Abstract
To solve the problem of embryonic lethality in conventional gene knockouts, site-specific recombinase (SSR) systems (Cre-loxP, Flp-FRT, and ΦC31) have been used for tissue-specific gene knockout. With the combination of an SSR system and inducible gene expression systems (tetracycline and tamoxifen), stage-specific knockout and transgenic expression can be achieved. The application of this “SSR+inducible” conditional tool to genomic manipulation can be extended in various ways. Alternatives to conditional gene targeting, such as conditional gene trapping, multipurpose conditional alleles, and conditional gene silencing, have been developed. SSR systems can also be used to construct precise disease models with point mutations and chromosomal abnormalities. With these exciting achievements, we are moving towards a new era in which the whole genome can be manipulated as we wish.
Keywords: Site-specific recombinase, Gene targeting, Gene trapping, Inducible systems, ΦC31 system
1. Introduction
After “the Human Genome Project” (HGP) transcribed the human genome into a huge book of “ATGC” (Adenine, Thymine, Guanine, Cytosine) (Watson and Cook-Deegan, 1991; Abramowicz, 2003), the post-genomic era began. The mission during this era is to interpret the encrypted book into one that makes sense to us. Gene knockout has been the most efficient method for the functional research of these genes, and is based on the simple understanding that “we do not know what we’ve got until we lose it.” However, if the loss is lethal during early embryonic development, all we can learn through conventional knockout is that the product of the gene is essential to the organism. Also, one gene product can function differently within different tissues or cell types. In this case, the simultaneous knockout of a gene within a whole organism is likely to produce complicated results, with too much noise to understand the specific functions of the gene product. Finally, the gene knockout strategy has been applied to the construction of animal disease models for human genetic disorders, but some human genetic diseases such as cancers develop in certain cell types or at certain developmental stages. A precise model for these diseases has to achieve tissue- and/or stage-specific control of these gene mutations. Thus, conditional gene knockout was developed. Using site-specific recombinase (SSR) systems, we can knock out the target gene within a particular cell type at certain developmental phases (conditional knockout). In this way, we are able to analyze the functions of most gene products. With the expansion of various new technologies in SSR systems, conditional gene manipulation has greatly surpassed conditional gene knockout. Now, we can manipulate the genome freely. In this review, we introduce SSR and inducible systems, and then extend into the applications of new gene knockout methods that have been developed through the effective use of SSR systems. We also discuss other SSR-related gene manipulation methods. Finally, we provide guidelines for gene function studies and a perspective for the future.
2. Tissue-specific knockout using Cre-loxP, Flp-FRT, or ΦC31 system
Some conventional knockout mutations lead to animal embryonic lethality; hence, SSRs and floxed alleles are used to circumvent this problem. SSRs, the enzymes catalyzing site-specific recombination within particular nucleotide sequences, are the key elements for conditional gene knockout. The most widely used systems are the Cre-loxP system from the bacteriophage P1 (Hoess et al., 1982) and the Flp-FRT system from Saccharomyces cerevisiae (Sternberg et al., 1981). In the Cre-loxP or Flp-FRT system, Cre or Flp recombinase recognizes the 34-bp nucleotide sequence named loxP or FRT and precisely catalyzes the homologous exchange between the two loxP or the two FRT sites, respectively. No nucleotides are gained or lost in this process, which has sufficient in vivo and in vitro efficiencies (Hoess et al., 1984). In such exchanges, no additional elements, except for some monovalent or divalent cations, are needed (Stark et al., 1992). The size of loxP is neither too large to interrupt the function of the genes, nor too small to occur randomly within the genome.
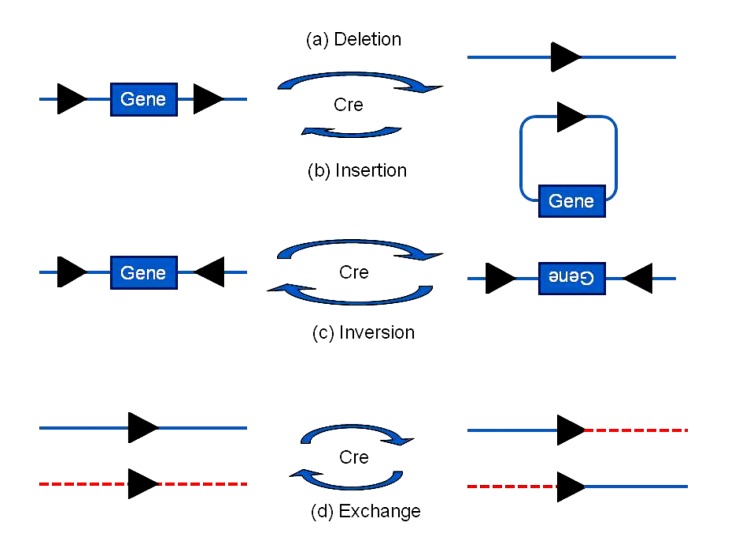
The Cre-loxP and Flp-FRT systems (Hoess et al., 1982; McLeod et al., 1986) enable the specific manipulation of DNA based on the direction and location of the two loxPs or FRTs (Fig. 1). Cre catalyzes the deletion of the DNA between the two loxPs when the two loxPs are in the same direction on one DNA molecule. When one loxP is on a linear DNA molecule and another loxP is on a circular DNA molecule, the circular DNA integrates into the linear DNA at the target. If two loxPs are in opposite directions, the fragment between them inverts. When one loxP is on a linear DNA and a second loxP is on another linear DNA, the two linear DNA molecules exchange a segment (similar to chromosomal rearrangement). Conditional gene knockout works using the excision function of the SSR systems. The two loxP or two FRT sites are inserted into the genome through gene targeting, flanking the gene of interest or an important exon (Gao et al., 2004). The DNA segment (DNA flanked by two loxP or two FRT sites) is catalytically excised by the recombinase (Fig. 2). To obtain a tissue-specific cut, a Cre transgenic mouse was produced by placing the Cre recombinase gene downstream of a tissue-specific promoter. By breeding tissue-specific Cre mice with floxed mice (produced by targeting a specific gene), tissue-specific knockout can be obtained. The first tissue-specific Cre line and genomic DNA modifications produced by the Cre-loxP system were developed by Orban et al. (1992). With the expansion of the tissue-specific Cre database, controlled gene manipulations can increasingly be achieved (Nagy et al., 2009). Researchers may search databases such as the Jackson Laboratory Cre Repository (http://cre.jax.org), the Cre-X-Mice database of the Nagy Laboratory (http://nagy.mshri.on.ca/cre_new/index.php), and the Mutant Mouse Regional Resource Center (http://www.mmrrc.org) to obtain access to the desired Cre-expressing lines, and cross these Cre mice with targeted mouse lines. Aside from tissue-specific knockout, the different Cre lines can be used for other types of genetic manipulation.
Fig. 1.
DNA manipulations controlled by Cre and loxP directions or locations
(a) If the two loxPs are in the same direction on one DNA molecule, the DNA between them will be deleted. (b) When one loxP is on a linear DNA while another is on a circular DNA, the circular DNA will integrate into the linear DNA at the target. Small circular DNA is easily lost in cells; hence, the reaction in (a) happens more easily than that in (b). (c) If two loxPs are opposite, the DNA fragment between them will be inverted. (d) The DNA molecules will exchange a segment if both loxPs lie on linear DNA molecules. The black triangles are loxP sites, and indicate the direction. The length of the arrows indicates the relative tendency of reactions. Modified from Nagy (2000)
Fig. 2.
