Abstract
In previous studies, we found that adenomatous polyposis coli (APC) blocks the base excision repair (BER) pathway by interacting with 5′-flap endonuclease 1 (Fen1). In this study, we identify the molecular features that contribute to the formation and/or stabilization of the APC/Fen1 complex that determines the extent of BER inhibition, and the subsequent accumulation of DNA damage creates mutagenic lesions leading to transformation susceptibility. We show here that APC binds to the nuclear localization sequence of Fen1 (Lys365Lys366Lys367), which prevents entry of Fen1 into the nucleus and participation in Pol-β-directed long-patch BER. We also show that levels of the APC/Fen1 complex are higher in breast tumors than in the surrounding normal tissues. These studies demonstrate a novel role for APC in the suppression of Fen1 activity in the BER pathway and a new biomarker profile to be explored to identify individuals who may be susceptible to the development of mammary and other tumors.
Introduction
The genomes of eukaryotic cells are under continuous attack by endogenous and exogenous sources. Endogenous DNA damage arises from chemical instability (depurination), oxygen free radicals, and misincorporation of nucleotides by DNA polymerases, whereas exogenous sources of DNA damage include environmental agents such as ultraviolet radiation, ionizing radiation, and various genotoxic agents [1,2]. Persistence of DNA damage can lead to genomic instability, apoptosis, or mutagenesis, which can eventually lead to carcinogenesis. Fortunately, the cell has developed various DNA damage checkpoints and DNA repair systems to deal with genomic insults.
DNA repair systems efficiently remove damaged DNA through several pathways. Abasic DNA lesions account for a large percentage of total DNA damage and are repaired through the base excision repair (BER) system. The BER system is composed of two subpathways: single-nucleotide BER, also referred to as short-patch (SP)-BER, and multinucleotide BER, also referred to as long-patch (LP)-BER. A DNA glycosylase recognizes the damaged base and cleaves the glycosidic bond between the sugar and base leaving an apurinic/apyrimidinic (AP) or abasic site [3,4]. DNA glycosylases can be monofunctional or bifunctional. In the case of a monofunctional DNA glycosylase, AP-endonuclease 1 (APE1) incises the 5′ end of the resulting AP site to generate the free 3′-hydroxyl required by the repair polymerase. Bifunctional glycosylases not only remove the aberrant base but also cleave the AP site on the 3′ side. The remaining deoxyribose phosphate (dRP) residue is then removed by APE1 [5]. Progress of the SP-BER or LP-BER pathway depends on the type of DNA damage. During SP-BER, the dRP lyase activity of DNA polymerase β (Pol-β), which is located in the 8-kDa domain of Pol-β, removes the 5′ dRP left behind by APE1, generating a 5′ phosphorylated nicked DNA strand [6]. Pol-β then uses its polymerase activity, which is located in its 31-kDa C-terminal domain, to insert the correct base at the site of damage. The remaining nick is sealed by DNA ligase I [7].
Mutations in the adenomatous polyposis coli (APC) gene are among the earliest events leading to the development of colorectal cancer [8,9]. The APC gene, located on chromosome 5q21–q22, consists of 8535 nt, and spans 21 exons. The APC gene encodes a 310-kDa protein, which is composed of 2843 amino acids in its common isoform [10]. It is present in both the cytoplasm and the nucleus. Approximately 75% of the coding sequence is located on exon 15, which seems to be the most common target for both germ line and somatic mutations of APC [11]. The mutation cluster region (MCR) is located within codons 1280 to 1513 and harbors more than 60% of all somatic mutations in the APC gene [12]. Mutations that occur in the MCR of APC result in a truncated protein that lacks most of the C-terminus required for proper APC function: the basic domain, which is the site of microtubule binding, and binding sites for EB1 and hDLG [13]. APC is also involved in Wnt signaling and β-catenin regulation, cell-cell adhesion, apoptosis, cell migration, chromosomal instability, and cell cycle control [10,14–17].
Fen1 is a structure-specific nuclease that is pivotal in maintaining genome stability through its involvement in DNA replication and repair. Fen1 is essential for survival as Fen1-knockout in mice is embryonic lethal. Mutations in Fen1 have been implicated in various diseases including Huntington disease, myotonic dystrophy, and cancer [18,19]. Indeed, Fen1 may be a novel tumor suppressor [20,21]. In addition to its 5′-flap endonuclease activity, Fen1 also possesses 5′-3′ exonuclease activity, which is required for the removal of displaced RNA-DNA primers during discontinuous lagging-strand replication and Okazaki fragment maturation [22,23]. Mutations in Fen1 increase genomic instability by causing chromosomal loss and accumulation of toxic flaps [24], whereas abnormal expression of Fen1 may cause defective BER and, thus, accumulation of mutations that can lead to genomic instability [25,26].
In previous studies, we demonstrated that APC interacts with the BER proteins Pol-β and Fen1 and that this interaction blocks BER through three different mechanisms: (i) APC blocks BER by interacting with Pol-β and blocking Pol-β-directed strand displacement synthesis in LP-BER; (ii) APC interacts with Pol-β and inhibits its deoxyribose phosphate (dRP) lyase activity, thus blocking SP-BER; and (iii) APC interacts with Fen1, inhibiting its 5′-flap endonuclease and 5′-3′ exonuclease activities, thus blocking LP-BER [27]. In previous studies, the effect of APC on BER was examined by using a 20-amino acid fragment of APC containing the DNA repair inhibitory (DRI) domain in an in vitro reconstituted BER system [27–30]. In this report, we provide more detail on how APC interacts with Fen1 and restricts its retention in cytoplasm to impair its participation in LP-BER in the nucleus of cultured breast and colon cancer cell lines. These findings suggest that in normal colon and breast epithelial cells the levels of wild-type APC levels are low and probably do not interfere with Fen1 and thus LP-BER activity. However, after treatment with environmental carcinogens cytoplasmic APC levels increase, resulting in the retention of Fen1 in the cytoplasm as well as sequestration of any nuclear Fen1 into the nucleolus. The combined reduction in Fen1 level and sequestration of nuclear Fen1 inhibit Fen1-mediated LP-BER. This inhibited or compromised LP-BER ultimately results in accumulation of mutations leading to the transformation of normal epithelial cells.
Materials and Methods
Yeast Cell Maintenance
The yeast strain PJ694-A was used for all yeast two-hybrid experiments. The yeast strain PJ69-4A (MATa trp1-901 leu2-3,112ura3-52 his3-200 gal4Δ gal80Δ GAL2-ADE2 LYS2::GAL1-HIS3 met2:: GAL7-lacZ) was grown on synthetic dropout medium lacking lysine (0.17% Difco yeast nitrogen base without amino acids, ammonium sulfate (5.0 g/L), complete supplemental amino acid mixture minus appropriate amino acids containing 2% glucose).
Preparation of Nuclear and Cytoplasmic Fractions
The nuclear extract and cytoplasmic fractions were prepared by the procedure described by Shapiro et al. [31]. APC, Fen1, lamin B, α-actin, and GAPDH protein levels were determined by Western blot analysis, as described in our previous studies [32].
In Vitro LP-BER Assay
The in vitro LP-BER assay reaction was composed of 2.5 nM APE1 nicked 32P-labeled 63-mer F-DNA, 5 µg of nuclear extract as a source of BER proteins, and 20 µM dNTPs. F-DNA is an AP-site analog (3-hydroxy-2-hydroxymethyltetrahydrofuran; noted as F) containing substrate, in which the F is inserted at the 24th position of the DNA oligomer [20]. The reaction was incubated at 37°C for 60 minutes after which the products were separated on a 15% acrylamide/7 M urea gel. The radioactive signals were visualized by autoradiography.
Immunohistochemistry of Breast and Colon Cancer Cell Lines in Culture
Human breast cancer cell line MB231 was plated into six-well plates containing sterilized coverslips. After 24 hours, cells were transfected for 24 hours with flag-tagged pAPC(1338), pAPC(1309), pAPC (1309mut), and pAPC(1211) plasmids using Lipofectamine transfection reagent (Invitrogen, Carlsbad, CA). One set of these cells without any transfection was treated for 24 hours with 25 µM benzo[a]pyrene (B[a]P). Similarly, HCT-116 and HCT-116-APC(KD) cells were treated with different concentrations of methyl methanesulfonate (MMS) for 24 hours. After transfection or treatment, cells were fixed with paraformaldehyde and incubated with antimouse Alexa 488 and antirabbit Alexa 594 antibodies (Invitrogen). Cells were washed and mounted onto slides using Vectashield mounting medium containing fluorescent 4,6-diamidino-2-phenylindole (DAPI) (Vector Laboratories, Burlingame, CA). Slides were viewed under a Zeiss upright microscope (Carl ZeissMicroscopy, Thornwood, NY) at wavelength 515 nm for Flag-APC and 640 nm for Fen1.
