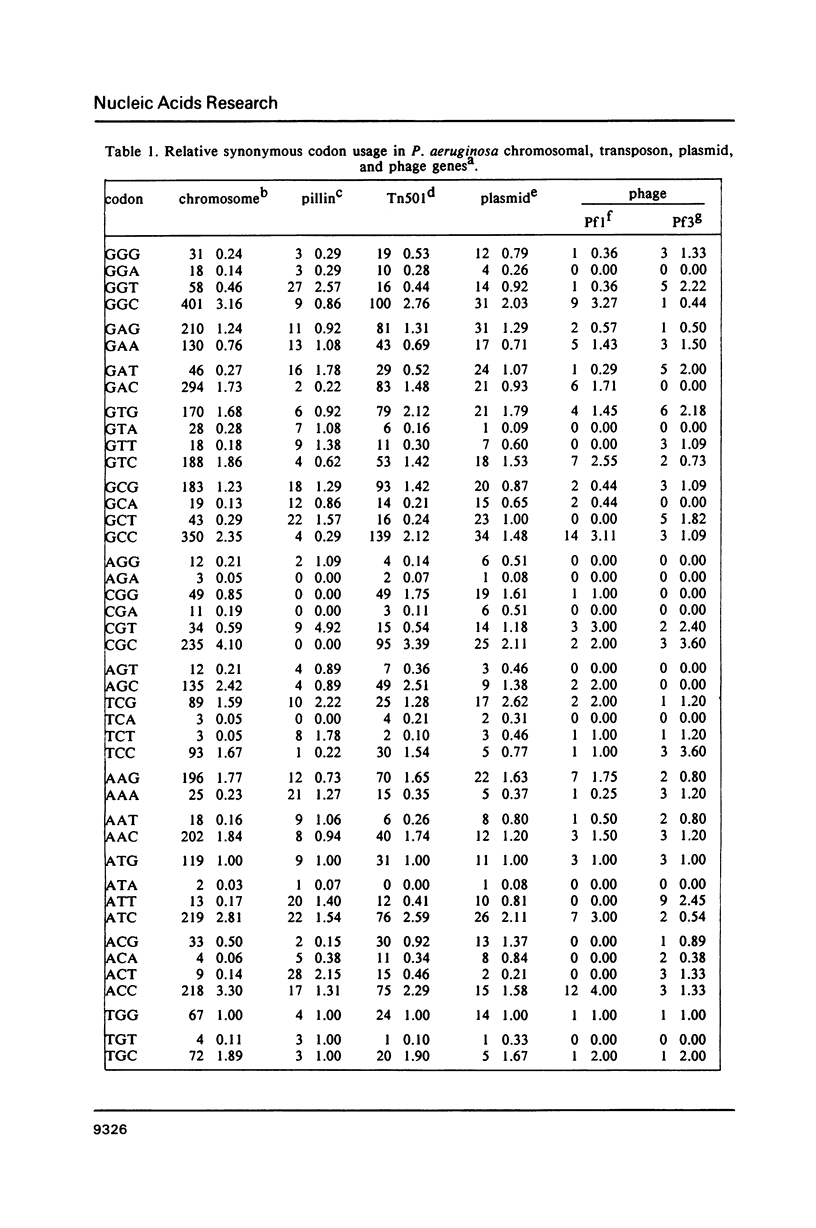
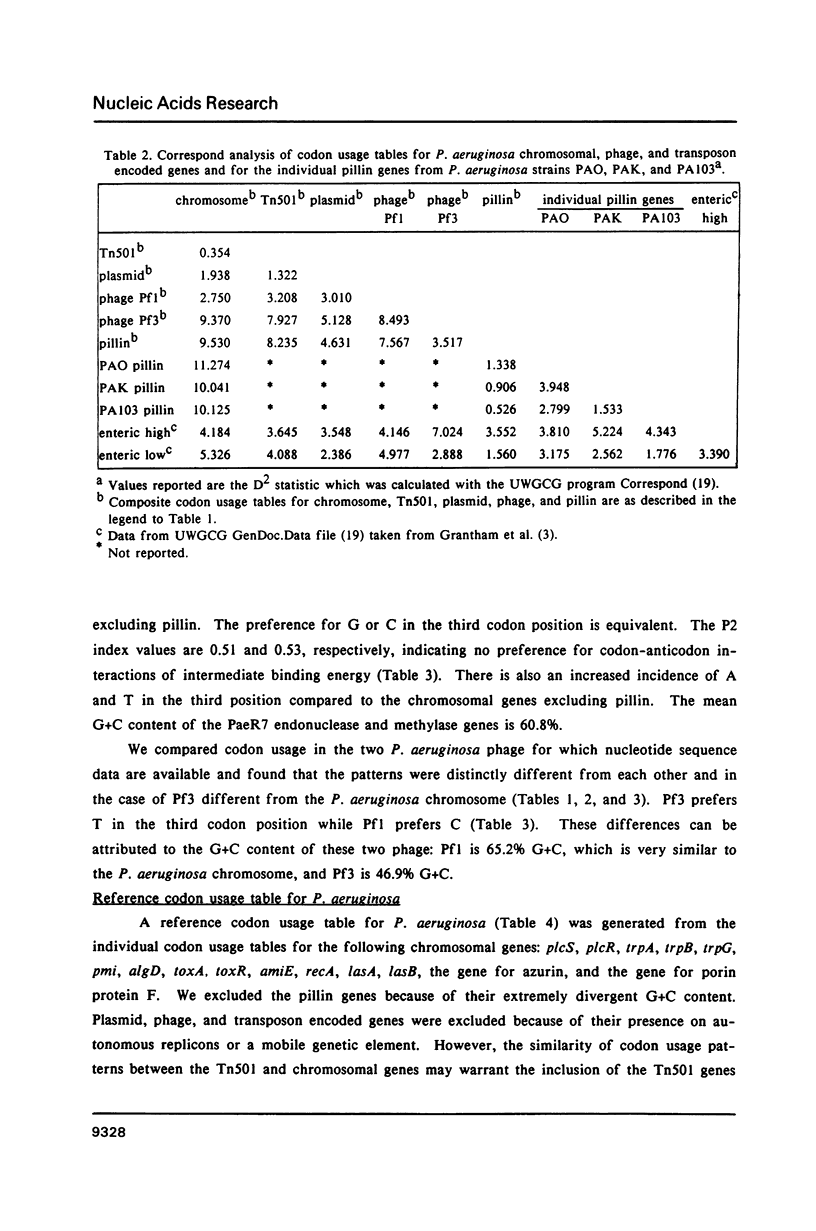
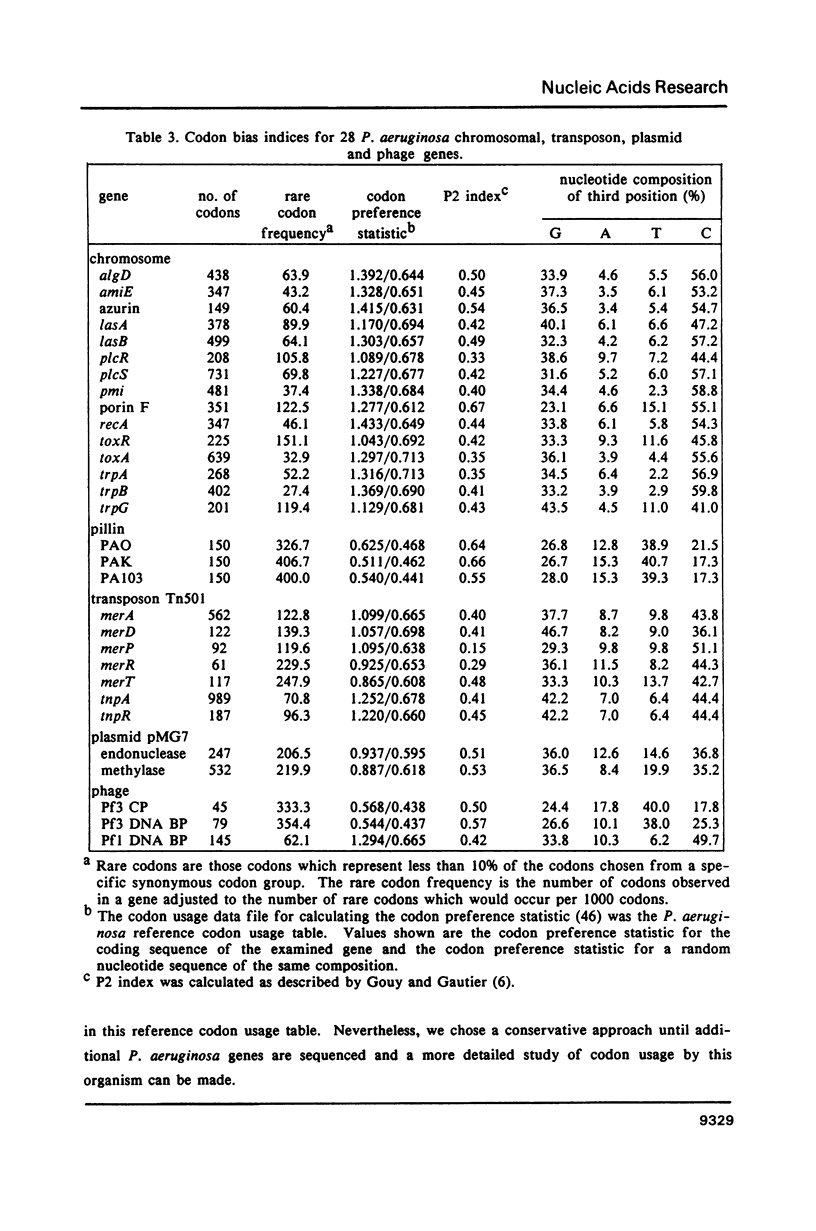
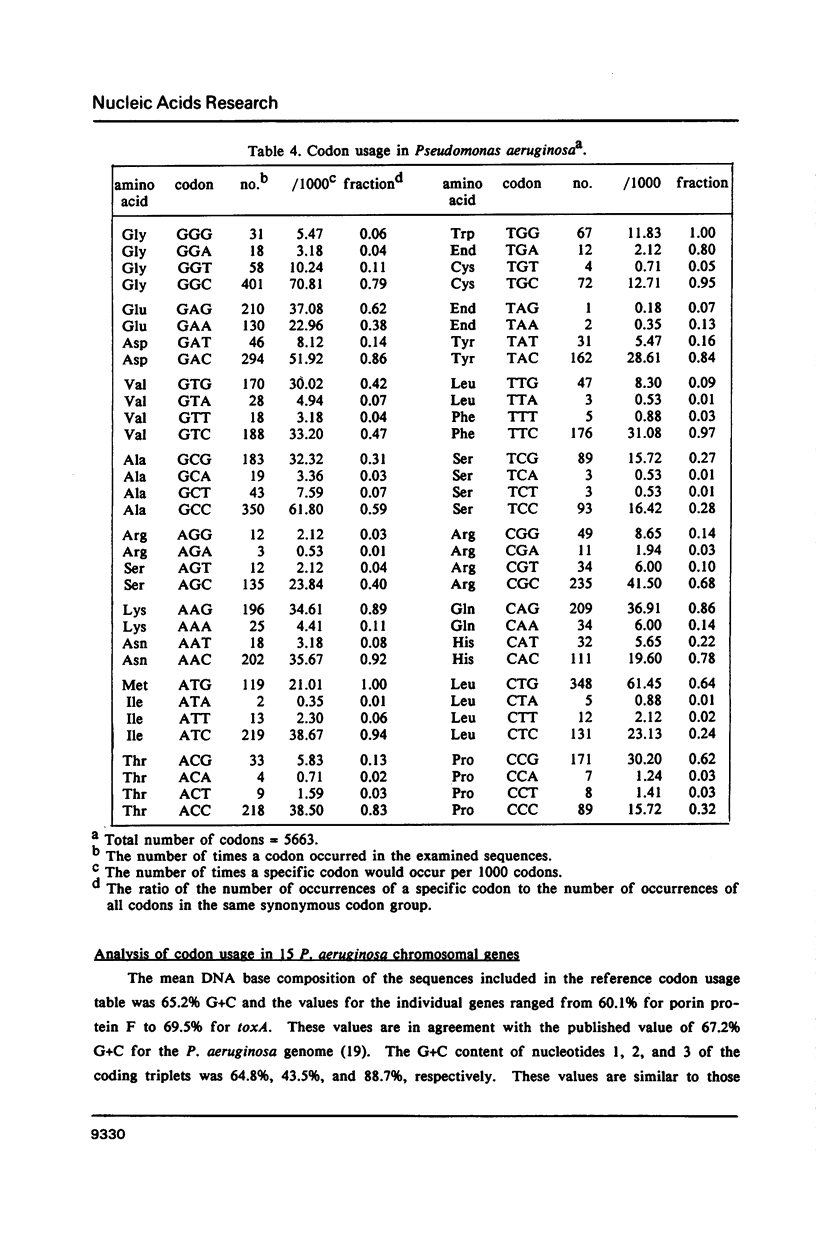
We have generated a codon usage table for Pseudomonas aeruginosa. Codon usage in P. aeruginosa is extremely biased. In contrast to E. coli and yeast, P. aeruginosa preferentially uses those codons within a synonymous codon group with the strongest predicted codon-anticodon interaction. We were unable to correlate a particular codon usage pattern with predicted levels of mRNA expressivity. The choice of a third base reflects the high guanine plus cytosine content of the P. aeruginosa genome (67.2%) and cytosine is the preferred nucleotide for the third codon position.
Full text
PDF








Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bennetzen J. L., Hall B. D. Codon selection in yeast. J Biol Chem. 1982 Mar 25;257(6):3026–3031. [PubMed] [Google Scholar]
- Bernardi G., Bernardi G. Codon usage and genome composition. J Mol Evol. 1985;22(4):363–365. doi: 10.1007/BF02115693. [DOI] [PubMed] [Google Scholar]
- Bever R. A., Iglewski B. H. Molecular characterization and nucleotide sequence of the Pseudomonas aeruginosa elastase structural gene. J Bacteriol. 1988 Sep;170(9):4309–4314. doi: 10.1128/jb.170.9.4309-4314.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bibb M. J., Bibb M. J., Ward J. M., Cohen S. N. Nucleotide sequences encoding and promoting expression of three antibiotic resistance genes indigenous to Streptomyces. Mol Gen Genet. 1985;199(1):26–36. doi: 10.1007/BF00327505. [DOI] [PubMed] [Google Scholar]
- Bibb M. J., Findlay P. R., Johnson M. W. The relationship between base composition and codon usage in bacterial genes and its use for the simple and reliable identification of protein-coding sequences. Gene. 1984 Oct;30(1-3):157–166. doi: 10.1016/0378-1119(84)90116-1. [DOI] [PubMed] [Google Scholar]