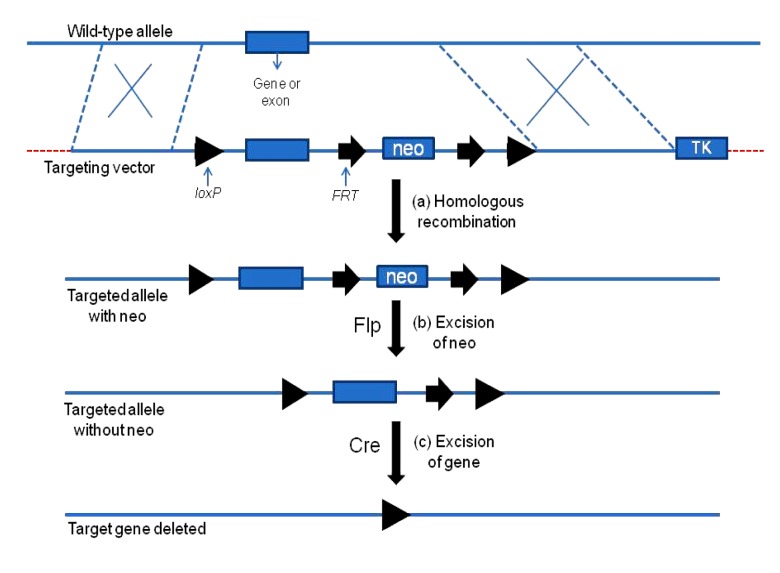
Conditional gene knockout
(a) In embryonic stem (ES) cells, the wild-type allele recombines with the targeting vectors on two homologous arms. This homologous recombination introduces two FRT sites flanking a selection marker neomycin-resistance gene (neo) and two loxP sites flanking the gene (or exon) of interest. (b) The Flp expression in the ES cells results in the excision of neo. (c) At the appropriate stage (usually in mice), the gene (or exon) is excised with the Cre expression. The rectangle (except TK) is the gene of interest (or exon). The black triangles indicate the loxP sites, and the black arrows indicate the FRT sites. TK: thymidine kinase gene. Modified from Guan et al. (2010)
Although powerful, the Cre-loxP and Flp-FRT systems remain imperfect. In theory, the recombination catalyzed by Cre or Flp is reversible. Occasionally, the equilibrium favors the undesired direction. For example, in site-specific insertion, the reaction is likely to cut out the already inserted site (Figs. 1a and 1b). Finding ways to control the reaction direction has become a big challenge for the Cre-loxP and Flp-FRT systems. Thus far, several methods have been developed, one of which involves limiting the reaction time with a heat-shock promoter (Huang et al., 1991; Morris et al., 1991) and introducing subtle mutations into the loxP or FRT sites to block the reverse reaction (Senecoff et al., 1988; Albert et al., 1995).
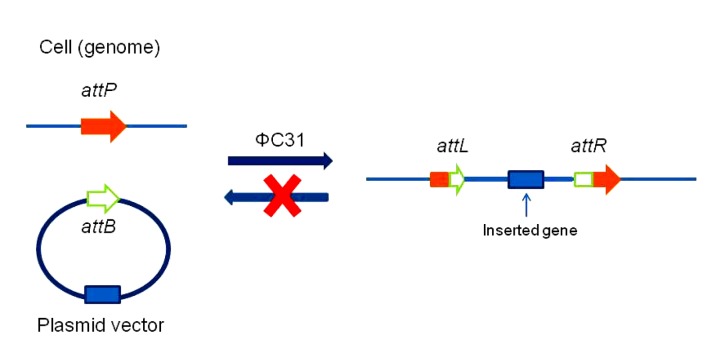
To address the reverse reaction problem, a third SSR system, the ΦC31 integrase system, was developed (Thyagarajan et al., 2001). The ΦC31 integrase was derived from the Streptomyces phage ΦC31 and catalyzes the recombination between the attP site (39 bp minimal size) and the attB site (34 bp minimal size), forming attL and attR sites (Fig. 3) (Groth et al., 2000). The ΦC31 integrase cannot catalyze recombination between the attL and attR sites (Thorpe and Smith, 1998; Thorpe et al., 2000). Therefore, the ΦC31 integrase can catalyze recombination only in a strictly controlled direction (Smith and Thorpe, 2002). The ΦC31 system has been successfully applied in site-directed insertion (Sharma et al., 2008) and gene targeting (Lister, 2010; Lu et al., 2011).
Fig. 3.
ΦC31 integrase system
The ΦC31 integrase catalyzes recombination between the attP and attB sites to form the attL and attR sites. The attL and attR sites are the hybrid sites, which are no longer recognized by the ΦC31 integrase. Modified from Groth et al. (2000)
In recent years, several pseudo-ΦC31 integrase target (attP) sites have been found in human and mouse cells. If a vector with an attB site is transfected into mammalian cells, a high frequency of integration occurs at these pseudo-attP sites (Thyagarajan et al., 2001; Ma et al., 2006). However, these pseudo sites in mammalian cells may not be a bad thing. Thyagarajan et al. (2001) has pointed out that these sites can be used as insertion sites in gene therapy and chromosomal engineering, and preclinical applications of the ΦC31 system have also been summarized (Calos, 2006).
3. Stage-specific knockout and transgenic expression using inducible gene expression systems
SSR systems are widely used and can be powerful tools for inactivating genes in specific tissues. However, sometimes we want to study gene function at a specific stage. Stage-specific knockout and transgenic expression systems require inducible expression of Cre or Flp. The inducible systems allow turning on the Cre or Flp expression when needed. To achieve stage-specific knockout and transgenic expression, inducible Cre-loxP and Flp-FRT systems have been developed. The first inducible system developed was a tetracycline system, the Tet-Off system (Gossen and Bujard, 1992). Other inducible systems were later developed, including the Tet-On system (Gossen et al., 1995), the tamoxifen system (Metzger et al., 1995), and hormone-inducible systems (Braselmann et al., 1993; Yao et al., 1993; Picard, 1994; No et al., 1996; Wang et al., 1999). These systems use an exogenous chemical to control the expression of a target gene. This control is reversible by providing or withdrawing the chemical. The two most frequently used inducible systems in stage-specific knockouts are the tetracycline system and the tamoxifen system.
3.1. Stage-specific knockout using tetracycline-inducible Cre-loxP or Flp-FRT system
The tetracycline-inducible system consists of the Tet-Off system (tetracycline-controlled transactivator protein (tTA) dependent) and the Tet-On system (reverse tetracycline-controlled transactivator protein (rtTA) dependent) (Gossen and Bujard, 1992; Gossen et al., 1995). In these two systems, expression of the target gene is dependent on the activity of an inducible transcriptional activator. The transcriptional activator is regulated reversibly by the inducing ligand tetracycline or tetracycline derivatives such as doxycycline (Dox).
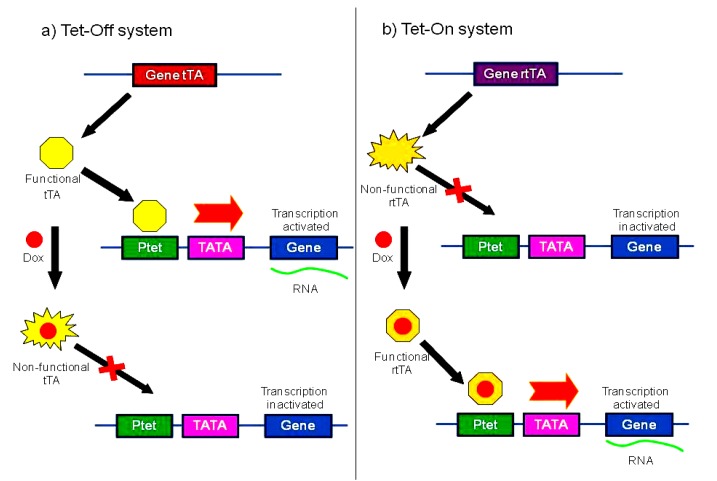
In the Tet-Off system (Fig. 4a), target gene expression is turned off with the inducing ligand tetracycline or Dox. tTA is an artificial transcription-activating fusion protein of a tetracycline repressor from Escherichia coli transposon Tn10 and the activation domain of the herpes simplex virus VP16 protein. tTA binds with a tTA-responsive promoter (Ptet) to drive the expression of a gene in the absence of tetracycline or Dox. After Dox is added, it binds with the transcription factor tTA. The Dox-tTA complex cannot bind with Ptet and target gene expression is stopped. In contrast, in the Tet-On system (Fig. 4b), target gene expression is turned on by the inducing ligand. The transcription factor rtTA does not bind with Ptet without tetracycline or Dox; hence, the gene under study is not expressed. After Dox is added, it binds with rtTA. The Dox-rtTA complex binds with Ptet and target gene expression is initiated.
Fig. 4.
Tetracycline-inducible system
(a) Tet-Off system. tTA is active without Dox (or tetracycline), and the gene of interest is expressed. With Dox (or tetracycline) treatment, tTA is inactivated, and the gene is no longer expressed. (b) Tet-On system. rtTA is inactive without Dox (or tetracycline), and the gene of interest is not expressed. With Dox (or tetracycline) treatment, rtTA is activated, and the gene is expressed
The Tet-Off and Tet-On systems are complementary (Kistner et al., 1996). Choosing one over the other depends on your strategy.