Immunohistochemistry of Breast Cancer and Normal Tissue Arrays
Human breast cancer tissue array slides were purchased fromBiomax, Inc (Rockville, MD). Slides contained both normal and breast cancer tissue embedded in paraffin. After deparaffinization, washed array slides were blocked with PBS containing 5% goat serum and 0.2% Triton X-100 and then incubated with mouse anti-Fen1 antibody (Abcam, Cambridge, MA) and rabbit anti-APC (Santa Cruz Biotechnology, Santa Cruz, CA) in PBS containing 5% goat serum and 0.2% Triton X-100 for 2 hours. The procedure for microscopic fluorescence signal detection was performed as described for cells in culture.
APC Protein Fragments
A 20-amino acid fragment of APC protein (1250-KVSSINQETIQTYCVEDTPI-1269), termed the DRI domain, was synthesized at the Protein Chemistry and Biomarkers core facility at the Interdisciplinary Center for Biotechnology Research of the University of Florida [28]. The sequence represents the DRI domain of wild-type APC (APCwt) or a mutated form of APC, APC(I-A,Y-A), in which amino acids Ile1259 and Tyr1262 were replaced with alanine (A). The amino acids Ile1259 and Tyr1262 of the DRI domain are the site of interaction with Pol-β and Fen1, located in the N-terminal region, and are spared by mutations of mutator cluster region (MCR) that result in truncation of the APC protein. Thus, most mutant APC proteins (those with an intact DRI domain), as well as wild-type APC, are capable of modulating BER [30]. We also cloned a longer APC protein expression (1–1309 aa) plasmid, which was generated from a natural nonsense mutation in colon cancer patients [33,34]. The 1-1309-aa protein fragment retains the DRI domain. We introduced mutations at residues I1259 and Y1262 of the DRI domain of this protein. An oligonucleotide containing FLAG and the N-terminal APC coding sequence were fused and inserted into the pcDNA5/FRT/TO vector (Invitrogen) using HindIII and XhoI sites (Table 1). This vector was used to introduce the APC (1–1309) coding plasmid into the BamHI and XhoI sites. BamHI and XhoI sites, which contained the start sequence of APC and the stop codon, respectively, were introduced in the APC complementary DNA (cDNA) by polymerase chain reaction (PCR) amplification. The upstream APC sequence was fused to the FLAG region and the oligonucleotide to introduce the stop codon in APC(1–1309) is shown in Table 1. The truncated APC construct carries a silent substitution at codon 1120G to introduce an NcoI site for efficient cloning. All PCR-amplified sequences were confirmed by direct sequencing.
Table 1.
Oligonucleotide Sequences for Different Primers and Substrates for Fen1.
| Experiment | Oligonucleotide Sequence |
| (1) TNT assay | |
| Fragment 1–115 | |
| Sense primer | 5′-TAATTTTAATACGACTCACTATAGGGAACAGCCACCATGGGAATTCAAGGCCTGGCC-3′ |
| Antisense primer | 5′-TTTTTTTTTTTTTTTTTTTTTTTTTTTTTACTGAGCCTGCTGCAGCTGCTTCTCTGCCTC 3′ |
| Fragment 78–192 | |
| Sense primer | 5′-TAATTTTAATACGACTCACTATGGGAACAGCCACCATGGGCATCAAGCCCGTGTAT-3′ |
| Antisense primer | 5′-TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTATCGCATTAGCACAGGGCTGCCGAACCT-3′ |
| Fragment 154–268 | |
| Sense primer | 5′-TAATTTTAATACGACTCACTATAGGGAACAGCCACCATGGATGCACCCAGTGAGGCA-3′ |
| Antisense primer | 5′-TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTAAGGGTACTTGTTGGGGTCAAGTCGCAC-3′ |
| Fragment 231–345 | |
| Sense primer | 5′-TAATTTTAATACGACTCACTATAGGGAACAGCCACCATGGGCAGTGACTACTGTGAGAGT-3′ |
| Antisense primer | 5′-TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTACTTGAAGAAATCATCCAGGCGGCCCTGGGT-3′ |
| Fragment 269–380 | |
| Sense primer | 5′-TAATTTTAATACGACTCACTATAGGGAACAGCCACCATGGTGCCAGAAAATTGGCTCCAC-3′ |
| Antisense primer | 5′-TTTTTTTTTTTTTTTTTTTTTTTATTTTCCCCTTTTAAACTTCCCTGCTGCCCCAGTCTTTGC-3′ |
| (2) Site-directed mutagenesis | |
| Set 1 mutant (Set 1 primer pair 1) | |
| Sense primer | 5′-GGCTCACTCTCTTCAGCTGCGCGCGCAGAGCCAGAACCCAAGGGATCC-3 |
| Antisense primer | 5′-GGATCCCTTGGGTTCTGGCTCTGCGCGCGCAGCTGAAGAGAGTGAGCC-3′ |
| Set 1 mutant (Set 1 primer pair 2) | |
| Sense primer | 5′-GGCTCACTCTCTTCAGCTGCGGCCGCAGAGCCAGAACCCAAGGGATCC-3′ |
| Antisense primer | 5′-GGATCCCTTGGGTTCTGGCTCTGCGGCCGCAGCTGAAGAGAGTGAGCC-3′ |
| Set 2 mutant (Set 2 primer pair 1) | |
| Sense primer | 5′-GAACCCAAGGGATCCACTGCGAAGGCCGCAAAGACTGGGGCAGCA-3′ |
| Antisense primer | 5′-TGCTGCCCCAGTCTTTGCGGCCTTCGCAGTGGATCCCTTGGGTTC-3′ |
| Set 2 (Set 2 primer pair 2) | |
| Sense primer | 5′-GAACCCAAGGGATCCACTGCGGCAGCCGCAAAGACTGGGGCAGCA-3′ |
| Antisense primer | 5′-TGCTGCCCCAGTCTTTGCGGCTGCCGCAGTGGATCCCTTGGGTTC-3′ |
| (3) FEN1 activity | |
| 23-mer upstream | 5′-CTAGATGCCTGCAGCTGATGCGC-3′ |
| 51-mer downstream | 5′-AACATTFTTTTTGTACGGATCCACGTGTACGGTACCGAGGGCGGGTCGACA-3′ |
| 63-mer complementary | 5′-AGATGCCTGCAGCTGATGCGCGGTACGGATCCACGTGTACGGTACCGAGGGCGGGTCGACA-3′ |
Structure of DNA Used for the Synthesis of Fen1 Protein Fragments for Transcription/Translation Assays
To identify the APC-binding site of Fen1, different Fen1 cDNA constructs were made for transcription/translation (TNT) assays. Fen1 protein fragments were expressed using the in vitro transcription/translation system and then the interaction of APC and Fen1 was analyzed by far-Western analysis. Primers for each protein fragment were designed to include a T7 promoter and a Kozak sequence in the sense primer and a stop codon and poly A tail in the antisense primer. Five fragments of Fen1 were designed (amino acids 1–115, 78–192, 154–268, 231–345, and 269–380). The primers used to generate the protein fragments are shown in Table 1.
Site-Directed Mutagenesis of Fen1
Two sets of Fen1 mutants (set 1: K354A/R355A/K356A and set 2: K365A/K366A/K367A) were generated using the QuickChange Site-directed mutagenesis kit (Stratagene, La Jolla, CA). The primer pairs used to generate set 1 and set 2 mutants are shown in Table 1.
Synthesis of Fen1 Protein Fragments, Fen1wt, and Fen1mut Proteins Using an In Vitro Transcription/Translation System
Fen1 protein fragments, NLS1mut, NLS2mut, and Fen1 wildtype proteins were synthesized using the TNT PCR Quick Kit from Promega (Madison, WI). The reaction was run per the instructions provided with the kit. The TNT-synthesized Fen1 protein fragments were analyzed with Western blot [23].
Far-Western Blot Analysis
APCwt and APC(I-A,Y-A) peptides were slot-blotted (0–15 µg) onto a polyvinylidene difluoride (PVDF) membrane (GE Healthcare, Piscataway, NJ) in a binding buffer containing 20 mM Tris-HCl at pH 7.4, 100 mM phosphate buffer at pH 7.4, 60 mM KCl, and 0.25% (vol/vol) Nonidet P-40 [27]. After blot analysis, the membrane was blocked with 5% (wt/vol) bovine serum albumin and washed three times with Tris-buffered saline with 0.025% (vol/vol) Tween-20 before incubating with in vitro TNT-synthesized human Fen1 protein fragments (described above), wild-type, NLS1mut [Fen1(K354A/K355A/K366A)], and NLS2mut [Fen1(K365A/K366A/K367A)] proteins. Binding was detected using mouse anti-Fen1 antibody (Novus Biologicals, Littleton, CO). The signals were detected using the enhanced chemiluminescence reagent (GE Healthcare).
Pull-down of Fen1 Wild-type and Fen1(K365A/K366A/K367A) Proteins from the Transcription/Translation Synthesis Reaction Mixture
Fen1wt and Fen1(K365A/K366A/K367A) mutant were pulled down by incubating with rabbit antihuman Fen1 antibody in immunoprecipitation buffer (30 mMHEPES at pH 7.5, 30mMKCl, 8mM MgCl2, 0.01% NP-40, and 2% glycerol) for 4 hours followed by the addition of 5% bovine serum albumin blocked Protein A Sepharose beads (Promega) for an additional 2 to 3 hours. Beads were washed three times with 30 mM HEPES at pH 7.5, 30 mM KCl, and 5% glycerol and then resuspended in reaction buffer and used for Fen1 activity assays or Western blot analysis.
