Abstract
A new thiol blocking reagent-methylsulfonyl benzothiazole was discovered. This reagent showed good selectivity and high reactivity for protein thiols.
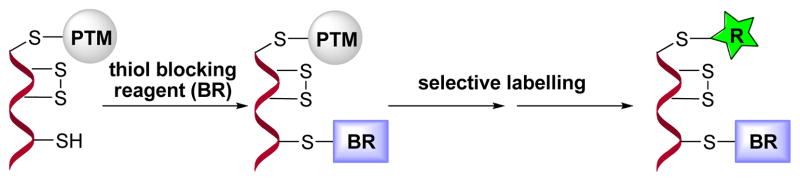
Protein cysteine residues are targets of numerous post-translational modifications (PTM) that are essential to maintain cell redox homeostasis as well as signalling.1 Modifications at cysteine residues are caused by their interaction with reactive oxygen/nitrogen species (ROS/RNS) in response to cellular oxidative damage.1a, 2 Most of these modifications, that comprise the thiol proteome, are reversible and encompass a range of functional groups with very distinctive chemistry including mixed disulfides (RS-SR′, SR′ low molecular weight thiols), nitrosothiols (RS-NO), sulfenic acids (RSOH), sulfinic acids (SO2H), sulfonic acids (SO3H), S-lipidation (palmytoylation, RS-COR) and perthiols (RS-SH). This diversity of functionality has possessed some difficulty in selectively determining each modification.3 Nevertheless many advances have been made in this field, in particular employing chemical methods to detect specific thiol modifications.3, 4 In these methods, a common step involves selective blocking of unmodified thiols (reduced thiols) (Scheme 1).
Scheme 1.
Chemical approach to study protein posttranslational modifications (PTM) at Cys.
To a great extent, the efficiency of these assays relies on the efficiency of the thiol blocking step. Many research efforts have been made to identify reagents that enable blocking or labelling of protein thiols with high selectivity and conversion yields.5 Among those, thiol-alkylation reagents such as iodoacetamides (IAM) and N-substituted maleimides (NSM) are by far the most commonly used and their reactivity profiles have been extensively studied. 6 It is known that under certain conditions, IAM and NSM can modify other reactive amino acids (e.g. Lys and His).7 As a consequence, it has been suggested that the selection of the thiol blocking reagent should not be arbitrary. Due to the disparate reactivity of various thiols influenced by their localization within the protein and physiological environment, one must consider the unique property of target protein and necessary experimental conditions to select proper thiol blocking agents.8
With all abovementioned, the development of new thiol blocking reagents that possess distinctive reactivity profiles from currently known compounds is needed. Our consideration in this subject was to explore molecules that could react with free thiols via nucleophilic aromatic substitution (NAS). We developed this idea from previous work in our laboratory that studied the reactions of 2-mercapto benzothiazole (2-SHBT) towards sulfenamides and alkyl disulfides.9 The results revealed that 2-SHBT was inert to disulfide exchange reactions, but showed significant reactivity against more reactive electrophiles, i.e. sulfenamides. In contrast, other aromatic thiols such as thiophenol, 2-mercapto pyridine, and 2-mercapto pyrimidine, were found reactive to both alkyl disulfides and sulfenamides. It should be noted that the disulfide exchange is a dynamic equilibrium and thus the progress of the reaction is controlled by both the electrophilicity/nucleophilicity of the starting disulfide/thiol pair as well as those generated. These results suggest that the electron withdrawing effect of benzothiazole ring decreases the reactivity of corresponding thiol and inhibits disulfide formation. We envisioned that by placing a leaving group at C-2 position, benzothiazole might be vulnerable for nucleophilic attack by thiols via NAS mechanism. With this idea in mind, we designed a series of experiments to examine whether benzothiazole moiety could be employed as an electrophilic trap for thiols and Cys residues. Here we report our results.