Stage-specific knockout can be achieved with a combination of SSR and Tet-On systems. Two mouse lines are needed for stage-specific knockout using tetracycline-inducible Cre-loxP or Flp-FRT system. The floxed animals are produced by targeting a specific gene in embryonic stem (ES) cells and generating mice. Another transgenic mouse line is needed in which the expression of Cre (or Flp) can be induced by tetracycline or Dox treatment. By crossing the Tet-On Cre (or Flp) line with a floxed mouse line, the target gene can be knocked out at the specific time when tetracycline or Dox is administered.
3.2. Stage-specific knockout using tamoxifen-inducible Cre-loxP system
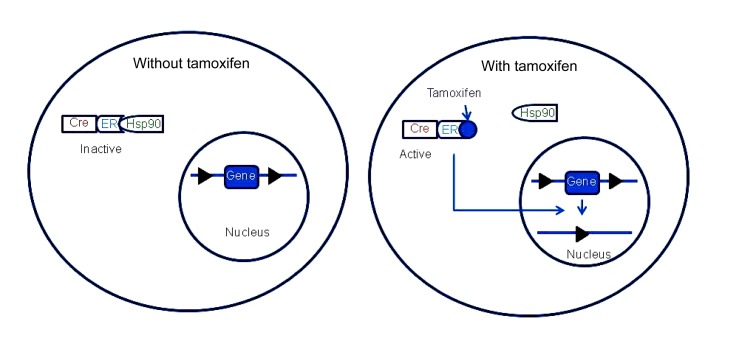
The tamoxifen system is the most commonly used inducible system for stage-specific knockout. Unlike the tetracycline system, which regulates the transcription of a gene, the tamoxifen system regulates gene function at the protein level (Logie and Stewart, 1995; Metzger et al., 1995). In this system, a mutated form of the ligand binding domain of the estrogen receptor (ER) is fused with Cre recombinase, forming Cre-ERT. The Cre-ERT protein no longer binds to 17β-estradiol, its natural ligand. Instead, it binds the synthetic ligands tamoxifen or 4-hydroxytamoxifen (Feil et al., 1996; Zhang et al., 1996). In the absence of tamoxifen, the Cre-ERT fusion protein is located in the cytoplasm and binds to heat-shock proteins such as Hsp90. In the presence of tamoxifen, Hsp90 is displaced from ER, which exposes the nuclear localization signal of the ER. Under the guidance of the nuclear localization signal, the Cre recombinase moves to the nucleus, where it executes its function (Fig. 5) (Chow et al., 2006; Tian et al., 2006). Newer versions of the ligand-dependent recombinase, Cre-ERT2 and ER-Cre-ER, have been developed, which are more efficient than Cre-ERT (Feil et al., 1997; Indra et al., 1999; Casanova et al., 2002; Matsuda and Cepko, 2007).
Fig. 5.
Tamoxifen-inducible system
In the absence of tamoxifen, Cre-ER binds with Hsp90, and is located in the cytoplasm. When tamoxifen is present, it binds with the ER, displacing Hsp90. The Cre-ER-tamoxifen complex translocates to the nucleus, in which Cre executes its function. The black triangles indicate loxP sites. Modified from Tian et al. (2006)
As in the tetracycline-inducible system, two mouse lines are needed for making stage-specific knockouts using the tamoxifen-inducible Cre-loxP system. In the floxed mouse, the gene of interest is flanked by two loxP sites. In the transgenic mouse line, the Cre-ERT is downstream of a ubiquitous or tissue-specific promoter, and can be activated by tamoxifen at the protein level. By crossing the Cre-ERT transgenic mouse line with the floxed mouse line, the target gene can be knocked out at the specific time when tamoxifen is administered.
3.3. Stage-specific gene expression in transgenic animals using an inducible Cre-loxP system
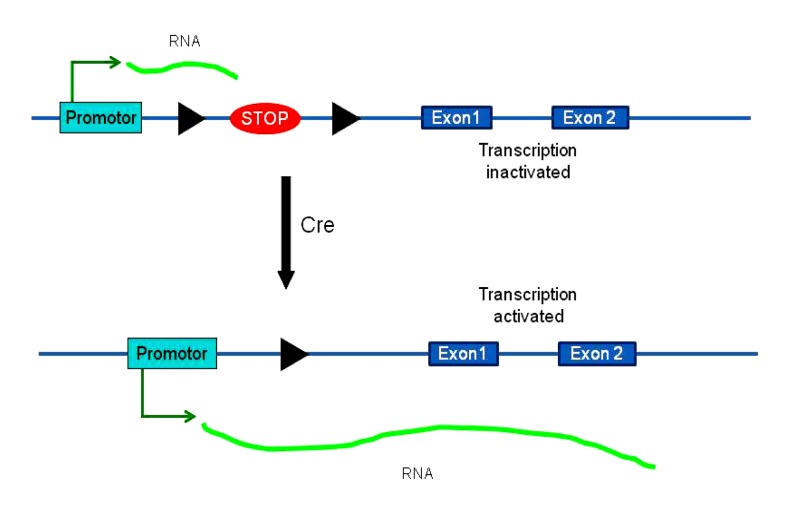
Transgene technology can be used to introduce a foreign gene into the genome of an organism and overexpress it. It is used mainly to study the function of the inserted gene or to produce large quantities of proteins for medical purposes. The gene transferred into transgenic animals may be controlled to express at a specific time and in a specific tissue using the inducible Cre-loxP system. In the construct for stage-specific gene expression in transgenic animals, a STOP is positioned between the ubiquitous promoter (CMV) and the gene of interest. This STOP is flanked by two loxP sites. Transgenic animals are produced using the above construct. The transgene is “off” at this stage because of the floxed STOP, but once the transgenic mice are crossed with a Cre line, Cre removes the STOP, and the transgene is turned on (Fig. 6) (Dragatsis and Zeitlin, 2001). If an inducible Cre is used, transgenic expression is achieved at a specific time and in a specific tissue.
Fig. 6.
Stage-specific gene expression in transgenic animals
The floxed STOP cassette is inserted between the promoter and the first exon of the gene. Transgenic animals may be generated using the construct, but exons following the STOP cassette are not transcribed. With the expression of Cre, the STOP cassette is cut out, restoring the normal expression of the gene. The black triangles indicate loxP sites. Modified from Dragatsis and Zeitlin (2001)
4. Conditional gene trapping
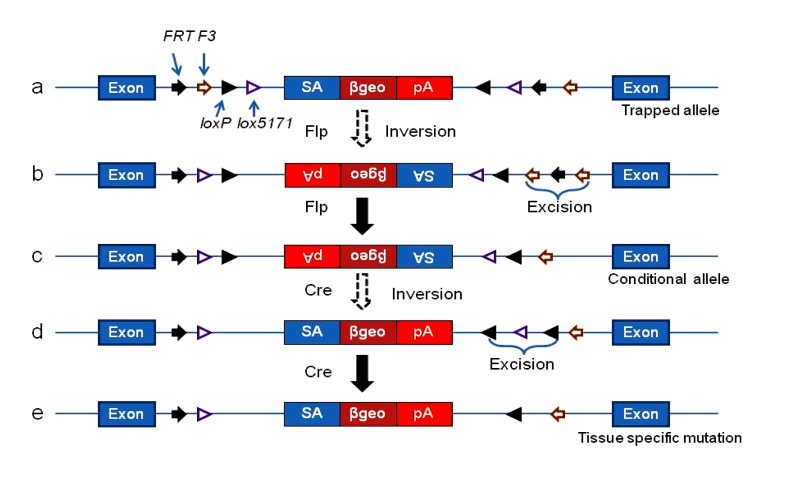
Aside from gene targeting, gene trapping is a widely used large-scale, high-throughput knockout method. Like conventional knockout via gene targeting, gene trapping also causes embryonic lethality. To circumvent this problem, a conditional gene trapping method was developed by the European Conditional Mouse Mutagenesis (EUCOMM) program. This method is based on the inversion function of the SSR system. The conditional gene trapping vector consists of a selection/reporter cassette flanked by four pairs of SSR sites in opposite orientations (Friedel et al., 2007). FRT and F3 are heterotypic recognition sites for Flp recombinase. F3 is a mutated version of FRT. Flp recombinase catalyzes the recombination between two FRT sites or between two F3 sites, but it cannot catalyze the recombination between an FRT site and an F3 site. LoxP and lox5171 are heterotypic recognition sites for Cre recombinase. Cre recombinase catalyzes the recombination between two loxP sites or between two lox5171 sites, but not between a loxP site and a lox5171 site. After transfection, this conditional gene trapping vector inserts into the intron of a gene, similar to the conventional trapping vectors, and leads to a mutation because the cassette is placed in the orientation for trapping genes. Knockout mice can be produced using these gene-trapped ES cell lines. Alternatively, this mutated allele can change into a conditional allele at the ES cell level by transiently expressing Flp recombinase in the ES cells. Flp recombinase inverts the selection/reporter cassette at either the FRT or the F3 site. This conditional allele is not mutagenic because the cassette is in the opposite orientation for trapping genes. A mouse line can be made using the ES cells carrying this conditional allele. The trapping vector may be re-inverted into the mutant orientation at loxP or lox5171 sites in some tissues by crossing the above mice with another tissue-specific Cre mouse line. Using this method, a tissue-specific knockout mouse line was generated (Fig. 7) (Schnütgen, 2006). This conditional trapping cassette traps any gene without modification, and may be used directly as a conventional trapping cassette to trap the genes in all tissues from the start of development. For genes that require conditional trapping, ES cells with this trapping vector can be treated with Flp to inactivate the trapping cassette. Mice can be generated using ES cells with this inactive trapping cassette. By crossing these mice with a tissue-specific Cre line, the trapping cassette is functionally restored and the target gene is trapped (knocked out).