- Bilofsky H. S., Burks C. The GenBank genetic sequence data bank. Nucleic Acids Res. 1988 Mar 11;16(5):1861–1863. doi: 10.1093/nar/16.5.1861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brammar W. J., Charles I. G., Matfield M., Liu C. P., Drew R. E., Clarke P. H. The nucleotide sequence of the amiE gene of Pseudomonas aeruginosa. FEBS Lett. 1987 May 11;215(2):291–294. doi: 10.1016/0014-5793(87)80164-3. [DOI] [PubMed] [Google Scholar]
- Brown N. L., Ford S. J., Pridmore R. D., Fritzinger D. C. Nucleotide sequence of a gene from the Pseudomonas transposon Tn501 encoding mercuric reductase. Biochemistry. 1983 Aug 16;22(17):4089–4095. doi: 10.1021/bi00286a015. [DOI] [PubMed] [Google Scholar]
- Brown N. L., Winnie J. N., Fritzinger D., Pridmore R. D. The nucleotide sequence of the tnpA gene completes the sequence of the Pseudomonas transposon Tn501. Nucleic Acids Res. 1985 Aug 12;13(15):5657–5669. doi: 10.1093/nar/13.15.5657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cameron G. N. The EMBL data library. Nucleic Acids Res. 1988 Mar 11;16(5):1865–1867. doi: 10.1093/nar/16.5.1865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Canters G. W. The azurin gene from Pseudomonas aeruginosa codes for a pre-protein with a signal peptide. Cloning and sequencing of the azurin gene. FEBS Lett. 1987 Feb 9;212(1):168–172. doi: 10.1016/0014-5793(87)81579-x. [DOI] [PubMed] [Google Scholar]
- Crawford I. P., Eberly L. Structure and regulation of the anthranilate synthase genes in Pseudomonas aeruginosa: I. Sequence of trpG encoding the glutamine amidotransferase subunit. Mol Biol Evol. 1986 Sep;3(5):436–448. doi: 10.1093/oxfordjournals.molbev.a040408. [DOI] [PubMed] [Google Scholar]
- Darzins A., Frantz B., Vanags R. I., Chakrabarty A. M. Nucleotide sequence analysis of the phosphomannose isomerase gene (pmi) of Pseudomonas aeruginosa and comparison with the corresponding Escherichia coli gene manA. Gene. 1986;42(3):293–302. doi: 10.1016/0378-1119(86)90233-7. [DOI] [PubMed] [Google Scholar]
