Abstract
Natural killer (NK) cells play a critical role in the control of HIV-1 infection, and NK cells that respond to HIV-1 peptides have been recently described. However, the mechanisms by which NK cells recognize HIV-1 antigens are not fully understood. We investigated NK cell activation in response to HIV-1 peptides during early and chronic HIV-1 clade B infection using a whole-blood assay and multiparameter flow cytometry. Antibody-mediated NK cell activation in response to HIV-1 peptides was not detected in HIV-1-uninfected individuals. In contrast, 79% of individuals with chronic infection and 22% of individuals with early infection had detectable gamma interferon (IFN-γ) NK cell responses to HIV-1 antigens (P < 0.00001). IFN-γ- and tumor necrosis factor alpha (TNF-α)-producing NK cells most frequently targeted Env gp120 (median of 4% and range of 0 to 31% of all NK cells). NK cells rarely targeted other HIV-1 proteins such as Gag, Pol, and Nef. Antibody-mediated NK cell responses to peptides mapped predominantly to Env protein, required the presence of plasma or plasma IgG, and resulted in lower CD16 expression on NK cells, suggesting an antibody-mediated activation of NK cells. Further studies are needed to assess the consequences of these antibody-mediated NK cell responses for HIV-1 disease progression and vaccine-induced protection from infection.
INTRODUCTION
Partial protection from HIV-1 acquisition observed in the recent RV144 Thai vaccine trial has renewed interest in the HIV field to identify correlates of protective immunity. Preliminary findings from RV144 suggest that factors other than neutralizing antibodies or virus-specific CD8+ T cell responses may have mediated the observed modest protection from infection (26). While the correlates of protection from infection in RV144 remain unclear and are under investigation, there has been speculation that protection might have been mediated by short-lived antibody responses (12, 23).
Natural killer (NK) cells are a crucial component of the innate immune response to viral infections and also participate in shaping the adaptive immune response through interactions with dendritic cells (5, 10, 28). NK cells might recognize HIV-1-infected cells directly through receptor-mediated interactions or indirectly by antibody cross-linking of CD16 Fc γ receptor (5). Recently, NK cell responses to HIV-1-derived peptides have been described (30, 31) and found to be associated with control of viremia in HIV-1-infected mothers (23) and protection from mother-to-child transmission of HIV-1 (23). Furthermore, recent studies suggested that these responses are mediated by antibodies (27) and can mediate sufficient immune pressure to drive viral escape by the selection of sequence mutations (8). However, studies by different groups have resulted in different conclusions regarding the precise mechanism by which NK cells recognize HIV-1 antigens and whether these responses are mediated through CD16 or other NK cell receptors, such as killer immunoglobulin-like receptors (KIR) (27, 29, 30).
To determine the frequency of NK cell responses to HIV-1 at different stages of HIV-1 infection and to better characterize those NK cells that do respond to HIV-1 antigens, we assessed NK cell responses to HIV-1 peptides in a large cohort of HIV-1-infected individuals. Strong antibody-mediated NK cell responses to Env were identified, in particular in individuals with chronic infection, and these responses depended on the presence of plasma IgG from HIV-1-infected individuals.
MATERIALS AND METHODS
Study subjects.
Seventy-four individuals with HIV-1 clade B infection and 15 HIV-uninfected control subjects were studied. Of the 74 HIV-1-infected subjects, 18 were in the early phase of infection (within 1 year of infection) and 56 were in the chronic phase of infection. Seven of the subjects (n = 7) with early infection were on highly active antiretroviral therapy (HAART) and had a low-to-undetectable plasma viral load (VL) (48 to 118 RNA copies/ml). Twenty-five subjects with chronic infection were on HAART (VL, 48 to 664 RNA copies/ml), 15 had untreated chronic infection with a median viral load of 14,600 RNA copies/ml (range, 4,200 to 515,502 RNA copies/ml), and the remaining 16 subjects had nonprogressive chronic infection and maintained low viral loads in the absence of HAART (VL, 48 to 2,430 RNA copies/ml) (Table 1). Acute infection was defined by signs and symptoms of acute retroviral syndrome in individuals with either a negative p24 enzyme-linked immunosorbent assay (ELISA) result or a positive p24 ELISA result but less than three bands in an HIV Western blot in the presence of plasma HIV-1 RNA that was detected by reverse transcriptase (RT) PCR. Early infected individuals in our cohort were identified within 12 months of infection. HIV controllers were individuals who maintained a plasma viral load below 2,000 copies/ml average in at least 3 determinations spanning at least a 12-month period off highly active antiretroviral therapy (HAART). The cohort was comprised of mostly Caucasian men (>90% of the participants were men). The individuals studied here were enrolled in this study during their scheduled routine visits at Massachusetts General Hospital (Boston, MA) and the Ragon Institute. The Institutional Review Board of Massachusetts General Hospital approved the study, and all participants gave informed consent for their participation in the study.
Table 1.
Characteristics of study subjects
| Patient group | No. of individuals | CD4 count (range) | Viral load count (range) |
|---|---|---|---|
| Early infection treated (ET) | 7 | 695 (399–1,153) | 48 (48–118) |
| Early infection untreated (EU) | 11 | 628 (285–1,058) | 18,500 (189–706,000) |
| Chronic infection treated (CT) | 25 | 631 (241–1,337) | 48 (48–664) |
| Chronic infection untreated (CU) | 15 | 565 (3–913) | 14,600 (4,200–515,502) |
| HIV controllers (HC) | 16 | 802 (392–1,331) | 81 (48–2,430) |
| HIV uninfected (HU) | 15 | N/Aa | N/Aa |
N/A, not available.
Synthetic HIV-1 peptides.