Fig. 7.
Conditional gene trapping in the EUCOMM
(a) Trapped allele. The trapping vector inserts into the genome of the ES cells. With a splice acceptor, a fusion transcript is formed between the upstream exons of the gene and the selection/reporter cassette. The transcription of the downstream exons is terminated, and results in the mutation of an endogenous gene. (b) Transient step for inverting the trapping vector with Flp recombinase. By expressing Flp recombinase in the ES cells, the selection/reporter cassette is inverted from a pair of F3 or FRT. Consequently, a pair of homotypic F3 or FRT sites is placed in the same orientation. Recombination between these two homotypic sites excises the other heterotypic site between them, and the selection/reporter cassette is locked against re-inversion. (c) Conditional allele. The inversion restores the gene function in the ES cells because the selection/reporter cassette is in the opposite orientation for trapping genes. (d, e) Transient step and trapped allele in specific tissues. By breeding the mouse from the ES cell line in (c) with a tissue-specific Cre mouse, the trapping vector is re-inverted, resulting in a conditional mutation in specific tissues depending on the Cre mouse line. The mechanism of re-inversion is the same as that of the inversion in the Flp-FRT/F3 system in (a) and (b). SA: splice acceptor; βgeo: a fusion of neo and β-gal; pA: poly A. Modified from Schnütgen (2006)
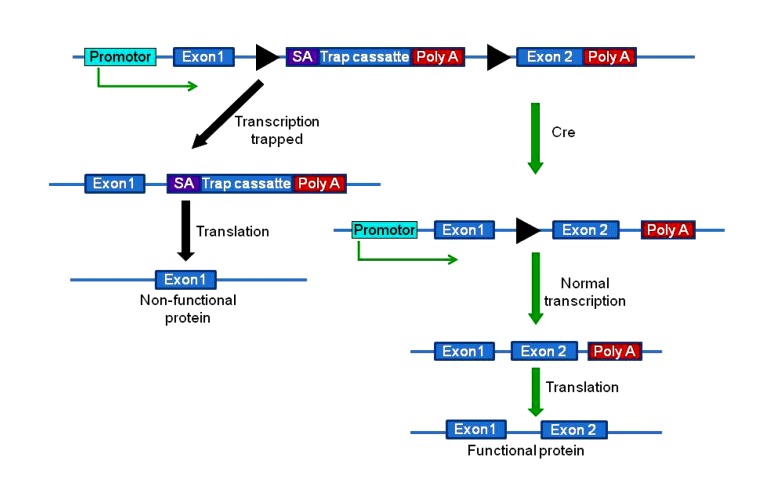
Another type of conditional gene trapping is called the “removable gene trap”, in which the trapping vector is flanked by two loxP sites (Fig. 8) (Ishida and Leder, 1999). Unlike Schnütgen (2006)’s strategy, Ishida and Leder (1999)’s method traps poly A signal but not promoters. An advantage of this “poly A trap” method is that it can disrupt silent genes in ES cells because the trap cassette has its own promoter. With the controlled expression of the Cre recombinase in certain tissues and at certain time points, the trapping vector can be excised from the gene, resulting in the restoration of normal gene expression. However, Shigeoka et al. (2005) found one problem of the “poly A trap” method: inefficient expression of the selection marker by nonsense-mediated messenger RNA (mRNA) decay (NMD). Thus, they developed a novel poly A trap strategy, unbiased poly A trapping (UPATrap), which suppresses NMD of the selectable-marker mRNA and permits the trapping of transcriptionally silent genes without a bias in the vector-integration site (Shigeoka et al., 2005).
Fig. 8.
Removable gene trap
A floxed trapping cassette is inserted within a gene (between exon1 and exon2 in this case). The transcription is “trapped”, and only exon1 is translated, resulting in a non-functional protein. When Cre is expressed, the trapping cassette is excised, and the gene resumes its normal transcription and translation. The black triangles indicate loxP sites. Modified from Ishida and Leder (1999)
5. Multipurpose conditional alleles
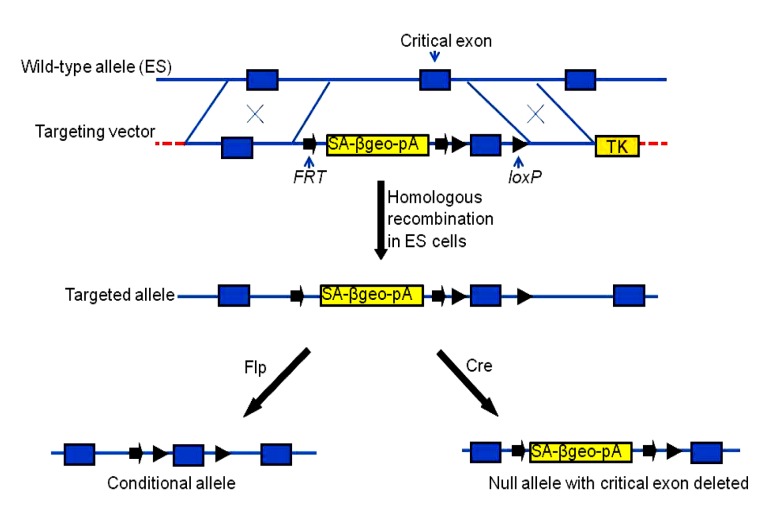
Conventional knockout, conditional knockout, and gene trapping are different methods for inducing null gene mutations in the genome. They are quite different in their manipulation. Each of these methods has advantages and disadvantages for studying gene function. Recently, a new method has been developed and used by the EUCOMM and the Knockout Mouse Project (KOMP; http://www.knockoutmouse.org). The method generates different alleles (gene trapping, conventional knockout, and conditional knockout) in one manipulation (Friedel et al., 2007; Skarnes et al., 2011). The vector for these multipurpose conditional alleles has a backbone similar to that for conditional knockout, but borrows some elements from the gene trapping vector. The selectable marker neomycin-resistance genes (neo) in the vector for conditional knockout was replaced with trapping elements that contain splice acceptor (SA), βgeo (a fusion of neo and β-gal), and a poly A signal. If the vector inserts into an intron of a gene, SA works to generate fusion transcripts of the selection marker and the exon upstream of the endogenous gene, resulting in a truncated protein from the endogenous gene. When the vector for targeted trapping is transfected into ES cells, homologous recombination occurs, resulting in an insertional mutation that resembles mutations produced through gene trapping. These ES cell lines can be used to produce knockout mice (mutant alleles by gene trapping). When gene trapping causes embryonic lethality, the ES cell lines can be treated with Flp recombinase. Flp deletes the trapping selection cassette flanked by the two FRT sites, and generates a conditional allele. The ES cell lines with this conditional allele can be used to produce conditional knockout mice. Alternatively, Cre recombinase can be transfected into the ES cells to delete critical exons, similar to conventional knockout (Fig. 9). However, this strategy appears to have two limitations: (1) in-frame expression of βgeo occurs with a probability of only one third and (2) expression of the selection marker depends on the promoter activity in ES cells, making it impossible to knock out silent genes in ES cells.