Synthesis and Labeling of the Fen1 Substrate
The Fen1 substrate for 5′-flap endonuclease activity was synthesized as described in Table 1 [35]. The 51-mer downstream oligonucleotide has a flap of 11 nt. The Fen1 substrate for 5′-3′ exonuclease activity was made similarly except that the 11-nt flap from the 51-mer oligonucleotide was removed. The 51- and 40-mer downstream oligonucleotides were radiolabeled at the 5′ end with [γ-32P]ATP using T4 polynucleotide kinase (New England Biolabs, Woburn, MA). After annealing with a 63-mer complementary template as described in Table 1, the labeled and annealed probes were purified by using a nick column (GE Healthcare). All the three oligonucleotides were annealed at a molar ratio of 1:1:1.
Fen1 Activity In Vitro
The assays for Fen1 5′-flap endonuclease and 5′-3′ exonuclease activities were performed in a final volume of 20 µl. The reaction mixture contained 30 mM HEPES at pH 7.5, 30 mM KCl, 8 mM MgCl2, 1 mM dithiothreitol, 200 µg/ml bovine serum albumin, and the indicated amounts of Fen1 or Fen1(K365A/K366A/K367A) mutant. Then, 2.5 nM 32P-labeled flapped-DNA or nicked-DNA substrates were added to the reaction mixture and incubated at 37°C for 30 minutes. Reactions were terminated with stop solution containing 0.4% (wt/vol) SDS and 5 mM EDTA. The DNA was recovered by phenol/chloroform extraction followed by ethanol precipitation. The 11-nt DNA product from the 5′-flap endonuclease activity and 1-nt product from the 5′-3′ exonuclease activity were separated on a 15% acrylamide and 7 M urea gel and quantified by electronic autoradiography (InstantImager; Packard Instrument Co, Meriden, CT). Statistical analysis (n = 3) was performed using a t-test in SigmaPlot 12 graphing software.
Results
Nuclear Extracts with Higher Levels of Endogenous APC Are Less Competent in LP-BER
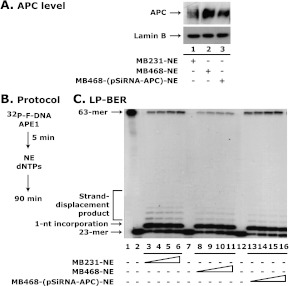
To establish the role of endogenous wild-type APC in blocking BER, we established an in vitro BER assay system with purified nuclear extracts (NEs) of MB231 and MB468 breast cancer cell lines. The MB231 cell line expresses a low level of APC, whereas the MB468 cell line expresses a high level of APC (Figure 1A, compare lane 1 with lane 2) [31]. We also knocked down the APC level in MB468 cells with pShRNA-APC [27,29,36,37]. We performed an LP-BER assay with these nuclear extracts (Figure 1B). Results of the LP-BER assay showed a protein concentration-dependent increase in LP-BER activity, which was higher withMB231-NE than MB468-NE (Figure 1C, compare lanes 3–6 with 8–11). Once we knocked-down APC in MB468 cells, the MB468-(pShRNA-APC)-NE exhibited higher LP-BER activity than MB468-NE (Figure 1C, compare lanes 8–11 with 13–16). These results clearly support our hypothesis that the presence of wildtype APC inhibits LP-BER.
Figure 1.
Nuclear extracts with increased APC protein levels have decreased LP-BER activity. (A) Western blot analysis of APC with purified nuclear extracts of MB231, MB468, and MB468-pSiRNA-APC cells. (B) Protocol for LP-BER with purified nuclear extracts. The in vitro LP-BER reaction was assembled by the addition of 5 µg of nuclear extract from pShRNA-APC knockdown and wild-type APC cells followed by the addition of 20 µM dNTPs and 32P-F-DNA substrate. (C) Autoradiogram of LP-BER. The nuclear extracts from these cell lines (0, 0.5, 1.0, 1.5, and 2.0 µg) were assembled with APE precut 32P-F-DNA and dNTPs and incubated for 90 minutes at 37°C. Lanes 1 and 2 show uncut and APE cut 63-mer 32P-labeled F-DNA. Data are representative of two independent experiments.
Overexpression of Truncated APC(1–1309) with and without Mutations at the DRI Domain Impairs Nuclear Localization of Fen1
In previous studies, we reported that the interaction of APC with Fen1 is necessary to block LP-BER [27]. In this study, we examined the mechanism by which APC interaction with Fen1 might perform this role. To gain a better understanding about the role of the functional interaction of APC with Fen1, we performed immunofluorescence analysis to examine colocalization in the MB231 breast cancer cell line, which expresses low levels of APC. We transiently overexpressed flag-tagged truncated APC(1–1309; amino acids 1–1309) protein. APC(1–1309) retains the DRI domain (amino acids 1245–1273), a critical region where Fen1 interacts with APC [27]. The APC(1–1309mut) contains alanine substitutions at amino acid residues I1259 and Y1262, which are important for the interaction with Fen1 [27]. Amino acids 1–1309 are a naturally occurring truncation of APC found in colorectal cancers [38].
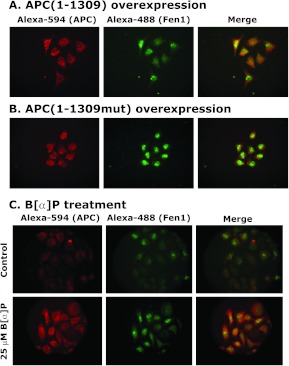
After 24 hours of transfection, colocalization of APC and Fen1 was determined by immunostaining with anti-APC and anti-Fen1 antibodies. We found a higher cytoplasmic and some nucleolar localization of overexpressed truncated APC proteins inMB231 cells compared with the control cells (Figure 2, A and B). Localization of APC(1-1309) in the cytoplasm and nucleolus was consistent with previous observations [33,39,40]. Punctate staining of APC(1–1309) can be seen in the cytoplasm, but it is not visible at the edges of the cells, probably because of the lack of the C-terminal end of APC(1–1309), which has the EB1-binding sequence necessary for microtubule association [41]. Fen1 showed decreased nucleolar and diffuse cytoplasmic staining with increased interaction with APC(1–1309) (Figure 2A).
Figure 2.
Colocalization of APC(1-1309) and Fen1 in MB231 cells after treatment with B[a]P. (A and B) Cells were transfected with flag-pAPC(1–1309) or flag-pAPC(1–1309mut) plasmids for 24 hours. MDA-MB231 cells were stained for APC (Alexa-594, red) and Fen1 (Alexa-488, green). The localization of flag-APC and Fen1 was determined by fluorescence microscopy. Images were merged to determine the colocalization of these proteins. Data are representative of three independent experiments. (C) Immunofluorescence analysis of wildtype APC and Fen1 colocalization in MB231 cells treated with B[a]P. Cells were treated with 25 µMB[a]P for 24 hours. The colocalization of endogenous APC and Fen1 was determined by fluorescence microscopy. Images were merged to determine the colocalization of these proteins. Data are representative of three independent experiments.
Treatment with DNA-Damaging Agent B[a]P and/or MMS Increases the Interaction of APC with Fen1 in the Cytoplasm and Impairs Nuclear Translocation in Breast and Colon Cancer Cell Lines
B[a]P is a potent breast carcinogen [42–44]. To mimic the APC overexpression conditions, we treated MB231 breast cancer cells with 25 µM B[a]P for 24 hours and then determined the colocalization of APC and Fen1. Results showed increased cytoplasmic levels of endogenous APC in B[a]P-treated cells compared with controls (Figure 2C, compare lower panel with upper panel). In control cells, Fen1 localization was diffuse and mainly localized in the nucleolus. However, after B[a]P treatment, the level of Fen1 increased in both cytoplasm and nucleolus with increased colocalization with APC (Figure 2C, compare lower panel with upper panel).
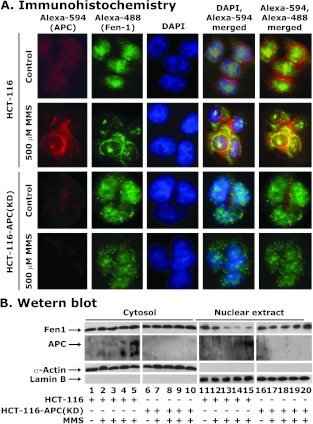
To further confirm our findings that APC blocks translocation of nuclear Fen1, we used colon cancer cell lines HCT-116 (expressing either wild-type APC) or HCT-116-APC(KD) (APC knockdown by means of stable expression of pShRNA-APC) [29]. These cells were treated with 500 µM of MMS, which causes methylation DNA damage that is repaired by BER [45]. After treatment, cells were processed either for immunofluorescence or for Western blot analysis. The immunofluorescence data with HCT-116 cells containing wild-type APC showed an increased colocalization of APC and Fen1 in the cytoplasm afterMMS treatment. The nuclear APC and Fen1 levels were reduced, and if present, they were localized in the nucleolus (Figure 3). Conversely, in HCT-116-APC(KD) cells, the translocation of nuclear Fen1 was not affected after MMS treatment (Figure 3), suggesting that, in the absence of APC, Fen1 is not sequestered in the cytoplasm.