Overlapping peptides (OLPs; 18-mers overlapping by 10 to 11 amino acids) spanning the entire HIV-1 clade B consensus sequence (Gag, Nef, Pol, gp120, gp41, Rev, Tat, Vpu, Vpr, Vif [RTVVV]) were arranged in pools. Responding pools were further tested in selected individuals to allow mapping of responding pools to individual peptides. The peptides were synthesized at the Massachusetts General Hospital peptide synthesis core facility on an automated peptide synthesizer (MBS 396; Advanced ChemTech).
HLA class I and KIR typing.
High-resolution HLA class I typing and KIR genotyping were performed as described previously (4).
Whole-blood intracellular cytokine staining (ICS) and NK cell degranulation assay.
The frequency of NK cells and CD8+ T cells producing gamma interferon (IFN-γ) and tumor necrosis factor alpha (TNF-α) was determined by stimulating whole blood with HIV-1 peptide pools as previously described (22). Briefly, 200 μl of whole blood collected in sodium heparin Vacutainer tubes was stimulated with Gag, Nef, Pol, gp120, gp41, and RTVVV peptide pools at a final concentration of 2 μg/ml in the presence of costimulatory anti-CD28 and anti-CD49d antibodies (BD Biosciences). A negative-control tube with whole blood alone and a positive-control tube with whole blood stimulated with phorbol myristate acetate (PMA)-ionomycin (AG Scientific) at a final concentration of 1 μg/ml and 0.5 μg/ml, respectively, were also included in each experiment. Brefeldin A (Sigma-Aldrich) was added to each sample at a final concentration of 10 μg/ml, and the samples were then mixed and incubated at 37°C for 6 h. After incubation, 20 μl of EDTA (BD Biosciences) was added to all tubes to arrest activation and to remove adherent cells. Samples were then transferred to fluorescence-activated cell sorter (FACS) tubes, red blood cells were lysed, and cells were washed and permeabilized as previously described (22) before staining.
CD8+ T cells and NK cells producing cytokines were assessed by staining with the following cocktail of antibodies: anti-CD8-phycoerythrin (PE) Cy5, CD56-fluorescein isothiocyanate (FITC), CD16-allophycocyanin (APC) Cy7, IFN-γ-APC, TNF-α-Alexa 700, CD19-Pacific Blue (all from BD Biosciences), CD3-PE Cy5.5, and CD14-Pacific Blue (Caltag). In addition, NK cell receptors were also assessed in separate experiments using the following antibodies: anti-NKp46-PE, CD158a-PE, CD158b-PE (all from BD Biosciences), NKG2A-PE, CD158e1/e2-PE, and NKG2D-PE (Beckman Coulter). CD57 and CD7 expression was also assessed using anti-CD57-PE and anti-CD7-PE (BD Biosciences). The above-mentioned individual markers CD7, CD57, KIR, NKG2D, NKG2A, NKp46, and CD16 were assessed to further characterize the responding NK cells. The tubes with antibodies were incubated for 1 h in the dark at room temperature. Cells were then washed twice and resuspended in phosphate-buffered saline (PBS) with paraformaldehyde before acquisition on the LSRII.
To assess the ability of NK cells to degranulate in response to HIV-1 peptides, a CD107a degranulation assay (3) was performed on 4 individuals who demonstrated high gp120 responses during previous screening assays. Whole blood was stimulated with the gp120 pool as detailed above in the presence of anti-CD107a PE Cy5 and 2.5 μg/ml of Golgi stop (both from BD Biosciences) and brefeldin A (10 μg/ml; Sigma-Aldrich). A negative-control tube with peripheral blood mononuclear cells (PBMCs) in R10 (10% fetal calf serum in RPMI medium) and a positive-control tube stimulated with MHC-devoid targets (K562 cell line; ATCC) at an effector-to-target cell (E:T) ratio of 10:1 was included in each degranulation assay experiment. Staining was carried out as described above.
Five hundred thousand events were acquired for whole-blood assays, and between 500,000 and 1,000,000 events were acquired for isolated PBMC assays and were analyzed using FlowJo software (TreeStar, Inc.). Initial gating was on the lymphocytes, followed by the removal of doublets by forward scatter height (FSC-H) versus forward scattered area (FSC-A). B cells and monocytes were eliminated by gating on CD19/CD14 in the Pacific Blue channel, respectively. Gates were set on CD3-negative cells, followed by gating on CD56- and CD16-positive cells for definition of NK cells. Gating on CD3-positive cells followed by gating on CD8+ cells allowed for definition of CD8+ T cells. To rule out the possibility that granulocytes and T cells could be mediating the response, granulocyte and T cell depletion assays were done from whole blood in preliminary assays (data not shown). A response 3 times above background (unstimulated controls) was considered a positive result in all whole blood and isolated PBMC assays.
ICS and NK cell degranulation assays on isolated PBMCs.
To compare the frequencies of NK cells and CD8+ T cells producing cytokines in whole blood versus isolated PBMCs, ICS was also performed on freshly isolated PBMCs from selected individuals. One million PBMCs were resuspended in R10 and were stimulated with peptide pools at a final concentration of 2 μg/ml in the presence of costimulatory antibodies anti-CD28 and anti-CD49d (BD Biosciences). Brefeldin A was added to the tubes (final concentration 10 μg/ml; Sigma Corp.) before incubation at 37°C for 6 h. Control tubes with PBMCs or PBMCs stimulated with PMA-ionomycin were included with each experiment. Following stimulation, cells were washed with 2 ml of wash buffer (1% PBS-fetal calf serum [FCS]; Sigma) and centrifuged at 1,500 rpm for 10 min before the addition of the following surface antibodies: anti-CD3 PE Cy5.5, CD14-Pacific Blue (Caltag), CD19-Pacific Blue, CD56-FITC, CD8-PE Cy5, and NKp46-PE or CD57-PE, CD7-PE, NKG2A-PE, NKG2D-PE, CD158a-PE, CD-158b PE, and CD158e1/e2-PE. Cells stained with antibodies were incubated for 20 min at room temperature in the dark. Following incubation, cells were washed and fixed (Fix & Perm A; Invitrogen) for 20 min in the dark before permeabilization (Fix & Perm B; Caltag) and intracellular staining with anti-IFN-γ-APC and TNF-α-Alexa 700 for 20 min in the dark. Cells were then washed twice and resuspended in 200 μl of PBS before acquisition on the LSRII.
