Abstract
Circadian rhythms are endogenous self-sustained oscillations with 24-hour periods that regulate diverse physiological and metabolic processes through complex gene regulation by “clock” transcription factors. The oral cavity is bathed by saliva, and its amount and content are modified within regular daily intervals. The clock mechanisms that control salivary production remain unclear. Our objective was to evaluate the expression and periodicity of clock genes in salivary glands. Real-time quantitative RT-PCR, in situ hybridization, and immunohistochemistry were performed to show circadian mRNA and protein expression and localization of key clock genes (Bmal1, Clock, Per1, and Per2), ion and aqua channel genes (Ae2a, Car2, and Aqp5), and salivary gland markers. Clock gene mRNAs and clock proteins were found differentially expressed in the serous acini and duct cells of all major salivary glands. The expression levels of clock genes and Aqp5 showed regular oscillatory patterns under both light/dark and complete-dark conditions. Bmla1 overexpression resulted in increased Aqp5 expression levels. Analysis of our data suggests that salivary glands have a peripheral clock mechanism that functions both in normal light/dark conditions and in the absence of light. This finding may increase our understanding of the control mechanisms of salivary content and flow.
Keywords: circadian clock, saliva, transcriptional regulation, circadian rhythms, Aqp5, expression pattern
Introduction
The oral cavity has a moist environment due to saliva, which is secreted from numerous major and minor glands in the oral cavity (Tucker and Miletich, 2010). The major, parotid, sublingual, and submandibular salivary glands (SGs) normally contribute over 90% of the total volume of unstimulated saliva (Edgar, 1990). SGs contain several cell types, including acinar cells, which are responsible for water and protein secretion, myoepithelial cells surrounding the acini and ducts, and ductal cells, which modulate the composition of the saliva (Gresik, 1994; Denny et al., 1997).
Saliva contains numerous enzymes, hormones, growth factors, immunoglobulins, and many other active molecules (Dawes, 1969; Levine, 1993). It has been shown that salivary flow rate follows circadian rhythms (Dawes, 1972). The secretion levels of salivary substances also follow circadian rhythms, which seem to be important for effective nutrition and self-defense (Dawes, 1974). These studies suggest that the SGs may contain a circadian clock to regulate the type, amount, and content of saliva. Indeed, the existence of such a clock mechanism has been demonstrated for other vital organs such as the kidney (Stow and Gumz, 2011). SGs and kidneys share physiological mechanisms involving a multitude of ion and water channels. Several genes that regulate fluid movement are highly expressed in both kidneys and SGs (Takata et al., 2004; Maeda et al., 2008). Little is known about the molecular mechanisms that control circadian saliva production and secretion. This study was undertaken to fill this important scientific gap.
A circadian clock is regulated by differential daily expression of 20 transcription factors called clock genes. There is a central clock in the brain and several peripheral clocks that can also act independently of the central clock. The central clock is composed of about 20,000 neurons, all of which express clock genes that oscillate in synchrony (Welsh et al., 1995; Ikeda, 2004). Clock genes are defined by a set of criteria that include rhythm in activity or amount as well as molecular evidence of a feedback mechanism (Williams and Sehgal, 2001; Myers et al., 2003). Clock genes transmit output signals that drive rhythms of gene expression in central and peripheral tissues. The most direct mechanism by which clock genes drive circadian gene expression is through regulation of promoter activity of clock-controlled genes. Clock genes can also indirectly drive circadian gene expression through regulation of promoter activity of clock-controlled genes, which in turn regulate the transcription of downstream genes at specific daily times (Kondratov et al., 2006).
Very recently, clock genes have begun to be discovered as key regulators of several diverse diseases, including cancer and diabetes (Krugluger et al., 2007; Mostafaie et al., 2009; Marcheva et al., 2010; Xia et al., 2010; Shimba et al., 2011). Saliva plays numerous protective roles for oral tissues maintenance (Mandel, 1989; Dowd, 1999). Adequate salivary flow and saliva content are directly related to health status. In contrast, reduced salivary flow rate is an indicator of xerostomia, which may be caused by SG disorders, such as Sjögren’s syndrome, and chronic sialoadenitis (Daniels and Fox, 1992; Mese and Matsuo, 2007). Reduced salivary flow is also found in cases of sialolithiasis, due to blockage of the ducts, and in post-radiation head and neck cancer patients (Rice, 1984; Mese and Matsuo, 2007). Many aspects of the biological mechanisms that result in xerostomia remain unclear. Regardless of the cause, patients with hyposalivation suffer from difficulties with swallowing and food mastication, dental caries, hampered speech, impaired taste, and nocturnal oral discomfort (Vissink et al., 2003a,b). Our study aims to increase understanding on how clock genes affect salivary flow rates and content. This knowledge may be applied in the future toward a better understanding of salivary gland physiology.