Figure 3.
Colocalization of wild-type APC and Fen1 in HCT-116 cells after treatment with MMS. (A) HCT-116 and HCT-116-APC(KD) cells were treated with 500 µM MMS for 24 hours. Cells were stained for APC (Alexa-594, red), Fen1 (Alexa-488, green), and heterochromatin (DAPI, blue). The localization of flag-APC, Fen1, and heterochromatin was determined by fluorescence microscopy. Images were merged to determine the colocalization of these proteins. Data are representative of three independent experiments. (B) Western blot analysis. Cells were treated with 0, 50, 100, 250, and 500 µM MMS for 24 hours. The cytoplasmic and nuclear fractions were prepared and used for the determination of APC and Fen1 levels. β-Actin and lamin B levels were determined for protein loading normalization and for identifying potential cross-contamination of the cytoplasmic and nuclear fraction preparations. Data are representative of two independent experiments.
Next, we confirmed the immunofluorescence data by Western blot analysis. We treated HCT-116 and HCT-116-APC(KD) cell lines with different concentrations of MMS and examined cytosol and nuclear extracts for APC and Fen1 protein levels after MMS treatment. APC protein level increased in a dose-dependent fashion with a greater increase in cytosol than in nucleus (Figure 3B, compare lanes 1–5 with lanes 11–15, respectively). Concomitantly, cytosolic Fen1 protein decreased in nuclear extracts but not in the cytoplasmic fraction (Figure 3B, compare lanes 1–5 with lanes 11–15, respectively). As expected, in HCT-116-APC(KD) cells the nuclear Fen1 level increased after MMS treatment compared with HCT-116 cells (Figure 3B, compare lanes 16–20 with lanes 11–15). These results indicate that APC interrupts the entry of Fen1 to the nucleus afterMMS treatment.
Reduced Levels of APC Increase Nuclear Fen1 Levels and LP-BER Activity
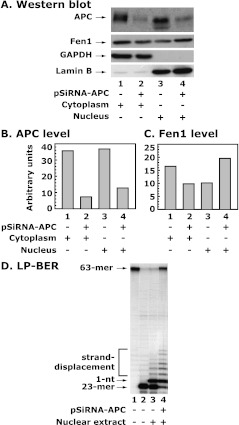
Because wild-type and truncated APC(1-1309) shuttles between cytoplasm and nucleus [33], overexpression of APC(1–1309) in transfected colon cancer cells may limit our conclusion with regard to colocalization. Therefore, we returned to MB468 breast cancer cell line, which expresses high levels of APC without transfection of foreign genetic material, depleted endogenous APC by knockdown with pSiRNA-APC, and prepared cytoplasmic and nuclear (combined nucleolar and nuclear plasma fraction) extracts. We determined the levels of APC and Fen1 by Western blot and performed BER assays (Figure 4). The results indicate that 1) transfection with pSiRNA-APC reduces the levels of APC in both the cytoplasm and nucleus (by 80% and 65%, respectively; Figure 4, A and B, compare lanes 1 and 3 with lanes 2 and 4), 2) the reduced level of APC was associated with altered levels of Fen1 in both fractions, with decreased Fen1 in the cytoplasm and increased Fen1 in the nucleus (Figure 4, A and C, compare lane 2 with lane 4), and 3) the increased nuclear Fen1 after APC knockdown was associated with increased Fen1-mediated LP-BER (Figure 4D, compare lanes 3 and 4). The decreased level of Fen1 in the cytoplasm of APC knockdown MB468 cell extract was evident and a significant increase in nuclear Fen1 was observed after APC knockdown (Figure 4, A and C, lanes 3 and 4). These results suggest that APC knockdown results in the decreased level of nuclear Fen1.
Figure 4.
APC knockdown increases Fen1 levels and activity in the nucleus. APC knockdown in MB468 breast cancer cells was accomplished by transfection with pSiRNA-APC, as shown in Figure 1. (A) The cytoplasmic and nuclear fractions were prepared and used for the determination of APC and Fen1 levels. GAPDH and lamin B protein levels were determined for protein loading normalization and for identifying potential cross-contamination of the cytoplasmic and nuclear fraction preparations. (B and C) Quantitative analysis of APC and Fen1 proteins levels. (D) The nuclear extract was also used for the determination of LP-BER. Lanes 1 and 2 show 32P-labeled 63-mer F-DNA and APE1 incised 23-mer product, respectively. Data shown are representative of two independent experiments.
Mapping of Fen1 for Interaction with APC
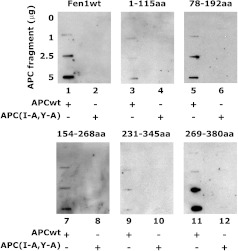
To identify the Fen1 amino acid residues that might be important for interaction with APC, we synthesized six different overlapping fragments: 1–380 aa (wild-type), 1–115 aa, 78–192 aa, 154–268 aa, 231–345 aa, and 269–380 aa. Synthesis of the protein fragments was confirmed by Western blot analysis. The interaction of Fen1 fragments with APC was determined by far-Western analysis. The Fen1 protein fragments of amino acids 1–380, 78–192, 154–268, and 269–380 interacted with the APCwt fragment but not with the APC(I-A,Y-A) mutant fragment (Figure 5). These results suggested that APC interacts with Fen1 fragments 78–192 and 154–268, which are part of Fen1's nuclease domain with an additional site outside the nuclease domain at residues 346–380.
Figure 5.
Interaction of APC and Fen1 protein fragments. APCwt and APC(I-A,Y-A) protein fragments were immobilized on a PVDF membrane and incubated with either wild-type or fragments of Fen1 protein as indicated. Equal amounts of TNT-synthesized proteins were used. Data are representative of three different determinations.
APC Interacts with the Nuclease Domain of Fen1
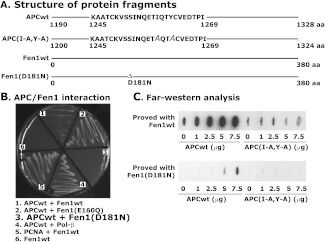
To observe the interaction of APC with Fen1 at amino acids 154–268 directly, we used the yeast two-hybrid (Y2H) assay. Because Fen1 amino acid residues Glu160 and Asp181 are important for its nuclease activity [46] and interaction with APC blocks this activity, we reasoned that the APC interaction might occur near these residues. We introduced mutations E160Q and D181N into Fen1 and assessed the mutant proteins for interaction with APC. Full-length and mutant Fen1 were cloned into the Y2H vector pGAD-C3. The APCwt fragment, which contained the DRI domain (amino acids 1190–1328), and the APC mutant (mut) fragment, which contained a mutated DRI domain (amino acids 1200–1324 with I1259A and Y1262A mutations) were cloned into the Y2H vector pGBDU-C3 (Figure 6A). The D181N mutation abolished Fen1-APC interaction; however, the E160Q mutated protein retained strong binding (Figure 6B). The Fen1(E160Q) mutant, which retains wild-type-like binding ability, but partial catalytic activity, exhibited strong APC binding, indicating that mutation of Fen1 at Glu160 retains the ability to interact with APC. The interaction of PCNA and Pol-β was used as a positive control, whereas the APC binding domain was used as a negative control to check for background colony formation in Y2Hassays. We verified the Y2H results by far-Western analysis. We purified His-tagged wild-type Fen1 and mutant Fen1(D181N) proteins and blotted them against wild-type (1250–1269 aa) and mutant (1250–1269 aa with I1259A and Y1262A substitutions) APC peptides. The results showed increased binding of Fen1 with wild-type (APCwt) but not with the APC(I-A, Y-A) fragment (Figure 6C). Fen1 (D181N) showed drastically reduced binding activity with APCwt and no binding activity with APC(I-A,Y-A) peptides (Figure 6C). These results indicate that Asp181 in the nuclease domain of Fen1 is critical for interaction with APC; however, the remaining poor binding exhibited by Fen1(D181N) indicated the presence of an additional binding site that might also be involved in the interaction with APC. These results show that APCwt interacts with Fen1 in vivo at specific residues.
Figure 6.
Interaction of APC with Fen1 by a yeast two-hybrid system. (A) Structure of APC and Fen1 constructs. The mutated amino acid residues I and Y with A are shown with italics. (B) Yeast two-hybrid analysis of APC and Fen1 interaction. (C) Far-Western analysis of APC and Fen1 showing that the D181N mutation in Fen1 reduces APC binding. Data are representative of three independent experiments.