The ability of NK cells to degranulate in response to HIV-1 peptides was assessed in a few individuals who demonstrated high responses at screening visits. One million freshly isolated PBMCs were resuspended in R10 and stimulated with the gp120 peptide pool. A negative-control tube with PBMCs in R10 and a positive-control tube stimulated with MHC-devoid targets (K562 cell line; ATCC) at an E:T ratio of 10:1 was included in each experiment. Anti-CD107a-PE Cy5 antibody (BD Biosciences) was added to all the tubes together with Golgi stop (2.5 μg/ml; BD Biosciences) and brefeldin A (10 μg/ml; Sigma-Aldrich). The samples were incubated for 6 h at 37°C and then processed for ICS staining from isolated PBMCs as described above. For PBMC experiments, PBMCs were either stimulated alone or stimulated with different dilutions of autologous plasma (1:1, 1:10, 1:5, and 1:100). A plasma dilution factor of 1:5 resulted in low background levels without loss of response and was thus used in subsequent degranulation/PBMC assays. Sample acquisition on the LSRII was performed as described above. We also assessed PBMCs in dimethyl sulfoxide (DMSO) as an additional control, and we detected no responses in tubes that were not stimulated with peptide.
IgG isolation from plasma.
To determine if immunoglobulin G (IgG) antibodies in plasma mediated the NK cell activation in response to HIV-1 peptides, IgG was isolated from 4 HIV-1-infected individuals using the Melon gel IgG spin purification kit (Thermo Scientific, Pierce Biotechnology) according to the manufacturer's instructions. Briefly, 500 μl of the subject's plasma diluted 1:10 in buffer was loaded on a purification support, mixed for 5 min to ensure antibody binding, and spun down for 1 min at 3,000 × g to elute the bound IgG. The concentration of collected IgG was measured on the NanoDrop 2000 spectrophotometer (Thermo Scientific) and stored at 4°C for downstream use. For PBMC and IgG ICS assays, IgG titrations ranging from 1 mg to 1 μg were done in PBMCs; a concentration of 20 μg/ml was found to give an optimal response and was used in subsequent assays.
Statistical analysis.
Viral loads and CD4 counts and HIV-specific NK cell and CD8+ T cell differences in cytokine production between different disease groups were compared using one-way analysis of variance (ANOVA); the significantly different groups identified were further compared using the Mann-Whitney test. Differences in degranulation and cytokine production from whole blood, PBMC, PBMC with plasma assays, and NK receptor profiles between different cytokine-producing and -nonproducing cells were compared using the Mann-Whitney test. Proportions of individuals with early and chronic infection making responses to peptides and KIR-HLA genotype combinations in responders and nonresponders were compared using Fisher's exact test.
RESULTS
Viral loads and CD4 counts in study participants.
We investigated the frequency of NK cell responses to HIV-1 antigens in a cohort of 74 HIV-1-infected adults, consisting of untreated and treated individuals with early or chronic infection. In addition, 15 HIV-1-uninfected healthy control subjects were examined (Table 1). There were no significant differences in CD4 counts between treated and untreated subjects with early and chronic infection; the CD4 counts in HIV-1 controllers were, however, significantly higher than in subjects with progressive infection (median of 802 and range of 392 to 1,331 versus median of 565 and range of 3 to 913; P = 0.003 by one-way ANOVA) (Table 1). Furthermore, viral loads were significantly lower in individuals with early or chronic infection undergoing HAART than untreated subjects (P < 0.0005 and P < 0.0001 in early infected and chronically infected subjects, respectively) or in HIV-1 controllers than untreated subjects with early or chronic HIV-1 infection (P < 0.0001 by one-way ANOVA) (Table 1).
Frequent targeting of HIV-1 Env by NK cells from HIV-1-infected individuals.
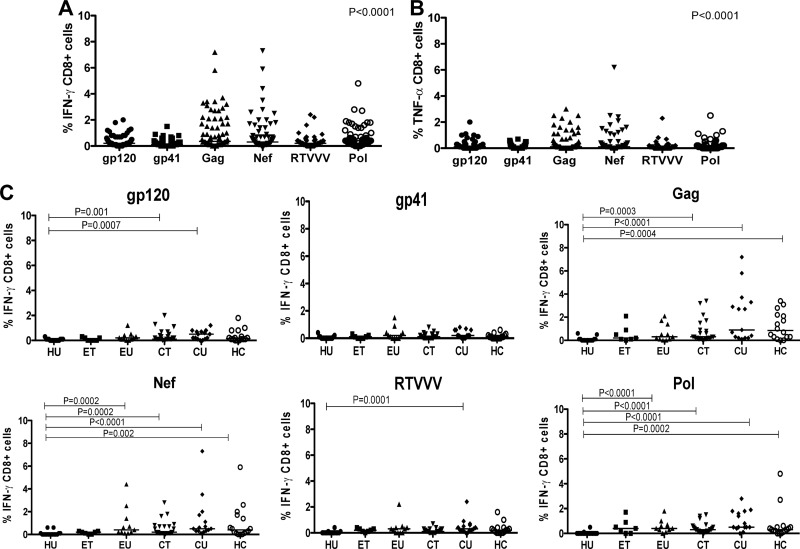
Of the 74 HIV-1-infected individuals studied, 49/74 (66%) mounted an NK cell response to at least one of the tested HIV-1 pools, with the majority of them (47/74) targeting at least the Env gp120 pool (Fig. 1; see also Fig. S1 in the supplemental material). No NK cell activation in response to HIV-1 peptides was detected in the 15 uninfected controls. NK cell responses to HIV-1 antigens were of a high magnitude (e.g., the frequency of gp120 IFN-γ responses in all individuals ranged from 0 to 31% with a median of 4%) and produced IFN-γ and TNF-α (Fig. 1A and B). HIV-1 Env gp120 and gp41 were the two HIV-1 proteins most frequently targeted by NK cells and were targeted significantly more frequently than Gag, Nef, RTVVV, and Pol (P < 0.0001 by one-way ANOVA) (Fig. 1A and B). While responses to Env gp120 were strong and frequent in individuals with chronic infection, irrespective of treatment status, NK cell responses to HIV-1 peptides were very rare and low in individuals with early infection (45/56 versus 4/18; P < 0.0001 by Fisher's exact test) (Fig. 1C). Analysis of the combined chronic infection and early infection groups showed that NK cell cytokine responses in whole blood to HIV-1 peptides were significantly higher in chronically infected subjects with progressive and nonprogressive HIV-1 infection than in individuals with early infection (P < 0.0001 and P = 0.04 for gp120 and gp41, respectively, by Fisher's exact test) (Fig. 1C). Interestingly, NK cell responses directed to Gag and Pol regions were rarely detected in the studied cohort. Together these data show frequent IFN-γ and TNF-α NK cell responses that dominantly targeted Env during chronic HIV-1 infection.