- Deretic V., Gill J. F., Chakrabarty A. M. Pseudomonas aeruginosa infection in cystic fibrosis: nucleotide sequence and transcriptional regulation of the algD gene. Nucleic Acids Res. 1987 Jun 11;15(11):4567–4581. doi: 10.1093/nar/15.11.4567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diver W. P., Grinsted J., Fritzinger D. C., Brown N. L., Altenbuchner J., Rogowsky P., Schmitt R. DNA sequences of and complementation by the tnpR genes of Tn21, Tn501 and Tn1721. Mol Gen Genet. 1983;191(2):189–193. doi: 10.1007/BF00334812. [DOI] [PubMed] [Google Scholar]
- Duchêne M., Schweizer A., Lottspeich F., Krauss G., Marget M., Vogel K., von Specht B. U., Domdey H. Sequence and transcriptional start site of the Pseudomonas aeruginosa outer membrane porin protein F gene. J Bacteriol. 1988 Jan;170(1):155–162. doi: 10.1128/jb.170.1.155-162.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gouy M., Gautier C. Codon usage in bacteria: correlation with gene expressivity. Nucleic Acids Res. 1982 Nov 25;10(22):7055–7074. doi: 10.1093/nar/10.22.7055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grantham R., Gautier C., Gouy M. Codon frequencies in 119 individual genes confirm consistent choices of degenerate bases according to genome type. Nucleic Acids Res. 1980 May 10;8(9):1893–1912. doi: 10.1093/nar/8.9.1893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grantham R., Gautier C., Gouy M., Jacobzone M., Mercier R. Codon catalog usage is a genome strategy modulated for gene expressivity. Nucleic Acids Res. 1981 Jan 10;9(1):r43–r74. doi: 10.1093/nar/9.1.213-b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grantham R., Gautier C., Gouy M., Mercier R., Pavé A. Codon catalog usage and the genome hypothesis. Nucleic Acids Res. 1980 Jan 11;8(1):r49–r62. doi: 10.1093/nar/8.1.197-c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gray G. L., Smith D. H., Baldridge J. S., Harkins R. N., Vasil M. L., Chen E. Y., Heyneker H. L. Cloning, nucleotide sequence, and expression in Escherichia coli of the exotoxin A structural gene of Pseudomonas aeruginosa. Proc Natl Acad Sci U S A. 1984 May;81(9):2645–2649. doi: 10.1073/pnas.81.9.2645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gribskov M., Devereux J., Burgess R. R. The codon preference plot: graphic analysis of protein coding sequences and prediction of gene expression. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):539–549. doi: 10.1093/nar/12.1part2.539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grosjean H., Fiers W. Preferential codon usage in prokaryotic genes: the optimal codon-anticodon interaction energy and the selective codon usage in efficiently expressed genes. Gene. 1982 Jun;18(3):199–209. doi: 10.1016/0378-1119(82)90157-3. [DOI] [PubMed] [Google Scholar]
- Hadero A., Crawford I. P. Nucleotide sequence of the genes for tryptophan synthase in Pseudomonas aeruginosa. Mol Biol Evol. 1986 May;3(3):191–204. doi: 10.1093/oxfordjournals.molbev.a040388. [DOI] [PubMed] [Google Scholar]
- Ikemura T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes. J Mol Biol. 1981 Feb 15;146(1):1–21. doi: 10.1016/0022-2836(81)90363-6. [DOI] [PubMed] [Google Scholar]
- Ikemura T. Correlation between the abundance of yeast transfer RNAs and the occurrence of the respective codons in protein genes. Differences in synonymous codon choice patterns of yeast and Escherichia coli with reference to the abundance of isoaccepting transfer RNAs. J Mol Biol. 1982 Jul 15;158(4):573–597. doi: 10.1016/0022-2836(82)90250-9. [DOI] [PubMed] [Google Scholar]
- Johnson K., Parker M. L., Lory S. Nucleotide sequence and transcriptional initiation site of two Pseudomonas aeruginosa pilin genes. J Biol Chem. 1986 Nov 25;261(33):15703–15708. [PubMed] [Google Scholar]
- Kagawa Y., Nojima H., Nukiwa N., Ishizuka M., Nakajima T., Yasuhara T., Tanaka T., Oshima T. High guanine plus cytosine content in the third letter of codons of an extreme thermophile. DNA sequence of the isopropylmalate dehydrogenase of Thermus thermophilus. J Biol Chem. 1984 Mar 10;259(5):2956–2960. [PubMed] [Google Scholar]