APC Interacts with the Nuclear Localization Signal Domains of Fen1
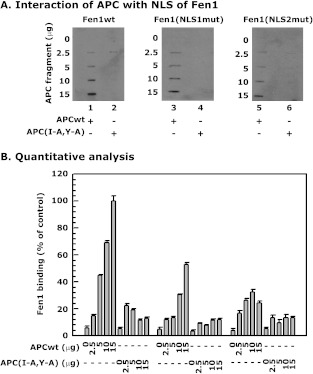
Next, we identified the residues in the Fen1 346–380 fragment that interacted with APC in the far-Western analysis (Figures 5 and 6). The most striking feature of these residues is that most of them form part of the Fen1 nuclear localization signal (NLS) [21]. Several other proteins including Werner protein (WRN), Bloom syndrome protein (BLM), and endonuclease G (EndoG) interact with Fen1 at residues 353–380 [47,48], which overlaps the region that interacts with APC, according to far-Western analysis [27]. On the basis of the previously mentioned information, we mutated two sets of Fen1 residues (NLS1 = Lys354Arg355Lys356 and NLS2 = Lys365Lys366Lys367) to determine whether they participated in the interaction of Fen1 with APC. Each set was chosen for its importance in nuclear localization, which was located in the same region as that identified for the interaction of Fen1 withWRN, BLM, and EndoG. Each amino acid was substituted with alanine and the Fen1 mutants were synthesized using the TNT-PCR Quick Kit. Binding of Fen1 (NLS1mut) and Fen1 (NLS2mut) was determined by far-Western analysis. Both Fen1 (NLS1mut) and Fen1 (NLS2mut) showed reduced binding to the APCwt peptide fragment (Figure 7, A and B). The remaining binding of APC to Fen1 (NLS1mut) and Fen1 (NLS2mut) could have been due to its interaction with the nuclease domain (Asp181). These results suggest that APC interacts with both the nuclease and the NLS domains of Fen1.
Figure 7.
Interaction of APC with NLS domains of Fen1. (A) Far-Western analysis of the interaction of APC with Fen1wt, Fen1 (NLS1mut), and Fen1 (NLS2mut) protein fragments. APCwt and APC(I-A,Y-A) peptides were immobilized on a PVDF membrane and incubated with either wild-type or fragments of Fen1 protein as indicated. (B) Percent binding of APCwt with Fen1 wild-type and mutant proteins. Percent binding was determined by using the density of APC and Fen1 wild-type binding band as 100% binding and comparing Fen1 mutant proteins and APC binding to the density of the APC and Fen1 wild-type band [(density of APCwt and Fen1 mutant binding/density of APCwt and Fen1 wild-type) x 100 = percent bound)]. The experiment was repeated twice.
Fen1 (NLS2mut) Retains Catalytic Activity
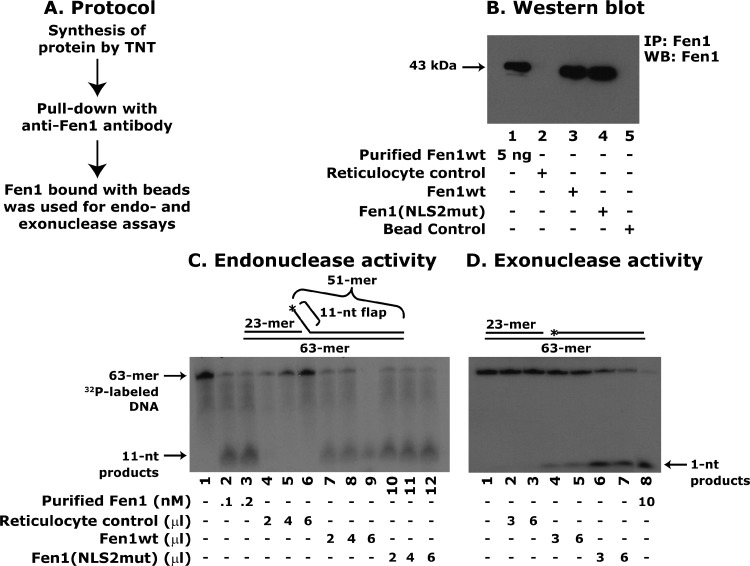
Fen1 (NLS2mut) bound APC to a lesser extent than Fen1 (NLS1mut). Because previous experiments showed that APC blocks Fen1 catalytic activity [27] we examined the effect of APC-Fen1 (NLS2mut) interaction on Fen1 catalytic activity using a 32P-labeled flapped substrate. To use the TNT-synthesized Fen1wt and Fen1 (NLS2mut) for these assays, the proteins were immunoprecipitated using an anti-Fen1 antibody (Figure 8A). Protein levels were determined by Western blot analysis (Figure 8B). Equal amounts of Fen1wt or Fen1 (NLS2mut) proteins bound with beads were then used in the endonuclease reaction. The 5′-flap endonuclease activity of Fen1wt and Fen1 (NLS2mut) was determined by the presence of an 11-nt product. Interestingly, the Fen1(NLS2mut) cleaved the 5′-flap endonuclease substrate as efficiently as the wild-type protein (Figure 8C, compare lanes 7–9 to lanes 10–12) indicating that mutations in Fen1 at K365/K366/K367 are not important for 5′-flap endonuclease activity.
Figure 8.
Fen1(NLS2mut) retains 5′-flap endonuclease and 5′-3′ exonuclease activities. (A) Schematic of protocol for the pull-down of Fen1 proteins from the TNT reaction mixture. (B) Western blot analysis of Fen1 proteins synthesized by TNT. (C) Autoradiogram depicting 5′-flap endonuclease activity. Equal amounts of TNT-synthesized proteins were used in these assays. Purified Fen1wt protein at 0.2 nM was used as a positive control. A 11-nt cleaved endonuclease product is shown with an arrow. (D) Autoradiogram depicting 5′-3′ exonuclease activity. A 1-nt cleaved exonuclease product is shown with an arrow. Equal amounts of TNT-synthesized proteins were used in these assays. Data are representative of three different determinations.
The 5′-3′ exonuclease activity of Fen1 (NLS2mut) was also tested. To determine the effect of APC on Fen1 exonuclease activity, a nicked DNA substrate was used. We labeled the 63-mer nicked substrate with γ-32P[ATP] as described previously [27]. Fen1wt and Fen1 (NLS2mut) proteins were pulled down fromthe TNT reaction mixture as described previously. Beads with bound Fen1wt or Fen1 (NLS2mut) were used to assay exonuclease activity, which was determined by the appearance of a 1-nt product. Surprisingly, the Fen1 (NLS2mut) had more 5′-3′ exonuclease activity than the wild-type protein (Figure 8D, compare lanes 4 and 5 to lanes 6 and 7). These results indicate that the mutations in Fen1 at amino acids Lys365Lys366Lys367 are important for regulating its catalytic activity and ensuring that Fen1 does not remove nucleotides adjacent to the primary cleavage site. Therefore, APC binding to Fen1 Lys365Lys366Lys367 may block Fen1 activity by inducing a conformational change. Alternatively, the large size of APC may create steric hindrance.
Clinical Significance of APC and Fen1 Interaction in Breast Cancer
We used Oncomine's gene search function to locate microarray studies of expression of APC and Fen1 in breast cancer (https://www.oncomine.com). Oncomine processes and normalizes each dataset independently. For differential expression analysis, we used t statistics with false discovery rates as a corrected measure of significance. We sorted the results based on each class of analysis, noted the significance number of the studies produced, and then created a box plot. These microarray values were preprocessed for normalization by Oncomine. We compared the data from different studies and found that both APC and Fen1 gene expressions were significantly increased in breast carcinoma in comparison to adjacent normal or benign tissues (data not shown). Although these results could have major implications for the role of APC and Fen1 in breast tumori-genesis, their utility is limited because the available data do not include information about variables, such as age and ethnicity, and confounding factors, such as smoking status, alcohol use, and epidemiologic health disparity information such as socioeconomic status.
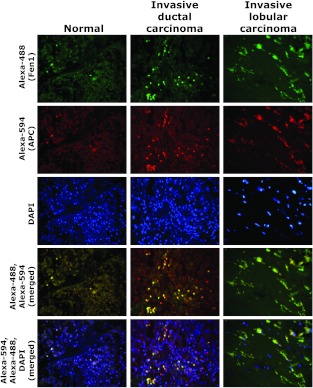
Then, we examined whether the increased APC and Fen1 mRNA levels were associated with increased protein levels and whether the proteins colocalized in breast tumor tissues. We obtained breast cancer tissue arrays from US Biomax, Inc (Rockville, MD) containing eight normal tissues, eight invasive ductal carcinoma, and four each of lobular and mucinous carcinoma. We performed multicolor labeling of normal and tumor samples using highly specific APC and Fen1 antibodies and determined their colocalization by fluorescence microscopy. Results showed increased colocalization of APC and Fen1 (yellow color in merged samples) in both invasive ductal carcinoma and invasive lobular carcinoma in comparison to normal breast tissues (Figure 9). Colocalization was increased in seven of the eight invasive ductal carcinomas and all four invasive lobular carcinomas; however, colocalization failed to occur in any of the mucinous carcinomas (data not shown). Although the number of samples examined in the present study was limited, the results warrant future studies with larger samples to establish a stronger link between APC/Fen1 interaction and DNA damage-induced breast carcinogenesis.
Figure 9.
APC/Fen1 interaction in human breast tumor tissues. Normal and breast tumor tissues were stained for APC (Alexa-594, red), Fen1 (Alexa-488, green), and nuclei (DAPI, blue). The localization of APC and Fen1 was determined by fluorescence microscopy. Images were merged to observe colocalization.