Fig 1.
Proteins targeted by antibody-mediated NK cell responses in HIV-1-infected individuals. NK cell IFN-γ (A) and TNF-α (B) responses to HIV-1 peptide pools in all studied HIV-1-infected individuals (n = 74) irrespective of disease or treatment status (C). P values in panels A and B indicate the overall P value obtained from comparison of all the HIV regions using the one-way ANOVA test, and subsequent comparisons between the groups were done using the Mann-Whitney test and are discussed in the Results section. Abbreviations on the x axis in panel C refer to different patient groups as outlined in Table 1.
Fig 2.
Proteins targeted by CD8+ T cell responses in HIV-1-infected individuals. CD8+ IFN-γ (A) and TNF-α (B) T cell responses to HIV-1 peptide pools in all studied HIV-1-infected individuals (n = 74) irrespective of disease or treatment status (C). P values in panels A and B indicate the overall P value obtained from comparison of all the HIV regions using the one-way ANOVA test, and subsequent comparisons between the groups were done using the Mann-Whitney test and are discussed in Results. Abbreviations on the x axis in panel C refer to different patient groups as outlined in Table 1.
NK cells and CD8+ T cells significantly differ in the targeted regions of HIV-1.
In contrast to the observed strong and frequent NK cell activation in response to HIV-1 Env (Fig. 1), IFN-γ CD8+ T cell responses directed against gp120 and gp41 were significantly lower in the same studied subjects (median of 0.2% and range of 0 to 2% for gp120 and median of 0.1% and range of 0 to 1.5% for gp41; P < 0.0001 and P = 0.0002, respectively, by the Mann-Whitney test). The same trend was observed for TNF-α CD8+ T cell responses. As expected based on previous studies (1, 16), CD8+ T cell responses directed against Gag, Nef, and Pol were significantly higher than Env-specific CD8+ T cell responses (P < 0.0001 by one-way ANOVA) (Fig. 2A and B). HIV-1-specific CD8+ T cell responses to Gag, Nef, and Pol were detectable in treated and untreated individuals with chronic HIV-1 infection and individuals with untreated early HIV-1 infection but only rarely in individuals that were treated during early HIV-1 infection and never in uninfected controls (Fig. 2C).
NK cells degranulate in response to HIV-1 peptides and require IgG in plasma.
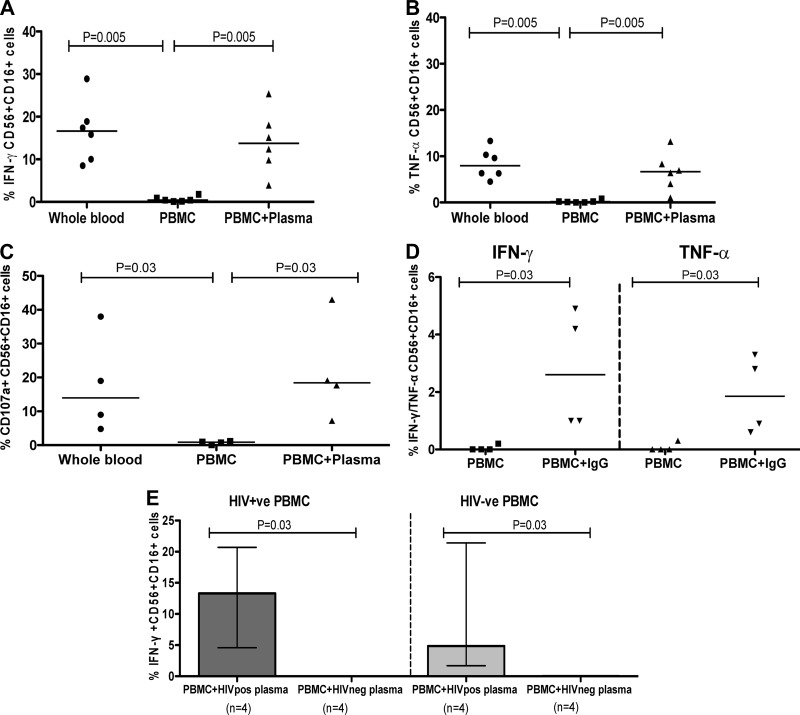
Our studies described above detecting NK cell activation in response to HIV-1 peptides in whole blood are in line with previous reports (9, 27, 30, 31). However, previous studies have come to different conclusions regarding the mechanisms responsible for these HIV-1-specific NK cell responses (27, 29). We therefore compared NK cell activation following stimulation with peptide pools in whole blood versus freshly isolated PBMCs in six subjects who demonstrated strong NK cell responses to HIV-1 antigens upon screening and returned for follow-up visits. Since gp120 was the dominant HIV-1 protein targeted in these study subjects, we focused on this protein for the subsequent studies. Remarkably, NK cells from isolated PBMCs did not produce any cytokines in response to gp120. However, NK cell responses to gp120 were restored in PBMCs from the same study subjects by the addition of autologous plasma, reaching levels of cytokine (IFN-γ and TNF-α) production similar to those measured in whole blood when equal amounts of plasma were added (Fig. 3A and B; see also Fig. S2 in the supplemental material). Similar results were noted upon degranulation assessment using the CD107a marker in a subset of 4 studied individuals (Fig. 3C).