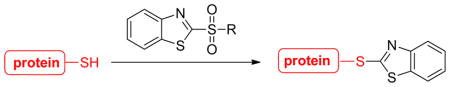
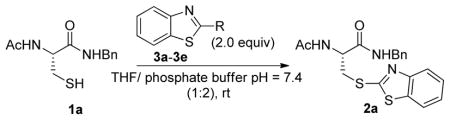
We first examined the reactivity of various benzothiazole substrates containing different leaving groups at C-2 position (Table 1). A cysteine derivative 1a was used as thiol model. In a typical experimental setting, to a solution of 1a in 1:2 THF/phosphate buffer (200 mM, pH=7.4) was added 2 equivalents of benzothiazole substrate respectively. The reaction was monitored by TLC. We tested several commercially available 2-halogenated benzothiazoles (3a–3c). These compounds are known to react with thiol at high temperature and under strong basic conditions.10 However, under mild and biologically mimic conditions, these substrates displayed very poor reactivity and only trace amount of the desired product 2a was formed. The reaction using 2-diazo substrate 3d resulted in a complicated mixture of products and only a small amount of 2a was produced (judging by TLC and crude NMR). Interestingly, 2-methylsulfonyl benzothiazole (MSBT) showed very high reactivity towards 1a, with almost quantitative formation of 2a within 20 min. To the best of our knowledge, this was the first example illustrating excellent reactivity of MSBT towards alkylthiols in aqueous solutions.
Table 1.
Reactions of benzothiazole substrates with model substrate 1a.
| |||
|---|---|---|---|
| entry | R | time | 2a % yield |
| 1 |
3a |
4 h | trace |
| 2 |
3b |
4 h | trace |
| 3 |
3c |
4 h | trace |
| 4 |
 3d |
20 min | < 20 |
| 5 |
 3e |
20 min | 95 |
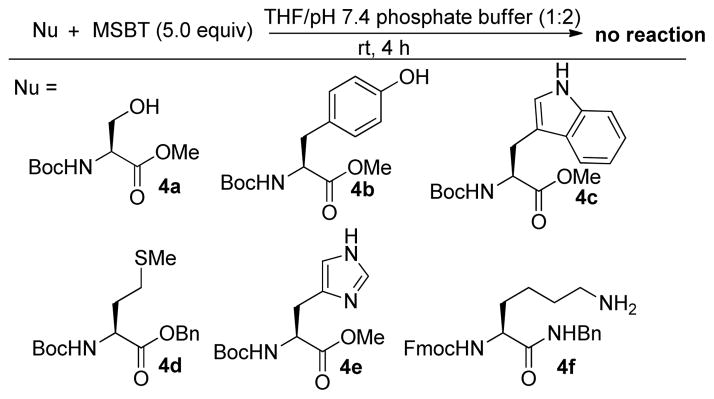
We next investigated whether the pronounced reactivity of MSBT 3e towards thiols could also occur with other potential nucleophilic species found in proteins. A series of amino acids derivatives (4a–4f) were then tested under the same conditions. As shown in Scheme 2, side chain functionalities of serine, tyrosine, tryptophan, and methionine are inert to MSBT (5 equiv.). In addition, lysine and histidine substrates (4e and 4f) did not react with MSBT to form any product (monitored by TLC), even after 4 hr. These outcomes were expectable as known reactions of MSBT with amines and alcohols require high temperature and/or strong basic media.11 Nevertheless, these results suggested that MSBT is a thiol-selective blocking reagent.
Scheme 2.
Control experiments
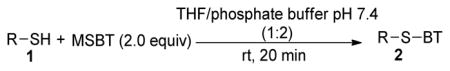
In order to explore the generality of MSBT mediated thiol-blocking reaction, a series of cysteine derivatives (1a–1g) were tested. As shown in Table 2, in all the cases the reaction went smoothly and the desired products were obtained in high yields. We did not observe any byproducts in these reactions.