Materials & Methods
Animals and Circadian Experiments
Male C57BL/6J mice (from 8 to 12 wks of age) were purchased from Jackson Laboratory (Bar Harbor, ME, USA). Only male mice were used, to minimize potential sex-related differences in hormonal differences over circadian rhythms. The animal use was approved by the Animal Welfare Committee of the University of Michigan. For circadian experiments, all time interval calculations were based on the indicated zeitgeber (ZT, an event that provides the settings for a biological clock), and 6:00 a.m. was considered ZT 0 (detailed in the Appendix).
Human Tissues
Samples of human salivary glands were obtained from patients of the University of Michigan Hospital (Ann Arbor, USA) with prior signed informed consent form. All the material used for this study was collected from biopsies necessary for the diagnosis of salivary gland pathologies. The study was conducted in accordance with IRB regulations.
RNA Isolation and RT-PCR
Total RNA was isolated from freshly excised submandibular SGs and kidney tissues at the same time-point, with the use of TRIzol (Invitrogen, Carlsbad, CA, USA), and a 2-µg quantity of RNA was reverse-transcribed with TaqMan reverse transcription reagents (Applied Biosystems, Branchbury, NJ, USA). The submandibular gland is a mixed gland that secretes the highest amount of unstimulated saliva and contains both serous and mucous cells, making the submandibular gland a good model for evaluating the differential expression of clock RNAs and proteins in salivary glands. The resulting cDNA was then amplified by RT-PCR or real-time RT-PCR (detailed in the Appendix) with specific primers (Appendix Table 2).
In situ Hybridization, Immunohistochemistry, and Imaging
In situ hybridization (ISH) and immunohistochemistry (IHC) of SG sections were performed (described in the Appendix). Briefly, paraffin mouse submandibular, parotid, and sublingual salivary glands were used. At the end, sections were photographed on an Olympus microscope. Sense probes of mouse Bmal1 and Per2 were used as negative controls of ISH. As IHC negative controls, both omission of primary antibodies and omission of secondary antibodies were used.
Cell Culture and Transfection Studies
HEK293 (human embryonic kidney 293) cells were plated in 6-well culture plates at 70% confluence and transfected with 2 μg of Bmal1-pCMV (kindly provided by Dr. Lei Yin, Department of Molecular and Integrative Physiology, University of Michigan) or empty pCMV plasmid with Lipofectamine 2000 (Invitrogen; detailed in the Appendix). After transfection, RNA was reverse-transcribed into cDNA and used for quantitative RT-PCR.
Statistical Analysis
Statistical analyses were performed with Student’s unpaired t test. Each experiment was performed at least twice, and the representative data are presented as means ± SD of at least 3 independent replicates.
Results
Detection Clock RNAs by RT-PCR
Aryl hydrocarbon receptor nuclear translocator-like (Arntl or Bmal1), clock homolog (mouse) (Clock), period homolog 1 (Drosophila) (Per1), and period homolog 2 (Drosophila) (Per2) mRNAs were detected in mouse submandibular SG extracts by conventional RT-PCR (Fig. 1), as well as in the kidney, which is a tissue known to be regulated by clock genes (positive control). However, submandibular gland protein C (Smgc), a specific marker of SGs, can be detected only in SGs but not in the kidney. All the PCR products were sequenced to confirm clock RNA expression.
Figure 1.
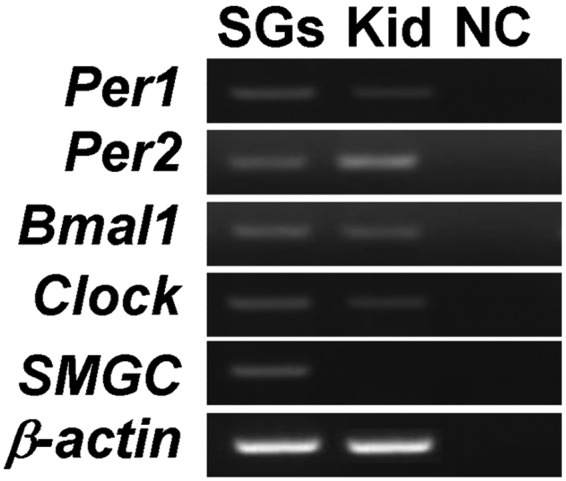
Clock gene RNAs (Per1, Per2, Bmal1, and Clock) were detected in both mouse SGs and the kidney. The SG marker (Smgc) was expressed only in SGs. SGs, salivary glands; kid, kidney; NC, negative control.