Fig 3.
NK cell responses to HIV-1 peptides require IgG or the presence of plasma. (A) Representative example of the antibody-mediated NK cell responses following stimulation with gp120 peptide pool in whole blood, isolated PBMCs in the absence of plasma, and PBMCs with autologous plasma added back at varying dilutions. Simultaneous measurement of IFN-γ-producing (A) and TNF-α-producing (B) NK cells to the gp120 peptide pool in six individuals—in whole blood, PBMCs without plasma, and PBMCs with plasma at a 1:5 dilution. (C) Degranulation assessment of NK cells in 4 individuals—in whole blood, PBMCs without plasma, and PBMCs with plasma at a 1:5 dilution. (D) Measurement of NK cell responses in infected individuals (n = 4)—in PBMCs with or without plasma IgG following stimulation with the gp120 peptide pool. (E) Assessment of antibody-mediated NK cell responses to the gp120 peptide pool using plasma and PBMCs derived from HIV-1-infected donors (n = 4) and healthy donors (n = 4).
To assess whether antibodies mediated this NK cell activation in response to peptides as previously suggested by others (26), we isolated IgG from plasma and determined cytokine responses to the gp120 pool in PBMCs with and without adding plasma-derived IgG. No NK cell activation in response to peptides was detected in PBMCs without IgG. In contrast, significant NK cell activation was observed in the presence of plasma IgG (P = 0.03 by the Mann-Whitney test) (Fig. 3D).
To further determine whether the plasma from HIV-1-infected individuals was sufficient to mediate NK cell responses against HIV-1 gp120 peptide pools, plasma transfer experiments were done using plasma from HIV-1-infected individuals and PBMCs from HIV-1-uninfected subjects and vice versa. Adding plasma from HIV-1-negative individuals to isolated PBMCs derived from HIV-1-infected or -uninfected individuals had no effect on the NK cell response to gp120 peptides. However, the NK cell response to peptides was restored when plasma from HIV-1-positive individuals was added back to isolated PBMCs in both HIV-infected and -uninfected individuals (Fig. 3E). Notably, the NK cell responses were lower when using PBMCs from HIV-1-uninfected individuals than PBMCs from HIV-1 individuals when plasma from HIV-1-positive individuals was added (median, 4.8% versus 13.3%); however, these differences were not statistically significant (Fig. 3E). Taken together, these data demonstrate that the NK cell response to HIV-1 peptides is mediated by plasma IgGs and that plasma derived from HIV-1-infected individuals is sufficient to trigger responses to HIV-1 gp120 peptide by NK cells from HIV-1-negative individuals.
Characterization of the individual HIV-1 peptides targeted by antibody-mediated NK cell activation in whole blood.
To determine the specific regions within HIV-1 targeted by NK cells, the individual 18-mer peptides triggering cytokine production by NK cells were identified in 5 subjects who demonstrated significant antibody-mediated NK cell responses to peptide pools upon screening and returned for follow-up visits. A total of 21 peptides were identified from whole blood in the 5 studied subjects, with NK cells in each study subject targeting a range of 2 to 10 individual peptides (Table 2). Most of the epitopes mapped (76%) were in the gp120 region of Env, and some peptides (e.g., OLP 301 and 316) were commonly targeted by NK cells in different individuals. In some instances, consecutive OLPs were targeted, and we were not able to determine whether a single epitope in the overlapping regions of the peptides was targeted or two independent epitopes were contained in the two peptides. These data demonstrate that multiple epitopes within HIV-1 Env can be mapped to activated NK cells in infected individuals.
Table 2.
Individual peptides targeted by NK cells from study subjects
| Patient no. | Epitope | HIV protein | Sequence | Magnitude (%) |
|---|---|---|---|---|
| 1 | OLP 126 | Tat | IVFIEYRKILRQRKIDRL | 6.2 |
| OLP 127 | Tat | ILRQRKIDRLIDRIRERA | 1.1 | |
| OLP 282 | Vpr | HFPRIWLHSLGQHIYETY | 1.0 | |
| OLP 301a | gp120 | NVTENFNMWKNNMVEQMH | 6.8 | |
| OLP 303 | gp120 | QMHEDIISLWDQSLKPCV | 2.5 | |
| OLP 315 | gp120 | SVITQACPKVSFEPIPIH | 0.7 | |
| OLP 316a | gp120 | KVSFEPIPIHYCAPAGFA | 2.2 | |
| OLP 329 | gp120 | NNNTRKSIHIGPGRAFY | 0.7 | |
| OLP 332 | gp120 | EIIGDIRQAHCNISRAKW | 2.2 | |
| OLP 363 | gp120 | LRAIEAQQHLQLTVWGI | 1.0 | |
| 2 | OLP 293 | gp120 | AAEQLWVTVYYGVPVWK | 8.0 |
| OLP 301a | gp120 | NVTENFNMWKNNMVEQMH | 14.2 | |
| 3 | OLP 91 | Nef | PEKEVLVWKFDSRLAFHH | 7.6 |
| OLP 108 | Rev | QILVESPAVLESGTKEE | 18.4 | |
| OLP 301a | gp120 | NVTENFNMWKNNMVEQMH | 9.3 | |
| OLP 316a | gp120 | KVSFEPIPIHYCAPAGFA | 4.3 | |
| OLP 332 | gp120 | EIIGDIRQAHCNISRAKW | 3.2 | |
| 4 | OLP 301a | gp120 | NVTENFNMWKNNMVEQMH | 6.1 |
| OLP 312 | gp120 | YALFYKLDVVPIDNDNTSY | 9.0 | |
| 5 | OLP 364 | gp120 | HLQLTVWGIKQLQARVL | 9.4 |
| OLP 365 | gp120 | GIKQLQARVLAVERYLK | 8.0 |
Peptides that were targeted by more than one individual.