- Konigsberg W., Godson G. N. Evidence for use of rare codons in the dnaG gene and other regulatory genes of Escherichia coli. Proc Natl Acad Sci U S A. 1983 Feb;80(3):687–691. doi: 10.1073/pnas.80.3.687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maeda K., Kneale G. G., Tsugita A., Short N. J., Perham R. N., Hill D. F., Petersen G. B. The DNA-binding protein of Pf1 filamentous bacteriophage: amino-acid sequence and structure of the gene. EMBO J. 1982;1(2):255–261. doi: 10.1002/j.1460-2075.1982.tb01156.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marrs C. F., Schoolnik G., Koomey J. M., Hardy J., Rothbard J., Falkow S. Cloning and sequencing of a Moraxella bovis pilin gene. J Bacteriol. 1985 Jul;163(1):132–139. doi: 10.1128/jb.163.1.132-139.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Misra T. K., Brown N. L., Fritzinger D. C., Pridmore R. D., Barnes W. M., Haberstroh L., Silver S. Mercuric ion-resistance operons of plasmid R100 and transposon Tn501: the beginning of the operon including the regulatory region and the first two structural genes. Proc Natl Acad Sci U S A. 1984 Oct;81(19):5975–5979. doi: 10.1073/pnas.81.19.5975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muto A., Kawauchi Y., Yamao F., Osawa S. Preferential use of A- and U-rich codons for Mycoplasma capricolum ribosomal proteins S8 and L6. Nucleic Acids Res. 1984 Nov 12;12(21):8209–8217. doi: 10.1093/nar/12.21.8209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muto A., Osawa S. The guanine and cytosine content of genomic DNA and bacterial evolution. Proc Natl Acad Sci U S A. 1987 Jan;84(1):166–169. doi: 10.1073/pnas.84.1.166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nicas T. I., Hancock R. E. Pseudomonas aeruginosa outer membrane permeability: isolation of a porin protein F-deficient mutant. J Bacteriol. 1983 Jan;153(1):281–285. doi: 10.1128/jb.153.1.281-285.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohama T., Yamao F., Muto A., Osawa S. Organization and codon usage of the streptomycin operon in Micrococcus luteus, a bacterium with a high genomic G + C content. J Bacteriol. 1987 Oct;169(10):4770–4777. doi: 10.1128/jb.169.10.4770-4777.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pasloske B. L., Finlay B. B., Paranchych W. Cloning and sequencing of the Pseudomonas aeruginosa PAK pilin gene. FEBS Lett. 1985 Apr 22;183(2):408–412. doi: 10.1016/0014-5793(85)80821-8. [DOI] [PubMed] [Google Scholar]