Localization of Clock Gene RNAs and Proteins in SGs
In situ hybridization results showed that Bmal1 and Per2 RNAs were detected strongly in the nucleus and cytoplasm of striated ducts and mucous acini, but weakly in serous acini cells (Figs. 2M, 2N). No positive staining was identified with a sense probe by in situ hybridization (data not shown). A similar expression pattern was found for clock proteins with IHC staining of mouse parotid (Figs. 2A, 2D, 2G, 2J), sublingual (Figs. 2B, 2E, 2H, 2K), and submandibular (Figs. 2C, 2F, 2I, 2L) SG sections. Four key clock proteins, Bmal1, Clock, Per1, and Per2, were found to be rarely expressed in serous acini, but their expression in mucous acini and striated ducts was very strong in submandibular SGs. In contrast, all 4 clock proteins were very strongly expressed in serous acini and mucous acini, as well as in striated ducts. We also tested human submandibular SG sections for PER1 expression. As in the mouse sections, strong expression was found in human mucous acini and striated ducts (Fig. 2O). Among the 4 clock proteins studied here, Clock and Per2 showed stronger expression levels than Bmal1 and Per1. Negative control sections (omission of the first antibody) showed absence of staining (data not shown).
Figure 2.
Clock gene products localization in SGs. Immunohistochemistry results showed that Bmal1 (A-C), Clock (D-F), Per1 (G-I), and Per2 (J-L) proteins were expressed in the nucleus of both acinar cells and in the epithelial cells of the ducts in mouse SGs. Expression was much stronger in duct cells. The same expression pattern was detected in human SGs (O). Bmal1 and Per1 RNAs were detected in the nuclear of serous acini and duct cells of mouse SGs by in situ hybridization (M, N). MA, mucous acini; SA, serous acini; SD, striated ducts; PSG, parotid salivary gland; SLSG, sublingual salivary gland; SMSG, submandibular salivary gland. Bars = 50 µm in A, B, D, E, G, H, J, K; = 20 µm in C, F, I, L, M-O.
Clock Genes Are Expressed in a Circadian Manner in Mouse SGs
We used real-time PCR to assess whether intestinal clock genes are expressed in a rhythmic circadian manner in mouse SGs under regular light/dark cycles. Rhythmic expression patterns were observed for two consecutive days for all clock RNAs studied in the SGs (Figs. 3A, 3C, 3E, 3G). Bmal1 and Clock RNAs showed highest expression levels at ZT 0 and lowest at ZT 12. In contrast, Per1 and Per2 RNAs showed strong expression levels at ZT 12 and low expression at ZT 0. The same expression pattern of individual clock gene RNAs was apparent on day 2 as on day 1, confirming the presence of a 24-hour cycle.
Figure 3.
Analysis of real-time PCR data showed that Clock, Bmal1, Per1, and Per2 RNAs are expressed in a rhythmic circadian manner in SGs under light/dark and dark/dark conditions (A-H). The mRNA levels are expressed as means of SE (n = 3 mice per time-point). All time interval calculations are based at the indicated zeitgeber (ZT, an event that provides the settings for a biological clock), and 6:00 a.m. was considered ZT 0.
To determine whether rhythmically expressed clock genes in the SGs were driven by the light-dark cycle, we kept the mice in constant darkness to eliminate the effect of light, with ad libitum access to food. Under dark/dark conditions, although clock gene RNAs showed reversed circadian expression, they still maintained rhythmic expression patterns in the SGs (Figs. 3B, 3D, 3F, 3H). Under dark/dark conditions, the circadian amplitude was decreased for Per1 and Per2 but not for Bmal1 and Clock.