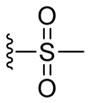
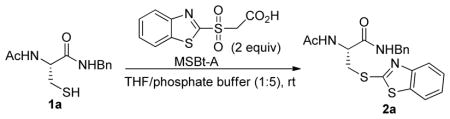
To further expand our understanding of the reactivity profile of MSBT, we studied the effect of pH on the reaction. This was driven by the fact that pKa values of protein thiols are variable within distinctive protein domains and pH fluctuates in different cell compartments. 12 Because the reactivity of many electrophiles towards SH depends on the concentration of thiolate, disruption in the pH will affect the equilibrium between thiol (RSH) and thiolate (RS−) and therefore change thiol blocking efficiency.13 To study this problem, we prepared a water soluble MSBT reagent (MSBT-A). This compound allowed us to study pH effects in high aqueous buffer containing systems. The results are summarized in Table 3. In acidic media (pH=6.2), MSBT-A reacted with 1a slowly and only a small amount of product was obtained after 20 min. At pH 7.0 or 7.4, the reaction went well and resulted the blocking product in good yield in 20 min. When pH was 9.0, the reactivity of 1a was greatly enhanced and the reaction was completed in a few minutes. Interestingly, when this reaction was performed in pure organic solvents such as pure THF, we did not observe the formation of 2a even after 1 h. With these results we concluded that the reactions between MSBT substrates and RSH largely rely on the thiolate concentration. Similar reactivity profiles were observed with both IAM and NSM derivatives. 13, 14
Table 3.
pH dependence of MSBT reaction
| ||
|---|---|---|
| pH | time | % yield |
| 6.2 | 20 min | 34 |
| 7.0 | 20 min | 87 |
| 7.4 | 20 min | 95 |
| 9.0 | 5 min | 95 |
We also tested the stability of the thiol-blocking adducts (RS-Bt) under common conditions used in protein labelling experiments. For example, tris-(2-carboxyethyl)phosphine (TCEP) or dithiothreitol (DTT) are often used for protein thiol reduction or quenching excess of thiol-blocking reagent. We found that RS-Bt adducts did not show any reaction or decomposition in the presence of excess of TCEP or DTT (see supporting information for details).
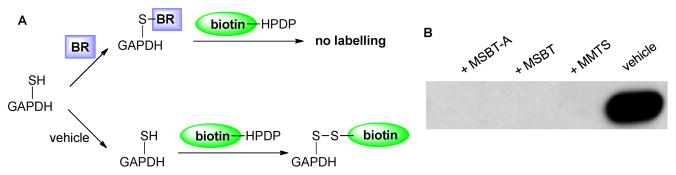
Finally we tested the capability of MSBT and MSBT-A in blocking protein thiol residues. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH), whose biological function has been shown to be mediated by its cysteine thiol modifications,15 was used as the model. Briefly (Fig 1A), reduced GADPH was treated with vehicle, MSBT, MSBT-A, or a common thiol blocking reagent methyl methanethiosulfonate (MMTS)16 and then excess reagents were removed by desalting. The protein sample was then exposed to a thiol labeling reagent N-[6-(biotinamido)hexyl]-3′-(2′-pyridyldithio)propionamide (biotin-HPDP). Biotin labelled GAPDH was deteced by non-reducing SDS-PAGE followed by western blot using anti-biotin antibody. As shown in Fig 1B, MSBT and MSBT-A pronounced excellent thiol blocking activity as MMTS. This result confirmed the efficiency of MSBT substrates in thiol specific blocking.
Figure 1.
Thiol blocking capability of MSBT and MSBT-A on GAPDH compared to MMTS. A) Schematic representation of the assay using blocking reagents (MSBT, MSBT-A, MMTS) and vehicle. B) Western blot results.
In summary, we have discovered a new reagent, MSBT, capable of blocking protein thiols selectively and effectively. As observed in other thiol blocking reagents, the reactivity profile of MSBT as a function of pH suggests that the rate of reaction depends on thiolate concentration. We expect MSBT substrates will find applications in protein chemistry.
Supplementary Material
Table 2.