- Pritchard A. E., Vasil M. L. Nucleotide sequence and expression of a phosphate-regulated gene encoding a secreted hemolysin of Pseudomonas aeruginosa. J Bacteriol. 1986 Jul;167(1):291–298. doi: 10.1128/jb.167.1.291-298.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Putterman D. G., Casadevall A., Boyle P. D., Yang H. L., Frangione B., Day L. A. Major coat protein and single-stranded DNA-binding protein of filamentous virus Pf3. Proc Natl Acad Sci U S A. 1984 Feb;81(3):699–703. doi: 10.1073/pnas.81.3.699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sano Y., Kageyama M. The sequence and function of the recA gene and its protein in Pseudomonas aeruginosa PAO. Mol Gen Genet. 1987 Jul;208(3):412–419. doi: 10.1007/BF00328132. [DOI] [PubMed] [Google Scholar]
- Sastry P. A., Finlay B. B., Pasloske B. L., Paranchych W., Pearlstone J. R., Smillie L. B. Comparative studies of the amino acid and nucleotide sequences of pilin derived from Pseudomonas aeruginosa PAK and PAO. J Bacteriol. 1985 Nov;164(2):571–577. doi: 10.1128/jb.164.2.571-577.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schad P. A., Iglewski B. H. Nucleotide sequence and expression in Escherichia coli of the Pseudomonas aeruginosa lasA gene. J Bacteriol. 1988 Jun;170(6):2784–2789. doi: 10.1128/jb.170.6.2784-2789.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. M., Li W. H. Codon usage in regulatory genes in Escherichia coli does not reflect selection for 'rare' codons. Nucleic Acids Res. 1986 Oct 10;14(19):7737–7749. doi: 10.1093/nar/14.19.7737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. M., Tuohy T. M., Mosurski K. R. Codon usage in yeast: cluster analysis clearly differentiates highly and lowly expressed genes. Nucleic Acids Res. 1986 Jul 11;14(13):5125–5143. doi: 10.1093/nar/14.13.5125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen B. F., Tai P. C., Pritchard A. E., Vasil M. L. Nucleotide sequences and expression in Escherichia coli of the in-phase overlapping Pseudomonas aeruginosa plcR genes. J Bacteriol. 