Downstream Targets of Clock Genes
We then investigated whether clock genes regulate aquaporin 5 (Aqp5), solute carrier family 4 (anion exchanger), member 2 (Slc4a2 or Ae2a), and carbonic anhydrase 2 (Car2) expression. By promoter sequence analysis, we found an E-box putative binding site on the promoter of Aqp5 and a RRE-box putative binding site on the Car2 promoter (Appendix Table 1). Preferential binding of clock transcription factors has been shown on both E-box and RRE-box promoter sequences. In contrast, we did not find a putative clock-binding site on the Ae2a gene promoter. We then investigated whether clock genes regulated Aqp5, Ae2a, and Car2 expression in SGs in vitro.
We then tested if the expression of key functional genes, i.e., Aqp5, Car2, and Ae2a, follows a circadian pattern in SGs. Under light/dark conditions, Aqp5 and Ae2a showed the same rhythmic expression patterns as clock genes (Figs. 4A, 4C). In contrast, Car2 expression did not follow a circadian pattern (data not shown). Of interest, under dark/dark conditions, only Aqp5, but not Ae2a, showed a reproducible circadian pattern (Figs. 4B, 4D, respectively). We focused our subsequent studies on Aqp5.
Figure 4.
Expression of potential downstream targets of clock genes. Analysis of real-time PCR data showed that Aqp5 RNA is expressed in a circadian manner in SGs under light/dark and dark/dark conditions (A, B). In contrast, Ae2a showed a circadian pattern only in light/dark conditions (C) but not in complete-dark conditions (D). The AQP5 mRNA level in HEK293 was up-regulated after Bmal1 overexpression. All time interval calculations were based at the indicated zeitgeber (ZT 0 was considered to be 2 hrs after cell-cycle synchronization). The experiments were repeated 3 times, and 1 representative experiment is presented (*p < 0.05; compared with 0 hr).
HEK293, which is a Human Embryonic Kidney cell line, was used for the transfection experiments. We performed real-time quantitative RT-PCR to evaluate whether clock genes and AQP5 are expressed in HEK293 cells. Our results showed that, as in SGs, HEK293 cells also express BMAL1, CLOCK, PER1, PER2, and AQP5 (data not shown). After Bmal1 overexpression by transfection (Fig. 4E) and cell-cycle synchronization, up-regulated AQP5 expression levels were found in HEK293 cells (Fig. 4F). Of interest, AQP5 up-regulation also followed a circadian pattern (Fig. 4F).
Discussion
Clock genes, which generate circadian rhythms, are expressed in a circadian manner in the suprachiasmatic nucleus of the brain and in various peripheral tissues. Here, we showed that clock gene RNAs and clock proteins were differentially detected in SG cells. Especially strong expression was detected in serous acinar and duct cells, which indicates that clock genes may play important roles in salivary flow and salivary electrolyte flux regulation. This is the first detailed characterization of clock genes and their products, by in situ hybridization and immunohistochemistry, in all 3 major SGs.
To confirm the existence of circadian regulation in SGs, we then evaluated the oscillatory profiles of key clock gene RNAs, including Per1, Per2, Clock, and Bmal1, in submandibular glands. All clock genes studied here showed robust circadian expression rhythms in SGs when mice were housed under 12:12-hour light-dark (L/D) cycle conditions. Our data are consistent with those from a previous study which reported the expression of clock genes in submandibular glands (Furukawa et al., 2005). In addition to that study, our study also characterized, for the first time, the cellular localization of clock proteins in all 3 major SGs. More importantly, our study demonstrated that clock gene oscillation persists even in dark-dark conditions (D/D), suggesting that the circadian clock mechanism of SGs may also function independently of the central clock located in the brain. These findings provide a major advancement in our knowledge of SG physiology.
Previous studies showed that clock genes are the key regulators of circadian gene expression and daily physiological functions in many organs (Huang et al., 2011; Lefta et al., 2011). However, the molecular mechanisms that control circadian salivary secretion are unclear. We hypothesized that the SGs contain a peripheral clock that regulates their functions through differential expression of clock genes, the genes responsible for maintaining circadian rhythms. This hypothesis is in accordance with our data showing that clock genes continue to be expressed at regular daily intervals, even under D/D conditions. We also postulated that clock genes may regulate the expression of key SG genes such as Aqp5, Ae2a, and Car2. Our promoter analysis revealed that 2 out of 3 key SG genes (i.e., Aqp5 and Car2) contain putative promoter binding sites for clock protein binding, supporting a regulatory role for clock genes in SGs. Consistently, Aqp5 expression showed a clear circadian pattern under both normal light/dark conditions and under complete-dark conditions. Furthermore, Bmla1 overexpression resulted in AQP5 RNA up-regulation, suggesting that Aqp5 is a key target of clock genes in SGs. In contrast, Ae2a and Car2 did not show a definitive circadian pattern of expression, suggesting that these 2 genes may not be direct targets of clock genes in SGs.