Reactions of MSBT with RSH substrates
| ||
|---|---|---|
| entry | R-SH | % yield |
| 1 |
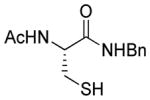 1a |
>95 |
| 2 |
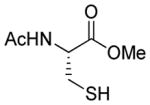 1b |
>95 |
| 3 |
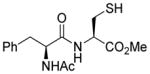 1c |
>95 |
| 4 |
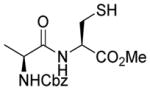 1d |
>95 |
| 5 |
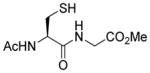 1e |
85 |
| 6 |
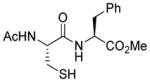 1f |
86 |
| 7 |
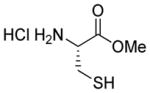 1g |
97 |
Acknowledgments
This work is supported in part by NIH (R01GM088226 to MX and R01CA131653 to JRL) and Burroughs Wellcome Fund (Collaborative Research Travel Grant to MX).
Footnotes
Supporting Information Available: Spectroscopic and analytical data and selected experimental procedures. This material is available free of charge via the Internet at http://pubs.acs.org
References
- 1.(a) Forrester MT, Stamler JS. Am J Respir Cell Mol Biol. 2007;36:135. doi: 10.1165/rcmb.2006-001ED. [DOI] [PubMed] [Google Scholar]; (b) Jones DP, Go YM. Curr Opinion Chem Biol. 2011;15:103. doi: 10.1016/j.cbpa.2010.12.014. [DOI] [PubMed] [Google Scholar]; (c) Jacob C, Giles GI, Giles NM, Sies H. Angew Chem Int Ed. 2003;42:4742. doi: 10.1002/anie.200300573. [DOI] [PubMed] [Google Scholar]
- 2.Selected reviews: Thamsen M, Jakob U. Curr Opinion Chem Biol. 2011;15:113. doi: 10.1016/j.cbpa.2010.11.013.Riederer BM. Curr Proteomics. 2009;6:51.
- 3.Chouchani ET, James AM, Fearnley IM, Lilley KS, Murphy MP. Curr Opinion Chem Biol. 2011;15:120. doi: 10.1016/j.cbpa.2010.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.For recent reviews: Leonard SE, Carroll KS. Curr Opinion Chem Biol. 2011;15:88. doi: 10.1016/j.cbpa.2010.11.012.Wang H, Xian M. Curr Opinion Chem Biol. 2011;15:32. doi: 10.1016/j.cbpa.2010.10.006.
- 5.(a) Hansen RE, Winther JR. Anal Biochem. 2009;394:147. doi: 10.1016/j.ab.2009.07.051. [DOI] [PubMed] [Google Scholar]; (b) Weerapana E, Simon GM, Cravatt BF. Nat Chem Biol. 2008;4:405. doi: 10.1038/nchembio.91. [DOI] [PMC free article] [PubMed] [Google Scholar]; (c) Wright SK, Viola RE. Anal Biochem. 1998;265:8. doi: 10.1006/abio.1998.2858. [DOI] [PubMed] [Google Scholar]; (d) Riener CK, Kada G, Gruber HJ. Anal Bioanal Chem. 2002;373:266. doi: 10.1007/s00216-002-1347-2. [DOI] [PubMed] [Google Scholar]; (e) Roberts DD, Lewis SD, Ballou DP, Olson ST, Shafer JA. Biochemisty. 1986;25:5595. doi: 10.1021/bi00367a038. [DOI] [PubMed] [Google Scholar]; (f) Kim Y, Ho SO, Gassman NR, Korlann Y, Landorf EV, Collart FR, Weiss S. Bioconjugate Chem. 2008;19:786. doi: 10.1021/bc7002499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.(a) Johnson I, Spence MTZ, editors. The Molecular Probes Handbook: A guide to Fluorescent Probes and Labeling Technologies. 11. Life Technologies Corporation; 2010. Chapter 2 Thiol-Reactive Probes; pp. 97–116. [Google Scholar]; (b) Hermanson GT. Bioconjugate Techniques. 2. Vol. 2008. Elsiever Inc; 2008. p. 1202. [Google Scholar]