Responding and nonresponding NK cells differ in their phenotypic characteristics.
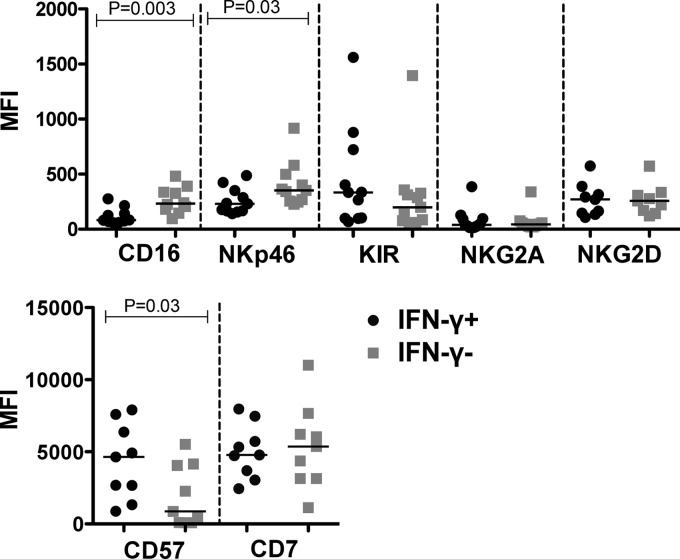
To further characterize the NK cells that responded to HIV-1 peptides, we assessed differences in the receptors expressed by NK cells that responded to HIV-1 peptides by cytokine production, and those NK cells that did not, in the same study subjects. The markers assessed included the activating NK cell receptors CD16, NKp46, NKG2D, the inhibitory receptors KIR and NKG2A, and CD57 and CD7 (Fig. 4). NK cells that produced IFN-γ in response to peptides had significantly lower expression of CD16 and NKp46 (P = 0.003 and P = 0.03, respectively, by the Mann-Whitney test). In addition, IFN-γ-producing NK cells expressed higher levels of CD57 (P = 0.03 by the Mann-Whitney test). In contrast, we did not observe any significant differences in the expression of KIR, NKG2A, NKG2D, or CD7 between cytokine-producing and nonresponding NK cells in the same individuals (Fig. 4). Together these data demonstrate that the NK cells that respond with cytokine production to peptides have reduced CD16 and NKp46 expression and a more mature/terminally differentiated phenotype (CD57 positive).
Fig 4.
Receptor expression profiles of cytokine-producing and -nonproducing NK cells. Receptor profiles and activation markers in IFN-γ-producing and -nonproducing NK cells in responder subjects following gp120 stimulation. Subjects were assessed for the following NK receptors and markers: CD16 (n = 10), NKp46, KIR (n = 11), NKG2A (n = 12), NKG2D, CD57, and CD7 (n = 9).
DISCUSSION
NK cell responses to HIV-1 peptides have been recently described by different groups studying HIV-1 clade B and clade C in chronically infected individuals (7, 8, 27, 29–31). However, some of these two groups have come to different conclusions regarding the mechanisms involved in mediating these peptide-specific NK cell responses, with some groups suggesting that NK cell responses to HIV-1 peptides depend on the presence of antibodies directed against HIV-1 (25, 27) and another group suggesting that KIR and HLA genotypes are most likely involved (29). In line with these previous observations, we demonstrate that NK cells can respond to HIV-1 peptides in a whole-blood assay. These responses depend on the presence of plasma IgG and result in low expression of CD16, suggesting a critical role of antibodies in mediating this activity. Furthermore, we demonstrate that these NK cell responses to HIV-1 peptides are of a low magnitude or absent in individuals during the first year of infection, when antibody titers have been described to be lower, but have higher frequencies and magnitudes in individuals with controlled or progressive chronic infection. Finally, we demonstrate that peptide-specific NK cell responses are mediated by a subset of terminally differentiated CD57+ NK cells. Taken together, these data demonstrate that NK cell responses to specific antigens can be induced in HIV-1 infection and are mediated by plasma IgG.
The antibody-mediated NK cell responses that were detected in our cohort dominantly targeted Env gp120 and gp41 to a lesser extent Nef and accessory RTVVV proteins and were very rarely detected in other regions of HIV-1 such as Gag and Pol. In contrast to our findings of rare Pol-specific responses, a recent study found Pol to be a significant target of antibody-dependent cellular cytotoxicity (ADCC) in a larger cohort of therapy-naïve subjects (13) and that these responses decreased with follow-up and did not correlate with disease progression (13). The mechanism of recognition of internal HIV-1 proteins and the consequences of these for HIV-1 disease progression remain an area of investigation. Notably, our study showed that NK cell responses to HIV-1 peptides were significantly higher in subjects with progressive and nonprogressive chronic HIV-1 infection than in individuals with early infection and were not affected by antiretroviral treatment. Taken together, these data further support an antibody-mediated NK cell response against HIV-1 that slowly develops during early infection and is more established in chronic infection. The slow development of the antibody-mediated NK cell responses in early infection noted in this study are in line with reports in infants with HIV-1 clade C infection (30). It is also possible that this delayed development in responses may be an indication of the restriction of responses to conformational epitopes that broaden to linear HIV peptides with disease progression, as recently suggested (8). In contrast to previous studies (14, 31), we found no association of the NK cell responses to peptides with markers of disease progression in our cohort, and responses in individuals with progressive and nonprogressive HIV-1 infection did not differ.