Fig. 9.
Multipurpose conditional alleles
The targeting vector contains the floxed gene (or exon) and a trapping cassette flanked by two FRT sites. After homologous recombination in ES cells, the gene is targeted. Gene expression is interrupted because of the trapping cassette. If the gene trap causes embryonic lethality, the ES cells can be transfected with Flp, and the trapping cassette is excised, resulting in a conditional allele. Alternatively, the ES cells may be transfected with Cre, and generate null alleles with the critical exon deleted as in the conventional knockout. Modified from Friedel et al. (2007)
6. Conditional gene silencing
Following the successful completion of the HGP, the goal of current biological research is to elucidate the function of every gene in the genome and find new therapeutic interventions for human diseases. Gene targeting is powerful for studying gene functions and making mouse models of human diseases. Gene knockout is used to make null mutations and perform loss-of-function studies for specific genes in mammals. However, even with the emergence of high-throughput gene knockout methods and several new resources, such as the International Knockout Mouse Consortium, the production of gene-targeted mice is still time-consuming and labor-intensive.
RNA interference strategies based on gene silencing via short interfering RNA (siRNA) molecules have been thriving in recent years. siRNAs are small, double-stranded segments of RNA that mediate the degradation of mRNAs transcribed from a certain gene (Montgomery et al., 1998). RNA interference (RNAi), a method for gene knockdown, can be used in both transgenic animals and tissue-cultured cells. An siRNA vector may be introduced into mammalian cells either by pronuclear injection in transgenic animals or by transfection in cultured cells.
By introducing the siRNA into the cells, gene expression can be blocked at the translational level (Montgomery et al., 1998). Its high efficiency in blocking expression and relatively cheap, simple process have established RNAi as an alternative method for studying gene function. However, as in conventional knockout, conventional RNAi cannot be used when the target gene is essential for cellular survival. For conditional gene silencing, the tetracycline-inducible system can be used to drive siRNA expression and knock down the target gene. Alternatively, the floxed STOP/Cre-loxP system may be used to down-regulate inducibly the target gene in certain tissues at a specific time.
7. Knocking in point mutations and making mouse models of human diseases
Thus far, knockout remains the most efficient method for studying gene functions and making animal models of human diseases. However, not all gene abnormalities are caused by the total absence of a protein. Point mutations in genes can alter certain amino acids so that normal protein functions are reduced or changed, and such mutations are very common in human diseases. In such circumstances, a simple gene knockout cannot simulate the phenotype of human disease (Cohen-Tannoudji and Babinet, 1998). To make a mouse model of a human disease caused by point mutations, a knock-in mouse line can be generated through gene targeting.
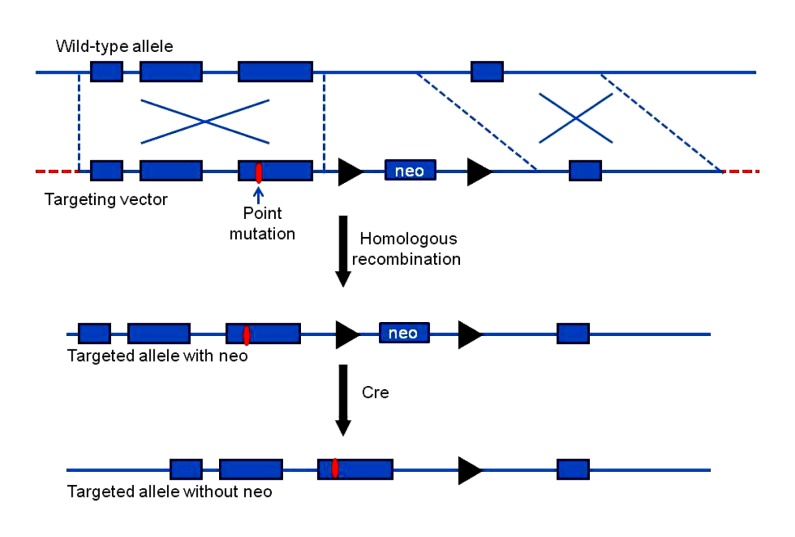
Three steps are needed for knocking a point mutation into a gene (Fig. 10). First, the point mutation is inserted into one homologous arm of the gene-targeting vector. Second, the mutation is introduced into the endogenous gene by homologous recombination in ES cells. Third, the floxed selection markers (neo flanked by two loxP sites) are removed by the Cre recombinase activity in the ES cells, or by breeding the mice produced from the targeted ES cells with another Cre-expressing mouse. In this system, Cre recombinase is used to remove the selectable marker and eliminate the potential side effects on gene function (Cohen-Tannoudji and Babinet, 1998).
Fig. 10.
Introducing point mutations into genes
Similar to gene knockout, the wild-type allele is recombined with the targeting vector with a designed point mutation and a floxed selection marker neo. The selection marker neo can be cut out by Cre, leaving the point mutation and one loxP site in the genome. The black triangles indicate loxP sites. Modified from Gao et al. (2004)
8. Chromosomal engineering
Besides loss-of-function or subtle mutations in a gene, genetic diseases can be caused by large-scale chromosomal abnormalities such as extensive deletions, insertions, translocations, or inversions. Diseases such as Down syndrome, Cri du chat syndrome, and certain cancers are caused by such chromosomal structural abnormalities. Cytogenetic imbalances exist in about 0.6% of newborn infants, 25.0% of miscarriages and stillbirths, and 50.0%–60.0% of first-trimester miscarriages (Shaffer and Lupski, 2000). Therefore, manipulating chromosomes is an important part of constructing genetic disease models.
The SSR system has made possible the generation of mouse models of human diseases associated with chromosomal rearrangements (Yu and Bradley, 2001). The system has been applied to chromosomal rearrangement because of its great potential for manipulating chromosomes: (1) Two loxP sites in the same direction cause a deletion; (2) Two loxP sites in opposite direction cause an inversion; (3) One loxP site on a chromosome and another on a circular DNA cause duplication; (4) Two loxP sites on different chromosomes cause a translocation (Ramirez-Solis et al., 1995; van Deursen et al., 1995).
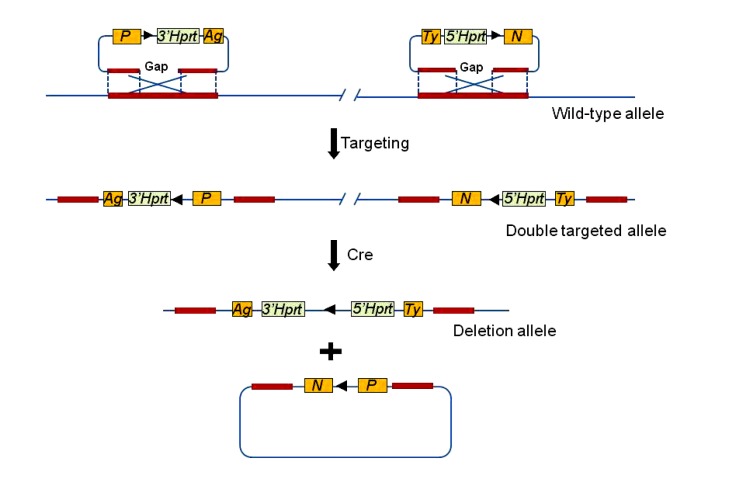
The Mutagenic Insertion and Chromosome Engineering Resource (MICER; http://www.sanger.ac.uk/resources/mouse/micer), a public resource, provides targeting vectors for insertional mutations. This type of vector can be used to create extensive deletions, insertions, translocations, or inversions on chromosomes (Adams et al., 2004). In MICER, two libraries were constructed by inserting genomic DNA fragments into two phage vectors (Fig. 11). The two vectors contain different antibiotic genes that can be used to screen out positive integration in mouse ES cells. Each vector can be used for one targeted integration in ES cells and this integration introduces a loxP site into the genome. The two introduced loxP sites can be in the same or in opposite orientations, and may be either on the same or on different chromosomes. By expressing Cre in the targeted ES cells with the two loxP sites, chromosomal rearrangements will occur. Mice can be generated using these chromosome-engineered ES cells. MICER lists information on the vector sequences and comprises more than 93 000 constructed, insertional targeting vectors from two libraries (5′Hprt and 3′Hprt). As well as using the Hprt marker, a new chromosome manipulating vector using a pair of fluorescent proteins has been developed (Uemura et al., 2010).