The saliva secreted by the SGs fulfills many functions to maintain the normal homeostasis of the oral cavity (Dawes, 1969). The major constituent of saliva is water. The SGs also secrete ions, mainly K(+) and HCO3(−), and re-absorb Na(+) and Cl(−). The volume and the composition of the oral fluid can vary during the day within regular time intervals and within every individual. Salivation can be either stimulated or reduced by several factors and diseases. Hyposalivation, for example, is directly related to high caries, xerostomia, and pain, and much research is focused on treating salivary abnormalities. However, the direct links between causes and effects in different salivary pathologies are unclear. Recently, abnormal expression of clock genes has been found in patients with reduced salivary flow (our own unpublished observations). We therefore postulate that a clock mechanism that implies direct or indirect regulation of key genes important for SG physiology may be altered in diseases with abnormal salivary flow. We are currently evaluating the role of clock genes in salivary gland pathologies.
In conclusion, oscillations of clock genes were first detected in the SGs under light/dark but also under complete-dark conditions. Analysis of our data supports the idea that the SGs contain a circadian clock mechanism. Understanding the role of clock genes in controlling salivary flow and electrolyte content will enhance our understanding of the regulation of salivary functions and will provide a foundation for subsequent studies that may elucidate the potential links between clock genes and salivary diseases.
Footnotes
This research was supported by funds provided by the Department of Orthodontics and Pediatric Dentistry, School of Dentistry, University of Michigan; NIH/NCI cancer center core grant P30 CA46592 (start up funds to SP); and NIH/NIDCR grant DE 018878 (PP).
The authors declare no conflicts of interest with respect to the authorship and/or publication of this article.
A supplemental appendix to this article is published electronically only at http://jdr.sagepub.com/supplemental.
References
- Daniels TE, Fox PC. (1992). Salivary and oral components of Sjögren’s syndrome. Rheum Dis Clin North Am 18:571-589 [PubMed] [Google Scholar]
- Dawes C. (1969). The effects of flow rate and duration of stimulation on the concentrations of protein and the main electrolytes in human parotid saliva. Arch Oral Biol 14:277-294 [DOI] [PubMed] [Google Scholar]
- Dawes C. (1972). Circadian rhythms in human salivary flow rate and composition. J Physiol 220:529-545 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dawes C. (1974). Rhythms in salivary flow rate and composition. Int J Chronobiol 2:253-279 [PubMed] [Google Scholar]
- Denny PC, Ball WD, Redman RS. (1997). Salivary glands: a paradigm for diversity of gland development. Crit Rev Oral Biol Med 8:51-75 [DOI] [PubMed] [Google Scholar]
- Dowd FJ. (1999). Saliva and dental caries. Dent Clin North Am 43: 579-597 [PubMed] [Google Scholar]
- Edgar WM. (1990). Saliva and dental health. Clinical implications of saliva: report of a consensus meeting. Br Dent J 169:96-98 [DOI] [PubMed] [Google Scholar]
- Furukawa M, Kawamoto T, Noshiro M, Honda KK, Sakai M, Fujimoto K, et al. (2005). Clock gene expression in the submandibular glands. J Dent Res 84:1193-1197 [DOI] [PubMed] [Google Scholar]
- Gresik EW. (1994). The granular convoluted tubule (GCT) cell of rodent submandibular glands. Microsc Res Tech 27:1-24 [DOI] [PubMed] [Google Scholar]
- Huang W, Ramsey KM, Marcheva B, Bass J. (2011). Circadian rhythms, sleep, and metabolism. J Clin Investig 121:2133-2141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ikeda M. (2004). Calcium dynamics and circadian rhythms in suprachiasmatic nucleus neurons. Neuroscientist 10:315-324 [DOI] [PubMed] [Google Scholar]