1987 Oct;169(10):4602–4607. doi: 10.1128/jb.169.10.4602-4607.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shields D. C., Sharp P. M. Synonymous codon usage in Bacillus subtilis reflects both translational selection and mutational biases. Nucleic Acids Res. 1987 Oct 12;15(19):8023–8040. doi: 10.1093/nar/15.19.8023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staden R., McLachlan A. D. Codon preference and its use in identifying protein coding regions in long DNA sequences. Nucleic Acids Res. 1982 Jan 11;10(1):141–156. doi: 10.1093/nar/10.1.141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theriault G., Roy P. H., Howard K. A., Benner J. S., Brooks J. E., Waters A. F., Gingeras T. R. Nucleotide sequence of the PaeR7 restriction/modification system and partial characterization of its protein products. Nucleic Acids Res. 1985 Dec 9;13(23):8441–8461. doi: 10.1093/nar/13.23.8441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trifonov E. N. Translation framing code and frame-monitoring mechanism as suggested by the analysis of mRNA and 16 S rRNA nucleotide sequences. J Mol Biol. 1987 Apr 20;194(4):643–652. doi: 10.1016/0022-2836(87)90241-5. [DOI] [PubMed] [Google Scholar]
- Wozniak D. J., Cram D. C., Daniels C. J., Galloway D. R. Nucleotide sequence and characterization of toxR: a gene involved in exotoxin A regulation in Pseudomonas aeruginosa. Nucleic Acids Res. 1987 Mar 11;15(5):2123–2135. doi: 10.1093/nar/15.5.2123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zalacain M., González A., Guerrero M. C., Mattaliano R. J., Malpartida F., Jiménez A. Nucleotide sequence of the hygromycin B phosphotransferase gene from Streptomyces hygroscopicus. Nucleic Acids Res. 1986 Feb 25;14(4):1565–1581. doi: 10.1093/nar/14.4.1565. [DOI] [PMC free article] [PubMed] [Google Scholar]