Fig. 11.
Generation of large-scope deletions using the two vectors in the MICER
The 5′Hprt and 3′Hprt vectors are used in two targeting steps to introduce the two loxP sites into the mouse genome. The 5′Hprt and 3′Hprt fragments are kept in the same orientation if a large-scale deletion is to be performed. The DNA between the two loxP sites is deleted by the excision of the Cre recombinase in the ES cells. After the excision, the 5′Hprt and 3′Hprt will form an intact Hprt gene, and positive ES cell clones can be selected by hypoxanthine, aminopterin, and thymidine (HAT). P: puromycin resistance gene; Ag: K14-Agouti transgene; N: neomycin resistance gene; Ty: tyrosinase minigene. The black triangles indicate loxP sites. Modified from Yu and Bradley (2001) and Klysik (2002)
9. Guidelines for functional study of genes
After researchers find a candidate gene for functional analysis, they usually begin by searching the ES cell lines in which the candidate gene has been trapped. Important trapping databases include the International Gene Trap Consortium (IGTC; http://www.genetrap.org), EUCOMM (http://www.eucomm.org), and the North America Conditional Mouse Mutagenesis Project (NorCOMM; http://www.norcomm.org). If no gene-trapped ES cell line is available, researchers may start constructing targeting vectors to target the gene in the ES cells. The conventional method is preferred because the mice with a targeted gene do not have to be crossed with a Cre-expression mouse, saving time for the research.
If the knockout mutation happens to be lethal, they have to use a conditional strategy. Researchers can still choose between gene targeting and trapping. Conditional gene targeting is a good choice because it features precision and a better chance of a true null allele. However, it has the disadvantage of a complicated procedure, and is time-consuming. Considerable time and effort could be saved if researchers can find the ES cell lines with the gene trapped with the conditional trapping cassette (EUCOMM). However, not every gene has been trapped conditionally despite the fast expansion of conditional trapping databases.
Researchers may choose the multipurpose targeted trapping if conventional gene knockout is likely to be lethal, the trapping ES cell is not available, or the risk of time-consuming conditional gene targeting is not acceptable. The targeted trapping ES cells can be used to produce mice directly as in conventional gene targeting. If the mice are born alive without a null allele, researchers may proceed with knocking out the targeted exon to ensure the complete loss of the gene. If the mice die during the embryonic stage, the targeted trapping ES cells may be transfected with Flp to change the allele into a conditional one. The mice produced with this conditional allele may be crossed with another Cre line to remove the targeted exon in some tissues or at certain stages. Another choice is gene silencing, which is fast for vector construction and mouse breeding. However, RNAi cannot silence a gene completely, and can knock down the expression of the targeted gene only to a certain extent.
10. Prospects
The HGP, one of the greatest accomplishments in the past century, has provided us with the sequence information of our genome. Presently, we are in a new stage of biological research, the post-genomic era. The main work in this era is to elucidate the function of each gene in our genome, and find new therapeutic interventions for human diseases. Comparative genomics has shown high homology between the mouse and human genomes. Hence, mice serve as ideal models for functional research of human genes. Currently, gene knockout is the most efficient way of studying gene functions. Information about the function of each gene is obtained by observing the phenotype of the whole animal. Knockout mice also serve as animal models of human diseases. We may use these models to study the mechanisms of the diseases and screen drugs for treatment.
Gene targeting and gene trapping are the two main methods for producing knockout mice. The first knockout mouse ES cells made by gene targeting were produced in 1987 (Doetschman et al., 1987; Thomas and Capecchi, 1987). In the last 20 years, different laboratories worldwide have generated a large number of knockout mice, which serve as crucial resources for biological research. Gene trapping is an alternative way to produce knockout mice. It is a random mutation technique (Kothary et al., 1988; Gossler et al., 1989; Abuin et al., 2007) and therefore not as specific as gene targeting. However, it has a high throughput and a large number of genes may be knocked out in a short period of time.
In 2003, the KOMP was proposed at a meeting in Banbury, Cold Spring Harbor, USA. Scientists discussed objectives following the completion of the HGP (Check, 2002; Marshall, 2002), and agreed unanimously to focus on discovering the functions of all mammalian genes. Thus, a genome-wide knockout mouse project is as important as the HGP (Austin et al., 2004). With the fast expansion of gene targeting and trapping programs (Guan et al., 2010), genetic functional studies based on gene targeting and trapping will reach saturation; i.e., most genes within our knowledge will be targeted or trapped, and their function analyzed. However, this saturation does not complete this endeavor, and much work has to be done after all genes in the mammalian genome have been knocked out. SSR systems will play critical roles in various conditional gene manipulations in the biological research after saturation is reached.
In the beginning, SSR systems were used for conditional gene knockout; now, they have more applications. Based on SSR systems, a method was developed for conditional gene silencing in mice. Conditional gene silencing can be used not only for knocking down gene expression but also for gene therapy. In addition to traditional gene therapy, conditional gene silencing has potential in treating cancers and viral infections (Kappel et al., 2006; Lee and Kumar, 2009). Using SSR systems, we may introduce point mutations into genes or engineer chromosomes so that mouse models for most human genetic diseases can be generated. SSR systems have other applications as well. For example, researchers may conveniently analyze gene interactions using inducible systems. By introducing different drug cocktails to an organism in which different genes are targeted by different systems, researchers can evaluate the interaction of the two genes with various drug formulations. For example, we may produce a mouse line in which gene A is controlled by the Tet-On system and gene B by the tamoxifen system. When either tetracycline or tamoxifen is used alone, the mouse does not exhibit a certain phenotype, whereas when used together, the mouse shows an obvious deficiency. Therefore, gene A and gene B work cooperatively, or have similar functions. In another case, when either tetracycline or tamoxifen is used alone, the mouse shows the same deficiency, genes A and B may function at different steps in the same catalytic pathway. Compared with the Cre-loxP and Flp-FRT systems, the ΦC31 integrase system has several unique applications. It is a powerful tool for site-directed insertion and shows potential for future use.
The SSR system was developed originally to avoid the problem of embryonic lethality in gene knockouts. Currently, the applications of this technology are expanding beyond this area, and are leading us to a “Cre-ative” era, in which the genes in the genome can be manipulated as desired.