Discussion
In this report, we present data showing that APC regulates LP-BER by controlling access of Fen1 to the nucleus as well as by sequestering any nuclear Fen1 into the nucleolus. APC interacts with Fen1 at Fen1's two nuclear localization sequences (NLS1 and NLS2), which provides the rationale for failure of the complex to transit the nuclear envelope. However, the NLS sequences serve dual roles, the second being to prevent Fen1 from acting as a nonspecific endonuclease. This is the first report of APC regulating Fen1's cellular localization, which is an extension of previous work that identified an interaction between APC and Fen1 [27,29,30].
Knockdown of wild-type APC with a pShRNA-APC plasmid increases the nuclear pool of Fen1 and results in increased LP-BER activity and decreases DNA damage [36]. Here we show, that in contrast, exposure to genotoxic agents (DNA alkylating agents) or carcinogens results in overexpression of APC and increased DNA damage. These results are consistent with plasmid-based LP-BER assay system in cell culture [36]. Thus, based on our present and previous results, we conclude that the increased levels of APC associate with and block the entry of Fen1 to the nucleus, resulting in decreased LP-BER.
In vitro studies with purified proteins showed that APC interacts with two regions of Fen1 nuclease: the nuclease domain involving Asp181 and NLS domains NLS1 and NLS2 at the carboxyl terminus of the protein. Mutation at the nuclease domain (D181N) abolished Fen1 catalytic activity [49], whereas mutations at the NLS1 and NLS2 domains had no effect on catalysis but impaired nuclear localization. Our results using fluorescence microscopy further suggested that the interaction of APC with the NLS1 and NLS2 of Fen1 impairs nuclear translocation and makes it unavailable for LP-BER.
APC forms complexes through distinct domains with a variety of proteins in addition to Fen1. APC complexes with topoisomerase IIa at APC residues 1000–1326; participates in the G2/M transition by directly interacting with DNA through C-terminal residues 2140–2421, which inhibits replication and prevents cell cycle progression into or through S-phase; complexes with DNA PKcs through APC residues 1441–2077, which enhances the early response to DSB repair; and interacts with Pol-β and Fen1 to promote LP-BER. The formation of different protein/protein complexes through different APC domains and the ability to sense DNA damage are reminiscent of p53, which also acts as a sensor of DNA damage and cellular stress [50]. Some APC activities are particularly relevant to specific cell types, as in the development of colon and colorectal cancer, which is associated with truncation of APC, leading to blockage of LP-BER and genomic instability.
Fen1, like APC, plays various roles in DNA repair, replication, and cell cycle control through interaction of different domains with different proteins or with DNA itself. For instance, Fen1 promotes elongation and maturation of Okazaki fragments through residues 337–345 and interacts with endonuclease G (EndoG) at residues 349–367, whereas interactions with the DNA repair proteins WRN, APE1, and NEIL-1 and the Rad9-Rad1-Hus1 checkpoint and PCNA complex occur at several C-terminal amino acids. The C-terminus contains the nuclear localization sequences NLS1 and NLS2 that are positioned between residues 354 and 367. Indeed, like APC and p53, Fen1 is a highly versatile protein that performs various cellular functions, depending on its interaction with different proteins.
Both Fen1 and APC are critical to preventing colon and breast cancer by sensing DNA damage and promoting LP-BER. The evidence for involvement in colon cancer prevention is extensive: APC plays a tumor suppression role that is lost as APC is truncated at amino acids 1441–2077 that block DSB DNA repair [51]. The evidence for preventing breast cancer is more subtle but, nevertheless, compelling. Epigenetic inactivation of APC due to DNA methylation is frequently present in both breast cancer cell lines and breast cancer tissue. Epigenetic inactivation of APC affects its tumor suppression function. It has been observed that disruption of the APC/β-catenin pathway play an important role in breast cancer, whereas loss of APC expression and up-regulation of β-catenin have been described in human breast cancer and breast cancer cells [52,53]. We have previously demonstrated that genotoxic agents (DNA alkylating agents) transcriptionally induce APC level [32]. However, the function of nuclear APC has not been elucidated. We predict that nuclear APC has a negative regulatory role in DNA repair. The increased cytosolic level of APC results in increased Fen1 interaction thereby reducing Fen1 translocation to the nucleus, which is essential for its role in LP-BER. However, increased nuclear APC level depletes the freely available nuclear Fen1 by binding to it and leads to impaired LP-BER.
Although genetic mutations of APC or β-catenin are rarely observed outside colonic tumors [14,54–56], loss of heterozygosity at APC locus has been observed in 63% of breast cancer cell lines [57]. In another study, 6% alteration in APC protein level in breast tumors was detected [53]. A targeted constitutive mutation in the Apc tumor suppressor gene has been shown to cause mammary, but not intestinal, tumorigenesis [58]. However, APC can interact with numerous proteins critical for genome stability; modification of these interactions is likely to play major roles in carcinogenesis including breast cancer [59]. In fact, results with a hypomorphic mutant allele of Apc (Apc1572T) in mouse suggests that APC regulates specific dosages of Wnt/β-catenin signaling activity differentially, thus affecting tissue homeostasis and initiating breast tumorigenesis [59]. Also, a model in which K14-cremediated Apc heterozygosity (in which Apc exon 14 is flanked by loxP sequences) implicates APC in the development of mammary adenocarcinoma in APCmin/+ mice [60]. Notably, these mutations are located in the C-terminus required for interaction with Fen1 and well beyond the MCR region of Apc mutations frequently found in mouse gastrointestinal tumors [61–64]. Mutated Apc, whether frame-shifted or truncated, typically retains the DRI domain (amino acids 1245–1273) and remains capable of interacting with Fen1 to block LP-BER.
Furthermore, Oncomine data analysis and our immunohistochemical analysis data clearly indicate an increased association of APC with Fen1 in breast carcinoma in comparison to normal breast tissue. We have shown that, after treatment with environmental carcinogens, Fen1 [37] and APC expression [32,36,65,66] increase in spontaneously immortalized normal human breast epithelial cells, human colon cancer cells, and mouse embryonic fibroblasts. Because both Fen1 and APC are tumor suppressors [14,20,67], an increase in their expression would be expected to protect against tumorigenesis; however, this does not seem to be the case in breast and lung tumorigenesis. Instead, the increased expression of Fen1 in breast and lung cancer cells may reflect either increased proliferation of cancer cells or a response to increased DNA damage [68,69]. Therefore, it is possible that the interaction of Fen1 and APC not only blocks LP-BER but also impairs each other's tumor suppressor functions involved in neoplastic transformation.
In conclusion, these results demonstrate that APC and Fen1 participate in DNA repair in both cultured cells and primary cultures and provide the first direct evidence for how carcinogens promote Fen1 retention in the cytoplasm as well as sequestration in the nucleolus that results in reduced LP-BER and accumulation of abasic lesions that can accelerate the malignant transformation. These studies form the framework for development of novel agents that target APC/Fen1 interaction and the BER pathway for the prevention of breast and colorectal cancer development.
Acknowledgments
The authors thank Silvia Tornaletti for her critical evaluation of the article.
Abbreviations
- APC
adenomatous polyposis coli
- DRI domain
DNA repair inhibitory domain
- Fen1
flap endonuclease 1
- LP-BER
long-patch base excision repair
- NLS
nuclear localization signal
- Pol-β
DNA polymerase β
Footnotes
Financial support for these studies was provided by grants from the National Institutes of Health (R01-CA097031 and R01-CA100247 to S.N. and RO1-CA98664 to T.I.) and Flight Attendant Medical Research Institute, Miami, FL (CIA-072088) to S.N. The authors declare no potential conflicts of interest with respect to the authorship and/or publication of this article.