Our data demonstrate that the NK cell-mediated responses to HIV-1 peptides were only detected in the presence of plasma or plasma-derived IgG and resulted in low CD16 expression on responding NK cells, suggesting that the response activity is dependent on the binding of the Fc portion of the antibodies in plasma to the CD16 Fcγ receptor on NK cells. Some additional, nonspecific plasma factors involved in the NK cell response to HIV-1 peptides have been suggested by other groups (30). In our study, IgG derived from the plasma of HIV-1-infected individuals was sufficient to reconstitute the response of NK cells to HIV-1 peptides. In transfer experiments, adding plasma from HIV-1-infected individuals to PBMCs from HIV-1-negative individuals, we also observed induction of NK cell responses to HIV-1 peptides, however, with a larger degree of heterogeneity. Transfer of plasma from HIV-1-positive individuals to healthy donor PBMCs resulted in NK cell responses to peptides in all individuals studied, but responses were lower than in the presence of autologous PBMCs in some cases. Further experiments will be needed to determine whether this might be due to the fact that NK cells from infected individuals have a higher baseline activation status than NK cells from uninfected individuals. Taken together, our data demonstrate the critical role that plasma-derived IgG play in mediating the NK cell response to HIV-1 peptides in vitro and are in line with previous studies by Stratov and others (27).
Furthermore, we demonstrate that NK cells that responded to HIV-1 peptides expressed CD57, a marker that has been recently described to identify more differentiated NK cells (20) and lower levels of NKp46. The CD57+ phenotype exhibited by activated NK cells in this study has been recently demonstrated to be associated with stronger lytic activity of NK cells when stimulated through the CD16 receptor (20). Similarly, reduced NKp46 expression of activated NK cells has also been described previously, but different studies have come to different conclusions (11, 21, 32).
The interaction between KIR and their HLA class I ligands during NK cell development has an important influence on the functionality of NK cells, a process referred to as “licensing” or “arming” (6, 17, 24). We therefore investigated whether the HLA and KIR genotypes accounted for the differences in individuals with NK cells that responded to HIV-1 peptides (responders), and those that did not (nonresponders), in our cohort, as previously suggested by others (15, 18, 19, 29). However, we observed no significant differences in HLA and KIR genotypes between the two groups after controlling for multiple comparisons (see Table S1 in the supplemental material), suggesting that mechanisms other than KIR and HLA genotypes might account for the differences in NK cell responsiveness observed in this cohort. Interestingly, a recent study reported that the effects of NK cell licensing on ADCC effector functions are abrogated during HIV-1 infection (25). Further studies on large and well-characterized cohorts will be required to understand the impact of KIR/HLA genotypes on NK cell functions, including ADCC function, in HIV-1 infection, the consequences of HIV-1 disease progression, and the ability of HIV-1 to evade NK cell-mediated immune pressure (2, 8).
To further characterize the regions within HIV-1 targeted by antibodies mediating NK cell activation, we mapped a total of 21 epitopes from 5 individuals using the whole-blood assay. Over 75% of these epitopes were located within Env, and some epitopes were commonly targeted by more than 1 individual, although in some cases the responses may have been mediated by one epitope (e.g., OLP 364/365). Recently, a total of 59 linear ADCC epitopes were mapped in 22 individuals using a similar approach, most of which were also in Env (8). The parts of gp120 specifically targeted and the functionality of different epitope-specific responses will need further investigation to identify those antibodies that can mediate protective responses and might reduce viral fitness (8).
To sum up, we assessed NK cell responses to HIV-1 peptides in individuals with treated or untreated early, progressive, and nonprogressive HIV-1 infection. We demonstrate that NK cells in chronically infected individuals responded more frequently and strongly to HIV-1 peptides than individuals with early infection and that these NK cell responses did not differ between individuals with progressive and nonprogressive chronic infection. NK cell responses to HIV-1 peptides mapped predominantly to the Env protein, depended on the presence of plasma IgG, resulted in low CD16 and NKp46 expression, and were mediated by CD57+ NK cells. Future studies will be required to determine the consequences of these NK cell responses against HIV-1 for the control or prevention of HIV-1 infection, in particular in the setting of vaccines that induce binding antibodies against HIV-1.
Supplementary Material
ACKNOWLEDGMENTS
This study was supported by the Doris Duke Charitable Foundation, the Ragon Institute of MGH, MIT, and Harvard, and the National Institutes of Health (grant no. P01 AI074415). C.F.T. was supported by the Columbia University-Southern African Fogarty AIDS International Training and Research Program (AITRP) through the Fogarty International Center, National Institutes of Health (grant no. 3 D43 TW000231 16S2). This project has been funded in whole or in part with federal funds from the National Cancer Institute, National Institutes of Health, under contract no. HHSN261200800001E. This research was supported in part by the Intramural Research Program of the National Institutes of Health, National Cancer Institute, Center for Cancer Research.
The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government.
Footnotes
Published ahead of print 11 April 2012
Supplemental material for this article may be found at http://jvi.asm.org/.