- 7.(a) Yang Z, Attygalle AB. J Mass Spectrom. 2007;42:233. doi: 10.1002/jms.1157. [DOI] [PubMed] [Google Scholar]; (b) Ying J, Claverul N, Sethraman M, Adachi T, Cohen RA. Free Rad Biol Med. 2007;43:1099. doi: 10.1016/j.freeradbiomed.2007.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hill BG, Reily C, Oh JH, Jonhson MS, Landar A. Free Rad Biol Med. 2009;47:675. doi: 10.1016/j.freeradbiomed.2009.06.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Pan J, Xian M. Chem Commum. 2011;47:352. doi: 10.1039/c0cc02076a. [DOI] [PubMed] [Google Scholar]
- 10.(a) Naya A, Kobayashi K, Ishikawa M, Ohwaki K, Saeki T, Noguchi K, Ohtake N. Chem Pharm Bull. 2003;51:697. doi: 10.1248/cpb.51.697. [DOI] [PubMed] [Google Scholar]; (b) Lohmann S, Andrews SP, Burke BJ, Smith MD, Attfield JP, Tanaka H, Kaneko K, Ley SV. Synlett. 2005;8:1291. [Google Scholar]; (c) Chapelon AM, Ollivier C, Santelli M. Tetrahedron Lett. 2006;47:2747. [Google Scholar]; Egi M, Liebeskind LS. Org Lett. 2003;5:801. doi: 10.1021/ol0273497. [DOI] [PubMed] [Google Scholar]
- 11.(a) Kayoda S, Takamura I, Suzuki N, Dohmori R. Chem Pharm Bull. 1976;24:136. doi: 10.1248/cpb.24.136. [DOI] [PubMed] [Google Scholar]; (b) Giesbrecht HE, Knight BJ, Tanguileg NR, Emerson CR, Blakemore PR. Synlett. 2010;3:374. [Google Scholar]; (c) Nomoto Y, Takai H, Ohno T, Kubo K. Chem Pharm Bull. 1991;39:352. doi: 10.1248/cpb.39.352. [DOI] [PubMed] [Google Scholar]; (d) De Kock H, Jonckers THM, Boonants PJGM, Last SJ, Baumeister JE, Van’tklooster GAE. PTC Int Appl 2007147884. [Google Scholar]
- 12.(a) Britto PJ, Knipling L, Wolff J. J Biol Chem. 2002;277:29108. doi: 10.1074/jbc.M204263200. [DOI] [PubMed] [Google Scholar]; (b) Phimister AJ, Lango J, Lee EU, Ernst-Russell MA, Takeshima H, Ma J, Allen PD, Pessah IN. J Biol Chem. 2007;282:8667. doi: 10.1074/jbc.M609936200. [DOI] [PubMed] [Google Scholar]
- 13.Schelte P, Boeckler C, Frisch B, Schuber F. Bioconjugate Chem. 2000;11:118. doi: 10.1021/bc990122k. [DOI] [PubMed] [Google Scholar]
- 14.(a) Lee CC, Samuels ER. Can J Chem. 1964;42:168. [Google Scholar]; (b) Bednar R. Biochemistry. 1990;29:3684. doi: 10.1021/bi00467a014. [DOI] [PubMed] [Google Scholar]
- 15.(a) Chakravarti R, Kulwan SA, Fox PL, Stuehr DJ. Proc Natl Acad Sci USA. 2010;107:18004. doi: 10.1073/pnas.1008133107. [DOI] [PMC free article] [PubMed] [Google Scholar]; (b) Kornberg MD, Sen N, Hara MR, Juluri KR, Nguyen JVK, Snowman AM, Law L, Hester LD, Snyder SH. Nat Cell Biol. 2010;12:1094. doi: 10.1038/ncb2114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Jaffrey SR. Methods Enzymol. 2005;396:105. doi: 10.1016/S0076-6879(05)96011-4. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.