Footnotes
Project supported by the National Natural Science Foundation of China (Nos. 30871436, 30973297, and 31171194), the National Basic Research Program (973) of China (No. 2010CB945002), the Ministry of Education of China (No. 200804220011), and Shandong Provincial Science and Technology Key Program (No. 2009GG10003039), China
References
- 1.Abramowicz M. The Human Genome Project in retrospect. Adv Genet. 2003;50:231–261. doi: 10.1016/S0065-2660(03)50011-6. [DOI] [PubMed] [Google Scholar]
- 2.Abuin A, Hansen G, Zambrowicz B. Gene trap mutagenesis. Handb Exp Pharmacol. 2007;178(1):129–147. doi: 10.1007/978-3-540-35109-2_6. [DOI] [PubMed] [Google Scholar]
- 3.Adams DJ, Biggs PJ, Cox T, Davies R, van der Weyden L, Jonkers J, Smith J, Plumb B, Taylor R, Nishijima I, et al. Mutagenic insertion and chromosome engineering resource (MICER) Nat Genet. 2004;36(8):867–871. doi: 10.1038/ng1388. [DOI] [PubMed] [Google Scholar]
- 4.Albert H, Dale EC, Lee E, Ow DW. Site-specific integration of DNA into wild-type and mutant lox sites placed in the plant genome. Plant J. 1995;7(4):649–659. doi: 10.1046/j.1365-313X.1995.7040649.x. [DOI] [PubMed] [Google Scholar]
- 5.Austin CP, Battey JF, Bradley A, Bucan M, Capecchi M, Collins FS, Dove WF, Duyk G, Dymecki S, Eppig JT, et al. The knockout mouse project. Nat Genet. 2004;36(9):921–924. doi: 10.1038/ng0904-921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Braselmann S, Graninger P, Busslinger M. A selective transcriptional induction system for mammalian cells based on Gal4-estrogen receptor fusion proteins. PNAS. 1993;90(5):1657–1661. doi: 10.1073/pnas.90.5.1657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Calos MP. The ΦC31 integrase system for gene therapy. Curr Gene Ther. 2006;6(6):633–645. doi: 10.2174/156652306779010642. [DOI] [PubMed] [Google Scholar]
- 8.Casanova E, Fehsenfeld S, Lemberger T, Shimshek DR, Sprengel R, Mantamadiotis T. ER-based double iCre fusion protein allows partial recombination in forebrain. Genesis. 2002;34(3):208–214. doi: 10.1002/gene.10153. [DOI] [PubMed] [Google Scholar]
- 9.Check E. Draft mouse genome makes public debut. Nature. 2002;417(6885):106. doi: 10.1038/417106a. [DOI] [PubMed] [Google Scholar]
- 10.Chow LML, Tian Y, Weber T, Corbett M, Zuo J, Baker SJ. Inducible Cre recombinase activity in mouse cerebellar granule cell precursors and inner ear hair cells. Dev Dyn. 2006;235(11):2991–2998. doi: 10.1002/dvdy.20948. [DOI] [PubMed] [Google Scholar]
- 11.Cohen-Tannoudji M, Babinet C. Beyond ‘knock-out’ mice: new perspectives for the programmed modification of the mammalian genome. Mol Hum Reprod. 1998;4(10):929–938. doi: 10.1093/molehr/4.10.929. [DOI] [PubMed] [Google Scholar]
- 12.Doetschman T, Gregg RG, Maeda N, Hooper ML, Melton DW, Thompson S, Smithies O. Targetted correction of a mutant HPRT gene in mouse embryonic stem cells. Nature. 1987;330(6148):576–578. doi: 10.1038/330576a0. [DOI] [PubMed] [Google Scholar]
- 13.Dragatsis I, Zeitlin S. A method for the generation of conditional gene repair mutations in mice. Nucleic Acids Res. 2001;29(3):E10. doi: 10.1093/nar/29.3.e10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Feil R, Brocard J, Mascrez B, LeMeur M, Metzger D, Chambon P. Ligand-activated site-specific recombination in mice. PNAS. 1996;93(20):10887–10890. doi: 10.1073/pnas.93.20.10887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Feil R, Wagner J, Metzger D, Chambon P. Regulation of Cre recombinase activity by mutated estrogen receptor ligand-binding domains. Biochem Biophys Res Commun. 1997;237(3):752–757. doi: 10.1006/bbrc.1997.7124. [DOI] [PubMed] [Google Scholar]
- 16.Friedel RH, Seisenberger C, Kaloff C, Wurst W. EUCOMM—the European conditional mouse mutagenesis program. Brief Funct Genomic Proteomic. 2007;6(3):180–185. doi: 10.1093/bfgp/elm022. [DOI] [PubMed] [Google Scholar]
- 17.Gao J, Wu X, Zuo J. Targeting hearing genes in mice. Mol Brain Res. 2004;132(2):192–207. doi: 10.1016/j.molbrainres.2004.06.035. [DOI] [PubMed] [Google Scholar]
- 18.Gossen M, Bujard H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. PNAS. 1992;89(12):5547–5551. doi: 10.1073/pnas.89.12.5547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Gossen M, Freundlieb S, Bender G, Muller G, Hillen W, Bujard H. Transcriptional activation by tetracyclines in mammalian cells. Science. 1995;268(5218):1766–1769. doi: 10.1126/science.7792603. [DOI] [PubMed] [Google Scholar]
- 20.Gossler A, Joyner AL, Rossant J, Skarnes WC. Mouse embryonic stem cells and reporter constructs to detect developmentally regulated genes. Science. 1989;244(4903):463–465. doi: 10.1126/science.2497519. [DOI] [PubMed] [Google Scholar]
- 21.Groth AC, Olivares EC, Thyagarajan B, Calos MP. A phage integrase directs efficient site-specific integration in human cells. PNAS. 2000;97(11):5995–6000. doi: 10.1073/pnas.090527097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Guan C, Ye C, Yang X, Gao J. A review of current large-scale mouse knockout efforts. Genesis. 2010;48(2):73–85. doi: 10.1002/dvg.20594. [DOI] [PubMed] [Google Scholar]
- 23.Hoess RH, Ziese M, Sternberg N. P1 site-specific recombination: nucleotide sequence of the recombining sites. PNAS. 1982;79(11):3398–3402. doi: 10.1073/pnas.79.11.3398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hoess R, Abremski K, Sternberg N. The nature of the interaction of the P1 recombinase Cre with the recombining site loxP . Cold Spring Harb Symp Quant Biol. 1984;49:761–768. doi: 10.1101/SQB.1984.049.01.086. [DOI] [PubMed] [Google Scholar]
- 25.Huang LC, Wood EA, Cox MM. A bacterial model system for chromosomal targeting. Nucleic Acids Res. 1991;19(3):443–448. doi: 10.1093/nar/19.3.443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Indra AK, Warot X, Brocard J, Bornert JM, Xiao JH, Chambon P, Metzger D. Temporally-controlled site-specific mutagenesis in the basal layer of the epidermis: comparison of the recombinase activity of the tamoxifen-inducible Cre-ERT and Cre-ERT2 recombinases. Nucleic Acids Res. 1999;27(22):4324–4327. doi: 10.1093/nar/27.22.4324. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ishida Y, Leder P. RET: a poly A-trap retrovirus vector for reversible disruption and expression monitoring of genes in living cells. Nucleic Acids Res. 1999;27(24):e35–e42. doi: 10.1093/nar/27.24.e35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kappel S, Matthess Y, Zimmer B, Kaufmann M, Strebhardt K. Tumor inhibition by genomically integrated inducible RNAi-cassettes. Nucleic Acids Res. 2006;34(16):4527–4536. doi: 10.1093/nar/gkl628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kistner A, Gossen M, Zimmermann F, Jerecic J, Ullmer C, Lübbert H, Bujard H. Doxycycline-mediated quantitative and tissue-specific control of gene expression in transgenic mice. PNAS. 1996;93(20):10933–10938. doi: 10.1073/pnas.93.20.10933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Klysik J. Mice and humans: chromosome engineering and its application to functional genomics. Acta Biochim Pol. 2002;49(3):553–569. [PubMed] [Google Scholar]
- 31.Kothary R, Clapoff S, Brown A, Campbell R, Peterson A, Rossant J. A transgene containing lacZ inserted into the dystonia locus is expressed in neural tube. Nature. 1988;335(6189):435–437. doi: 10.1038/335435a0. [DOI] [PubMed] [Google Scholar]
- 32.Lee SK, Kumar P. Conditional RNAi: towards a silent gene therapy. Adv Drug Deliv Rev. 2009;61(7-8):650–664. doi: 10.1016/j.addr.2009.03.016. [DOI] [PubMed] [Google Scholar]