References
- 1.Cheng KC, Loeb LA. Genomic stability and instability: a working paradigm. Curr Top Microbiol Immunol. 1997;21:5–18. doi: 10.1007/978-3-642-60505-5_2. [DOI] [PubMed] [Google Scholar]
- 2.Houtgraaf JH, Versmissen J, van der Giessen WJ. A concise review of DNA damage checkpoints and repair in mammalian cells. Cardiovasc Revasc Med. 2006;7:165–172. doi: 10.1016/j.carrev.2006.02.002. [DOI] [PubMed] [Google Scholar]
- 3.Lindahl T, Nyberg B. Rate of depurination of native deoxyribonucleic acid. Biochemistry. 1972;11:3610–3618. doi: 10.1021/bi00769a018. [DOI] [PubMed] [Google Scholar]
- 4.Doetsch PW, Cunningham RP. The enzymology of apurinic/apyrimidinic endonucleases. Mutat Res. 1990;236:173–201. doi: 10.1016/0921-8777(90)90004-o. [DOI] [PubMed] [Google Scholar]
- 5.Huffman JL, Sundheim O, Tainer JA. DNA base damage recognition and removal: new twists and grooves. Mutat Res. 2005;7:55–76. doi: 10.1016/j.mrfmmm.2005.03.012. [DOI] [PubMed] [Google Scholar]
- 6.Matsumoto Y, Kim K. Excision of deoxyribose phosphate residues by DNA polymerase β during DNA repair. Science. 1995;269:699–702. doi: 10.1126/science.7624801. [DOI] [PubMed] [Google Scholar]
- 7.Mitra S, Boldogh I, Izumi T, Hazra TK. Complexities of the DNA base excision repair pathway for repair of oxidative DNA damage. Environ Mol Mutagen. 2001;38:180–190. doi: 10.1002/em.1070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bodmer WF, Bailey CJ, Bodmer J, Bussey HJ, Ellis A, Gorman P, Lucibello FC, Murday VA, Rider SH, Scambler P, et al. Localization of the gene for familial adenomatous polyposis on chromosome 5. Nature. 1987;328:614–616. doi: 10.1038/328614a0. [DOI] [PubMed] [Google Scholar]
- 9.Powell SM, Zilz N, Beazer-Barclay Y, Bryan TM, Hamilton SR, Thibodeau SN, Vogelstein B, Kinzler KW. APC mutations occur early during colorectal tumorigenesis. Nature. 1992;359:235–237. doi: 10.1038/359235a0. [DOI] [PubMed] [Google Scholar]
- 10.Narayan S, Roy D. Role of APC and DNA mismatch repair genes in the development of colorectal cancers. Mol Cancer. 2003;2:41. doi: 10.1186/1476-4598-2-41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Beroud C, Soussi T. APC gene: database of germline and somatic mutations in human tumors and cell lines. Nucleic Acids Res. 1996;24:121–124. doi: 10.1093/nar/24.1.121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Miyoshi Y, Nagase H, Ando H, Horii A, Ichii S, Nakatsuru S, Aoki T, Miki Y, Mori T, Nakamura Y. Somatic mutations of the APC gene in colorectal tumors: mutation cluster region in the APC gene. Hum Mol Genet. 1992;1:229–233. doi: 10.1093/hmg/1.4.229. [DOI] [PubMed] [Google Scholar]
- 13.Morrison EE. The APC-EB1 interaction. Adv Exp Med Biol. 2009;656:41–50. doi: 10.1007/978-1-4419-1145-2_4. [DOI] [PubMed] [Google Scholar]
- 14.Kinzler KW, Vogelstein B. Lessons from hereditary colorectal cancer. Cell. 1996;87:159–170. doi: 10.1016/s0092-8674(00)81333-1. [DOI] [PubMed] [Google Scholar]
- 15.Aoki K, Taketo MM. Adenomatous polyposis coli (APC): a multifunctional tumor suppressor gene. J Cell Sci. 2007;120:3327–3335. doi: 10.1242/jcs.03485. [DOI] [PubMed] [Google Scholar]
- 16.Rustagi AK. The genetics of hereditary colon cancer. Genes Dev. 2007;21:2525–2538. doi: 10.1101/gad.1593107. [DOI] [PubMed] [Google Scholar]
- 17.Brocardo M, Henderson B. APC shuttling to the membrane, nucleus and beyond. Trends Cell Biol. 2008;18:587–596. doi: 10.1016/j.tcb.2008.09.002. [DOI] [PubMed] [Google Scholar]
- 18.Zheng L, Dai H, Zhou M, Li M, Singh P, Qiu J, Tsark W, Huang Q, Kernstine K, Zhang X, et al. Fen1 mutations result in autoimmunity, chronic inflammation and cancers. Nat Med. 2007;13:812–819. doi: 10.1038/nm1599. [DOI] [PubMed] [Google Scholar]
- 19.Zheng L, Jia J, Finger LD, Guo Z, Zer C, Shen B. Functional regulation of FEN1 nuclease and its link to cancer. Nucleic Acids Res. 2011;39:781–794. doi: 10.1093/nar/gkq884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Henneke G, Friedrich-Heineken E, Hubscher U. Flap endonuclease 1: a novel tumour suppressor protein. Trends Biochem Sci. 2003;28:384–390. doi: 10.1016/S0968-0004(03)00138-5. [DOI] [PubMed] [Google Scholar]
- 21.Shen B, Singh P, Liu R, Qiu J, Zheng L, Finger LD, Alas S. Multiple but dissectible functions of FEN-1 nucleases in nucleic acid processing, genome stability and diseases. BioEssays. 2005;27:717–729. doi: 10.1002/bies.20255. [DOI] [PubMed] [Google Scholar]
- 22.Goulian M, Richards SH, Heard CJ, Bigsby BM. Discontinuous DNA synthesis by purified mammalian proteins. J Biol Chem. 1990;265:18461–18471. [PubMed] [Google Scholar]
- 23.Schweitzer JK, Livingston DM. Expansions of CAG repeat tracts are frequent in a yeast mutant defective in Okazaki fragment maturation. Hum Mol Genet. 1998;7:69–74. doi: 10.1093/hmg/7.1.69. [DOI] [PubMed] [Google Scholar]
- 24.Xu H, Zheng L, Dai H, Zhou M, Hua Y, Shen B. Chemical-induced cancer incidence and underlying mechanisms in Fen1 mutant mice. Oncogene. 2011;30:1072–1081. doi: 10.1038/onc.2010.482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Liu Y, Kao HI, Bambara RA. Flap endonuclease 1: a central component of DNA metabolism. Annu Rev Biochem. 2004;73:589–615. doi: 10.1146/annurev.biochem.73.012803.092453. [DOI] [PubMed] [Google Scholar]
- 26.Shibata Y, Nakamura T. Defective flap endonuclease 1 activity in mammalian cells is associated with impaired DNA repair and prolonged S phase delay. J Biol Chem. 2002;277:746–754. doi: 10.1074/jbc.M109461200. [DOI] [PubMed] [Google Scholar]
- 27.Jaiswal AS, Balusu R, Armas ML, Kundu CN, Narayan S. Mechanism of adenomatous polyposis coli (APC)-mediated blockage of long-patch base excision repair. Biochemistry. 2006;45:15903–15914. doi: 10.1021/bi0607958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Narayan S, Jaiswal AS, Balusu R. Tumor suppressor APC blocks DNA polymerase β-dependent strand displacement synthesis during long patch but not short patch base excision repair and increases sensitivity to methylmethane sulfonate. J Biol Chem. 2005;280:6942–6949. doi: 10.1074/jbc.M409200200. [DOI] [PubMed] [Google Scholar]
- 29.Balusu R, Jaiswal AS, Armas ML, Bloom LB, Narayan S. Structure/function analysis of the interaction of adenomatous polyposis coli with DNA polymerase β and its implications for base excision repair. Biochemistry. 2007;46:13961–13974. doi: 10.1021/bi701632e. [DOI] [PubMed] [Google Scholar]
- 30.Jaiswal AS, Narayan S. A novel function of adenomatous polyposis coli (APC) in regulating DNA repair. Cancer Lett. 2008;271:272–280. doi: 10.1016/j.canlet.2008.06.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Shapiro DJ, Sharp PA, Wahli WW, Keller MJ. A high-efficiency HeLa cell nuclear transcription extract. DNA. 1988;7:47–55. doi: 10.1089/dna.1988.7.47. [DOI] [PubMed] [Google Scholar]
- 32.Narayan S, Jaiswal AS. Activation of adenomatous polyposis coli (APC) gene expression by the DNA-alkylating agent N-methyl-N′-nitro-N-nitrosoguanidine requires p53. J Biol Chem. 1997;272:30619–30622. doi: 10.1074/jbc.272.49.30619. [DOI] [PubMed] [Google Scholar]
- 33.Henderson BR. Nuclear-cytoplasmic shuttling of APC regulates β-catenin subcellular localization and turnover. Nat Cell Biol. 2000;2:653–660. doi: 10.1038/35023605. [DOI] [PubMed] [Google Scholar]
- 34.Neufeld KL. Nuclear APC. Adv Exp Med Biol. 2009;656:13–29. doi: 10.1007/978-1-4419-1145-2_2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jaiswal AS, Narayan S. Assembly of the base excision repair complex on abasic DNA and role of adenomatous polyposis coli on its functional activity. Biochemistry. 2011;50:1901–1909. doi: 10.1021/bi102000q. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kundu CN, Balusu R, Jaiswal AS, Gairola CG, Narayan S. Cigarette smoke condensate-induced level of adenomatous polyposis coli blocks long-patch base excision repair in breast epithelial cells. Oncogene. 2007;26:1428–1438. doi: 10.1038/sj.onc.1209925. [DOI] [PubMed] [Google Scholar]