- Kondratov RV, Shamanna RK, Kondratova AA, Gorbacheva VY, Antoch MP. (2006). Dual role of the CLOCK/BMAL1 circadian complex in transcriptional regulation? Faseb J 20:530-532 [DOI] [PubMed] [Google Scholar]
- Krugluger W, Brandstaetter A, Kallay E, Schueller J, Krexner E, Kriwanek S, et al. (2007). Regulation of genes of the circadian clock in human colon cancer: reduced period-1 and dihydropyrimidine dehydrogenase transcription correlates in high-grade tumors. Cancer Res 67:7917-7922 [DOI] [PubMed] [Google Scholar]
- Lefta M, Wolff G, Esser KA. (2011). Circadian rhythms, the molecular clock, and skeletal muscle. Curr Top Dev Biol 96:231-271 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levine MJ. (1993). Development of artificial salivas. Crit Rev Oral Biol Med 4:279-286 [DOI] [PubMed] [Google Scholar]
- Maeda S, Kuwahara S, Ito H, Tanaka K, Hayakawa T, Seki M. (2008). Expression and localization of aquaporins in the kidney of the musk shrew (Suncus murinus). J Histochem Cytochem 56:67-75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mandel ID. (1989). The role of saliva in maintaining oral homeostasis. J Am Dent Assoc 119:298-304 [DOI] [PubMed] [Google Scholar]
- Marcheva B, Ramsey KM, Buhr ED, Kobayashi Y, Su H, Ko CH, et al. (2010). Disruption of the clock components CLOCK and BMAL1 leads to hypoinsulinaemia and diabetes. Nature 466:627-631 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mese H, Matsuo R. (2007). Salivary secretion, taste and hyposalivation. J Oral Rehabil 34:711-723 [DOI] [PubMed] [Google Scholar]
- Mostafaie N, Kallay E, Sauerzapf E, Bonner E, Kriwanek S, Cross HS, et al. (2009). Correlated downregulation of estrogen receptor beta and the circadian clock gene Per1 in human colorectal cancer. Mol Carcinog 48:642-647 [DOI] [PubMed] [Google Scholar]
- Myers EM, Yu J, Sehgal A. (2003). Circadian control of eclosion: interaction between a central and peripheral clock in Drosophila melanogaster. Curr Biol 13:526-533 [DOI] [PubMed] [Google Scholar]
- Rice DH. (1984). Advances in diagnosis and management of salivary gland diseases. West J Med 140:238-249 [PMC free article] [PubMed] [Google Scholar]
- Shimba S, Ogawa T, Hitosugi S, Ichihashi Y, Nakadaira Y, Kobayashi M, et al. (2011). Deficient [sic] of a clock gene, brain and muscle Arnt-like protein-1 (BMAL1), induces dyslipidemia and ectopic fat formation. PLoS One 6:e25231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stow LR, Gumz ML. (2011). The circadian clock in the kidney. J Am Soc Nephrol 22:598-604 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takata K, Matsuzaki T, Tajika Y. (2004). Aquaporins: water channel proteins of the cell membrane. Prog Histochem Cytochem 39:1-83 [DOI] [PubMed] [Google Scholar]
- Tucker AS, Miletich I. (2010). Salivary glands: development, adaptations and disease. 1st ed. Basel: S. Karger Publishers [Google Scholar]
- Vissink A, Jansma J, Spijkervet FK, Burlage FR, Coppes RP. (2003a). Oral sequelae of head and neck radiotherapy. Crit Rev Oral Biol Med 14:199-212 [DOI] [PubMed] [Google Scholar]
- Vissink A, Burlage FR, Spijkervet FK, Jansma J, Coppes RP. (2003b). Prevention and treatment of the consequences of head and neck radiotherapy. Crit Rev Oral Biol Med 14:213-225 [DOI] [PubMed] [Google Scholar]
- Welsh DK, Logothetis DE, Meister M, Reppert SM. (1995). Individual neurons dissociated from rat suprachiasmatic nucleus express independently phased circadian firing rhythms. Neuron 14:697-706 [DOI] [PubMed] [Google Scholar]
- Williams JA, Sehgal A. (2001). Molecular components of the circadian system in Drosophila. Annu Rev Physiol 63:729-755 [DOI] [PubMed] [Google Scholar]
- Xia HC, Niu ZF, Ma H, Cao SZ, Hao SC, Liu ZT, et al. (2010). Deregulated expression of the Per1 and Per2 in human gliomas. Can J Neurol Sci 37:365-370 [DOI] [PubMed] [Google Scholar]