REFERENCES
- 1. Addo MM, et al. 2003. Comprehensive epitope analysis of human immunodeficiency virus type 1 (HIV-1)-specific T-cell responses directed against the entire expressed HIV-1 genome demonstrate broadly directed responses, but no correlation to viral load. J. Virol. 77:2081–2092 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Alter G, et al. 2011. HIV-1 adaptation to NK-cell-mediated immune pressure. Nature 476:96–100 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Alter G, Malenfant JM, Altfeld M. 2004. CD107a as a functional marker for the identification of natural killer cell activity. J. Immunol. Methods 294:15–22 [DOI] [PubMed] [Google Scholar]
- 4. Alter G, et al. 2007. Differential natural killer cell-mediated inhibition of HIV-1 replication based on distinct KIR/HLA subtypes. J. Immunol. Methods 204:3027–3036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Altfeld M, Fadda L, Frleta D, Bhardwaj N. 2011. DCs and NK cells: critical effectors in the immune response to HIV-1. Nat. Rev. Immunol. 11:176–186 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Anfossi N, et al. 2006. Human NK cell education by inhibitory receptors for MHC class I. Immunity 25:331–342 [DOI] [PubMed] [Google Scholar]
- 7. Chung A, Rollman E, Johansson S, Kent SJ, Stratov I. 2008. The utility of ADCC responses in HIV infection. Curr. HIV Res. 6:515–519 [DOI] [PubMed] [Google Scholar]
- 8. Chung AW, et al. 2011. Immune escape from HIV-specific antibody-dependent cellular cytotoxicity (ADCC) pressure. Proc. Natl. Acad. Sci. U. S. A. 108:7505–7510 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Chung AW, et al. 2011. Activation of NK cells by ADCC responses during early HIV infection. Viral Immunol. 24:171–175 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Fauci AS, Mavilio D, Kottilil S. 2005. NK cells in HIV infection: paradigm for protection or targets for ambush. Nat. Rev. Immunol. 5:835–843 [DOI] [PubMed] [Google Scholar]
- 11. Fogli M, et al. 2004. Significant NK cell activation associated with decreased cytolytic function in peripheral blood of HIV-1-infected patients. Eur. J. Immunol. 34:2313–2321 [DOI] [PubMed] [Google Scholar]
- 12. Haynes BF, Liao HX, Tomaras GD. 2010. Is developing an HIV-1 vaccine possible? Curr. Opin. HIV AIDS. 5:362–367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Isitman G, Chung AW, Navis M, Kent SJ, Stratov I. 2011. Pol as a target for antibody dependent cellular cytotoxicity responses in HIV-1 infection. Virology 412:110–116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Johansson SE, et al. 2011. NK cell function and antibodies mediating ADCC in HIV-1-infected viremic and controller patients. Viral Immunol. 24:359–368 [DOI] [PubMed] [Google Scholar]
- 15. Khakoo SI, et al. 2004. HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science 305:872–874 [DOI] [PubMed] [Google Scholar]
- 16. Kiepiela P, et al. 2004. Dominant influence of HLA-B in mediating the potential co-evolution of HIV and HLA. Nature 432:769–775 [DOI] [PubMed] [Google Scholar]
- 17. Kim S, et al. 2005. Licensing of natural killer cells by host major histocompatibility complex class I molecules. Nature 436:709–713 [DOI] [PubMed] [Google Scholar]
- 18. Kim S, et al. 2008. HLA alleles determine differences in human natural killer cell responsiveness and potency. Proc. Natl. Acad. Sci. U. S. A. 105:3053–3058 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Lanier LL. 2005. NK cell recognition. Annu. Rev. Immunol. 23:225–274 [DOI] [PubMed] [Google Scholar]
- 20. Lopez-Vergès S, et al. 2010. CD57 defines a functionally distinct population of mature NK cells in the human CD56dimCD16+ NK-cell subset. Blood 116:3865–3874 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Mavilio D, et al. 2005. Characterization of CD56-/CD16+ natural killer (NK) cells: a highly dysfunctional NK subset expanded in HIV-infected viremic individuals. Proc. Natl. Acad. Sci. U. S. A. 102:2886–2891 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Meddows-Taylor S, Shalekoff S, Kuhn L, Gray GE, Tiemessen CT. 2007. Development of a whole blood intracellular cytokine staining assay for mapping CD4(+) and CD8(+) T-cell responses across the HIV-1 genome. J. Virol. Methods 144:115–121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Munier CM, Andersen CR, Kelleher AD. 2011. HIV vaccines: progress to date. Drugs 71:387–414 [DOI] [PubMed] [Google Scholar]
- 24. Orr MT, Lanier LL. 2011. Natural killer cell licensing during viral infection. Advances in experimental medicine and biology. 780:37–44 [DOI] [PubMed] [Google Scholar]
- 25. Parsons MS, et al. 2012. HIV infection abrogates the functional advantage of natural killer cells educated through KIR3DL1/HLA-Bw4 interactions to mediate anti-HIV antibody-dependent cellular cytotoxicity. J. Virol. 86:4488–4495 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Rerks-Ngarm S, et al. 2009. Vaccination with ALVAC and AIDSVAX to prevent HIV-1 infection in Thailand. N. Engl. J. Med. 361:2209–2220 [DOI] [PubMed] [Google Scholar]
- 27. Stratov I, Chung A, Kent SJ. 2008. Robust NK cell-mediated human immunodeficiency virus (HIV)-specific antibody-dependent responses in HIV-infected subjects. J. Virol. 82:5450–5459 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Tasca S, et al. 2003. Escape of monocyte-derived dendritic cells of HIV-1 infected individuals from natural killer cell-mediated lysis. AIDS 17:2291–2298 [DOI] [PubMed] [Google Scholar]
- 29. Tiemessen CT, et al. 2011. Natural killer cell responses to HIV-1 peptides are associated with more activating KIR genes and HLA-C genes of the C1 allotype. J. Acquir. Immune Defic. Syndr. 57:181–189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Tiemessen CT, et al. 2009. Cutting edge: unusual NK cell responses to HIV-1 peptides are associated with protection against maternal-infant transmission of HIV-1. J. Immunol. 182:5914–5918 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Tiemessen CT, et al. 2010. Natural killer cells that respond to human immunodeficiency virus type 1 (HIV-1) peptides are associated with control of HIV-1 infection. J. Infect. Dis. 202:1444–1453 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Wong AH, et al. 2010. Alterations in natural killer cell receptor profiles during HIV type 1 disease progression among chronically infected South African adults. AIDS Res. Hum. Retroviruses 26:459–469 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.