- 37.Kundu CN, Balusu R, Jaiswal AS, Narayan S. Adenomatous polyposis coli-mediated hypersensitivity of mouse embryonic fibroblast cell lines to methylmethane sulfonate treatment: implication of base excision repair pathways. Carcinogenesis. 2007;28:2089–2095. doi: 10.1093/carcin/bgm125. [DOI] [PubMed] [Google Scholar]
- 38.Smith KJ, Johnson KA, Bryan TM, Hill DE, Markowitz S, Willson JK, Paraskeva C, Petersen GM, Hamilton SR, Vogelstein B, et al. The APC gene product in normal and tumor cells. Proc Natl Acad Sci USA. 1993;90:2846–2850. doi: 10.1073/pnas.90.7.2846. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Neufeld KL, White RL. Nuclear and cytoplasmic localizations of the adenomatous polyposis coli protein. Proc Natl Acad Sci USA. 1997;94:3034–3039. doi: 10.1073/pnas.94.7.3034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Galea MA, Eleftheriou A, Henderson BR. ARM domain-dependent nuclear import of adenomatous polyposis coli protein is stimulated by the B56α subunit of protein phosphatase 2A. J Biol Chem. 2001;276:45833–45839. doi: 10.1074/jbc.M107149200. [DOI] [PubMed] [Google Scholar]
- 41.Su LK, Burrell M, Hill DE, Gyuris J, Brent R, Wiltshire R, Trent J, Vogelstein B, Kinzler KW. APC binds to the novel protein EB1. Cancer Res. 1995;55:2972–2977. [PubMed] [Google Scholar]
- 42.Hecht SS. Tobacco smoke carcinogens and breast cancer. Environ Mol Mutagen. 2002;39:119–126. doi: 10.1002/em.10071. [DOI] [PubMed] [Google Scholar]
- 43.Russo J, Tahin Q, Lareef MH, Hu YF, Russo IH. Neoplastic transformation of human breast epithelial cells by estrogens and chemical carcinogens. Environ Mol Mutagen. 2002;39:254–263. doi: 10.1002/em.10052. [DOI] [PubMed] [Google Scholar]
- 44.Han X, Naeher LP. A review of traffic-related air pollution exposure assessment studies in the developing world. Environ Internat. 2006;32:106–120. doi: 10.1016/j.envint.2005.05.020. [DOI] [PubMed] [Google Scholar]
- 45.Sobol RW, Wilson SH. Mammalian DNA β-polymerase in base excision repair of alkylation damage. Prog Nucleic Acid Res Mol Biol. 2001;68:57–74. doi: 10.1016/s0079-6603(01)68090-5. [DOI] [PubMed] [Google Scholar]
- 46.Shen B, Nolan JP, Sklar LA, Park MS. Essential amino acids for substrate binding and catalysis of human flap endonuclease 1. J Biol Chem. 1996;271:9173–9176. doi: 10.1074/jbc.271.16.9173. [DOI] [PubMed] [Google Scholar]
- 47.Sharma S, Sommers JA, Gary RK, Friedrich-Heineken E, Hubscher U, Brosh RM., Jr The interaction site of flap endonuclease-1 with WRN helicase suggests a coordination of WRN and PCNA. Nucleic Acids Res. 2005;33:6769–6781. doi: 10.1093/nar/gki1002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Guo Z, Chavez V, Singh P, Finger LD, Hang H, Hegde ML, Shen B. Comprehensive mapping of the C-terminus of flap endonuclease-1 reveals distinct interaction sites for five proteins that represent different DNA replication and repair pathways. J Mol Biol. 2008;377:679–690. doi: 10.1016/j.jmb.2007.10.074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Panda H, Jaiswal AS, Corsino PE, Armas ML, Law BK, Narayan S. Amino acid Asp181 of 5′-flap endonuclease 1 is a useful target for chemotherapeutic development. Biochemistry. 2009;48:9952–9958. doi: 10.1021/bi9010754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Levine AJ, Lane DP. The p53 Family. Singapore, Singapore: Cold Spring Harbor Press, Agency for Science and Technology Research; 2010. [Google Scholar]
- 51.Kouzmenko AP, Takeyama K, Kawasaki Y, Akiyama T, Kato S. Truncation mutations abolish chromatin-associated activities of adenomatous polyposis coli. Oncogene. 2008;27:4888–4899. doi: 10.1038/onc.2008.127. [DOI] [PubMed] [Google Scholar]
- 52.Ho KY, Kalle WH, Lo TH, Lam WY, Tang CM. Reduced expression of APC and DCC gene protein in breast cancer. Histopathology. 1999;35:249–256. doi: 10.1046/j.1365-2559.1999.00725.x. [DOI] [PubMed] [Google Scholar]
- 53.Jönsson M, Borg A, Nilbert M, Andersson T. Involvement of adenomatous polyposis coli (APC)/β-catenin signaling in human breast cancer. Eur J Cancer. 2000;36:242–248. doi: 10.1016/s0959-8049(99)00276-2. [DOI] [PubMed] [Google Scholar]
- 54.Morin PJ. β-Catenin signaling and cancer. BioEssays. 1999;21:1021–1030. doi: 10.1002/(SICI)1521-1878(199912)22:1<1021::AID-BIES6>3.0.CO;2-P. [DOI] [PubMed] [Google Scholar]
- 55.Polakis P. Wnt signaling and cancer. Genes Dev. 2000;14:1837–1851. [PubMed] [Google Scholar]
- 56.Schlosshauer PW, Brown SA, Eisinger K, Yan Q, Guglielminetti ER, Parsons R, Ellenson LH, Kitajewski J. APC truncation and increased β-catenin levels in a human breast cancer cell line. Carcinogenesis. 2000;21:1453–1456. [PubMed] [Google Scholar]
- 57.Thompson AM, Morris RG, Wallace M, Wyllie AH, Steel CM, Carter DC. Allele loss from 5q21 (APC/MCC) and 18q21 (DCC) and DCC mRNA expression in breast cancer. Br J Cancer. 1993;68:64–68. doi: 10.1038/bjc.1993.287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Gaspar C, Franken P, Molenaar L, Breukel C, van der Valk M, Smits R, Fodde R. A targeted constitutive mutation in the APC tumor suppressor gene underlies mammary but not intestinal tumorigenesis. PLoS Genet. 2009;5:1–13. doi: 10.1371/journal.pgen.1000547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Hattrup CL, Fernandez-Rodriguez J, Schroeder JA, Hansson GC, Gendler SJ. MUC1 can interact with adenomatous polyposis coli in breast cancer. Biochem Biophys Res Commun. 2004;316:364–369. doi: 10.1016/j.bbrc.2004.02.072. [DOI] [PubMed] [Google Scholar]
- 60.Kuraguchi M, Ohene-Baah NY, Sonkin D, Bronson RT, Kucherlapati R. Genetic mechanisms in Apc-mediated mammary tumorigenesis. PLoS Genet. 2009;5:1–14. doi: 10.1371/journal.pgen.1000367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Moser AR, Luongo C, Gould KA, McNeley MK, Shoemaker AR, Dove WF. ApcMin: a mouse model for intestinal and mammary tumorigenesis. Eur J Cancer. 1995;31A:1061–1064. doi: 10.1016/0959-8049(95)00181-h. [DOI] [PubMed] [Google Scholar]
- 62.Moser AR, Mattes EM, Dove WF, Lindstrom MJ, Haag JD, Gould MN. ApcMin, a mutation in the murine Apc gene, predisposes to mammary carcinomas and focal alveolar hyperplasias. Proc Natl Acad Sci USA. 1993;90:8977–8981. doi: 10.1073/pnas.90.19.8977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Kuraguchi M, Edelmann W, Yang K, Lipkin M, Kucherlapati R, Brown AM. Tumor-associated Apc mutations in Mlh1-/- Apc1638N mice reveal a mutational signature of Mlh1 deficiency. Oncogene. 2000;19:5755–5763. doi: 10.1038/sj.onc.1203962. [DOI] [PubMed] [Google Scholar]
- 64.Kuraguchi M, Yang K, Wong E, Avdievich E, Fan K, Kolodner RD, Lipkin M, Brown AM, Kucherlapati R, Edelmann W. The distinct spectra of tumor-associated Apc mutations in mismatch repair-deficient Apc1638N mice define the roles of MSH3 and MSH6 in DNA repair and intestinal tumorigenesis. Cancer Res. 2001;61:7934–7942. [PubMed] [Google Scholar]
- 65.Jaiswal AS, Narayan S. p53-dependent transcriptional regulation of the APC promoter in colon cancer cells treated with DNA alkylating agents. J Biol Chem. 2001;276:18193–18199. doi: 10.1074/jbc.M101298200. [DOI] [PubMed] [Google Scholar]
- 66.Jaiswal AS, Balusu R, Narayan S. 7,12-Dimethylbenzanthracene-dependent transcriptional regulation of adenomatous polyposis coli (APC) gene expression in normal breast epithelial cells is mediated by GC-box binding protein Sp3. Carcinogenesis. 2006;27:252–261. doi: 10.1093/carcin/bgi225. [DOI] [PubMed] [Google Scholar]
- 67.Parshad R, Price FM, Bohr VA, Cowans KH, Zujewski JA, Sanford KK. Deficient DNA repair capacity, a predisposing factor in breast cancer. Br J Cancer. 1996;74:1–5. doi: 10.1038/bjc.1996.307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Singh P, Yang M, Dai H, Yu D, Huang Q, Tan W, Kernstine KH, Lin D, Shen B. Overexpression and hypomethylation of flap endonuclease 1 gene in breast and other cancers. Mol Cancer Res. 2008;6:1710–1717. doi: 10.1158/1541-7786.MCR-08-0269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Sato M, Girard L, Sekine I, Sunaga N, Ramirez RD, Kamibayashi C, Minna JD. Increased expression and no mutation of the flap endonuclease (FEN1) gene in human lung cancer. Oncogene. 2003;22:7243–7246. doi: 10.1038/sj.onc.1206977. [DOI] [PubMed] [Google Scholar]