- 33.Lister JA. Transgene excision in zebrafish using the phiC31 integrase. Genesis. 2010;48(2):137–143. doi: 10.1002/dvg.20613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Logie C, Stewart AF. Ligand-regulated site-specific recombination. PNAS. 1995;92(13):5940–5944. doi: 10.1073/pnas.92.13.5940. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lu J, Maddison LA, Chen W. PhiC31 integrase induces efficient site-specific excision in zebrafish. Transgenic Res. 2011;20(1):183–189. doi: 10.1007/s11248-010-9394-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ma QW, Sheng HQ, Yan JB, Cheng S, Huang Y, Chen-Tsai Y, Ren ZR, Huang SZ, Zeng YT. Identification of pseudo attP sites for phage ΦC31 integrase in bovine genome. Biochem Biophys Res Commun. 2006;345(3):984–988. doi: 10.1016/j.bbrc.2006.04.145. [DOI] [PubMed] [Google Scholar]
- 37.Marshall E. Genome sequencing. Public group completes draft of the mouse. Science. 2002;296(5570):1005. doi: 10.1126/science.296.5570.1005b. [DOI] [PubMed] [Google Scholar]
- 38.Matsuda T, Cepko CL. Controlled expression of transgenes introduced by in vivo electroporation. PNAS. 2007;104(3):1027–1032. doi: 10.1073/pnas.0610155104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.McLeod M, Craft S, Broach JR. Identification of the crossover site during FLP-mediated recombination in the Saccharomyces cerevisiae plasmid 2 microns circle. Mol Cell Biol. 1986;6(10):3357–3367. doi: 10.1128/mcb.6.10.3357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Metzger D, Clifford J, Chiba H, Chambon P. Conditional site-specific recombination in mammalian cells using a ligand-dependent chimeric Cre recombinase. PNAS. 1995;92(15):6991–6995. doi: 10.1073/pnas.92.15.6991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Montgomery MK, Xu S, Fire A. RNA as a target of double-stranded RNA-mediated genetic interference in Caenorhabditis elegans . PNAS. 1998;95(26):15502–15507. doi: 10.1073/pnas.95.26.15502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Morris AC, Schaub TL, James AA. FLP-mediated recombination in the vector mosquito, Aedes aegypti . Nucleic Acids Res. 1991;19(21):5895–5900. doi: 10.1093/nar/19.21.5895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Nagy A. Cre recombinase: the universal reagent for genome tailoring. Genesis. 2000;26(2):99–109. doi: 10.1002/(SICI)1526-968X(200002)26:2<99::AID-GENE1>3.3.CO;2-2. [DOI] [PubMed] [Google Scholar]
- 44.Nagy A, Mar L, Watts G. Creation and use of a Cre recombinase transgenic database. Methods Mol Biol. 2009;530:365–378. doi: 10.1007/978-1-59745-471-1_19. [DOI] [PubMed] [Google Scholar]
- 45.No D, Yao TP, Evans RM. Ecdysone-inducible gene expression in mammalian cells and transgenic mice. PNAS. 1996;93(8):3346–3351. doi: 10.1073/pnas.93.8.3346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Orban PC, Chui D, Marth JD. Tissue- and site-specific DNA recombination in transgenic mice. PNAS. 1992;89(15):6861–6865. doi: 10.1073/pnas.89.15.6861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Picard D. Regulation of protein function through expression of chimaeric proteins. Curr Opin Biotechnol. 1994;5(5):511–515. doi: 10.1016/0958-1669(94)90066-3. [DOI] [PubMed] [Google Scholar]
- 48.Ramirez-Solis R, Liu P, Bradley A. Chromosome engineering in mice. Nature. 1995;378(6558):720–724. doi: 10.1038/378720a0. [DOI] [PubMed] [Google Scholar]
- 49.Schnütgen F. Generation of multipurpose alleles for the functional analysis of the mouse genome. Brief Funct Genomic Proteomic. 2006;5(1):15–18. doi: 10.1093/bfgp/ell009. [DOI] [PubMed] [Google Scholar]
- 50.Senecoff JF, Rossmeissl PJ, Cox MM. DNA recognition by the FLP recombinase of the yeast 2 mu plasmid: a mutational analysis of the FLP binding site. J Mol Biol. 1988;201(2):405–421. doi: 10.1016/0022-2836(88)90147-7. [DOI] [PubMed] [Google Scholar]
- 51.Shaffer LG, Lupski JR. Molecular mechanisms for constitutional chromosomal rearrangements in humans. Annu Rev Genet. 2000;34:297–329. doi: 10.1146/annurev.genet.34.1.297. [DOI] [PubMed] [Google Scholar]
- 52.Sharma N, Moldt B, Dalsgaard T, Jensen TG, Mikkelsen JG. Regulated gene insertion by steroid-induced ΦC31 integrase. Nucleic Acids Res. 2008;36(11):e67. doi: 10.1093/nar/gkn298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Shigeoka T, Kawaichi M, Ishida Y. Suppression of nonsense-mediated mRNA decay permits unbiased gene trapping in mouse embryonic stem cells. Nucleic Acids Res. 2005;33(2):e20. doi: 10.1093/nar/gni022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Skarnes WC, Rosen B, West AP, Koutsourakis M, Bushell W, Iyer V, Mujica AO, Thomas M, Harrow J, Cox T, et al. A conditional knockout resource for the genome-wide study of mouse gene function. Nature. 2011;474(7351):337–342. doi: 10.1038/nature10163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Smith MC, Thorpe HM. Diversity in the serine recombinases. Mol Microbiol. 2002;44(2):299–307. doi: 10.1046/j.1365-2958.2002.02891.x. [DOI] [PubMed] [Google Scholar]
- 56.Stark WM, Boocock MR, Sherratt DJ. Catalysis by site-specific recombinases. Trends Genet. 1992;8(12):432–439. doi: 10.1016/0168-9525(92)90327-Z. [DOI] [PubMed] [Google Scholar]
- 57.Sternberg N, Hamilton D, Austin S, Yarmolinsky M, Hoess R. Site-specific recombination and its role in the life cycle of bacteriophage P1. Cold Spring Harb Symp Quant Biol. 1981;45:297–309. doi: 10.1101/SQB.1981.045.01.042. [DOI] [PubMed] [Google Scholar]
- 58.Thomas KR, Capecchi MR. Site-directed mutagenesis by gene targeting in mouse embryo-derived stem cells. Cell. 1987;51(3):503–512. doi: 10.1016/0092-8674(87)90646-5. [DOI] [PubMed] [Google Scholar]
- 59.Thorpe HM, Smith MC. In vitro site-specific integration of bacteriophage DNA catalyzed by a recombinase of the resolvase/invertase family. PNAS. 1998;95(10):5505–5510. doi: 10.1073/pnas.95.10.5505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Thorpe HM, Wilson SE, Smith MC. Control of directionality in the site-specific recombination system of the Streptomyces phage ΦC31. Mol Microbiol. 2000;38(2):232–241. doi: 10.1046/j.1365-2958.2000.02142.x. [DOI] [PubMed] [Google Scholar]
- 61.Thyagarajan B, Olivares EC, Hollis RP, Ginsburg DS, Calos MP. Site-specific genomic integration in mammalian cells mediated by phage ΦC31 integrase. Mol Cell Biol. 2001;21(12):3926–3934. doi: 10.1128/MCB.21.12.3926-3934.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Tian Y, James S, Zuo J, Fritzsch B, Beisel KW. Conditional and inducible gene recombineering in the mouse inner ear. Brain Res. 2006;1091(1):243–254. doi: 10.1016/j.brainres.2006.01.040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Uemura M, Niwa Y, Kakazu N, Adachi N, Kinoshita K. Chromosomal manipulation by site-specific recombinases and fluorescent protein-based vectors. PLoS One. 2010;5(3):e9846. doi: 10.1371/journal.pone.0009846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.van Deursen J, Fornerod M, van Rees B, Grosveld G. Cre-mediated site-specific translocation between nonhomologous mouse chromosomes. PNAS. 1995;92(16):7376–7380. doi: 10.1073/pnas.92.16.7376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Wang Y, Tsai SY, O′Malley BW. Antiprogestin regulable gene switch for induction of gene expression in vivo. Methods Enzymol. 1999;306:281–294. doi: 10.1016/S0076-6879(99)06018-8. [DOI] [PubMed] [Google Scholar]
- 66.Watson JD, Cook-Deegan RM. Origins of the Human Genome Project. FASEB J. 1991;5(1):8–11. doi: 10.1096/fasebj.5.1.1991595. [DOI] [PubMed] [Google Scholar]
- 67.Yao TP, Forman BM, Jiang Z, Cherbas L, Chen JD, McKeown M, Cherbas P, Evans RM. Functional ecdysone receptor is the product of EcR and Ultraspiracle genes. Nature. 1993;366(6454):476–479. doi: 10.1038/366476a0. [DOI] [PubMed] [Google Scholar]
- 68.Yu Y, Bradley A. Engineering chromosomal rearrangements in mice. Nat Rev Genet. 2001;2(10):780–790. doi: 10.1038/35093564. [DOI] [PubMed] [Google Scholar]
- 69.Zhang Y, Riesterer C, Ayrall AM, Sablitzky F, Littlewood TD, Reth M. Inducible site-directed recombination in mouse embryonic stem cells. Nucleic Acids Res. 1996;24(4):543–548. doi: 10.1093/nar/24.4.543. [DOI] [PMC free article] [PubMed] [Google Scholar]